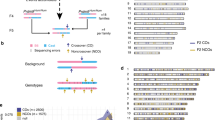
Abstract
Meiotic recombination differs between males and females; however, when and how these differences are established is unknown. Here we identify extensive sex differences at the initiation of recombination by mapping hotspots of meiotic DNA double-strand breaks in male and female mice. Contrary to past findings in humans, few hotspots are used uniquely in either sex. Instead, grossly different recombination landscapes result from up to fifteen-fold differences in hotspot usage between males and females. Indeed, most recombination occurs at sex-biased hotspots. Sex-biased hotspots seem to be partly determined by chromosome structure, and DNA methylation, which is absent in females at the onset of meiosis, has a substantial role. Sex differences are also evident later in meiosis as the rate at which meiotic breaks are repaired as crossovers differs between males and females in distal regions. The suppression of distal crossovers may help to minimize age-related aneuploidy that arises owing to cohesion loss during dictyate arrest in females.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing data are archived at the Gene Expression Omnibus (GEO) under accession GSE99921.
References
Lenormand, T., Engelstädter, J., Johnston, S. E., Wijnker, E. & Haag, C. R. Evolutionary mysteries in meiosis. Phil. Trans. R. Soc. Lond. B 371, 20160001 (2016).
Tortereau, F. et al. A high density recombination map of the pig reveals a correlation between sex-specific recombination and GC content. BMC Genomics 13, 586 (2012).
Khil, P. P., Smagulova, F., Brick, K. M., Camerini-Otero, R. D. & Petukhova, G. V. Sensitive mapping of recombination hotspots using sequencing-based detection of ssDNA. Genome Res. 22, 957–965 (2012).
Brick, K., Smagulova, F., Khil, P., Camerini-Otero, R. D. & Petukhova, G. V. Genetic recombination is directed away from functional genomic elements in mice. Nature 485, 642–645 (2012).
Smagulova, F., Brick, K., Pu, Y., Camerini-Otero, R. D. & Petukhova, G. V. The evolutionary turnover of recombination hot spots contributes to speciation in mice. Genes Dev. 30, 266–280 (2016).
Smagulova, F. et al. Genome-wide analysis reveals novel molecular features of mouse recombination hotspots. Nature 472, 375–378 (2011).
Pratto, F. et al. DNA recombination. Recombination initiation maps of individual human genomes. Science 346, 1256442–1256442 (2014).
Bishop, D. K., Park, D., Xu, L. & Kleckner, N. DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69, 439–456 (1992).
Peters, H. Migration of gonocytes into the mammalian gonad and their differentiation. Phil. Trans. R. Soc. Lond. B 259, 91–101 (1970).
Kong, A. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010).
Bhérer, C., Campbell, C. L. & Auton, A. Refined genetic maps reveal sexual dimorphism in human meiotic recombination at multiple scales. Nat. Commun. 8, 14994 (2017).
Cole, F. et al. Mouse tetrad analysis provides insights into recombination mechanisms and hotspot evolutionary dynamics. Nat. Genet. 46, 1072–1080 (2014).
Halldorsson, B. V. et al. The rate of meiotic gene conversion varies by sex and age. Nat. Genet. 48, 1377–1384 (2016).
Lenzi, M. L. et al. Extreme heterogeneity in the molecular events leading to the establishment of chiasmata during meiosis I in human oocytes. Am. J. Hum. Genet. 76, 112–127 (2005).
Perry, J., Palmer, S., Gabriel, A. & Ashworth, A. A short pseudoautosomal region in laboratory mice. Genome Res. 11, 1826–1832 (2001).
Lange, J. et al. The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167, 695–708.e16 (2016).
Kauppi, L. et al. Numerical constraints and feedback control of double-strand breaks in mouse meiosis. Genes Dev. 27, 873–886 (2013).
Shao, Z., Zhang, Y., Yuan, G. C., Orkin, S. H. & Waxman, D. J. MAnorm: a robust model for quantitative comparison of ChIP-Seq data sets. Genome Biol. 13, R16 (2012).
Davies, B. et al. Re-engineering the zinc fingers of PRDM9 reverses hybrid sterility in mice. Nature 530, 171–176 (2016).
Baudat, F., Imai, Y. & de Massy, B. Meiotic recombination in mammals: localization and regulation. Nat. Rev. Genet. 14, 794–806 (2013).
Narasimhan, V. M. et al. Health and population effects of rare gene knockouts in adult humans with related parents. Science 352, 474–477 (2016).
Walker, M. et al. Affinity-seq detects genome-wide PRDM9 binding sites and reveals the impact of prior chromatin modifications on mammalian recombination hotspot usage. Epigenetics Chromatin 8, 31 (2015).
Tease, C. & Hultén, M. A. Inter-sex variation in synaptonemal complex lengths largely determine the different recombination rates in male and female germ cells. Cytogenet. Genome Res. 107, 208–215 (2004).
Gruhn, J. R., Rubio, C., Broman, K. W., Hunt, P. A. & Hassold, T. Cytological studies of human meiosis: sex-specific differences in recombination originate at, or prior to, establishment of double-strand breaks. PLoS One 8, e85075 (2013).
Liu, E. Y. et al. High-resolution sex-specific linkage maps of the mouse reveal polarized distribution of crossovers in male germline. Genetics 197, 91–106 (2014).
Hunt, P. & Hassold, T. Female meiosis: coming unglued with age. Curr. Biol. 20, R699–R702 (2010).
de Boer, E., Jasin, M. & Keeney, S. Local and sex-specific biases in crossover vs. noncrossover outcomes at meiotic recombination hot spots in mice. Genes Dev. 29, 1721–1733 (2015).
de Boer, E., Stam, P., Dietrich, A. J. J., Pastink, A. & Heyting, C. Two levels of interference in mouse meiotic recombination. Proc. Natl Acad. Sci. USA 103, 9607–9612 (2006).
Yamada, S. et al. Genomic and chromatin features shaping meiotic double-strand break formation and repair in mice. Cell Cycle 16, 1870–1884 (2017).
Seisenberger, S. et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 48, 849–862 (2012).
Wang, H. et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 22, 1680–1688 (2012).
Bourc’his, D. & Bestor, T. H. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431, 96–99 (2004).
Vlachogiannis, G. et al. The Dnmt3L ADD domain controls cytosine methylation establishment during spermatogenesis. Cell Rep. 10, 944–956 (2015).
Zamudio, N. et al. DNA methylation restrains transposons from adopting a chromatin signature permissive for meiotic recombination. Genes Dev. 29, 1256–1270 (2015).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Collings, C. K., Waddell, P. J. & Anderson, J. N. Effects of DNA methylation on nucleosome stability. Nucleic Acids Res. 41, 2918–2931 (2013).
Persikov, A. V., Osada, R. & Singh, M. Predicting DNA recognition by Cys2His2 zinc finger proteins. Bioinformatics 25, 22–29 (2009).
Hammoud, S. S. et al. Chromatin and transcription transitions of mammalian adult germline stem cells and spermatogenesis. Cell Stem Cell 15, 239–253 (2014).
Kafer, G. R. et al. 5-hydroxymethylcytosine marks sites of DNA damage and promotes genome stability. Cell Rep. 14, 1283–1292 (2016).
Jain, D. et al. rahu is a mutant allele of Dnmt3c, encoding a DNA methyltransferase homolog required for meiosis and transposon repression in the mouse male germline. PLoS Genet. 13, e1006964 (2017).
Lin, I.-H., Chen, Y.-F. & Hsu, M.-T. Correlated 5-hydroxymethylcytosine (5hmC) and gene expression profiles underpin gene and organ-specific epigenetic regulation in adult mouse brain and liver. PLoS One 12, e0170779 (2017).
Brick, K., Pratto, F., Sun, C.-Y., Camerini-Otero, R. D. & Petukhova, G. Analysis of meiotic double-strand break initiation in mammals. Methods Enzymol. 601, 391–318(2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Petukhova, G. V., Romanienko, P. J. & Camerini-Otero, R. D. The Hop2 protein has a direct role in promoting interhomolog interactions during mouse meiosis. Dev. Cell 5, 927–936 (2003).
Liang, K. & Keleş, S. Normalization of ChIP-seq data with control. BMC Bioinformatics 13, 199 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Teng, M. & Irizarry, R. A. Accounting for GC-content bias reduces systematic errors and batch effects in ChIP-seq data. Genome Res. 27, 1930–1938 (2017).
Derrien, T. et al. Fast computation and applications of genome mappability. PLoS One 7, e30377 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Baker, C. L., Walker, M., Kajita, S., Petkov, P. M. & Paigen, K. PRDM9 binding organizes hotspot nucleosomes and limits Holliday junction migration. Genome Res. 24, 724–732 (2014).
Oey, H., Isbel, L., Hickey, P., Ebaid, B. & Whitelaw, E. Genetic and epigenetic variation among inbred mouse littermates: identification of inter-individual differentially methylated regions. Epigenetics Chromatin 8, 54 (2015).
dos Santos, C. O., Dolzhenko, E., Hodges, E., Smith, A. D. & Hannon, G. J. An epigenetic memory of pregnancy in the mouse mammary gland. Cell Rep. 11, 1102–1109 (2015).
Bonn, S. et al. Tissue-specific analysis of chromatin state identifies temporal signatures of enhancer activity during embryonic development. Nat. Genet. 44, 148–156 (2012).
Acknowledgements
We thank P. Hsieh for critical feedback and the NIDDK genomics core and NHLBI flow cytometry core for assistance. This work used the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov). This research was supported by NIGMS grant R01GM084104 (G.V.P.), March of Dimes Foundation grant 1-FY13-506 (G.V.P.) and by the NIDDK Intramural Research Program (R.D.C.-O.).
Reviewer information
Nature thanks S. Keeney, A. Pendas and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
K.B. performed in silico analyses. S.T.-S., F.S., K.-W.G.L., Y.P. and F.P. performed DMC1-SSDS experiments in male mice. F.S. performed DMC1 SSDS in females. F.P. and K.B. performed H3K4me3 ChIP–seq followed by bisulfite sequencing. K.-W.G.L., F.P. and K.B. performed sorting of ovary nuclei. G.L. performed H3K4me3 ChIP–seq in ovary. S.T.-S. performed DMC1 SSDS and H3K4me3 ChIP–seq in Dnmt3l−/− mice. K.B. wrote the manuscript. R.D.C.-O. and G.V.P. supervised the study. All authors contributed to experimental design and critiqued the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sample details and quality metrics for DSB maps.
a, The signal portion of tags is calculated for all samples at hotspots (HS) identified in the T1 sample. Sample identifiers are in panel g. b, c, Hotspots identified in each respective sample (b) or hotspots in the combined T1/O1 superset (c). Peak calling was not performed for N1 (see Methods). d, The estimated library size (x) was inferred using bisection root finding for f(x) = (1 − NNR/x) − exp(Ntot/x), 104 ≤ x ≤ 1012. NNR, number of unique fragments; Ntot = total number of fragments. e, The number of hotspots identified in each sample. f, The number of ssDNA fragments sequenced for each sample. g, Details of SSDS samples. Sample N1 was generated from Hop2−/− mice using 2 × 105 cells. This sample was run in single-end mode, and not processed through the ssDNA pipeline. Previously published samples are referenced by the GEO accession number. Note that the use of the SC 8973 (c20) and anti-DMC1-GP1 antibodies gave indistinguishable SSDS results in males (Fig. 2c, Extended Data Fig. 2a). h, Details of samples from H3K4me3 ChIP–seq. i, Details of publicly available datasets used.
Extended Data Fig. 2 Most DSB hotspots are used in both male and female meiosis.
a, The maximum reciprocal overlap between hotspots in each sample was calculated using the central ± 200 bp of hotspots. b, Hotspots exclusively found in either sex are weak. Hotspots were split into those found in both O1 and T1 (both; grey), O1 but not T1 (female-only; pink) and T1 but not O1 (male-only; blue). c, Female-only hotspots are weak in females, relative to shared hotspots. d, Male-only hotspots are weak in males, relative to shared hotspots. e, The O2 SSDS correlates better with hotspot strength in ovary (O1) than in testis (T1). Only hotspots that are detected in both samples are shown for each comparison. Note that the correlation between hotspot strength in ovary samples (Spearman’s R2O2O1 = 0.76) is not as high as that between replicates of SSDS in males (minimum Spearman’s R2 ≥ 0.9; Fig. 2). f, Noise in SSDS estimates can fully explain this diminished correlation between ovary-derived SSDS maps. We generated a series of downsampled O1 SSDS datasets to test whether reducing the SPoT value would reduce the maximum possible R2 value. For each simulated dataset, signal reads were randomly chosen from the uniquely mapping in-hotspot ssDNA fragments of the O1 DMC1 SSDS sample. Background fragments were randomly chosen from all uniquely mapping ssDNA fragments of the O1 input DNA SSDS sample. Samples at different SPoTs were then generated by varying the number of signal and background-derived reads (SPoT = signal/(signal + background)). The number of fragments was matched to the number of uniquely mapping fragments in O2. Ten replicate samples were generated for each SPoT, and the correlation coefficient (Spearman’s R2) with the original O1 SSDS sample was calculated. The magenta lines indicate the expected maximum R2 for a sample with a SPoT matching that of O2. The expected maximum R2 is very close to the observed R2. Thus, noise in SSDS estimates can reduce the R2 to within the observed range for a sample of this quality.
Extended Data Fig. 3 SSDS signal at hotspots is narrower in ovaries than in spermatocytes.
a, SSDS coverage is a measure of DMC1-bound ssDNA either side of each meiotic DSB. In a population of meiocytes, DSBs will occur in a several hundred nucleotide window around the hotspot centre (orange rectangle). To assess coverage, we first convert the position of each SSDS fragment into the distance along the ssDNA from the hotspot centre. Merging the top and bottom strand fragments in this way increases coverage twofold and minimizes the influence of asymmetric gaps and fluctuations in coverage. Coverage at each hotspot was normalized by the maximum value at the hotspot to prevent strong hotspots from dominating the average profile. The average normalized coverage across all hotspots was then calculated. b, c, DSB hotspots identified in females (ovary sample O1) (b) and males (testis sample T1) (c) were each split into three bins by strength. Coverage was calculated for all nine male and two female samples for each set. The SSDS signal is narrower for all female samples compared to male samples. The difference is particularly pronounced at stronger hotspots, in which coverage estimates are most accurate. At the widest point, the mean male and female profiles diverge by approximately 0.4 kb. d, We also examined the model-based analysis of ChIP–seq (MACS)-determined hotspot boundaries to further negate the possibility that the average profiles in b and c are not a reflection of the population. By this metric, the mean hotspot width estimated from male samples (1,759 ± 73 bp; mean (solid blue line) ± s.e.m. (dashed blue lines); n = 9) is significantly wider than the mean width of hotspots in female samples (1,490 ± 89 bp; mean (solid pink line) ± s.e.m. (dashed pink lines); n = 2) (P = 0.0007; t-test). Because sequencing quality and sample SPoT can affect width estimates, we processed each sample as follows: we reduced the SPoT of each sample to that of the lowest quality sample (O2; see Methods), considering only uniquely mapping and high quality (Q > 30) ssDNA type 1 fragments. We then reduced all samples to have the same number of fragments as the smallest. On these datasets, we performed peak calling and retained only DSB hotspots that were called in all samples (n = 1,975). e, Potential mechanistic explanations for the difference in SSDS signal between males and females. These differences may manifest in all meiocytes or in sub-populations. Notably, we see no evidence of shape differences at hotspots in sub-populations of spermatocytes (data not shown).
Extended Data Fig. 4 Most meiotic DSBs occur at sex-biased hotspots.
a, Quantification of SSDS fold change at sex biased and unbiased hotspots. The percentages show the percentage of hotspots in each category with a given absolute fold change. b, The hotspots in the testis/ovary superset were split into quintiles by strength in either females (left) or males (right). In both sexes, over 60% of the strongest hotspot subset exhibit sex-biased DSB formation. This is a proxy for the true amount of sex-biased DSB formation. In progressively weaker hotspot sets, fewer biased hotspots are detected. One outlier is the set of weak male hotspots. This set contains many female-biased default hotspots that form independently of PRDM9. c, We quantified the total in-hotspot SSDS signal at female-biased, unbiased and male-biased hotspots in the two ovary-derived samples and in the nine testis samples. In all cases, over half of the in-hotspot sequencing tags (referred to as total DSBs) occur at sex-biased hotspots. Hotspots biased towards usage in females are enriched in ovary samples, while those biased towards male usage are enriched in testis-derived samples.
Extended Data Fig. 5 Sex biased hotspots are consistent across replicates and are defined before DSB formation.
Hotspot strength was calculated at all autosomal hotspots from the merged O1/T1 DSB maps. The strength of hotspots was re-calculated in two testis (T1 and T2) and two ovary (O1 and O2) maps. Female-biased (pink), unbiased (grey) and male-biased (blue) hotspots were determined by comparing the T1 and O1 maps. These hotspots are coloured the same in all panels. a, Sex-biased hotspots are distributed as expected when comparing the O1 and T1 DSB maps. These data are also plotted in Fig. 2d, e. b, Sex-biased hotspots exhibit the same sex-biases in the O2 sample. c, d, Sex-biased hotspots exhibit no biased usage between samples derived from mice of the same sex. e–g, Sex biases that precede DSB formation were studied by performing H3K4me3 ChIP–seq in FACS-purified fetal oocytes at 15.5 d.p.c. (see Methods). The H3K4me3 signal at hotspots was quantified and compared to existing maps of H3K4me3 in juvenile mouse testis52. e, The H3K4me3 signal at hotspots is tightly correlated in replicate samples from mouse testis (Te1 = H3KP1, Te2 = H3KP2; Extended Data Fig. 1i). f, Similar to what we observe when examining the SSDS signal at DSB hotspots, there is extensive variation in the H3K4me3 signal at hotspots between male and female meiosis. This indicates that sex biases are established before DSB formation. Sex biases determined using SSDS remain broadly conserved when we compare H3K4me3 in females to males. g, H3K4me3 at hotspots is better correlated with SSDS from the respective sex. h–j, Sex biases in H3K4me3 ChIP–seq parallel the differences in the SSDS signal. H3K4me3 ChIP–seq coverage is shown in the top panels; testis (H3KP1; blue) and ovary (H3OV; pink). The middle panel shows the log2 fold difference (M) between the SSDS signal in testis (T1) and ovary (O1). SSDS coverage for these samples is shown in the bottom panels. Grey boxes represent DSB hotspot positions. To allow for quantitative cross comparison, the coverage in each sample is normalized by the median signal strength at DSB hotspots in that sample. The genomic coordinates of each window are given underneath.
Extended Data Fig. 6 Clustering of sex-biased hotspots.
a–c, SSDS coverage at a subset of biased hotspot clusters. The female-biased (a) and unbiased (b) clusters are those shown in Fig. 3a. c, Because no male-biased clusters are depicted in Fig. 3a, eight clusters were randomly chosen. SSDS coverage for testis (T1; blue) and ovary (O1; pink) are shown for each cluster. To allow for quantitative cross comparison, coverage in each sample is normalized by the median hotspot strength. Grey boxes represent DSB hotspot positions. d, Genomic patterning of sex-biased DSB hotspots. Female-biased (left; pink) and male-biased (right; blue) hotspot clusters on all autosomes. Biased hotspots do not exhibit particular spatial patterning, aside from a slight enrichment of female-biased hotspots at the q-arm telomere. e, The physical size of hotspot clusters scales with the number of hotspots per cluster. It therefore seems unlikely that clustering results from a physical size constraint imposed by sex-specific chromatin structure. Notably, however, the presence of such a size constraint may be masked by the presence of a large number of clusters that occur by chance. Semi-transparent box plots show the expected size distribution for randomly distributed clusters (n = 1,000 bootstraps). Clusters of three male-biased hotspots are marginally smaller than expected. There are no significant differences for clusters of other sizes. f, Similar proportions of PRDM9-defined and default hotspots occur in clusters. Hotspots in clusters of ≥2 consecutive hotspots of the same type were counted.
Extended Data Fig. 7 Differing patterns of DNA methylation at sex-biased hotspots.
a, Mean DNA methylation40 at the putative PrBS (grey bar) of female-biased (pink), unbiased (grey) and male-biased (blue) hotspots. Note that this panel is also shown in Fig. 4a. b, Heat map rows depict methylation at individual hotspots. Note that the density of methylation appears higher at unbiased hotspots because rows are more densely spaced.
Extended Data Fig. 8 DNA methylation at PrBSs is present across tissues and absent in the female germ line.
The pattern of DNA methylation is very similar across cell types and between the sexes. Hotspots are split by the magnitude of sex bias (SSDSO1/SSDST1) into seven sets. Sets are ranked from most female-biased (pink; left) to most male-biased (blue; right) by fold change. Methylation signal is binarized such that methylation >0% is considered methylated. Thus, the proportion of all hotspots with methylated cytosine at each position is shown. Variations in the magnitude of the signal may be expected for technical reasons. Plots are anchored by the C57BL/6 PrBS (grey area). Only hotspots with a single PRDM9-binding site are used (see Methods). a, Plot of ±100 bp to show methylation flanking the PrBS for female-biased hotspots. b, Plot of ±15 bp to show methylation at the PrBS for male-biased hotspots. Methylation data are from whole-genome bisulfite sequencing (WGBS) in tissue derived from whole testis (Te) in 13 d.p.p. mice40, from WGBS after H3K4me3 ChIP–seq in whole adult testis (S/c(Tgt)); Extended Data Fig. 1h), from WGBS in elutriated spermatocytes38 (S/c(Elu)), from WGBS in spermatogonia38 (Gonia), from WGBS in tissue from male liver53 (Liv(M)), from WGBS in tissue from female parous basal differentiated mammary gland cells54 (Mamm(F)) and from WGBS in sorted primordial germ cells (PGCs) at 13.5 and 16.5 d.p.c. (PGC13.5(F) and PGC16.5(F), representing earlier and later meiotic prophase I populations, respectively)30. WGBS in female PGCs captures the methylation status of the genome in oocytes during meiosis30.
Extended Data Fig. 9 Dual role of DNA methylation at hotspots in defining sex biases.
DNA methylation has a dual role in modulating sex-biased DSB formation. Left, at female-biased hotspots, DNA methylation in the region flanking the PrBS can suppress PRDM9 binding. Thus, in males, the use of these PrBS is reduced, resulting in a female-biased hotpsot. Methylated CpG dinucleotides (in males) are schematically shown as filled black circles. Right, at male-biased hotspots, DNA methylation at CpGs appears to favour PRDM9 binding and DSB formation. This results in a relatively strong DSB hotspot in males, but a relatively weak hotspot in females, in which DNA methylation at these sites is absent.
Extended Data Fig. 10 Hotspot strength variation in Dnmt3l−/− mice.
a, b, CpG methylation is partly reduced at PRDM9 binding sites in mice lacking functional DNMT3L. We compared WGBS data from Dnmt3lA/A (Dnmt3lD124A/D124A) (A/A) and matched wild-type mice33. The ±100-bp region around the PRDM9-binding sites was examined. a–f, Hotspots were split either by sex-bias (female-biased, F, pink; unbiased, U, grey; male-biased, M, blue) (a, c, e) or into quintiles by the fold change between the O1 and T1 SSDS samples (most female-biased (F) on left to most male-biased (M) on right) (b, d, f). The percentage decrease in the mean DNA methylation signal in Dnmt3lA/A mice for each set is shown. DNA methylation is reduced 5–7%. c–f, The usage of sex-biased hotspots is altered in mice in which DNA methylation is reduced (Dnmt3l−/−). c, Left, the log2 fold change between the tags per million normalized signal at hotspots in Dnmt3l−/− and wild-type (T1) male mice is shown. Right, to control for spermatocyte population changes resulting from meiotic arrest, we compare to experiments in Hop2−/− males instead of wild-type. HOP2 is essential for stabilizing recombination intermediates and mice lacking functional HOP2 exhibit spermatogenic arrest after DSB formation. Hotspots overlapping gene promoters or default hotspots are excluded as the non-PRDM9 derived H3K4me3 signal would confound these analyses. Furthermore, only hotspots detected in all samples being compared were analysed to remove spurious potential background correlation (c–f; nhotspots = 9,137). P values for all comparisons are shown (Wilcoxon test). c, The SSDS signal at female-biased hotspots is significantly increased in Dnmt3l−/−− mice compared to male-biased hotspots or unbiased hotspots. The strength of male-biased hotspots is relatively decreased. d, This is also seen when we simply split hotspots by fold change. e, H3K4me3 at female-biased hotspots is significantly increased in Dnmt3l−/− mice compared to male-biased hotspots. H3K4me3 signal at each hotspot was calculated as the sum of overlapping H3K4me3 peak strengths. This is a proxy for DSB hotspot strength, because PRDM9 trimethylates histone H4 lysine 3 before DSB formation. f, This is more apparent when we split hotspots into quintiles by sex-bias, probably because H3K4me3 at hotspots is a weak signal.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1: Oocyte nuclei can be isolated by FACS using a combination of DNA content and SCP3 immunofluorescence. We first gated to retain only single nuclei (not shown). We then gated 4C nuclei using DAPI (92,000 < DAPI signal <170,000; vertical lines), and using an oocyte-specific marker, SCP3 (SCP signal >10,000). a, We defined the SCP3 gate using an aliquot of our sample to which the primary antibody was not added (Secondary only). This estimates the background fluorescence from the secondary antibody. b, A distinct population of oocytes is identifiable (SCP3-positive, 4C nuclei; orange). >90% of the post-sort fraction was validated as SCP3 positive using immunofluorescence microscopy
Supplementary data
This file contains a table of DSB hotspots and associated metadata, and a table guide.
Source data
Rights and permissions
About this article
Cite this article
Brick, K., Thibault-Sennett, S., Smagulova, F. et al. Extensive sex differences at the initiation of genetic recombination. Nature 561, 338–342 (2018). https://doi.org/10.1038/s41586-018-0492-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0492-5
Keywords
This article is cited by
-
Mapping crossover events of mouse meiotic recombination by restriction fragment ligation-based Refresh-seq
Cell Discovery (2024)
-
DEtail-seq is an ultra-efficient and convenient method for meiotic DNA break profiling in multiple organisms
Science China Life Sciences (2023)
-
TOPOVIBL-REC114 interaction regulates meiotic DNA double-strand breaks
Nature Communications (2022)
-
3D chromatin remodelling in the germ line modulates genome evolutionary plasticity
Nature Communications (2022)
-
Two telomeric ends of acrocentric chromosome play distinct roles in homologous chromosome synapsis in the fetal mouse oocyte
Chromosoma (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.