Abstract
Adipocyte development and differentiation have an important role in the aetiology of obesity and its co-morbidities1,2. Although multiple studies have investigated the adipogenic stem and precursor cells that give rise to mature adipocytes3,4,5,6,7,8,9,10,11,12,13,14, our understanding of their in vivo origin and properties is incomplete2,15,16. This is partially due to the highly heterogeneous and unstructured nature of adipose tissue depots17, which has proven difficult to molecularly dissect using classical approaches such as fluorescence-activated cell sorting and Cre–lox lines based on candidate marker genes16,18. Here, using the resolving power of single-cell transcriptomics19 in a mouse model, we reveal distinct subpopulations of adipose stem and precursor cells in the stromal vascular fraction of subcutaneous adipose tissue. We identify one of these subpopulations as CD142+ adipogenesis-regulatory cells, which can suppress adipocyte formation in vivo and in vitro in a paracrine manner. We show that adipogenesis-regulatory cells are refractory to adipogenesis and that they are functionally conserved in humans. Our findings point to a potentially critical role for adipogenesis-regulatory cells in modulating adipose tissue plasticity, which is linked to metabolic control, differential insulin sensitivity and type 2 diabetes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Müller, S., Kulenkampff, E. & Wolfrum, C. in Metabolic Control (ed. Herzig, S.) 251–263 (Springer International Publishing, Cham, 2015).
Rosen, E. D. & Spiegelman, B. M. What we talk about when we talk about fat. Cell 156, 20–44 (2014).
Crisan, M. et al. A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 3, 301–313 (2008).
Tang, W. et al. White fat progenitor cells reside in the adipose vasculature. Science 322, 583–586 (2008).
Vishvanath, L. et al. Pdgfrβ+ mural preadipocytes contribute to adipocyte hyperplasia induced by high-fat-diet feeding and prolonged cold exposure in adult mice. Cell Metab. 23, 350–359 (2016).
Zannettino, A. C. W. et al. Multipotential human adipose-derived stromal stem cells exhibit a perivascular phenotype in vitro and in vivo. J. Cell. Physiol. 214, 413–421 (2008).
Cai, X., Lin, Y., Hauschka, P. V. & Grottkau, B. E. Adipose stem cells originate from perivascular cells. Biol. Cell 103, 435–447 (2011).
Gupta, R. K. et al. Zfp423 expression identifies committed preadipocytes and localizes to adipose endothelial and perivascular cells. Cell Metab. 15, 230–239 (2012).
Jiang, Y., Berry, D. C., Tang, W. & Graff, J. M. Independent stem cell lineages regulate adipose organogenesis and adipose homeostasis. Cell Reports 9, 1007–1022 (2014).
Tran, K.-V. et al. The vascular endothelium of the adipose tissue gives rise to both white and brown fat cells. Cell Metab. 15, 222–229 (2012).
Majka, S. M. et al. De novo generation of white adipocytes from the myeloid lineage via mesenchymal intermediates is age, adipose depot, and gender specific. Proc. Natl Acad. Sci. USA 107, 14781–14786 (2010).
Billon, N. et al. The generation of adipocytes by the neural crest. Development 134, 2283–2292 (2007).
Sowa, Y. et al. Adipose stromal cells contain phenotypically distinct adipogenic progenitors derived from neural crest. PLoS ONE 8, e84206 (2013).
Su, X. et al. Fascia origin of adipose cells. Stem Cells 34, 1407–1419 (2016).
Berry, R., Jeffery, E. & Rodeheffer, M. S. Weighing in on adipocyte precursors. Cell Metab. 19, 8–20 (2014).
Sanchez-Gurmaches, J. & Guertin, D. A. Adipocyte lineages: tracing back the origins of fat. Biochim. Biophys. Acta 1842, 340–351 (2014).
Cristancho, A. G. & Lazar, M. A. Forming functional fat: a growing understanding of adipocyte differentiation. Nat. Rev. Mol. Cell Biol. 12, 722–734 (2011).
Jeffery, E. et al. Characterization of Cre recombinase models for the study of adipose tissue. Adipocyte 3, 206–211 (2014).
Kolodziejczyk, A. A., Kim, J. K., Svensson, V., Marioni, J. C. & Teichmann, S. A. The technology and biology of single-cell RNA sequencing. Mol. Cell 58, 610–620 (2015).
Rodeheffer, M. S., Birsoy, K. & Friedman, J. M. Identification of white adipocyte progenitor cells in vivo. Cell 135, 240–249 (2008).
Hudak, C. S. et al. Pref-1 marks very early mesenchymal precursors required for adipose tissue development and expansion. Cell Reports 8, 678–687 (2014).
Vignali, D. A. A., Collison, L. W. & Workman, C. J. How regulatory T cells work. Nat. Rev. Immunol. 8, 523–532 (2008).
Rennert, R. C. et al. Microfluidic single-cell transcriptional analysis rationally identifies novel surface marker profiles to enhance cell-based therapies. Nat. Commun. 7, 11945 (2016).
Shook, B. et al. The role of adipocytes in tissue regeneration and stem cell niches. Annu. Rev. Cell Dev. Biol. 32, 609–631 (2016).
Gubelmann, C. et al. Identification of the transcription factor ZEB1 as a central component of the adipogenic gene regulatory network. eLife 3, e03346 (2014).
Johansson, T. et al. Building a zoo of mice for genetic analyses: A comprehensive protocol for the rapid generation of BAC transgenic mice. genesis 48, 264–280 (2010).
Rosenwald, M., Perdikari, A., Rülicke, T. & Wolfrum, C. Bi-directional interconversion of brite and white adipocytes. Nat. Cell Biol. 15, 659–667 (2013).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44, D710–D716 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Andrews, T. S. & Hemberg, M. Modelling dropouts allows for unbiased identification of marker genes in scRNASeq experiments. Preprint at https://www.biorxiv.org/content/early/2016/07/21/065094 (2016).
Van Der Maaten, L. J. P. & Hinton, G. E. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Kiselev, V. Y. et al. SC3: consensus clustering of single-cell RNA-seq data. Nat. Methods 14, 483–486 (2017).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
Scialdone, A. et al. Computational assignment of cell-cycle stage from single-cell transcriptome data. Methods 85, 54–61 (2015).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Pradhan, R. N. et al. Dissecting the brown adipogenic regulatory network using integrative genomics. Sci. Rep. 7, 42130 (2017).
Alpern, D., Gardeux, V., Russeil, J. & Deplancke, B. Time- and cost-efficient high-throughput transcriptomics enabled by Bulk RNA Barcoding and sequencing. Preprint at https://www.biorxiv.org/content/early/2018/01/30/256594 (2018).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Drost, H.-G. & Paszkowski, J. Biomartr: genomic data retrieval with R. Bioinformatics 33, 1216–1217 (2017).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kelder, T. et al. WikiPathways: building research communities on biological pathways. Nucleic Acids Res. 40, D1301–D1307 (2012).
The GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Acknowledgements
We thank J. Auwerx, W. Chen, E. Dorcey, N. Gheldof, O. Naveiras and K. Schoonjans for constructive discussions and carefully reading the manuscript. This research was supported by the Human Frontier Science Program LT001032/2013 (to P.C.S.), the Swiss National Science Foundation Grants (#31003A_162735 and 31003A_162887), the Kristian Gerhard Jebsen Foundation for Metabolic Research (to B.D.) and by institutional support from the Swiss Federal Institute of Technology in Lausanne (EPFL) and Zürich (ETH). We thank the EPFL FCCF, BIOP (especially R. Guiet and O. Burri), HCF and GECF for cell sorting, imaging, histology and sequencing support, as well as the VITAL-IT platform (University of Lausanne) for computational support.
Reviewer information
Nature thanks O. Stegle and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
B.D., C.W., P.C.S., M.Z. and H.D. designed the study and wrote the manuscript. P.C.S. conducted all analyses related to transcriptomics. N.A., J.R., H.D. and M.Z. performed the single-cell experiments. H.D. and M.Z. conducted all FACS, qPCR, cell culture and related imaging assays. H.D. and W.S. performed all experiments related to transplants. H.D. performed all siRNA-based knockdown experiments. M.Z. and J.R. conducted all histological assays. C.C., K.-U.S. and G.S. provided access to human samples and helped with processing them. M.Z. and D.A. performed all experiments related to human cells. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
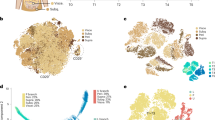
Extended Data Fig. 1 ScRNA-seq reveals the heterogeneity of ASPCs (Fluidigm C1).
a, Number of aligned reads per cell for each of the three Fluidigm C1 (C1) scRNA-seq experiments. R1, n = 74; R2, n = 71; R3, n = 63 single cells from three independent biological replicates, each derived from a pool of mice. b, Correlations (corr, Spearman’s rho) between expression in single cells across all genes or only the artificial RNA spike-ins (ERCC). c, Correlation (Spearman’s rho) between expression in merged single cell (per individual biological replicate, R1–R3) and bulk population (n = 4 biological replicates) samples. d, Number of expressed genes/sample in each of the three biological scRNA-seq replicates (R1–R3), as well as the merged individual replicates (SC, n = 3) or the bulk population control samples (bulk, n = 4). e, Expression of ubiquitous Actb, negative (Ptprc and Pecam1) and positive (Cd34, Ly6a and Itgb1) FACS markers. f, g, Pathways47 significantly enriched (f) and top 10 significantly enriched GTEx48 tissue samples (g) among the top 200 genes specific to one of the three populations (P1, green; P2, red; P3, blue). Full enrichment results are listed in Supplementary Table 3. h, t-SNE 2D maps of all analysed cells (C1), highlighting the expression of the stem cell marker Cd34 and the adipogenic marker Fabp4 (black). i, Bean plots showing the distribution of adipogenic scores across cells belonging to one of the three populations (P1, n = 83; P2, n = 96; P3, n = 29 cells). **P ≤ 0.01, Wilcoxon rank-sum test. j, Stem cell (lower panel) and adipogenic (upper panel) score distributions across all analysed cells, highlighting a single outlier cell not expressing—or expressing very low levels of—Itgb1 (CD29), Cd34 and Ly6a (SCA1) as well as very high levels of the mature adipogenic markers Adipoq, Restn and Cidec. k, Correlation (Spearman’s rho) between the expression of 12 (pre-)adipogenesis-related genes and the expression of Fabp4; significant (Bonferroni multiple testing adjusted P ≤ 0.01) correlations in black. l, t-SNE 2D maps of all analysed cells, highlighting the expression of RFP and various genes previously used to mark adipogenic precursors or pre-adipocytes (black); microscopic RFP status also displayed. In e, h, j, l, n = 208 cells from three independent biological experiments, each based on a pool of mice.
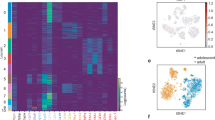
Extended Data Fig. 2 ScRNA-seq reveals the heterogeneity of ASPCs (10x Genomics).
a, b, Number of aligned reads per cell (a) and number of expressed genes per cell (b) for the scRNA-seq experiment performed using the 10x Genomics Chromium instrument. c, Expression of ubiquitous Actb, negative (Ptprc and Pecam1), positive (Cd34, Ly6a and Itgb1) FACS markers and adipocyte markers (Adipoq, Retn and Cidec). d, Number of cells in each of the four 10x Genomics populations (G1–G4). e, Percentage of the top 100 marker genes of the 10x Genomics populations (G1 and G4, G2, and G3) that overlap the top 100 marker genes of the C1 cell populations (P1–P3). f, Bean plots showing the distribution of adipogenic scores across cells that belong to one of the four 10x Genomics populations (G1–G4). Only P values for G2 comparisons are marked. g, Box plots showing the distribution of log(normalized expression) values of pre-adipogenic and ASPC markers across the four 10x Genomics populations (G1–G4). *P ≤ 0.05, **P ≤ 0.01, Wilcoxon rank-sum test. In e–g, G1, n = 699; G2, n = 664; G3, n = 122; and G4, n = 319 cells. h, t-SNE 2D maps of all analysed cells (10x Genomics), highlighting the expression of the pre-adipogenic markers Pdgfra and Pdgfrb (black). In c, h, n = 1,804 cells from one biological experiment based on a pool of mice.
Extended Data Fig. 3 Three ASPC subpopulations with distinct adipogenic differentiation capacity.
a, Box plots showing the distribution of log(normalized expression) values for surface markers with available FACS-grade antibodies selected for subpopulation follow-up: CD55 and IL13RA1 (P1, G1 and G4), VAP1 and ADAM12 (P2 and G2) and ABCG1 (P3 and G3). *P ≤ 0.05, **P ≤ 0.01, Wilcoxon rank-sum test. P1, n = 83; P2, n = 96; P3, n = 29; G1, n = 699; G2, n = 664; G3, n = 122; and G4, n = 319 cells. b, FACS-based sorting strategy to isolate the three identified populations. c, qPCR-based expression fold-changes (log2(fold-change (FC) marker-positive versus marker-negative ASPCs)) for a panel of P1-, P2- and P3-specific genes. d, Microscopy images of distinct ASPC fractions after adipogenic differentiation; CD55+ (P1 or G1 and G4), VAP1+ (P2 or G2), and ABCG1+ (P3 or G3). Nuclei are stained with DAPI (blue) and lipids with LD540 (yellow). Scale bar, 50 μm. e, Bean plots showing the distribution of the fraction of differentiated cells (Fr. diff.) per each ASPC subpopulation shown in d, n = 4 independent wells. M−, marker negative; M+, marker positive. *P ≤ 0.05, **P ≤ 0.01, t-test. f, qPCR-based expression fold-changes (log2(fold-change, marker-positive versus marker-negative ASPCs)) upon adipogenic differentiation for a panel of adipogenic marker genes. g, Bean plots showing the distribution of the mean nuclei number for the four differentiated ASPC fractions shown in d, e and Fig. 2c, d. n = 4 or 5 independent wells. In d, e, g, the experiments were repeated independently three times, yielding similar results; representative images are shown. All panels, population 1 (P1, green), population 2 (P2, red), population 3 (P3, blue).
Extended Data Fig. 4 Aregs (CD142+ABCG1+ ASPCs) show adipogenic inhibitory capacity.
a, Mean and s.d. of number of nuclei per experiment; x axis represents the percentage of Lin−SCA1+CD142+ABCG1+ cells (P3, Aregs) mixed in with Lin−SCA1+CD142−ABCG1− cells; n = 4 independent wells. b, Schematic of sample collection aiming at a detailed characterization of CD142+ ASPCs versus CD142− ASPCs and all ASPCs, after sorting (D0), upon plating (5 h), after culturing (D1) and after adipogenic differentiation (D12). c, Number of aligned reads (top) and number of expressed genes per sample (bottom) across barcoded RNA-seq samples. d, Correlation (Spearman’s rho) between the barcoded mRNA-seq samples after sorting, and the merged scRNA-seq P3 cells; n = 4 biological replicates (top). Example of the correlation (Spearman’s rho) between the log(normalized expression) estimates across merged scRNA-seq P3 cells and one barcoded mRNA-seq replicate (bottom). e, Number of genes (×103) that were significantly differentially expressed (FDR 0.05, fold-change 2) in all comparisons between all ASPCs, CD142+ ASPCs and CD142− ASPCs, after sorting (D0), upon plating (5 h), after culture (D1) and after adipogenic differentiation (D12) (left) as well as upon adipogenic induction (D0 versus D12, D0–D12, and 5 h versus D12, 5 h–D12), in all ASPCs, CD142+ ASPCs and CD142− ASPCs (right). f, Venn diagram showing overlaps between significantly differentially expressed genes (FDR 0.05, fold-change 2) in CD142+ ASPCs (+) versus CD142− ASPCs (−) and all ASPCs (ASPC), after sorting (D0) and after adipogenic differentiation (D12). g, Heat map of the expression (row-wise z-scores of log(normalized expression), blue-to-red) of genes significantly differentially expressed (FDR 0.05, fold-change 2) in CD142+ ASPCs versus CD142− ASPCs and versus all ASPCs upon adipogenic differentiation (D12). Adipogenic marker genes are highlighted. h, i, Pathways47 (h) and top 10 significantly enriched GTEx tissue samples (i) that are significantly enriched among the genes displayed in f. j, Venn diagram showing overlaps between significantly differentially expressed genes (FDR 0.05, fold-change 2) in CD142+ ASPCs versus CD142− ASPCs at all four time points that were assessed. k, Heat map of the expression (row-wise z-scores of log(normalized expression), blue-to-red) of genes that were expressed at significantly higher levels (FDR 0.05, fold-change 2) in CD142+ ASPCs versus CD142− ASPCs after sorting (D0). Genes encoding transcription factors (TFs) and secreted factors (Secr.) are highlighted. l, Top 10 significantly enriched GTEx tissue samples among the genes that were expressed at significantly higher levels (FDR 0.05, fold-change 2) in CD142+ ASPCs versus CD142− ASPCs after sorting, plating and culture (D0, 5 h and D1, respectively) (left) as well as after sorting only (D0) (right). m, Pathways47 significantly enriched among the genes displayed in k. For h, i, l, m, full enrichment results are listed in Supplementary Table 12. n, Heat map of the expression (row-wise z-scores of log(normalized expression), blue-to-red) of a panel of endothelial marker genes. In g, k, n, n = 4–8, 4 biological replicates, 1–3 independent wells each.
Extended Data Fig. 5 The adipogenic inhibitory capacity of Aregs is paracrine.
a, Microscopy images (after adipogenic differentiation) of mouse ASPCs co-cultured in transwells with all ASPCs, CD142− ASPCs or CD142+ ASPCs. Nuclei are stained with Hoechst (blue) and lipids with Bodipy (yellow). Scale bar, 50 μm. The experiments were repeated independently two times, yielding similar results; representative images are shown. b, Bean plots showing the extent of differentiation (arbitrary units, a.u.) per each ASPC fraction shown in a, as well as an additional independent biological replicate. **P ≤ 0.01, Wilcoxon rank-sum test. Experiment S1: ASPCs and CD142− ASPCs, n = 8; CD142+ ASPCs, n = 7 fields of view; experiment S2: ASPCs and CD142− ASPCs, n = 5; CD142+ ASPCs, n = 7 fields of view. c, qPCR-measured gene expression (Rel. expr.) of knockdowns of Fgf12, Rtp3, Spink2 (n = 4) and Vit (n = 6) in total SVF. Two biological replicates, 2 or 3 independent wells each. d, Microscopy images of plated SVF cells with knockdowns of Fgf12, Rtp3, Spink2 and Vit, as well as control (scramble siRNA) knockdowns, after adipogenesis. Nuclei are stained with DAPI (blue) and lipids are stained with LD540 (yellow). Scale bars, 50 μm. e, Bean plots showing the distribution of the fraction of differentiated SVF cells with knockdowns of Fgf12, Rtp3, Spink2 and Vit, as well as control knockdowns. f, Bean plots showing the distribution of the mean number of nuclei for the differentiated fractions shown in e. In e, f, for Fgf12, Rtp3 and Spink2 in experiment S1: n = 7, 2 biological replicates, 3 or 4 independent wells each; in experiment S2: n = 8, 2 biological replicates, 4 independent wells each; for Vit: n = 6, 2 biological replicates, 3 independent wells each. g, qPCR-measured gene expression of Fgf12, Rtp3, Spink2 and Vit in Aregs of the experiments shown in Fig. 2h, i; n = 6, 2 biological replicates, 3 independent wells each. h, Bean plots showing the distribution of the fraction of differentiated cells, as measured in the CD142- ASPCs underneath Aregs with the knockdowns of Fgf12, Rtp3, Spink2, and control; n = 6, 2 biological replicates, 3 independent wells each (independent replication of the experiment shown in Fig. 2h, i). i, Bean plots showing the distribution of the mean number of nuclei for the differentiated fractions shown in Fig. 2h, i (experiment S1) and Extended Data Fig. 5h (experiment S2). n = 6, 2 biological replicates, 3 independent wells. j, Expression of adipogenic markers measured using qPCR in the CD142− ASPCs underneath Aregs with knockdown of genes shown in Fig. 2h, i; n = 6: 2 biological replicates, 3 independent wells each; Experiments were repeated two times, yielding similar results. In c–j, *P ≤ 0.05, **P ≤ 0.01, t-test.
Extended Data Fig. 6 Aregs and their inhibitory capacity are conserved in humans.
a, FACS-based strategy to isolate human (one individual is shown) ex vivo CD142+ ASPCs and CD142− ASPCs. b, qPCR-based expression fold-changes (log2(fold change)) of CD142+ ASPCs versus CD142− ASPCs for F3 and ABCG1 after sorting (left) and F3, PPARG and FABP4 after differentiation (right). n = 3 biological replicates (distinct individuals). *P ≤ 0.05, one-sided paired t-test. c, FACS-based strategy to isolate the human (one individual is shown) in vitro CD142+ ASPCs and CD142− ASPCs. d, Venn diagram showing overlaps between significantly differentially expressed genes (FDR 0.1, fold-change 2) in CD142+ ASPCs versus CD142− ASPCs and all ASPCs after adipogenic differentiation (D12). e, f, Pathways47 (e) and top 10 GTEx tissue samples (f) significantly enriched among the genes that were expressed at significantly higher levels (FDR 0.1, fold-change 2) in CD142− ASPCs versus CD142+ ASPCs after adipogenic differentiation (Fig. 3e). g, Heat map of the expression (row-wise z-scores of log(normalized expression), blue-to-red) of genes that were expressed at significantly higher levels (FDR 0.1, fold-change 2) in CD142+ ASPCs versus CD142− ASPCs after adipogenic differentiation (D12). h, i, Pathways47 (h) and top 10 GTEx tissue samples (i) significantly enriched among the genes that were expressed at significantly higher levels (FDR 0.1, fold change 2) in CD142+ ASPCs versus CD142− ASPCs after adipogenic differentiation (D12) and displayed in g. For e, f, h, i, full enrichment results are provided in Supplementary Table 16. In d–i, n = 4 biological replicates (distinct individuals). j, Microscopy images (after adipogenic differentiation) of human ex vivo ASPCs co-cultured with mouse Areg- (CD142+) or Areg-depleted (CD142−) ASPCs. Nuclei are stained with Hoechst (blue) and lipids with Bodipy (yellow). Scale bar, 50 µm. The experiment was performed once. k, Bean plots showing the extent of adipogenic differentiation (arbitrary units, a.u.) per each ASPC fraction shown in j. n = 35 fields of view. All panels except b: *P ≤ 0.05, **P ≤ 0.01, Wilcoxon rank-sum test.
Extended Data Fig. 7 Aregs locate proximal to the vasculature and inhibit adipogenesis in vivo.
a–g, Microscopy images of the in vivo localization of CD31 (orange), CD142 (green) and SCA1 (pink) markers in mouse subcutaneous white adipose tissue. Nuclei are stained with Hoechst (blue). Scale bar, 50 μm. The experiments were repeated independently at least three times, yielding similar results; representative images are shown. Negative control (a) or tissue cryosections stained with the indicated secondary antibodies only (b). In c, perfused (left) versus non-perfused (right) tissue stained for the indicated markers. In d, arrows indicate CD142+ around big vessels within the adipose parenchyma (left), outside the lymph nodes (middle) and within individual cells (right); in e–g, arrows indicate individual cells presenting both CD142 and SCA1 staining (white). h, Total SVF and Areg-depleted SVF cells (CD142−ABCG1−) were implanted into the subcutaneous adipose depot in Matrigel. Microscopy images of Matrigel plugs with total SVF or CD142−ABCG1− SVF cells. Scale bar, 100 μm. Individuals 3 to 5 are shown, and individuals 1 and 2 are presented in Fig. 4g. i, Mean and s.d. of the percentage of mature adipocytes developed within the plugs composed of either Lin−SCA1+ or Lin−SCA1+CD142− ASPCs (n = 7 biological replicates) analysed using CellProfiler25. *P ≤ 0.01, paired t-test. Mean and s.d. are shown. j, Microscopy images of Matrigel plugs corresponding to i. Scale bar, 100 μm. k, Microscopy images of Matrigel plugs stained with isolectin GS-IB4 (green) and Hoechst (blue) corresponding to i. Scale bar, 100 μm. l, Quantification of vascularisation within the plugs, composed of either Lin−SCA1+ or Lin−SCA1+CD142− ASPCs (n = 6 biological replicates) analysed using CellProfiler25.
Extended Data Fig. 8 Supplementary methods for the scRNA-seq analyses.
a, b, t-SNE 2D maps of all analysed cells (Fluidigm C1), highlighting the four subpopulations (P1–P4) obtained when clustering analysis was performed with k = 4 (a) and cells belonging to one of three biological replicates (b) (pink, R1; green, R2; blue, R3). c, For each of the four clusters shown in a, the number of cells stemming from each biological replicate is highlighted. d, Silhouette analysis results for k = 3 and k = 4. e, Venn diagram showing the overlap between the 527 significantly differentially expressed genes identified using M3Drop in the Letter, and the 1,827 genes showing high variability (HVG). f, For each cluster shown in Fig. 1b, we determined the number of cells contained in the alternative clustering obtained by considering 1,827 highly variable genes for SC3 analysis. The vast majority of cells are similarly attributed, irrespective of gene set choice. g, Percentage of the top 100 marker genes of the three subpopulations described in the Letter that overlap with the top 100 marker genes of the subpopulations derived from the analysis based on highly variable genes. h, t-SNE 2D maps of all analysed cells (C1) considering only the highly variable genes, highlighting the three subpopulations (P1–P3) identified by cluster analysis in the Letter (M3Drop, right) and using the highly variable genes (HVG, left). In a, b, h, n = 208 cells from three independent biological experiments, each based on a pool of mice. i, Marker gene expression across all 10x Genomics cells, including a housekeeping gene (Actb), the negative FACS markers Ptprc (Cd45) and Pecam1 (Cd31), the positive FACS markers Cd34, Ly6a (Sca1) and Itgb1 (Cd29), and the mature adipogenic markers Adipoq, Retn and Cidec. j, t-SNE 2D maps of all 10x Genomics cells highlighting the expression of Xist and the cell division marker Mki67 (black). k, Sum of squares for distinct number of clusters (2 to 15) in the 10x Genomics data. In i, j, n = 2,919 cells from one biological experiment based on a pool of mice, before G1, Xist, Krt18 or Krt19 and Epcam filtering. l, Silhouette width for distinct number of clusters (2 to 13) in the 10x Genomics data. m, Percentage of the top 100 marker genes of the 10x Genomics populations (G1–G4) that overlaps with the top 100 marker genes of the populations (P1–P3) derived from the C1 data based on the highly variable genes.
Supplementary information
Supplementary Information
This file contains a guide to Supplementary Tables 1-19
Supplementary Data
This zipped file contains Supplementary Tables 1-19 – see Supplementary Information document for full guide
Source Data
Rights and permissions
About this article
Cite this article
Schwalie, P.C., Dong, H., Zachara, M. et al. A stromal cell population that inhibits adipogenesis in mammalian fat depots. Nature 559, 103–108 (2018). https://doi.org/10.1038/s41586-018-0226-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0226-8
This article is cited by
-
PDGFRβ + cell HIF2α is dispensable for white adipose tissue metabolic remodeling and hepatic lipid accumulation in obese mice
Lipids in Health and Disease (2024)
-
Unveiling the functional heterogeneity of cytokine-primed human umbilical cord mesenchymal stem cells through single-cell RNA sequencing
Cell & Bioscience (2024)
-
Reduced plasma GDF10 levels are positively associated with cholesterol impairment and childhood obesity
Scientific Reports (2024)
-
Crosstalk of human coronary perivascular adipose-derived stem cells with vascular cells: role of tissue factor
Basic Research in Cardiology (2024)
-
Characteristic and fate determination of adipose precursors during adipose tissue remodeling
Cell Regeneration (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.