Abstract
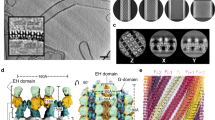
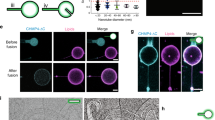
Membrane fission is a fundamental process in the regulation and remodelling of cell membranes. Dynamin, a large GTPase, mediates membrane fission by assembling around, constricting and cleaving the necks of budding vesicles1. Here we report a 3.75 Å resolution cryo-electron microscopy structure of the membrane-associated helical polymer of human dynamin-1 in the GMPPCP-bound state. The structure defines the helical symmetry of the dynamin polymer and the positions of its oligomeric interfaces, which were validated by cell-based endocytosis assays. Compared to the lipid-free tetramer form2, membrane-associated dynamin binds to the lipid bilayer with its pleckstrin homology domain (PHD) and self-assembles across the helical rungs via its guanine nucleotide-binding (GTPase) domain3. Notably, interaction with the membrane and helical assembly are accommodated by a severely bent bundle signalling element (BSE), which connects the GTPase domain to the rest of the protein. The BSE conformation is asymmetric across the inter-rung GTPase interface, and is unique compared to all known nucleotide-bound states of dynamin. The structure suggests that the BSE bends as a result of forces generated from the GTPase dimer interaction that are transferred across the stalk to the PHD and lipid membrane. Mutations that disrupted the BSE kink impaired endocytosis. We also report a 10.1 Å resolution cryo-electron microscopy map of a super-constricted dynamin polymer showing localized conformational changes at the BSE and GTPase domains, induced by GTP hydrolysis, that drive membrane constriction. Together, our results provide a structural basis for the mechanism of action of dynamin on the lipid membrane.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
30 October 2018
In Figs. 2b and 3d of this Letter, the labels ‘Dynamin 1’ and ‘Overlay’ were inadvertently swapped. This has been corrected online.
References
Ferguson, S. M. & De Camilli, P. Dynamin, a membrane-remodelling GTPase. Nat. Rev. Mol. Cell Biol. 13, 75–88 (2012).
Reubold, T. F. et al. Crystal structure of the dynamin tetramer. Nature 525, 404–408 (2015).
Chappie, J. S. et al. A pseudoatomic model of the dynamin polymer identifies a hydrolysis-dependent powerstroke. Cell 147, 209–222 (2011).
Heymann, J. A. W. & Hinshaw, J. E. Dynamins at a glance. J. Cell Sci. 122, 3427–3431 (2009).
Sundborger, A. C. & Hinshaw, J. E. Dynamins and BAR proteins—safeguards against cancer. Crit. Rev. Oncog. 20, 475–484 (2015).
Sun, Y. & Tien, P. From endocytosis to membrane fusion: emerging roles of dynamin in virus entry. Crit. Rev. Microbiol. 39, 166–179 (2013).
Harper, C. B., Popoff, M. R., McCluskey, A., Robinson, P. J. & Meunier, F. A. Targeting membrane trafficking in infection prophylaxis: dynamin inhibitors. Trends Cell Biol. 23, 90–101 (2013).
Daumke, O. & Praefcke, G. J. K. Mechanisms of GTP hydrolysis and conformational transitions in the dynamin superfamily. Biopolymers 105, 580–593 (2016).
Sundborger, A. C. & Hinshaw, J. E. Regulating dynamin dynamics during endocytosis. F1000Prime Rep. 6, 85 (2014).
Ford, M. G. J., Jenni, S. & Nunnari, J. The crystal structure of dynamin. Nature 477, 561–566 (2011).
Faelber, K. et al. Crystal structure of nucleotide-free dynamin. Nature 477, 556–560 (2011).
Sundborger, A. C. et al. A dynamin mutant defines a superconstricted prefission state. Cell Reports 8, 734–742 (2014).
Mattila, J.-P. et al. A hemi-fission intermediate links two mechanistically distinct stages of membrane fission. Nature 524, 109–113 (2015).
Antonny, B. et al. Membrane fission by dynamin: what we know and what we need to know. EMBO J. 35, 2270–2284 (2016).
Kozlovsky, Y. & Kozlov, M. M. Membrane fission: model for intermediate structures. Biophys. J. 85, 85–96 (2003).
Gao, S. et al. Structure of myxovirus resistance protein a reveals intra- and intermolecular domain interactions required for the antiviral function. Immunity 35, 514–525 (2011).
Damke, H., Binns, D. D., Ueda, H., Schmid, S. L. & Baba, T. Dynamin GTPase domain mutants block endocytic vesicle formation at morphologically distinct stages. Mol. Biol. Cell 12, 2578–2589 (2001).
Larson, B. T., Sochacki, K. A., Kindem, J. M. & Taraska, J. W. Systematic spatial mapping of proteins at exocytic and endocytic structures. Mol. Biol. Cell 25, 2084–2093 (2014).
Grassart, A. et al. Actin and dynamin2 dynamics and interplay during clathrin-mediated endocytosis. J. Cell Biol. 205, 721–735 (2014).
Srinivasan, S. et al. A noncanonical role for dynamin-1 in regulating early stages of clathrin-mediated endocytosis in non-neuronal cells. PLoS Biol. 16, e2005377 (2018).
Sochacki, K. A., Dickey, A. M., Strub, M.-P. & Taraska, J. W. Endocytic proteins are partitioned at the edge of the clathrin lattice in mammalian cells. Nat. Cell Biol. 19, 352–361 (2017).
Petrache, H. I. et al. Structure and fluctuations of charged phosphatidylserine bilayers in the absence of salt. Biophys. J. 86, 1574–1586 (2004).
Mitra, K., Ubarretxena-Belandia, I., Taguchi, T., Warren, G. & Engelman, D. M. Modulation of the bilayer thickness of exocytic pathway membranes by membrane proteins rather than cholesterol. Proc. Natl Acad. Sci. USA 101, 4083–4088 (2004).
Chen, Y.-J., Zhang, P., Egelman, E. H. & Hinshaw, J. E. The stalk region of dynamin drives the constriction of dynamin tubes. Nat. Struct. Mol. Biol. 11, 574–575 (2004).
Zhang, P. & Hinshaw, J. E. Three-dimensional reconstruction of dynamin in the constricted state. Nat. Cell Biol. 3, 922–926 (2001).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
He, S. & Scheres, S. H. W. Helical reconstruction in RELION. J. Struct. Biol. 198, 163–176 (2017).
Shaikh, T. R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nat. Protocols 3, 1941–1974 (2008).
Scheres, S. H. W. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Rost, B. PHD: predicting one-dimensional protein structure by profile-based neural networks. Methods Enzymol. 266, 525–539 (1996).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. 86, 2.9.1–2.9.37 (2016).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Taylor, M. J., Perrais, D. & Merrifield, C. J. A high precision survey of the molecular dynamics of mammalian clathrin-mediated endocytosis. PLoS Biol. 9, e1000604 (2011).
Acknowledgements
This work was supported by the NIDDK and NHLBI NIH Intramural Research Program. This work used the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy (NRAMM) located at the New York Structural Biology Center (supported by grants from the Simons Foundation (349247), NYSTAR, and the NIH National Institute of General Medical Sciences (GM103310) with additional support from the Agouron Institute (F00316) and NIH (S10 OD019994-01)), the NHLBI light microscopy core facility, the NHLBI flow cytometry core facility and the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov). We thank P. Flicker, J. Kehr, J. Morin-Leisk, A. Shuali and A. Sundborger for discussions; P. Dagur for technical assistance with flow cytometry; and B. Carragher and C. Potter for technical assistance with data collection at NRAMM.
Reviewer information
Nature thanks S. Scheres and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
L.K. and J.E.H. designed the research; L.K. and J.E.H. prepared protein samples; L.K., H.W., W.J.R. and J.E.H. collected cryo-EM data; L.K., S.F., A.D.K., H.W., B.C. and J.E.H. processed and analysed the data; K.A.S. and J.W.T. designed and performed cell-based assays; M.-P.S. generated all constructs for cell-based assays; L.K. and J.E.H. wrote the paper; all authors were asked to comment on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM parameters and data analysis.
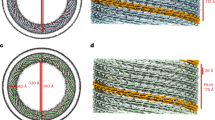
a, Diameter distribution of dynamin tubes in the absence or presence of GMPPCP or GTP. Scale bar, 200 nm. Each experiment was repeated three independent times with similar results. b, Local resolution and helical parameters. R, rise; T, twist; SPT, subunits per helical turn. c, Gold standard FSC curves of the dynamin 3D maps. d, Model-to-map FSC curves. Dotted blue line indicates gold standard resolution at 0.5 FSC.
Extended Data Fig. 2 Cryo-EM data collection and processing flowchart.
Starting from the top, the flowchart details the pathways of three separate samples (red, green and blue) of dynamin protein through imaging and processing. The samples were imaged by two different microscopes, and then three different conformational states were selected manually. Each of these states was processed separately by Spider and then Relion. The red stars in the polymer diameter histograms indicate the diameters of the particles used for the final reconstructions. See Methods for details.
Extended Data Fig. 3 Dynamin helical tubes and their Fourier transforms.
a, b, Representative cryo-EM images (left) of two dynamin polymers in different conformations. Each experiment was repeated three independent times with similar results. Right, Fourier transforms of the polymer images. The strong layer lines associated with the pitch (red arrows) are highlighted. The distance of the layer lines from the meridian (m), highlighted by dotted black lines, indicate that the dynGTP polymer is a two-start helix, whereas the dynGMPPCP polymer is a one-start helix. c, d, Power spectra of 2D class averages from dynGTP (c) and dynGMPPCP (d), highlighting the differences between a two-start and one-start helix. e, f, Sections of the dynGMPPCP (e) and dynGTP (f) maps starting from the outside and going to the middle section. A top middle section looking down the centre of the tubes is shown in the top left panels.
Extended Data Fig. 4 Functional cell-based assays probing dynamin mutants and structural comparison of the BSE hinge.
a, Transferrin uptake of additional interface 3 mutants. Wild-type and K44A are shown for comparison. Mean ± propagated s.d. from n = 3 biological replicates are shown. Also shown are single GFP+ biological replicates that are each background subtracted and referenced to mean values (grey dots). Experiment was repeated and trends verified with n = 2 biologically independent experiments. b, Comparison of α5G and α2B helices from dynGMPPCP (cryo-bent and cryo-unbent, respectively) and available crystal structures, including dynamin bound to GMPPCP (PCP 1 and 2, PDB ID: 3ZYC), dynamin bound to GDP-AlF (GDP-AlF 1 and 2, PDB ID: 2X2E) and dynamin bound to GDP (GDP 1 and 2, PDB ID: 5D3Q). The number after each structure (1 or 2) represents the first or the second member of the dynamin dimer presented in each structure.
Extended Data Fig. 5 FACS gating.
a–h, To test the robustness of our gating, we used either side-scattering (a–d) or DAPI (e–h) to isolate single cells in two different experimental replicates. In both cases, single cells were isolated from debris (a, e). Single cells were then separated into GFP– and GFP+ gates (b, c, f, g). Controls lacking transfection or transferrin uptake (b, f) informed gating choices. Inhibitory mutants exhibited a characteristic dip in transferrin fluorescence in the GFP+ cells (c, g). The exact fraction of transferrin uptake was dependent on GFP+ gating choices (d, h) while qualitative trends were consistent. Mean ± propagated s.d. from n = 3 biological replicates are shown. Also shown are single GFP+ biological replicates that are each background subtracted and referenced to mean values (grey dots).
Extended Data Fig. 6 Comparison of dynGMPPCP and dynAPO at interface G2.
a, A large swing in the BSE of the tetramer in the apo conformation (PDB ID: 5A3F) disrupts interface G2. The apo tetramer and cryo-EM dynGMPPCP structure (GMPPCP-bound conformation) were aligned by the stalk. Curved arrow indicates the movement of the G domain. Domains are coloured green for GTPase, pink for BSE, blue for stalk in monomer B, and purple for GTPase, orange for BSE, tan for stalk in monomer A. b, Interface G2, coloured as above, in cryo-EM map of dynGMPPCP (grey mesh).
Supplementary information
Rights and permissions
About this article
Cite this article
Kong, L., Sochacki, K.A., Wang, H. et al. Cryo-EM of the dynamin polymer assembled on lipid membrane. Nature 560, 258–262 (2018). https://doi.org/10.1038/s41586-018-0378-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0378-6
Keywords
This article is cited by
-
SynDLP is a dynamin-like protein of Synechocystis sp. PCC 6803 with eukaryotic features
Nature Communications (2023)
-
Brominated lipid probes expose structural asymmetries in constricted membranes
Nature Structural & Molecular Biology (2023)
-
Molecular mechanics underlying flat-to-round membrane budding in live secretory cells
Nature Communications (2022)
-
Cryo-electron tomography reveals structural insights into the membrane remodeling mode of dynamin-like EHD filaments
Nature Communications (2022)
-
Event-triggered STED imaging
Nature Methods (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.