Abstract
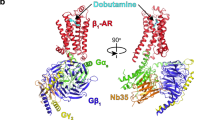
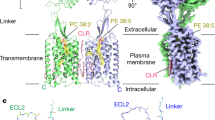
G-protein-coupled receptors (GPCRs) form the largest family of receptors encoded by the human genome (around 800 genes). They transduce signals by coupling to a small number of heterotrimeric G proteins (16 genes encoding different α-subunits). Each human cell contains several GPCRs and G proteins. The structural determinants of coupling of Gs to four different GPCRs have been elucidated1,2,3,4, but the molecular details of how the other G-protein classes couple to GPCRs are unknown. Here we present the cryo-electron microscopy structure of the serotonin 5-HT1B receptor (5-HT1BR) bound to the agonist donitriptan and coupled to an engineered Go heterotrimer. In this complex, 5-HT1BR is in an active state; the intracellular domain of the receptor is in a similar conformation to that observed for the β2-adrenoceptor (β2AR)3 or the adenosine A2A receptor (A2AR)1 in complex with Gs. In contrast to the complexes with Gs, the gap between the receptor and the Gβ-subunit in the Go–5-HT1BR complex precludes molecular contacts, and the interface between the Gα-subunit of Go and the receptor is considerably smaller. These differences are likely to be caused by the differences in the interactions with the C terminus of the Go α-subunit. The molecular variations between the interfaces of Go and Gs in complex with GPCRs may contribute substantially to both the specificity of coupling and the kinetics of signalling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
García-Nafría, J., Lee, Y., Bai, X., Carpenter, B. & Tate, C. G. Cryo-EM structure of the adenosine A2A receptor coupled to an engineered heterotrimeric G protein. eLife 7, e35946 (2018).
Liang, Y. L. et al. Phase-plate cryo-EM structure of a class B GPCR–G protein complex. Nature 546, 118–123 (2017).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Zhang, Y. et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 546, 248–253 (2017).
Syrovatkina, V., Alegre, K. O., Dey, R. & Huang, X. Y. Regulation, signaling, and physiological functions of G proteins. J. Mol. Biol. 428, 3850–3868 (2016).
Oldham, W. M. & Hamm, H. E. Structural basis of function in heterotrimeric G proteins. Q. Rev. Biophys. 39, 117–166 (2006).
Flock, T. et al. Universal allosteric mechanism for Gα activation by GPCRs. Nature 524, 173–179 (2015).
Venkatakrishnan, A. J. et al. Molecular signatures of G protein-coupled receptors. Nature 494, 185–194 (2013).
Oldham, W. M. & Hamm, H. E. Heterotrimeric G protein activation by G protein-coupled receptors. Nat. Rev. Mol. Cell Biol. 9, 60–71 (2008).
Nehme, R. et al. Mini-G proteins: novel tools for studying GPCRs in their active conformation. PLoS One 12, e0175642 (2017).
Masuho, I. et al. Distinct profiles of functional discrimination among G proteins determine the actions of G protein-coupled receptors. Sci. Signal. 8, ra123 (2015).
McCorvy, J. D. & Roth, B. L. Structure and function of serotonin G protein-coupled receptors. Pharmacol. Ther. 150, 129–142 (2015).
Berger, M., Gray, J. A. & Roth, B. L. The expanded biology of serotonin. Annu. Rev. Med. 60, 355–366 (2009).
Wacker, D. et al. Structural features for functional selectivity at serotonin receptors. Science 340, 615–619 (2013).
Wang, C. et al. Structural basis for molecular recognition at serotonin receptors. Science 340, 610–614 (2013).
Yin, W. et al. Crystal Structure of the human 5-HT1B serotonin receptor bound to an inverse agonist. Cell Discovery 4, 12 (2018).
Albert, P. R. & Tiberi, M. Receptor signaling and structure: insights from serotonin-1 receptors. Trends Endocrinol. Metab. 12, 453–460 (2001).
Ring, A. M. et al. Adrenaline-activated structure of β2-adrenoceptor stabilized by an engineered nanobody. Nature 502, 575–579 (2013).
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011).
Ballesteros, J.A., Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors Methods Neurosci. 25, 366–428 (1995).
Carpenter, B., Nehme, R., Warne, T., Leslie, A. G. & Tate, C. G. Structure of the adenosine A2A receptor bound to an engineered G protein. Nature 536, 104–107 (2016).
Lebon, G., Warne, T. & Tate, C. G. Agonist-bound structures of G protein-coupled receptors. Curr. Opin. Struct. Biol. 22, 482–490 (2012).
Bae, H., Cabrera-Vera, T. M., Depree, K. M., Graber, S. G. & Hamm, H. E. Two amino acids within the α4 helix of Gαi1 mediate coupling with 5-hydroxytryptamine1B receptors. J. Biol. Chem. 274, 14963–14971 (1999).
Venkatakrishnan, A. J. et al. Diverse activation pathways in class A GPCRs converge near the G protein-coupling region. Nature 536, 484–487 (2016).
Flock, T. et al. Selectivity determinants of GPCR–G protein binding. Nature 545, 317–322 (2017).
Kobilka, B. K. & Deupi, X. Conformational complexity of G protein-coupled receptors. Trends Pharmacol. Sci. 28, 397–406 (2007).
Scheerer, P. et al. Crystal structure of opsin in its G protein-interacting conformation. Nature 455, 497–502 (2008).
Grundmann, M. & Kostenis, E. Temporal bias: time-encoded dynamic GPCR signaling. Trends Pharmacol. Sci. 38, 1110–1124 (2017).
Lane, J. R., May, L. T., Parton, R. G., Sexton, P. M. & Christopoulos, A. A kinetic view of GPCR allostery and biased agonism. Nat. Chem. Biol. 13, 929–937 (2017).
Isberg, V. et al. GPCRdb: an information system for G protein-coupled receptors. Nucleic Acids Res. 44, D356–D364 (2016).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, https://doi.org/10.7554/eLife.18722 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr D 73, 496–502 (2017).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Burnley, T., Palmer, C. M. & Winn, M. Recent developments in the CCP-EM software suite. Acta Crystallogr D 73, 469–477 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. 60, 2126–2132 (2004).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D. 67, 355–367 (2011).
Lebedev, A. A. et al. JLigand: a graphical tool for the CCP4 template-restraint library. Acta Crystallogr. D. 68, 431–440 (2012).
Nicholls, R. A., Long, F. & Murshudov, G. N. Low-resolution refinement tools in REFMAC5. Acta Crystallogr. D. 68, 404–417 (2012).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. 66, 12–21 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. 66, 213–221 (2010).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Acknowledgements
This work was funded by a grant from the European Research Council (EMPSI 339995), Heptares Therapeutics and core funding from the Medical Research Council (MRC U105197215). We thank J. Espinosa and L. Renault for their help with data collection at NeCEN; S. Scheres and P. da Fonseca for useful discussions and C. Savva and G. Cannone for microscopy technical support.
Reviewer information:
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.N. performed receptor expression, purification and complex formation. P.C.E. expressed and purified mini-Go and Gβγ. R.N. and J.G.-N. performed cryo-grid preparation. J.G.-N. performed cryo-EM data collection, data processing and model building. J.G.-N. and C.G.T. carried out structure analysis and manuscript preparation. C.G.T. analysed data and managed the project. The manuscript was written by C.G.T and J.G.-N., and included contributions from all the authors.
Corresponding author
Ethics declarations
Competing interests
C.G.T. is a shareholder, consultant and member of the Scientific Advisory Board of Heptares Therapeutics, who also partly funded this work.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM single particle reconstruction of the 5-HT1BR–Go complex structure.
a, Representative micrograph (magnification 75,000×, defocus −0.6 μm) of the 5-HT1BR–Go complex collected using a Titan Krios with the Falcon III detector and Volta phase plate. b, Representative 2D class averages of the 5-HT1BR–Go complex. c, FSC curve of the final reconstruction showing an overall resolution of 3.8 Å using the gold-standard FSC of 0.143. Both masked and unmasked FSC curves are shown to highlight the lack of masking artefacts. d, Final reconstruction coloured by subunit. Inset shows a magnified view of the weak density for ICL3. The magnified region corresponds to a map sharpened with B = –50 to remove noise from lower density levels. e, Local resolution estimation of the 5-HT1BR map as calculated by Resmap.
Extended Data Fig. 2 Cryo-EM map quality and model validation.
a, Transmembrane helices of 5-HT1BR. b, The α5-helix of Go. c, Donitriptan and the neighbouring side chains in the orthosteric binding site. d, FSC of the refined model versus the map (green curve) and FSCwork/FSCtest validation curves (blue and red curves, respectively).
Extended Data Fig. 3 Flow chart of data processing.
Micrographs were collected during nine sessions on the Titan Krios (either 24 h or 48 h) and each session was processed independently. The number of images and particles from one 48-h session is indicated on the flowchart as a guide. At the bottom of the figure, the final number of particles is shown. Each dataset was corrected separately for drift, beam-induced motion and radiation damage. After CTF estimation, particles were picked using a Gaussian blob and submitted to either one or two rounds of reference-free 2D classification (see Methods). A 3D classification was performed on the selected particles using an ab initio model generated from ten thousand particles. Classification was performed in parallel in three and four classes. The models with best features were refined on their own; if there were two classes of similar high quality, these were then re-refined together (the resolution of the models refers to the resolution after refinement and calculation of gold-standard FSC = 0.143). The set of particles that obtained the best map quality and resolution were saved and merged with the best particles from other datasets. A final model with 730,118 particles was refined and achieved a global resolution of 3.78 Å.
Extended Data Fig. 4 Modelling quality of the 5-HT1BR structure.
a, Amino acid sequence of the 5-HT1BR construct used for the cryo-EM structure determination. Residues are coloured according to how they have been modelled. Black, good density allows the side chain to be modelled; red, limited density for the side chain, therefore the side chain has been truncated to Cβ; blue, no density observed and therefore the residue was not modelled. Regions highlighted in grey represent the transmembrane α-helices, and amphipathic helix 8 is highlighted in yellow. b, Model of 5-HT1BR showing the Cα positions of amino acid residues with poor density (spheres) and unmodelled regions (dotted lines).
Extended Data Fig. 6 Comparison of the amino acid sequences of the α-subunits of Go and Gs.
Diamonds above the sequences indicate amino acid residues in Gαs in which the side chains make atomic contacts to residues in β2AR (β2 con) or A2AR (2A con). Ovals indicate amino acid residues in Gαs in which only the main chain atoms make contacts with the receptor. Secondary structural elements are indicated as grey bars with positions numbered according to the CGN numbering system.
Extended Data Fig. 7 Similarity of Gα structures and the difference poses of the α5-helices in Gαo and Gαs coupled to receptors.
a, The structures of the α-subunits in complex with 5-HT1BR, β2AR3 and A2AR1 were superimposed over the whole of their sequence in Pymol. Blue, Gαo coupled to 5-HT1BR; green, Gαs coupled to A2AR; Gαs coupled to β2AR. b, 5-HT1BR (blue), β2AR3 (green) and A2AR1 (red) were superimposed based on H3, H5 and H6. Two different views are shown with the red arrows indicating differences in orientation of Gαs and Gαo.
Extended Data Fig. 8 Alignment of the amino acid sequences of Go and Gi α-subunits.
Residues in grey correspond to the α-helical region that does not make contact with GPCRs and was deleted during the construction of mini-Go. Secondary structural elements are depicted as grey bars with the CGN numbers shown to aid comparisons. Amino acids are highlighted as follows: pink, stabilizing residues required to generate mini-Go; yellow; residues in Gαo that are different from residues conserved in all three Gαi sequences; blue, residues that are non-conserved in Gαi sequences. #, the affinity tag on mini-Go used for purification (MGHHHHHHENLYFQG).
Extended Data Fig. 9 Comparison of the α5-helices of Gs and Go.
The α5 helices in the cryo-EM structures of A2AR–Gs (carbon, green) and 5-HT1BR–Go (carbon, light blue) were aligned (in Pymol) along their whole sequence and displayed in different poses: cartoon depiction (a); Gs (green spheres), Go (blue sticks) (b); Go (blue spheres), Gs (green sticks) (c).
Supplementary information
Rights and permissions
About this article
Cite this article
García-Nafría, J., Nehmé, R., Edwards, P.C. et al. Cryo-EM structure of the serotonin 5-HT1B receptor coupled to heterotrimeric Go. Nature 558, 620–623 (2018). https://doi.org/10.1038/s41586-018-0241-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0241-9
This article is cited by
-
Binding kinetics drive G protein subtype selectivity at the β1-adrenergic receptor
Nature Communications (2024)
-
Prediction of dynamic allostery for the transmembrane domain of the sweet taste receptor subunit, TAS1R3
Communications Biology (2023)
-
Ligand and G-protein selectivity in the κ-opioid receptor
Nature (2023)
-
GPCRome-wide analysis of G-protein-coupling diversity using a computational biology approach
Nature Communications (2023)
-
Structural insights into the human niacin receptor HCA2-Gi signalling complex
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.