Abstract
Histone post-translational modifications (PTMs) are associated with epigenetic states that form the basis for cell-type-specific gene expression1,2. Once established, histone PTMs can be maintained by positive feedback involving enzymes that recognize a pre-existing histone modification and catalyse the same modification on newly deposited histones. Recent studies suggest that in wild-type cells, histone PTM-based positive feedback is too weak to mediate epigenetic inheritance in the absence of other inputs3,4,5,6,7. RNA interference (RNAi)-mediated histone H3 lysine 9 methylation (H3K9me) and heterochromatin formation define a potential epigenetic inheritance mechanism in which positive feedback involving short interfering RNA (siRNA) amplification can be directly coupled to histone PTM positive feedback8,9,10,11,12,13,14. However, it is not known whether the coupling of these two feedback loops can maintain epigenetic silencing independently of DNA sequence and in the absence of enabling mutations that disrupt genome-wide chromatin structure or transcription15,16,17. Here, using the fission yeast Schizosaccharomyces pombe, we show that siRNA-induced H3K9me and silencing of a euchromatic gene can be epigenetically inherited in cis during multiple mitotic and meiotic cell divisions in wild-type cells. This inheritance involves the spreading of secondary siRNAs and H3K9me3 to the targeted gene and surrounding areas, and requires both RNAi and H3K9me, suggesting that the siRNA and H3K9me positive-feedback loops act synergistically to maintain silencing. By contrast, when maintained solely by histone PTM positive feedback, silencing is erased by H3K9 demethylation promoted by Epe1, or by interallelic interactions that occur after mating to cells containing an expressed allele even in the absence of Epe1. These findings demonstrate that the RNAi machinery can mediate transgenerational epigenetic inheritance independently of DNA sequence or enabling mutations, and reveal a role for the coupling of the siRNA and H3K9me positive-feedback loops in the protection of epigenetic alleles from erasure.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Margueron, R. & Reinberg, D. Chromatin structure and the inheritance of epigenetic information. Nat. Rev. Genet. 11, 285–296 (2010).
Moazed, D. Mechanisms for the inheritance of chromatin states. Cell 146, 510–518 (2011).
Ragunathan, K., Jih, G. & Moazed, D. Epigenetic inheritance uncoupled from sequence-specific recruitment. Science 348, 1258699 (2015).
Audergon, P. N. et al. Restricted epigenetic inheritance of H3K9 methylation. Science 348, 132–135 (2015).
Wang, X. & Moazed, D. DNA sequence-dependent epigenetic inheritance of gene silencing and histone H3K9 methylation. Science 356, 88–91 (2017).
Laprell, F., Finkl, K. & Müller, J. Propagation of Polycomb-repressed chromatin requires sequence-specific recruitment to DNA. Science 356, 85–88 (2017).
Coleman, R. T. & Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of a Drosophila HOX gene. Science 356, eaai8236 (2017).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Motamedi, M. R. et al. Two RNAi complexes, RITS and RDRC, physically interact and localize to noncoding centromeric RNAs. Cell 119, 789–802 (2004).
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Sienski, G. et al. Silencio/CG9754 connects the Piwi–piRNA complex to the cellular heterochromatin machinery. Genes Dev. 29, 2258–2271 (2015).
Yu, Y. et al. Panoramix enforces piRNA-dependent cotranscriptional silencing. Science 350, 339–342 (2015).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Luteijn, M. J. & Ketting, R. F. PIWI-interacting RNAs: from generation to transgenerational epigenetics. Nat. Rev. Genet. 14, 523–534 (2013).
Iida, T., Nakayama, J. & Moazed, D. siRNA-mediated heterochromatin establishment requires HP1 and is associated with antisense transcription. Mol. Cell 31, 178–189 (2008).
Yu, R., Jih, G., Iglesias, N. & Moazed, D. Determinants of heterochromatic siRNA biogenesis and function. Mol. Cell 53, 262–276 (2014).
Kowalik, K. M. et al. The Paf1 complex represses small-RNA-mediated epigenetic gene silencing. Nature 520, 248–252 (2015).
Allshire, R. C., Javerzat, J. P., Redhead, N. J. & Cranston, G. Position effect variegation at fission yeast centromeres. Cell 76, 157–169 (1994).
Thakurta, A. G., Gopal, G., Yoon, J. H., Kozak, L. & Dhar, R. Homolog of BRCA2-interacting Dss1p and Uap56p link Mlo3p and Rae1p for mRNA export in fission yeast. EMBO J. 24, 2512–2523 (2005).
Jih, G. et al. Unique roles for histone H3K9me states in RNAi and heritable silencing of transcription. Nature 547, 463–467 (2017).
Grewal, S. I. & Klar, A. J. Chromosomal inheritance of epigenetic states in fission yeast during mitosis and meiosis. Cell 86, 95–101 (1996).
Gaydos, L. J., Wang, W. & Strome, S. H3K27me and PRC2 transmit a memory of repression across generations and during development. Science 345, 1515–1518 (2014).
Trewick, S. C., Minc, E., Antonelli, R., Urano, T. & Allshire, R. C. The JmjC domain protein Epe1 prevents unregulated assembly and disassembly of heterochromatin. EMBO J. 26, 4670–4682 (2007).
Ronshaugen, M. & Levine, M. Visualization of trans-homolog enhancer–promoter interactions at the Abd-B Hox locus in the Drosophila embryo. Dev. Cell 7, 925–932 (2004).
Chen, J. L. et al. Enhancer action in trans is permitted throughout the Drosophila genome. Proc. Natl Acad. Sci. USA 99, 3723–3728 (2002).
Wong, K. H. & Struhl, K. The Cyc8–Tup1 complex inhibits transcription primarily by masking the activation domain of the recruiting protein. Genes Dev. 25, 2525–2539 (2011).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Halic, M. & Moazed, D. Dicer-independent primal RNAs trigger RNAi and heterochromatin formation. Cell 140, 504-516 (2010).
Acknowledgements
We thank members of the Moazed laboratory for discussions and K. Connolly, N. Iglesias, G. Jih and G. Shipkovenska for comments on the manuscript. R.Y. was partially supported by a National Institutes of Health (NIH) training grant (T32, GM96911). This work was supported by a grant from the NIH to D.M. (GM072805). D.M. is an investigator of the Howard Hughes Medical Institute.
Reviewer information
Nature thanks S. Jia and M. Zaratiegui for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.Y. and D.M. conceived the project; R.Y. and X.W. performed experiments; R.Y. designed and performed experiments examining the endogenous ade6+ locus and made the discovery of homologous erasure; X.W. designed and performed experiments examining non-endogenous ade6+ transgenes, comparing hairpin versus cen siRNA triggers and deletion of vtc4+ and rpl3402+, and repeated several experiments; R.Y. analysed all the sequencing libraries; R.Y. and D.M. wrote the paper; all authors discussed results and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
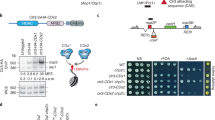
Extended Data Fig. 1 siRNA-induced H3K9me3 at the euchromatic ade6+ locus and flanking region.
a, Summary of pathways and factors involved in mRNA 3′-end processing and its coupling to nuclear export. Chp1 and Ago1 are subunits of the RITS. Uap56 and Mlo3 are TREX (transcription–export) complex subunits, and associate with Dss1 to mediate mRNA export. Tho1 is a subunit of the THO/TREX complex, which is responsible for recruiting other subunits of the TREX complex to mRNA. Mex67 is an orthologue of human NXF1, a critical mRNA export receptor, and associates with Nxt1, another mRNA export factor. Puf6 is an mRNA 3′-UTR-binding protein that has been shown to associate with Mlo3. Rhn1 is involved in RNA polymerase II transcriptional termination. Nab2 is a poly(A)-binding protein. The Paf1 complex is required for transcriptional elongation and 3′ end processing, and mutations in Paf1 subunits allow siRNA-mediated heterochromatin formation and silencing of euchromatic genes. Nup132 is a component of the nuclear pore complex (NPC) that has been linked to mRNA export factors. Bdf2 is a histone-binding protein that has been shown to inhibit the spreading of centromeric heterochromatin. Epe1 is a JmjC-domain containing putative demethylase that also promotes spreading of heterochromatin. Mst2 is a histone aceyltransferase. See main text for references. b–c, Around 1,000 cen::ade6+ mst2∆ (b) or cen::ade6+ leo1∆ (c) cells were plated on low adenine medium (Low Ade). Most cells formed white colonies, indicating expression of endogenous ade6+, although approximately 2% of mst2∆ (b) and 12% of leo1∆ (c) cells formed red or pink colonies, indicating silencing of endogenous ade6+ (white arrow). Upon replating, the resulting red colonies formed mostly red colonies, indicating efficient maintenance of the silent state. Experiment was performed twice with similar results for each. d–e, ChIP–qPCR assays showing mean ± s.d. H3K9me2 levels at the vtc4+ locus, which is located next to the ade6+ gene, in the indicated mutant cells on the basis of two (d) or three (e) independent clones. P values are based on a two-tailed Student's t-test comparing the indicated mutants to wild-type cells. f–h, ChIP–qPCR assays showing mean ± s.d. RNA polymerase II occupancy at the ade6+ (purple) or vtc4+ (blue) locus in mlo3∆ or leo1∆ clones that have not been selected for silencing (99.5% or 88% white, respectively). P values are based on a two-tailed Student's t-test comparing the indicated mutants to cen+ mlo3∆ (f) or to wild-type cells (g). In h, either an ade6+ ON (W) or ade6+ OFF (R) colony from each of the two clones was picked for analysis. P values are based on a two-tailed Student's t-test comparing the indicated red to white cells for each clone. Three biological replicates were used per sample.
Extended Data Fig. 2 The acquired ade6+ silent allele is stable in the absence of the cen::ade6+ siRNA trigger and the mlo3∆ enabling mutation.
a, ade6+ OFF progeny of the cross in Fig. 2b with the indicated genotypes were plated on low adenine medium. See Fig. 2c for related results. b–e, Biological replicates of the crosses shown in Fig. 2d–g. cen::ade6+ mlo3∆ ade6+ OFF cells were crossed to cen+ mlo3+ ade6BC+ ON cells with deletions of key RNAi components (b–d) or H3K9 methyltransferase Clr4 (e) and RSA was performed. All ade6+ OFF progeny were RNAi+ and clr4+. Bars indicate number of ade6+ OFF meiotic progeny for each genotype.
Extended Data Fig. 3 The siRNA driver locus is critical for establishing a heritable silent epiallele.
a, cen::ade6+ leo1∆ (top) or ade6+ hairpin (HP) leo1∆ (bottom) cells were plated on low adenine medium. Approximately 12% of cen::ade6+ leo1∆ (top) and 100% of ade6+ HP leo1∆ (bottom) cells formed red or pink colonies, indicating silencing of endogenous ade6+. Experiment repeated twice. b, c, cen::ade6+ ade6+ OFF leo1∆ cells (b) or ade6+ HP ade6+ OFF leo1∆ (c) cells were crossed to cen+ ade6BC+ ON cells and RSA was performed. Number of progeny matching each indicated genotype and phenotype is shown. 80 red and 80 white colonies were genotyped using PCR. Results reflect two independent crosses. d, siRNA sequencing showing limited secondary siRNA generation in ade6+ HP leo1∆ ade6+ OFF cells, compared with more extensive secondary siRNA spreading to neighbouring genes bub1+ and vtc4+ in cen::ade6+ leo1∆ ade6+ OFF cells. Shaded areas represent sequence identity to ade6+ HP (top three rows) or cen::ade6+ (bottom three rows). Two independent clones shown for each experimental sample. e, f, H3K9me2 (e) and H3K9me3 (f) ChIP–seq reads mapping to the endogenous ade6+ locus in cells with the indicated genotypes and expression states. Shaded areas represent sequence identity to ade6+ HP (top three rows) or cen::ade6+ (bottom three rows). Two independent clones are shown for each experimental sample. g, h, ChIP–qPCR assays showing differences in H3K9me2 (g) or H3K9me3 (h) levels in cen::ade6+ leo1∆ ade6+ OFF and ade6+ HP leo1∆ ade6+ OFF cells at ade6+ (purple) and vtc4+ (blue). Data are mean ± s.d. from three biological replicates. P values are based on two-tailed Student's t-test comparing leo1∆ cells to appropriate wild-type cells. On the right, control ChIP–qPCR at dg repeats.
Extended Data Fig. 4 cis inheritance of the acquired ade6+ silencing and its stable propagation over multiple meiotic generations.
a, Top, ade6BC OFF cells were crossed to ade6+ ON cells and tetrad dissection was performed on low adenine medium. Bottom, genotyping using allele-specific PCRs demonstrated the 2:2 segregation of the OFF and ON states and cis transmission of each state. Experiment performed once, but see Fig. 3a for the reciprocal cross. b, ade6+ OFF progeny of repeated ade6+ OFF × ade6BC+ ON crosses were selected and crossed again, showing stability of the ade6+ OFF allele over five meiotic generations. n, number of meiotic progeny analysed. Independent replicate of the cross shown in Fig. 3c.
Extended Data Fig. 5 Induction of H3K9me and siRNAs at the endogenous ade6+ locus.
a, H3K9me2 ChIP–seq reads mapping to the endogenous ade6+ locus in cells with the indicated genotypes and expression states. Shaded area indicates the region of sequence identity with cen::ade6+. b, c, H3K9me2 (b) and H3K9me3 (c) ChIP–seq reads mapping to the dg and dh repeats of centromere 1 (dg1 and dh1, respectively) in cells with the indicated genotypes and phenotypes. d, Zoomed in view of sRNA-seq reads at the endogenous ade6+ locus shown in Fig. 3e. Shaded area indicates cen::ade6+ homology. Two to three independent clones were sequenced each for ON and OFF meiotic progeny.
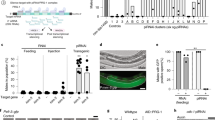
Extended Data Fig. 6 vtc4+ and rpl3402+ are critical for inheritance of silencing at the endogenous ade6+ locus.
a, Schematic of the siRNA driver cen::ade6+ locus on chromosome 1 (upper) and the endogenous ade6+ locus (lower) in which the vt4+ and rpl3402+ genes were replaced with the ura4+ gene (vtc4Δ rpl3402∆::ura4+). b, Frequency of silencing establishment at endogenous ade6+ in cen::ade6+ leo1∆ vtc4Δ rpl3402∆ cells. Experiments were repeated twice. c, cen::ade6+ leo1∆ vtc4Δ rpl3402∆ ade6+ OFF cells were crossed to cen+ leo1+ ade6BC+ ON cells and RSA was performed to test the epigenetic maintenance of the OFF state in the absence of the cen::ade6+ siRNA driver and the leo1∆ enabling mutation. The number of progeny matching each genotype and phenotype are shown. 80 white and 80 red progeny were genotyped. Results reflect two independent crosses.
Extended Data Fig. 7 Heritable silencing of a KanR-ade6+ transgene inserted at four different genomic loci.
a–d, Schematic diagrams of KanR-ade6+ insertions at the mal1+ (a), efm3+ (b), meu10+ (c), and mrp1+ (d) loci. e–h, KanR-ade6+ OFF cells of the indicated genotypes were crossed to cen1+ ade6-M210 cells and RSA was performed. The number of progeny matching each genotype and phenotype are shown. Total red or white colonies genotyped are indicated for each cross. For red cells, the first number indicates kanamycin-resistant colonies (indicating presence of the KanR-ade6+ OFF transgene) and second number indicates total red colonies (the remainder of which possess the endogenous ade6-M210 mutant allele). i, cen+ KanR-ade6+ mlo3+ leo+ progeny of the crosses in e–h were plated on media containing low adenine to test for stability of each epiallele during mitotic growth. Experiments were performed once. j, ChIP–qPCR assay showing enrichment of H3K9me3 at KanR-ade6+ epialleles in the cen+ mlo3+ leo1+ progeny of the crosses in e–h. Data are mean ± s.d. from three biological replicates. k, l, efm3::KanR-ade6+ OFF (k) or meu10::KanR-ade6+ OFF (l) cells were crossed to ade6-M216 ago1∆ (left) or clr4∆ (right) and RSA was performed. All ade6+ OFF progeny were ago1+ and clr4+. Bars indicate the number of ade6+ OFF meiotic progeny for each genotype. Total red or white colonies genotyped are indicated for each cross. For red cells, the first number indicates kanamycin-resistant colonies (reflecting the presence of the KanR-ade6+ OFF transgene) and the second number indicates the total red colonies (the remainder of which possess the ade6-M210 allele).
Extended Data Fig. 8 Heritable silencing of KanR-ade6+ transgenes correlates with local secondary siRNA generation.
a–c, Zoomed-in (upper) or zoomed-out (lower) sRNA-seq reads mapping to the indicated KanR-ade6+ transgenes in cen+ mlo3+ leo1+ meiotic progeny of the crosses shown in Extended Data Fig. 7e–g. In c, two different red clones are shown, corresponding to clone 1 (upper) and clone 2 (lower) in Extended Data Fig. 7i. In these clones, the magnitude of locally hopped siRNAs (not mapping to KanR-ade6+) correlates with the magnitude and efficiency of inherited silencing. d, Zoomed-in view of sRNA-seq reads mapping to the endogenous ade6+ locus. Sequencing was performed once, but for meu10::KanR-ade6+ OFF, which represented a strong silent epiallele, small RNAs from two independent clones (1 and 2) were analysed.
Extended Data Fig. 9 Erasure of the silent state during diploid formation and meiosis requires interallelic pairing.
a, Biological replicate of the cross shown in Fig. 4a. cen::ade6+ mlo3∆ ade6+ OFF cells were crossed to cen+ ago1∆ epe1∆ ade6+ ON cells and RSA was performed. Number and phenotype of mlo3+ progeny with the indicated genotypes and phenotypes (OFF, red; ON, white) are shown. mlo3∆ progeny were excluded. n, number of meiotic progeny analysed. b, Mating of a ura4∆::10XtetO-ade6+ epe1∆ OFF allele with an identical OFF allele, diploid formation and sporulation (meiosis) produces progeny in which the OFF state is maintained at a high frequency (47%). Experiments were repeated twice. c, Mating of the ura4∆::10XtetO-ade6+ epe1∆ OFF allele with a genetically identical ON allele results in erasure of the OFF state in nearly all of the resulting meiotic progeny. Experiments repeated twice. d, Partial disruption of pairing by replacement of ura4∆::10XtetO-ade6+ with ura4+ partially restores the epigenetic maintenance of the OFF state. Experiment performed once.
Extended Data Fig. 10 Strategy for siRNA-induced silencing at the ura4∆::ade6+ locus.
a, The ura4+ coding sequence was replaced with ade6+ to generate a ura4Δ::ade6+ allele in cen::ade6+ leo1Δ ade6-M210 cells. Formation of red colonies on low adenine medium indicated ura4Δ::ade6+ silencing. b, cen::ade6+ leo1Δ ura4Δ::ade6+ OFF cells were crossed to cen+ ura4Δ::ade6+ ON cells to demonstrate that the resulting ura4Δ::ade6+ OFF state is stable in the absence of the cen::ade6+ siRNA trigger or the leo1Δ enabling mutation. c, d, Crosses showing that the ura4Δ::ade6+ OFF epigenetic state depends on Ago1 (c) and Clr4 (d). e, Cross for generating an epe1Δ ura4Δ::ade6+ OFF epiallele (top) and comparison of RNAi-independent (TetR–Clr4-I-induced) and RNAi-dependent (cen::ade6+-induced) ade6+ OFF epialleles. The same results were obtained with independent clones.
Supplementary Information
Supplementary Figure 1
The uncropped PCR genotyping gels for panels used in Figure 2a, Figure 3a, and Extended Data Figure 4a.
Supplementary Table 1
A list of fission yeast S. pombe strains used in this study.
Supplementary Table 2
A list of oligonucleotides used in this study.
Rights and permissions
About this article
Cite this article
Yu, R., Wang, X. & Moazed, D. Epigenetic inheritance mediated by coupling of RNAi and histone H3K9 methylation. Nature 558, 615–619 (2018). https://doi.org/10.1038/s41586-018-0239-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0239-3
This article is cited by
-
Bacterial N4-methylcytosine as an epigenetic mark in eukaryotic DNA
Nature Communications (2022)
-
CHD7 regulates bone-fat balance by suppressing PPAR-γ signaling
Nature Communications (2022)
-
Molecular mechanisms of transgenerational epigenetic inheritance
Nature Reviews Genetics (2022)
-
Mechanisms of chromatin-based epigenetic inheritance
Science China Life Sciences (2022)
-
A Review on Epigenetic Inheritance of Experiences in Humans
Biochemical Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.