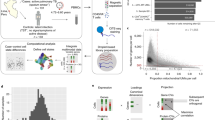
Abstract
Most infections with Mycobacterium tuberculosis (Mtb) manifest as a clinically asymptomatic, contained state, known as latent tuberculosis infection, that affects approximately one-quarter of the global population1. Although fewer than one in ten individuals eventually progress to active disease2, tuberculosis is a leading cause of death from infectious disease worldwide3. Despite intense efforts, immune factors that influence the infection outcomes remain poorly defined. Here we used integrated analyses of multiple cohorts to identify stage-specific host responses to Mtb infection. First, using high-dimensional mass cytometry analyses and functional assays of a cohort of South African adolescents, we show that latent tuberculosis is associated with enhanced cytotoxic responses, which are mostly mediated by CD16 (also known as FcγRIIIa) and natural killer cells, and continuous inflammation coupled with immune deviations in both T and B cell compartments. Next, using cell-type deconvolution of transcriptomic data from several cohorts of different ages, genetic backgrounds, geographical locations and infection stages, we show that although deviations in peripheral B and T cell compartments generally start at latency, they are heterogeneous across cohorts. However, an increase in the abundance of circulating natural killer cells in tuberculosis latency, with a corresponding decrease during active disease and a return to baseline levels upon clinical cure are features that are common to all cohorts. Furthermore, by analysing three longitudinal cohorts, we find that changes in peripheral levels of natural killer cells can inform disease progression and treatment responses, and inversely correlate with the inflammatory state of the lungs of patients with active tuberculosis. Together, our findings offer crucial insights into the underlying pathophysiology of tuberculosis latency, and identify factors that may influence infection outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
30 October 2018
The spelling of author Qianting Yang was corrected; the affiliation of author Stephanus T. Malherbe was corrected; and graphs in Fig. 4b and c were corrected owing to reanalysis of the data into the correct timed intervals.
References
Houben, R. M. & Dodd, P. J. The global burden of latent tuberculosis infection: a re-estimation using mathematical modelling. PLoS Med. 13, e1002152 (2016).
Shea, K. M., Kammerer, J. S., Winston, C. A., Navin, T. R. & Horsburgh, C. R. Jr. Estimated rate of reactivation of latent tuberculosis infection in the United States, overall and by population subgroup. Am. J. Epidemiol. 179, 216–225 (2014).
WHO. Global Tuberculosis Report. http://www.who.int/tb/publications/global_report/en/ (2017).
Spitzer, M. H. & Nolan, G. P. Mass cytometry: single cells, many features. Cell 165, 780–791 (2016).
Mahomed, H. et al. Predictive factors for latent tuberculosis infection among adolescents in a high-burden area in South Africa. Int. J. Tuberc. Lung Dis. 15, 331–336 (2011).
Takenami, I. et al. Blood cells and interferon-gamma levels correlation in latent tuberculosis infection. ISRN Pulmonol. 2013, 256148 (2013).
Joosten, S. A. et al. Patients with tuberculosis have a dysfunctional circulating B-cell compartment, which normalizes following successful treatment. PLoS Pathog. 12, e1005687 (2016).
Lu, L. L., et al. A functional role for antibodies in tuberculosis. Cell 167, 433–443 (2016).
Ruvinsky, I. & Meyuhas, O. Ribosomal protein S6 phosphorylation: from protein synthesis to cell size. Trends Biochem. Sci. 31, 342–348 (2006).
Ueda, Y., Kondo, M. & Kelsoe, G. Inflammation and the reciprocal production of granulocytes and lymphocytes in bone marrow. J. Exp. Med. 201, 1771–1780 (2005).
Shih, C. H., van Eeden, S. F., Goto, Y. & Hogg, J. C. CCL23/myeloid progenitor inhibitory factor-1 inhibits production and release of polymorphonuclear leukocytes and monocytes from the bone marrow. Exp. Hematol. 33, 1101–1108 (2005).
Schölkopf, B., Smola, A. J., Williamson, R. C. & Bartlett, P. L. New support vector algorithms. Neural Comput. 12, 1207–1245 (2000).
Vallania, F. et al. Leveraging heterogeneity across multiple data sets increases accuracy of cell-mixture deconvolution and reduces biological and technical biases. Preprint at https://biorxiv.org/content/early/2017/10/20/206466 (2017).
Malherbe, S. T. et al. Persisting positron emission tomography lesion activity and Mycobacterium tuberculosis mRNA after tuberculosis cure. Nat. Med. 22, 1094–1100 (2016).
Zak, D. E. et al. A blood RNA signature for tuberculosis disease risk: a prospective cohort study. Lancet 387, 2312–2322 (2016).
Scriba, T. J. et al. Sequential inflammatory processes define human progression from M. tuberculosis infection to tuberculosis disease. PLoS Pathog. 13, e1006687 (2017).
Thompson, E. G. et al. Host blood RNA signatures predict the outcome of tuberculosis treatment. Tuberculosis 107, 48–58 (2017).
Esin, S. & Batoni, G. Natural killer cells: a coherent model for their functional role in Mycobacterium tuberculosis infection. J. Innate Immun. 7, 11–24 (2015).
Junqueira-Kipnis, A. P. et al. NK cells respond to pulmonary infection with Mycobacterium tuberculosis, but play a minimal role in protection. J. Immunol. 171, 6039–6045 (2003).
Feng, C. G. et al. NK cell-derived IFN-γ differentially regulates innate resistance and neutrophil response in T cell-deficient hosts infected with Mycobacterium tuberculosis. J. Immunol. 177, 7086–7093 (2006).
Kwan, C. K. & Ernst, J. D. HIV and tuberculosis: a deadly human syndemic. Clin. Microbiol. Rev. 24, 351–376 (2011).
Scully, E. & Alter, G. NK cells in HIV disease. Curr. HIV/AIDS Rep. 13, 85–94 (2016).
Mansour, I., Doinel, C. & Rouger, P. CD16+ NK cells decrease in all stages of HIV infection through a selective depletion of the CD16+CD8+CD3− subset. AIDS Res. Hum. Retroviruses 6, 1451–1457 (1990).
Xie, X., Li, F., Chen, J. W. & Wang, J. Risk of tuberculosis infection in anti-TNF-α biological therapy: from bench to bedside. J. Microbiol. Immunol. Infect. 47, 268–274 (2014).
Nocturne, G. et al. Impact of anti-TNF therapy on NK cells function and on immunosurveillance against B-cell lymphomas. J. Autoimmun. 80, 56–64 (2017).
Bruns, H. et al. Anti-TNF immunotherapy reduces CD8+ T cell-mediated antimicrobial activity against Mycobacterium tuberculosis in humans. J. Clin. Invest. 119, 1167–1177 (2009).
Berry, M. P. et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature 466, 973–977 (2010).
Maertzdorf, J. et al. Human gene expression profiles of susceptibility and resistance in tuberculosis. Genes Immun. 12, 15–22 (2011).
Maertzdorf, J. et al. Functional correlations of pathogenesis-driven gene expression signatures in tuberculosis. PLoS ONE 6, e26938 (2011).
Kaforou, M. et al. Detection of tuberculosis in HIV-infected and -uninfected African adults using whole blood RNA expression signatures: a case–control study. PLoS Med. 10, e1001538 (2013).
Anderson, S. T. et al. Diagnosis of childhood tuberculosis and host RNA expression in Africa. N. Engl. J. Med. 370, 1712–1723 (2014).
Bloom, C. I. et al. Detectable changes in the blood transcriptome are present after two weeks of antituberculosis therapy. PLoS ONE 7, e46191 (2012).
Verhagen, L. M. et al. A predictive signature gene set for discriminating active from latent tuberculosis in Warao Amerindian children. BMC Genomics 14, 74 (2013).
Cai, Y. et al. Increased complement C1q level marks active disease in human tuberculosis. PLoS ONE 9, e92340 (2014).
Ottenhoff, T. H. et al. Genome-wide expression profiling identifies type 1 interferon response pathways in active tuberculosis. PLoS ONE 7, e45839 (2012).
Tientcheu, L. D. et al. Differential transcriptomic and metabolic profiles of M. africanum- and M. tuberculosis-infected patients after, but not before, drug treatment. Genes Immun. 16, 347–355 (2015).
Lee, S. W. et al. Gene expression profiling identifies candidate biomarkers for active and latent tuberculosis. BMC Bioinformatics 17, S3 (2016).
Leipold, M. D. & Maecker, H. T. Phenotyping of live human PBMC using CyTOF™ mass cytometry. Bio Protoc. 5, e1382 (2015).
Fernandez, R. & Maecker, H. Cytokine-stimulated phosphoflow of PBMC using CyTOF mass cytometry. Bio Protoc. 5, e1496 (2015).
Bruggner, R. V., Bodenmiller, B., Dill, D. L., Tibshirani, R. J. & Nolan, G. P. Automated identification of stratifying signatures in cellular subpopulations. Proc. Natl Acad. Sci. USA 111, E2770–E2777 (2014).
Amir, E. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013).
Somanchi, S. S., McCulley, K. J., Somanchi, A., Chan, L. L. & Lee, D. A. A novel method for assessment of natural killer cell cytotoxicity using image cytometry. PLoS ONE 10, e0141074 (2015).
Salinas-Jazmín, N., Hisaki-Itaya, E. & Velasco-Velázquez, M. A. A flow cytometry-based assay for the evaluation of antibody-dependent cell-mediated cytotoxicity (ADCC) in cancer cells. Methods Mol. Biol. 1165, 241–252 (2014).
Haynes, W. A. et al. Empowering multi-cohort gene expression analysis to increase reproducibility. Pac. Symp. Biocomput. 22, 144–153 (2017).
Sweeney, T. E., Braviak, L., Tato, C. M. & Khatri, P. Genome-wide expression for diagnosis of pulmonary tuberculosis: a multicohort analysis. Lancet Respir. Med. 4, 213–224 (2016).
Khatri, P. et al. A common rejection module (CRM) for acute rejection across multiple organs identifies novel therapeutics for organ transplantation. J. Exp. Med. 210, 2205–2221 (2013).
Acknowledgements
We thank R.-P. Sekaly, A. Filali-Mouhim and K. Ghneim for transcriptional analysis of the Mtb acquisition subcohort of the Adolescent Cohort Study; E. Long, C. Blish for advice and the P815 mouse cell line and K562 human cell line; A. Kasmar for critically reading the manuscript. This work was supported by the Bill and Melinda Gates Foundation (T.J.S., P.K., Y.-h.C.), the National Institutes of Health AI127128 (Y.-h.C.), AI109662, AI057229 and AI125197 (P.K.), K12 5K12HL120001 (F.V.), 5T32AI07290-31 (R.R.C.), VirBio (P.K.), the Natural Science Foundation of China (81525016, 81772145) and the Science and Technology Project of Shenzhen (JSGG20160427104724699) (X.C.).
Reviewer information
Nature thanks T. H. M. Ottenhoff and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.R.C. and Y.-h.C. conceptualized the study; F.V. and P.K. established immunoStates and performed cell-type deconvolution of transcriptome data. R.R.C. performed experiments and analysis of South African cohorts and overall data analysis. F.D., A.P.-N., V.R., S.T.M. and T.J.S. provided samples and analysed South African cohorts of individuals who progressed from LTBI to active disease. Q.Y. and X.C. performed FACS analysis of the Chinese cohort. T.J.S. and E.N. designed and oversaw the Mtb acquisition subcohort of the Adolescent Cohort Study. S.T.M., K.R., G.W. and J.W. provided lung pathology results from the Catalysis cohort. T.J.S. and W.H. established the South African Tuberculosis cohorts. M.M.D., C.J.L.A. and T.J.S. contributed to project design and interpretation. R.R.C. and Y.-h.C. wrote the manuscript with input from T.J.S., P.K., J.W., F.V., M.M.D. and A. P-N. P.K. designed and oversaw all computational analyses. Y.-h.C. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Broad alterations of peripheral immune cell distributions in LTBI.
PBMCs from 14 latently infected and 14 uninfected participants of a South African adolescent cohort were characterized using CyTOF with antibody panel 1 (Supplementary Table 2), followed by Citrus analysis and clustering. This unsupervised hierarchical clustering analysis produced a branching structure (dendrogram) that allowed the grouping of total live cells into known immune cell compartments (contoured). Cell clusters are represented as nodes (circles) in this Citrus-derived circular dendrogram, which delineates lineage relationships that were identified from the data. Cluster granularity (that is, cell subset specificity) increases from the centre of the diagram to the periphery. a, Annotation of cluster hierarchy plots based on surface marker expression. The expression intensity of each marker used for cell population characterization is overlaid per cluster on the Citrus circular dendrogram and is indicated, independently for each marker, by the coloured gradient for which the range corresponds to the arcsinh-transformed expression of the median marker expression measured across all Citrus clusters. For each marker, we also provide a dot plot graph demonstrating the marker labelling in the manually gated indicated population. b, Citrus plots showing, based on cell-surface protein expression, clusters (in red, designated A–F) that exhibit significantly different abundances (SAM analysis with FDR < 1%) between the uninfected and latently infected individuals. Individual cell clusters are mapped to well-established, gross-cell types: B cells (CD19+), CD8+ αβ T cells (CD3+TCRβ+CD8+), CD4+ αβ T cells (CD3+TCRβ+CD4+), γδ T cells (CD3+TCRδ+), monocytes (CD3−CD19−CD33+CD14+HLA-DR+), NK cells (CD3−CD19−CD14−HLA-DR−CD16+CD56bright/dim), identifiable by annotated shaded background groupings. c, The phenotype and the composition of cells in each of the stratifying cell subsets (A–F), identified by Citrus analysis. d, Percentages of NK cells and B cells determined by manual gating of 20 additional samples using CyTOF antibody panel 2 (left; Supplementary Table 2) and 32 samples using flow cytometry (right). e, Percentages of CD4+ αβ T cells, CD8+ αβ T cells and γδ T cells in uninfected controls and latently infected individuals, analysed by CyTOF (n = 24 per group; top) and flow cytometry (n = 16 per group; bottom). Throughout, P values were derived using a Mann–Whitney U-test. Mean and error bars representing the 95% confidence intervals are shown for each comparison.
Extended Data Fig. 2 Enhanced effector function response in LTBI.
a, Cell subsets, shown as red nodes in a Citrus-derived circular dendrogram and designated as 1–5, were identified as significantly different in abundance (SAM analysis at FDR < 1%) based on CyTOF analysis of effector and cell-surface molecule expression on PBMCs (antibody panel 1, Supplementary Table 2) from uninfected controls and individuals with LTBI (n = 14 per group) after 4-h PMA and ionomycin stimulation. Mapping of individual cell clusters to established, gross-cell types are identified by annotated shaded background groupings. b, Expression intensity of selected effector molecules is indicated by the coloured gradient for which the range corresponds to the arcsinh-transformed expression of the median marker expression measured across all Citrus clusters. c, Effector molecule expression and the composition of cells in each of the stratifying cell clusters (1–5), identified by Citrus analysis. d, viSNE analysis of GZMB expression level in immune-cell subsets, representative of 14 uninfected and 14 individuals with LTBI (the colour gradient corresponds to the arcsinh-transformed expression level). e, Quantification of intracellular GZMB expression level in NK cells, CD8+ αβ T cells and γδ T cells in uninfected controls and individuals with LTBI (n = 14 per group). P values were derived using a Mann–Whitney U-test. Mean and error bars representing the 95% confidence intervals are shown for each comparison. f, Dot plots from CyTOF analysis of CD16+GZMBhigh cells within each lymphocyte subset, representative of 14 uninfected controls and 14 individuals with LTBI. g, Gating strategy for ADCC. ADCC was measured using NK-resistant P815 cells, which were either coated with antibody (2.4G2) or left uncoated (control), and labelled with the intracellular dye CFSE, followed by the DNA dye 7AAD. CFSE+7AAD+ cells were defined as dead target cells.
Extended Data Fig. 3 Alterations in plasma protein levels in LTBI.
The relative levels of plasma proteins (Supplementary Table 3), shown on a log2 scale, between uninfected controls and individuals with LTBI (n = 27 per group). Plasma proteins that were present at significantly higher levels (a) and significantly lower levels (b) in individuals with LTBI. Plasma protein quantification was performed using the proximity extension assay. P values were derived using a unpaired two-tailed Student’s t-test. Mean and error bars representing the 95% confidence intervals are shown for each comparison.
Extended Data Fig. 4 Changes in frequencies of peripheral B cell subsets in LTBI, active TB and after treatment.
Forest plots for estimated frequencies of B cell subsets: naive B cells, memory B cells and plasma cells. a, Comparison between the uninfected state (n = 189) and LTBI (n = 145). b, Comparison between LTBI (n = 409) and active TB (n = 543). c, Comparison between active TB (n = 76) and end-of-treatment (n = 97). Cohort GSE identifiers are listed on the left. In the plots, boxes represent the standardized mean difference in estimated cellular proportions in a cohort between two comparison groups. The size of the box is proportional to the sample size of a given cohort. Lines indicate the 95% confidence interval of the corresponding effect sizes. Diamonds indicate the summary effect size (Summary), obtained by integrating the effect sizes from individual cohorts. The width of the diamond corresponds to its 95% confidence interval. The P values and q values for the summary effect sizes are shown above each plot.
Extended Data Fig. 5 Changes in frequencies of peripheral T cell subsets, monocytes and granulocytes in LTBI, active TB and after treatment.
Forest plots for estimated frequencies of CD4+ αβ T cells, CD8+ αβ T cells, monocytes and granulocytes. a, Comparison between the uninfected state (n = 189) and LTBI (n = 145). b, Comparison between LTBI (n = 409) and active TB (n = 543). c, Comparison between active TB (n = 76) and end-of-treatment (n = 97). Boxes represent the standardized mean difference in estimated cellular proportions in a cohort between two comparison groups. The size of the box is proportional to the sample size of a given cohort. Lines indicate the 95% confidence interval of the corresponding effect sizes. Diamonds indicate the summary effect size (Summary), obtained by integrating the effect sizes from individual cohorts. The width of the diamond corresponds to its 95% confidence interval. The P values and q values for the summary effect sizes are shown above each plot.
Extended Data Fig. 6 Comparison of the frequencies of peripheral NK cells, B cells and T cells between uninfected controls and patients with active TB.
Forest plots comparing changes in the levels of NK cells, B cells and T cells between uninfected individuals (n = 191) and patients with active TB (n = 178). Boxes represent the standardized mean difference in estimated cellular proportions in a cohort between two comparison groups. The size of the box is proportional to the sample size of a given cohort. Lines indicate the 95% confidence interval of the corresponding effect sizes. Diamonds indicate the summary effect size (Summary), obtained by integrating the effect sizes from individual cohorts. The width of the diamond corresponds to its 95% confidence interval. The P values and q values for the summary effect sizes are shown above each plot.
Extended Data Fig. 7 Trajectories of different immune cell populations from the acquisition of Mtb infection to end-of-treatment.
Changes in the frequency distribution patterns of different peripheral leukocyte populations (a) and B and T cell subpopulations (b) at the different stages of infection. Lines indicate cumulative effect size scores starting from a healthy baseline level up to treatment of active TB disease. Error bars indicate corresponding standard errors.
Extended Data Fig. 8 Correlation between peripheral NK cell percentage and lung inflammation.
Correlation plot showing the relationship between estimated peripheral NK cell frequencies in patients with active TB at week 4 after treatment initiation and total glycolytic activity index (TGAI) of the lung measured by PET–CT imaging at the corresponding time point. The line represents the best fit and the shaded area the 95% confidence interval. NK cell frequencies were determined by deconvolution.
Extended Data Fig. 9 Synchronization of the adolescent cohort who underwent QuantiFERON conversion following Mtb acquisition
. To identify changes in peripheral NK cell frequencies after acquisition of Mtb infection by cell-mixture deconvolution analysis, the timescale of the gene expression dataset (GSE116014) was realigned according to the time of first infection diagnosis instead of study enrolment, allowing the identification of gene-expression profiles obtained before infection diagnosis. Each individual is represented by a horizontal bar. The length of the bar represents the number of days between study enrolment and diagnosis with Mtb infection. During follow-up, each individual transitioned from an uninfected state (blue) to infected state (brown), that is, underwent QFT conversion. The black circles represent time points for which gene-expression data were available. Pre-infection (Pre) data (180–360 days) were compared to data obtained at the time of infection diagnosis or the nearest time point after diagnosis (Post) (0–360 days).
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-6 and a Supplementary Table Guide.
Rights and permissions
About this article
Cite this article
Roy Chowdhury, R., Vallania, F., Yang, Q. et al. A multi-cohort study of the immune factors associated with M. tuberculosis infection outcomes. Nature 560, 644–648 (2018). https://doi.org/10.1038/s41586-018-0439-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0439-x
Keywords
This article is cited by
-
NK cell phenotypic profile during active TB in people living with HIV-evolution during TB treatment and implications for bacterial clearance and disease severity
Scientific Reports (2023)
-
Immune evasion and provocation by Mycobacterium tuberculosis
Nature Reviews Microbiology (2022)
-
The immunoregulatory landscape of human tuberculosis granulomas
Nature Immunology (2022)
-
Aberrant cytokine milieu and signaling affect immune cell phenotypes and functions in tuberculosis pathology: What can we learn from this phenomenon for application to inflammatory syndromes?
Cellular & Molecular Immunology (2021)
-
Mycobacterium tuberculosis-specific CD4 T cells expressing CD153 inversely associate with bacterial load and disease severity in human tuberculosis
Mucosal Immunology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.