Abstract
A short, 14-amino-acid segment called SP1, located in the Gag structural protein1, has a critical role during the formation of the HIV-1 virus particle. During virus assembly, the SP1 peptide and seven preceding residues fold into a six-helix bundle, which holds together the Gag hexamer and facilitates the formation of a curved immature hexagonal lattice underneath the viral membrane2,3. Upon completion of assembly and budding, proteolytic cleavage of Gag leads to virus maturation, in which the immature lattice is broken down; the liberated CA domain of Gag then re-assembles into the mature conical capsid that encloses the viral genome and associated enzymes. Folding and proteolysis of the six-helix bundle are crucial rate-limiting steps of both Gag assembly and disassembly, and the six-helix bundle is an established target of HIV-1 inhibitors4,5. Here, using a combination of structural and functional analyses, we show that inositol hexakisphosphate (InsP6, also known as IP6) facilitates the formation of the six-helix bundle and assembly of the immature HIV-1 Gag lattice. IP6 makes ionic contacts with two rings of lysine residues at the centre of the Gag hexamer. Proteolytic cleavage then unmasks an alternative binding site, where IP6 interaction promotes the assembly of the mature capsid lattice. These studies identify IP6 as a naturally occurring small molecule that promotes both assembly and maturation of HIV-1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
29 August 2018
In this Letter, the Protein Data Bank (PDB) accessions were incorrectly listed as ‘6BH5, 6BHT and 6BHS’ instead of ‘6BHR, 6BHT and 6BHS’; this has been corrected online.
References
Gross, I. et al. A conformational switch controlling HIV-1 morphogenesis. EMBO J. 19, 103–113 (2000).
Schur, F. K. et al. An atomic model of HIV-1 capsid-SP1 reveals structures regulating assembly and maturation. Science 353, 506–508 (2016).
Wagner, J. M. et al. Crystal structure of an HIV assembly and maturation switch. eLife 5, e17063 (2016).
Keller, P. W., Adamson, C. S., Heymann, J. B., Freed, E. O. & Steven, A. C. HIV-1 maturation inhibitor bevirimat stabilizes the immature Gag lattice. J. Virol. 85, 1420–1428 (2011).
Wang, M. et al. Quenching protein dynamics interferes with HIV capsid maturation. Nat. Commun. 8, 1779 (2017).
Letcher, A. J., Schell, M. J. & Irvine, R. F. Do mammals make all their own inositol hexakisphosphate? Biochem. J. 416, 263–270 (2008).
Campbell, S. et al. Modulation of HIV-like particle assembly in vitro by inositol phosphates. Proc. Natl Acad. Sci. USA 98, 10875–10879 (2001).
Datta, S. A. K. et al. Interactions between HIV-1 Gag molecules in solution: an inositol phosphate-mediated switch. J. Mol. Biol. 365, 799–811 (2007).
Munro, J. B. et al. A conformational transition observed in single HIV-1 Gag molecules during in vitro assembly of virus-like particles. J. Virol. 88, 3577–3585 (2014).
von Schwedler, U. K. et al. Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J. 17, 1555–1568 (1998).
Accola, M. A., Strack, B. & Göttlinger, H. G. Efficient particle production by minimal Gag constructs which retain the carboxy-terminal domain of human immunodeficiency virus type 1 capsid-p2 and a late assembly domain. J. Virol. 74, 5395–5402 (2000).
Loewus, F. A. & Murthy, P. P. N. Myo-inositol metabolism in plants. Plant Sci. 150, 1–19 (2000).
van Galen, J. et al. Interaction of GAPR-1 with lipid bilayers is regulated by alternative homodimerization. Biochim. Biophys. Acta 1818, 2175–2183 (2012).
Ouyang, Z., Zheng, G., Tomchick, D. R., Luo, X. & Yu, H. Structural basis and IP6 requirement for Pds5-dependent cohesin dynamics. Mol. Cell 62, 248–259 (2016).
Macbeth, M. R. et al. Inositol hexakisphosphate is bound in the ADAR2 core and required for RNA editing. Science 309, 1534–1539 (2005).
Chang, Y. F., Wang, S. M., Huang, K. J. & Wang, C. T. Mutations in capsid major homology region affect assembly and membrane affinity of HIV-1 Gag. J. Mol. Biol. 370, 585–597 (2007).
von Schwedler, U. K., Stray, K. M., Garrus, J. E. & Sundquist, W. I. Functional surfaces of the human immunodeficiency virus type 1 capsid protein. J. Virol. 77, 5439–5450 (2003).
Melamed, D. et al. The conserved carboxy terminus of the capsid domain of human immunodeficiency virus type 1 gag protein is important for virion assembly and release. J. Virol. 78, 9675–9688 (2004).
Rihn, S. J. et al. Extreme genetic fragility of the HIV-1 capsid. PLoS Pathog. 9, e1003461 (2013).
Ganser-Pornillos, B. K., von Schwedler, U. K., Stray, K. M., Aiken, C. & Sundquist, W. I. Assembly properties of the human immunodeficiency virus type 1 CA protein. J. Virol. 78, 2545–2552 (2004).
Ganser-Pornillos, B. K., Cheng, A. & Yeager, M. Structure of full-length HIV-1 CA: a model for the mature capsid lattice. Cell 131, 70–79 (2007).
Pornillos, O., Ganser-Pornillos, B. K. & Yeager, M. Atomic-level modelling of the HIV capsid. Nature 469, 424–427 (2011).
Jacques, D. A. et al. HIV-1 uses dynamic capsid pores to import nucleotides and fuel encapsidated DNA synthesis. Nature 536, 349–353 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2013).
Malakhov, M. P. et al. SUMO fusions and SUMO-specific protease for efficient expression and purification of proteins. J. Struct. Funct. Genomics 5, 75–86 (2004).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Chang, L. J., Urlacher, V., Iwakuma, T., Cui, Y. & Zucali, J. Efficacy and safety analyses of a recombinant human immunodeficiency virus type 1 derived vector system. Gene Ther. 6, 715–728 (1999).
Gipson, B., Zeng, X., Zhang, Z. Y. & Stahlberg, H. 2dx-user-friendly image processing for 2D crystals. J. Struct. Biol. 157, 64–72 (2007).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Pornillos, O. et al. X-ray structures of the hexameric building block of the HIV capsid. Cell 137, 1282–1292 (2009).
Pornillos, O., Ganser-Pornillos, B. K., Banumathi, S., Hua, Y. & Yeager, M. Disulfide bond stabilization of the hexameric capsomer of human immunodeficiency virus. J. Mol. Biol. 401, 985–995 (2010).
Jorgensen, W. L. & Jenson, C. Temperature dependence of TIP3P, SPC, and TIP4P water from NPT Monte Carlo simulations: seeking temperatures of maximum density. J. Comput. Chem. 19, 1179–1186 (1998).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38, 27–28 (1996).
Fletcher, R. & Reeves, C. M. Function minimization by conjugate gradients. Comput. J. 7, 149–154 (1964).
Sun, W. & Yuan, Y.-X. Optimization Theory and Methods: Nonlinear Programming (Springer US, 2006)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Shaw, D. E. et al. In Proc. Intl Conf. High Performance Computing, Networking, Storage and Analysis 41–53 (IEEE Press, New Orleans, 2014).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ(1) and χ(2) dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Vanommeslaeghe, K. et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Lippert, R. A. et al. Accurate and efficient integration for molecular dynamics simulations at constant temperature and pressure. J. Chem. Phys. 139, 164106 (2013).
Shan, Y., Klepeis, J. L., Eastwood, M. P., Dror, R. O. & Shaw, D. E. Gaussian split Ewald: A fast Ewald mesh method for molecular simulation. J. Chem. Phys. 122, 54101 (2005).
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Feller, S. E., Zhang, Y. H., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation—the Langevin Piston Method. J. Chem. Phys. 103, 4613–4621 (1995).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Acknowledgements
We thank J. Briggs for discussions and reading of the manuscript. This work was supported by the National Institutes of Health (NIH) grants R01-GM107013 (V.M.V.), R01-GM105684 (G. W. Feigenson), P30-GM110758 and P50-GM082251 (J.R.P.), R01-AI129678 (O.P. and B.K.G.-P.), U54-GM103297 (O.P.), and R01-GM110776 (M.C.J.). F.K.M.S. was supported by Deutsche Forschungsgemeinschaft grant BR 3635/2-1 awarded to J. A. G. Briggs. J.M.W. was supported by NIH postdoctoral fellowship grant F32-GM115007. Anton computer time was provided by the Pittsburgh Supercomputing Center (PSC) through NIH grant R01-GM116961. The Anton machine at PSC was generously made available by D. E. Shaw Research. This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation (NSF) grant number OCI-1053575. Specifically, it used the Bridges system, which is supported at PSC by NSF award number ACI-1445606. Some of The EM work was conducted at the Molecular Electron Microscopy Core facility at the University of Virginia.
Reviewer information
Nature thanks E. Freed and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.A.D. performed protein purification and in vitro assembly. F.K.M.S. did comparative analyses of cryo-EM and crystal structure data. K.K.Z, J.M.W., B.K.G.-P. and O.P. carried out crystallization trials and structure determination. B.K.G.-P. performed 2D cryo-EM. J.R.P. and C.X. performed all-atom MD simulations. T.D.L., C.L.R. and M.C.J., performed cell biology and virology. The manuscript was written primarily by R.A.D., J.R.P., B.K.G.-P., O.P. and V.M.V. The project was originally conceived by R.A.D., with input from all authors throughout experimentation and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Effect of acidic molecules on immature s-CANC assembly.
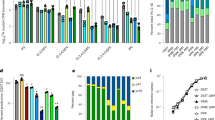
a, Representative negative-stain electron microscopy images. Scale bars, 200 nm. The experiment was repeated twice with similar results. b, Number of immature VLPs per 55 µm2. n = 5, mean shown above box plots; centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values.
Extended Data Fig. 2 s-CANC and s-CASP1 VLPs.
a–c, Representative negative-stain electron microscopy images of s-CANC (a), s-CASP1 (b) and s-CA (c) proteins assembled in the absence of GT25 and in the presence of the indicated IP6 concentrations. Scale bars, 200 nm. d, Diameters of immature VLPs; mean diameter above plot; n below plot. Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values.
Extended Data Fig. 3 Comparison of the HIV-1 Gag cryo-EM structure with the CACTDSP1–IP6 crystal structure.
a, The crystal structure of CACTDSP1 bound to IP6 (cyan) was superimposed on a previously described model of the CA-SP1 segment build into cryo-EM densities of immature HIV-1 particles (PDB 5L93, orange2). Note the close correspondence in K359 rotamers, which were modelled independently in the two structures. For visualization purposes, only one of the six possible IP6 conformations is displayed. b, RMSD calculations of the crystal structure and PDB 5L93. For full-length (residues 149–237) and CA-SP1 (residues 223–237), the RMSDs were calculated only for the atoms that were modelled in both maps. If a sidechain was not modelled, the entire residue was omitted from the calculation. The overall agreement of the models is very high, indicating that the crystal structure corresponds well with conformations found in the virus. c, The CACTDSP1 bound to IP6 (orange and red, respectively) was fitted into two previously published cryo-EM densities2 from VLPs collected from cells (EMD-2706 and EMD-4017). Both maps are shown at 8.8 Å, which is the resolution of the lower resolved map, EMD-2706. In the zoomed insets, only the density corresponding to IP6 is shown. Matching of models and maps and RMSD calculations were performed in Chimera.
Extended Data Fig. 4 Interpretation of the IP6 density in the immature CACTDSP1 hexamer structure.
a, Top and side views of the unbiased mFo–DFc difference density (blue mesh, 2σ) ascribed to the bound IP6. Shown are six IP6 molecules docked in six rotationally equivalent positions, consistent with the six-fold rotational symmetric density. b, Top view of the docked IP6 molecules within the CACTDSP1 hexamer. Unbiased mFo–DFc difference densities (blue mesh) are also shown for both the bound IP6 and sidechains of Lys290 (green) and Lys359 (cyan). Density for Lys359 is more pronounced, which we interpret to mean that this residue adopts a more restricted range of rotamers for binding IP6.
Extended Data Fig. 5 Quantification of wild-type and mutant HIV s-CANC assembly at pH 6 and pH 8.
a, c, Number of immature (purple) and mature (orange) VLPs per 55 μm2 without (−) and with (+) 10 μM IP6 at pH 6 and pH 8. Mean above and n below box plots. Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values. b, d, Representative negative stain electron microscopy images of wild-type and mutant s-CANC assembly in the absence (−) and presence (+) of 10 μM IP6 at pH 6 and pH 8. Scale bar, 400 nm. Repeated three times with similar results. e, Infectivity relative to wild-type virus of IP6 binding residues mutated to alanine and CA residue numbering in parenthesis. Error bars represent s.d., individual data points represented as dots; from four independent experiments.
Extended Data Fig. 6 IP6 modulates the stability of the 6HB.
a, Structural changes observed after 2 μs of molecular dynamics simulations of CACTDSP1 with and without bound IP6. b, RMSDs of the ligand-bound and unbound forms of the CACTDSP1 hexamer. c, RMSFs of the central hexamer during the simulation. The RMSF was averaged over the six central monomers; dashed line shows the s.d. for each residue.
Extended Data Fig. 7 Quantification of mature HIV-1 CA assembly and VLP diameter at pH 6.
a, Example of CA assembly in the absence of IP6 or mellitic acid. b, c, Representative negative-stain electron microscopy images of assemblies induced by IP6 (b) and mellitic acid (c). Scale bars, 200 nm. Tubes (T), cones (C), and other (O) morphologies are marked by coloured arrowheads. a–c, Repeated four times with similar results. d, Number of assembled CA tubes (blue), cones (orange) and other (green) per 55 μm2 at increasing IP6 concentrations. Mean shown above plots, n = 5. e, Number of assembled tubes (blue), cones (orange) and other (green) per 55 μm2 at increasing mellitic acid concentrations. Mean shown above and n below box plots. f, Representative images of mature VLPs assembled with IP5 and IP6 at 50 mM NaCl. Scale bars, 100 nm. Repeated three times with similar results. g, Number of CA VLPs per 10 µm2 without and with IP3, IP4, IP5, and IP6. Mean shown above, n = 5. d, e, g, Centre lines show medians; box limits indicate 25th and 75th percentiles as determined by R software; whiskers extend to minimum and maximum values.
Extended Data Fig. 8 Crystal structure of IP6 bound to the mature CA hexamer.
a, b, Top view (a) and side view (b) of a second CA hexamer crystal structure (P212121 space group) showing the protein in yellow ribbons and unbiased mFo–DFc difference density in blue mesh, contoured at 2.5σ. c, Close-up view showing IP6 densities both above and below the ring of Arg18 residues (magenta).
Supplementary information
Supplementary Figure 1
FACs gating strategy. a, Events were plotted along forward scatter (FSC) and side scatter (SSC) axises using FlowJo. Events with the right morphology were gated as "Live" cells and the position of the gate was copied onto all samples. b, Events from the "Live" gate were isolated and plotted along GFP and RFP intensity axes. The "Non-Fluorescent" gate was created based on a HEK293FT fluorescence negative sample (plot not shown) and the gate was copied onto all isolated "Live" samples. The "GFP-Positive" gate was created based on a HEK293FT GFP positive sample (plot not shown) and the gate was copied onto all isolated "Live" samples. Representative comparison of two cell types transduced with HIV Env deficient virus with a GFP reporter (HEK293FT = WT and IPPK KO = HEK293FT with inositol-pentakisphosphate 2-Kinase knocked out).
Video 1: 2-μs trajectories of IP6-unbound (left) and bound (right) CACTDSP1 models.
The protein hexamer is shown in cartoon representation and IP6 molecule is in stick representation.
Rights and permissions
About this article
Cite this article
Dick, R.A., Zadrozny, K.K., Xu, C. et al. Inositol phosphates are assembly co-factors for HIV-1. Nature 560, 509–512 (2018). https://doi.org/10.1038/s41586-018-0396-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0396-4
This article is cited by
-
HIV-1 capsids enter the FG phase of nuclear pores like a transport receptor
Nature (2024)
-
HIV-1 is dependent on its immature lattice to recruit IP6 for mature capsid assembly
Nature Structural & Molecular Biology (2023)
-
Enrich and switch: IP6 and maturation of HIV-1 capsid
Nature Structural & Molecular Biology (2023)
-
Molecular architecture and conservation of an immature human endogenous retrovirus
Nature Communications (2023)
-
Structural basis of HIV-1 maturation inhibitor binding and activity
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.