Abstract
The expression of co-inhibitory receptors, such as CTLA-4 and PD-1, on effector T cells is a key mechanism for ensuring immune homeostasis. Dysregulated expression of co-inhibitory receptors on CD4+ T cells promotes autoimmunity, whereas sustained overexpression on CD8+ T cells promotes T cell dysfunction or exhaustion, leading to impaired ability to clear chronic viral infections and diseases such as cancer1,2. Here, using RNA and protein expression profiling at single-cell resolution in mouse cells, we identify a module of co-inhibitory receptors that includes not only several known co-inhibitory receptors (PD-1, TIM-3, LAG-3 and TIGIT) but also many new surface receptors. We functionally validated two new co-inhibitory receptors, activated protein C receptor (PROCR) and podoplanin (PDPN). The module of co-inhibitory receptors is co-expressed in both CD4+ and CD8+ T cells and is part of a larger co-inhibitory gene program that is shared by non-responsive T cells in several physiological contexts and is driven by the immunoregulatory cytokine IL-27. Computational analysis identified the transcription factors PRDM1 and c-MAF as cooperative regulators of the co-inhibitory module, and this was validated experimentally. This molecular circuit underlies the co-expression of co-inhibitory receptors in T cells and identifies regulators of T cell function with the potential to control autoimmunity and tumour immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Anderson, A. C., Joller, N. & Kuchroo, V. K. Lag-3, Tim-3, and TIGIT: co-inhibitory receptors with specialized functions in immune regulation. Immunity 44, 989–1004 (2016).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511 (2016).
Maaten, L. H. G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Fitzgerald, D. C. et al. Suppression of autoimmune inflammation of the central nervous system by interleukin 10 secreted by interleukin 27-stimulated T cells. Nat. Immunol. 8, 1372–1379 (2007).
Awasthi, A. et al. A dominant function for interleukin 27 in generating interleukin 10-producing anti-inflammatory T cells. Nat. Immunol. 8, 1380–1389 (2007).
Stumhofer, J. S. et al. Interleukins 27 and 6 induce STAT3-mediated T cell production of interleukin 10. Nat. Immunol. 8, 1363–1371 (2007).
Zhu, C. et al. An IL-27/NFIL3 signalling axis drives Tim-3 and IL-10 expression and T-cell dysfunction. Nat. Commun. 6, 6072 (2015).
Hirahara, K. et al. Interleukin-27 priming of T cells controls IL-17 production in trans via induction of the ligand PD-L1. Immunity 36, 1017–1030 (2012).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
Burton, B. R. et al. Sequential transcriptional changes dictate safe and effective antigen-specific immunotherapy. Nat. Commun. 5, 4741 (2014).
Mayo, L. et al. IL-10-dependent Tr1 cells attenuate astrocyte activation and ameliorate chronic central nervous system inflammation. Brain 139, 1939–1957 (2016).
Castellino, F. J. et al. Mice with a severe deficiency of the endothelial protein C receptor gene develop, survive, and reproduce normally, and do not present with enhanced arterial thrombosis after challenge. Thromb. Haemost. 88, 462–472 (2002).
Sakuishi, K. et al. Targeting Tim-3 and PD-1 pathways to reverse T cell exhaustion and restore anti-tumor immunity. J. Exp. Med. 207, 2187–2194 (2010).
Peters, A. et al. Podoplanin negatively regulates CD4+ effector T cell responses. J. Clin. Invest. 125, 129–140 (2015).
Mackay, L. K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016).
Apetoh, L. et al. The aryl hydrocarbon receptor interacts with c-Maf to promote the differentiation of type 1 regulatory T cells induced by IL-27. Nat. Immunol. 11, 854–861 (2010).
Giordano, M. et al. Molecular profiling of CD8 T cells in autochthonous melanoma identifies Maf as driver of exhaustion. EMBO J. 34, 2042–2058 (2015).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Capaldi, A. P. et al. Structure and function of a transcriptional network activated by the MAPK Hog1. Nat. Genet. 40, 1300–1306 (2008).
Karwacz, K. et al. Critical role of IRF1 and BATF in forming chromatin landscape during type 1 regulatory cell differentiation. Nat. Immunol. 18, 412–421 (2017).
Sen, D. R. et al. The epigenetic landscape of T cell exhaustion. Science 354, 1165–1169 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Chen, L. & Flies, D. B. Molecular mechanisms of T cell co-stimulation and co-inhibition. Nat. Rev. Immunol. 13, 227–242 (2013).
Wende, H. et al. The transcription factor c-Maf controls touch receptor development and function. Science 335, 1373–1376 (2012).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Shalek, A. K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308-1323 (2016).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Blondel, V. D., Guillaume, J. L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. https://doi.org/10.1088/1742-5468/2008/10/P10008 (2008).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Smyth, G. K. in Bioinformatics and Computational Biology Solutions using R and Bioconductor. Statistics for Biology and Health (eds. Gentleman, R. et al.) 397–420 (Springer, New York, 2005).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Lopes, C. T. et al. Cytoscape Web: an interactive web-based network browser. Bioinformatics 26, 2347–2348 (2010).
Acknowledgements
We thank M. Collins for discussions, D. Kozoriz, J. Xia and Z. Chen for technical advice, S. Riesenfeld for computational advice, N. Paul and J. Keegan for CyTOF, and L. Gaffney for artwork. This work was supported by grants from the National Institutes of Health, the American Cancer Society, the Melanoma Research Alliance, the Klarman Cell Observatory at the Broad Institute, and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
N.C., A.M., P.R.B., A.C.A., O.R.-R., A.R. and V.K.K. designed the experiment; N.C., A.M., S.K., J.N., C.D.B., P.R.B., J.D.B. and A.R. developed analytical tools; N.C., A.M., T.K., N.A., J.N., N.D.M., M.S.K., C.W., H.Z., T.L., Y.E. and P.R.B. performed experiments; A.M. and M.S. performed computational analysis. N.C. and A.M. wrote the original draft of the paper and P.R.B., A.C.A., A.R. and V.K.K. reviewed and edited the paper; A.C.A., A.R. and V.K.K. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
A.C.A. is a member of the SAB for Potenza Therapeutics and Tizona Therapeutics. V.K.K. has an ownership interest and is a member of the SAB for Potenza Therapeutics and Tizona Therapeutics. A.C.A.’s and V.K.K.’s interests were reviewed and managed by the Brigham and Women’s Hospital and Partners Healthcare in accordance with their conflict of interest policies. A.R. is an SAB member for Thermo Fisher and Syros Pharmaceuticals and is a consultant for Driver Group.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 CyTOF analysis of co-inhibitory and co-stimulatory receptor co-expression in TILs.
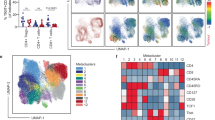
a, TILs were collected from B16F10 melanoma tumour-bearing wild-type and IL-27RA-knockout mice from Fig. 1b and analysed using CyTOF (5,000 cells from each). CyTOF data were analysed using viSNE. Applying k-means clustering with k = 9 on the CyTOF data resulted in a clear distinction between clusters 1, 2, 3 and 4. Polygons indicating clusters 1 and 2 (in CD8+ T cells), and 3 and 4 (in CD4+ T cells) are shown. Individual panels show expression of the indicated markers. b, Pie charts show the distribution of wild-type or IL-27RA-knockout CD8+ and CD4+ TILs in clusters 1 and 2 (C1 and C2) of CD8+ TILs and clusters 3 and 4 (C3 and C4) of CD4+ TILs as defined in Fig. 1d. c, Independent data of wild-type and IL-27RA-knockout TILs samples from that shown in Fig. 1 (5,000 cells from each). Applying k-means clustering with k = 7 on the CyTOF data resulted in a clear distinction between clusters 1, 2, 3 and 4. Polygons indicating clusters 1 and 2 (in CD8+ T cells), and 3 and 4 (in CD4+ T cells) are shown. d, viSNE plot highlighting the distribution of cells from wild-type (blue) and IL-27RA-knockout (red) mice in CD8+ TILs clusters 1 and 2 and CD4+ TILs clusters 3 and 4. Pie charts show the distribution of wild-type or IL-27RA-knockout CD8+ and CD4+ TILs in each cluster.
Extended Data Fig. 2 IL-27 induces multiple co-inhibitory receptors on CD4+ and CD8+ T cells.
a, Naive T cells from wild-type or IL-27RA-knockout mice were stimulated in vitro with anti-CD3/CD28 in the presence or absence of IL-27. Expression of co-inhibitory receptors was determined by flow cytometry. Representative data of three biologically independent experiments are shown. b, Expression of PD-1, TIM-3, LAG-3, TIGIT and IL-10 on CD8+ TILs obtained from wild-type and IL-27RA-knockout mice bearing B16F10 melanoma was determined by flow cytometry. Thy1.1-IL-10 reporter mice crossed with wild-type and IL-27RA-knockout mice were used for IL-10 expression analysis. Representative data of three biologically independent experiments are shown.
Extended Data Fig. 3 scRNA-seq expression analysis of wild-type and IL-27RA-knockout TILs.
a, TILs were obtained from B16F10 melanoma tumour-bearing wild-type (707 and 825 for CD4+ and CD8+, respectively) and IL-27RA-knockout (376 and 394 for CD4+ and CD8+, respectively) mice as in Fig. 1e. t-SNE plot shows the presence of wild-type and IL-27RA-knockout CD4+ and CD8+ TILs as indicated. b, Clustering using the Louvain–Jaccard method (40 nearest neighbours and 13 principal components37). c, The composition of each cluster in terms of total number (c) and percentage (d) of wild-type (red) and IL-27RA-knockout (blue) cells. *P < 0.05, **P < 0.01, ***P < 0.001, one sample t-test. e, Projection of the IL-27 co-inhibitory module signature on the scRNA-seq data. The contour plot marks the region of highly expressing cells by taking into account only those cells that have an expression value above the mean. f, Violin and box plots displaying the distribution of the IL-27 co-inhibitory module signature score compared between wild-type (72 and 98 for CD4+ and CD8+, respectively) and IL-27RA-knockout (85 and 77 for CD4+ and CD8+, respectively) cells in clusters 4 and 5 (CD4+ and CD8+, respectively). *P = 0.01, one-sided t-test. The top and bottom hinges in the boxplot correspond to the first and third quartiles, and the horizontal line denotes the median.
Extended Data Fig. 4 Overlap of the IL-27-induced gene program with signatures from four states of T cell impairment, tolerance and dysfunction.
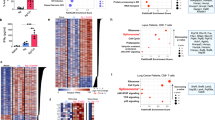
a, Pearson correlation between wild-type CD4+ and CD8+ T cells for the 1,201 genes that were differentially expressed between wild-type CD4+ T cells stimulated in the presence or absence of IL-27 (FDR < 0.05). b, Expression profile of 118 differentially expressed genes (from a) encoding cell-surface receptors and cytokines are shown as a heat map. c, The IL-27-induced gene program (1,201 genes) was compared to T cell signatures obtained from four states of T cell non-responsiveness. Number of overlapping genes between the IL-27 gene program and each signature is depicted. ***P < 0.001, hypergeometric test: nasal anti-CD3 4.7 × 10−21; cancer 1.2 × 10−33; antigen-specific tolerance 4 × 10−14; and viral exhaustion 1.7 × 10−26. d, P-value statistics for the significance of the overlap between the IL-27-induced gene program (1,201) and genes induced in other states of T cell non-responsiveness using WilcoxGST and camera. e, Gene signatures from c were sub-sampled and projected onto the CD8+ single-cell TIL data. Changes were quantified by randomly selecting decreasing subsets of genes from each signature and calculating the average silhouette width of cells that scored high for the different generated signatures based on Euclidian distance between the principal component values used to generate the t-SNE plot. The top and bottom hinges in the boxplot correspond to the first and third quartiles, and the horizontal line denotes the median (Methods). f, Panels I–V, t-SNE plots of the 588 CD8+ single-cell TILs (dots) obtained from wild-type mice bearing B16F10 melanoma tumour. Cells are coloured by their signature score. The score reflects the relative average expression of the genes in the overlap of the IL-27 gene signature with the signatures for each of the indicated states of T cell non-responsiveness. Panel VI is a projection of a signature of the differentially expressed genes between CD8+ TILs from wild-type and IL-27RA-knockout mice bearing B16F10 melanomas (Methods). The contour plot marks the region of highly scored cells by taking into account only those cells that have a signature score above the mean score. Cells that had a statistically significant score (adjusted P < 0.05) are marked by a plus symbol (Methods).
Extended Data Fig. 5 Characterization of the role of PDPN and PROCR in CD8+ TILs.
a, PDPN and PROCR protein and mRNA expression was determined in T cells from wild-type and IL-27RA-knockout mice stimulated with anti-CD3/CD28 in the presence or absence of IL-27. CD4+ cells were analysed at 96 h and CD8+ cells at 72 h. Data are mean ± s.e.m. from representative flow cytometry and qPCR data from biologically independent animals. b, Representative flow cytometry data of three independent experiments showing PDPN and PROCR expression in PD-1+TIM-3+CD8+ and PD-1−TIM-3−CD8+ TILs obtained from wild-type and IL-27RA-knockout mice bearing B16F10 melanoma. c, TILs from wild-type mice bearing B16F10 melanoma were stimulated with phorbol myristate acetate and ionomycin. Cytokine production in PROCR+ or PROCR− CD8+ TILs is shown. Thy1.1-IL-10 reporter mice were used for IL-10 expression analysis. Data are mean ± s.e.m. from biologically independent animals. *P < 0.05, **P < 0.01, paired t-test. d, CD8+ T cells (5 × 105) from wild-type or Procrd/d mice were transferred along with 1 × 106 wild-type CD4+ T cells to RAG1-knockout mice (n = 5). On day 2, 5 × 105 B16F10 cells were implanted. Data are mean ± s.e.m. *P < 0.05, repeated measures ANOVA, Sidak’s multiple comparisons test. e, TILs were obtained from wild-type and PDPN cKO mice bearing B16F10 melanoma and stained for the expression of IL-7RA. Representative flow cytometry data from three independent animals. f, Summary data of IL-7RA expression are from biologically independent animals. Data are mean ± s.e.m. *P < 0.05, one-sided t-test.
Extended Data Fig. 6 PRDM1 is a candidate regulator of the co-inhibitory module.
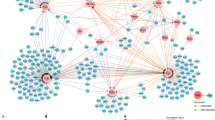
a, The log2 fold change in RNA levels between naive CD4+ or CD8+ T cells simulated with or without IL-27. Data are from two independent experiments. Transcription factors that are part of the IL-27 co-inhibitory module are shown (differentially expressed transcription factors were annotated as genes with FDR-corrected ANOVA < 0.05). b, Transcription factors that are both in the IL-27 co-inhibitory module and are also overexpressed in clusters 4 and 5 in the single-cell data (clusters that were enriched for the IL-27 signature; Extended Data Fig. 3e, f). Differentially expressed genes between clusters 4 and 5 and the rest of the clusters were determined using binomcount.test (binomial distribution, Methods). The log effect corresponds to log proportion of expressing cells and the P value is calculated by the probability of finding n or more cells positive for the gene in clusters 4 and 5 given the fraction in the rest of the clusters. c, t-SNE plot of Fig. 1e showing the expression of PRDM1 in wild-type (707 and 825 for CD4+ and CD8+, respectively) and IL-27RA-knockout (376 and 394 for CD4+ and CD8+, respectively) cells. d, Normalized RNA expression levels of PRDM1 in PD-1−TIM-3− (n = 3) and PD-1+TIM-3+ (n = 3) CD8+ TILs. Data are mean ± s.e.m. ***P = 0.0004, two-sided t-test. e, Network model based on RNA-seq gene expression data of naive CD8+ T cells from Prdm1fl/fl (WT) or Cd4crePrdm1fl/fl (PRDM1 cKO) mice stimulated in the presence of IL-27 and actual binding events (ChIP–seq data for PRDM1)19. Green arrows designate genes upregulated by PRDM1, red arrows designate genes downregulated by PRDM1, and dashed grey arrows denote binding events.
Extended Data Fig. 7 Genomic tracks surrounding the co-inhibitory molecules.
a–d, LAG-3 (a), PD-1 (b), TIGIT (c) and TIM-3 (d) with overlay of ChIP–seq data of PRDM116 and c-MAF19 and ATAC-seq data of naive CD4+ cells induced with IL-27 for 72 h and ATAC-seq data of CD8+ T cells 27 days after chronic viral infection22. Regions of binding sites common to both PRDM1 and c-MAF are indicated by the dotted rectangles. e, Luciferase activity in 293T cells transfected with pGL4.23 luciferase reporters for depicted enhancers of TIM-3 together with empty vector (control), constructs encoding PRDM1, c-MAF or both. Firefly luciferase activity was measured 48 h after transfection and is presented relative to constitutive Renilla luciferase activity. Data are mean ± s.e.m. from biologically independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, one-way ANOVA and Tukey’s multiple comparisons test.
Extended Data Fig. 8 Immune characterization of PRDM1 cKO, c-MAF cKO and PRDM1/c-MAF cDKO before and after tumour challenge.
a, Analysis of steady-state immune system in wild-type, c-MAF cKO, PRDM1 cKO and PRDM1/c-MAF cDKO. Summary data for CD4, CD8, FOXP3, CD44, CD62L and CD69 expression in spleen from wild-type, c-MAF cKO, PRDM1 cKO and PRDM1/c-MAF cDKO mice. Data are mean ± s.e.m. from biologically independent animals. *P < 0.05, **P < 0.01, ****P < 0.0001, one-way ANOVA and Tukey’s multiple comparisons test. b, Co-inhibitory receptor expression in CD4+ TILs from PRDM1/c-MAF cDKO mice. Top, representative flow cytometry data from three independent experiments for TILs from wild-type and PRDM1/c-MAF cDKO stained for PD-1, TIM-3, TIGIT, PDPN and PROCR expression. Bottom, summary data. Data are mean ± s.e.m. from biologically independent animals. *P < 0.05, two-sided t-test. c, Top, representative flow cytometry data from three independent experiments showing cytokine production from CD8+ TILs from wild-type and cDKO bearing B16F10 melanoma. Bottom, summary data. Data are mean ± s.e.m. from biologically independent animals. *P < 0.05, two-sided t-test. d, Co-inhibitory receptor expression on CD8+ TILs sorted from B16-OVA-bearing RAG1-knockout mice that were transferred with PRDM1/c-MAF cDKO (n = 4) or wild-type (n = 4) CD4+ and CD8+ T cells as indicated. Data are mean ± s.e.m. from biologically independent animals. *P < 0.05, one-way ANOVA and Tukey’s multiple comparisons test. e, RAG1-knockout mice were transferred with either wild-type or cDKO CD4+ and CD8+ (2:1 CD4:CD8 ratio), followed by subcutaneous injection of MC38-OVA. Data are mean ± s.e.m. ****P < 0.0001, repeated measures ANOVA, Sidak’s multiple comparisons test. On day 14 after tumour implantation, mice were euthanized and TILs, spleen and draining lymph nodes were obtained. f, The frequency of antigen-specific CD8+ T cells in the draining lymph nodes of mice in e.
Extended Data Fig. 9 Examination of additive and non-additive (synergistic) effects of PRDM1 and c-MAF.
a, A heat map showing all 940 differentially expressed genes between wild-type (n = 5) and cDKO (PRDM1/c-MAF, n = 4) mice, and their expression in single knockout (PRDM1 control n = 7, PRDM1 knockout n = 3, c-MAF control n = 4 and c-MAF-knockout n = 3) mice. The red markings at the top indicate genes on expression of which the two knockouts have a statistically significant (P < 0.05) non-additive effect in the cDKO (149 out of 940 differentially expressed genes). b, Volcano plot of the analysis in a for global gene expression. Genes whose expression in the two single knockouts have a statistically significant (P <0.05) non-additive effect in the cDKO (1,144 out of 12,906 genes) and had an absolute coefficient value > 1 (779 out of 1,144) are shown in orange.
Extended Data Fig. 10 Comparison of gene expression between PRDM1/c-MAF cDKO TILs and CD8+ TILs populations from wild-type mice.
a, Barcode enrichment plot displaying two gene sets in a ranked gene list. The ranked gene list was defined as fold change in gene expression between PRDM1/c-MAF cDKO and wild-type CD8+ TILs. The three gene sets consist of differentially expressed genes between: PD-1+TIM-3+CD8+ (n = 3) and PD-1−TIM-3−CD8+ (n = 3) TILs, PD-1+TIM-3+CD8+ (n = 3) TILs and memory CD8+ (n = 3), and PD-1+TIM-3−CD8+ (n = 3) and PD-1−TIM-3−CD8+ TILs. b, This analysis was followed by four statistical tests (one-sample Kolmogorov–Smirnov test, mean-rank gene set test (WilcoxGST), hypergeometric, and competitive gene set test accounting for inter-gene correlation) for enrichment of these signatures in the cDKO expression profile. c, Wild-type versus cDKO volcano plot. Green indicates genes that were upregulated in the PD-1−TIM-3−CD8+ (double negative) TILs and red indicates genes that were upregulated in the PD-1+TIM3+CD8+ (double positive) TILs. d, Wild-type versus cDKO volcano plot. Red indicates genes that were upregulated in PD-1+CXCR5+CD8+ T cells and green indicates genes that were upregulated in PD-1+CXCR5−CD8+ T cells in chronic LCMV infection23. e, A t-SNE plot of the 588 CD8+ TILs obtained from wild-type mice bearing B16F10 melanoma tumours, coloured by the relative signature score for the co-inhibitory module (272 genes, Supplementary Table 2), the cDKO signature (shown in g), and the PD-1+CXCR5+CD8+ T cell signature from chronic virus infection23. The contour plot marks the region of highly scored cells by taking into account only those cells that have a signature score above the mean.
Supplementary information
Supplementary Figure 1
Flow cytometry gating strategy
Supplementary Table 1
CyTOF panel.
Supplementary Table 2
IL-27 co-inhibitory gene module.
Supplementary Table 3
IL27/cancer custom Nanostring codeset.
Supplementary Table 4
A compiled list of cytokines, adhesion, aggregation, chemotaxis and other cell surface molecules.
Supplementary Table 5
Data for tumor sizes.
Rights and permissions
About this article
Cite this article
Chihara, N., Madi, A., Kondo, T. et al. Induction and transcriptional regulation of the co-inhibitory gene module in T cells. Nature 558, 454–459 (2018). https://doi.org/10.1038/s41586-018-0206-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0206-z
This article is cited by
-
Diffuse large B-cell lymphoma: the significance of CD8+ tumor-infiltrating lymphocytes exhaustion mediated by TIM3/Galectin-9 pathway
Journal of Translational Medicine (2024)
-
Targeting the NF-κB pathway as a potential regulator of immune checkpoints in cancer immunotherapy
Cellular and Molecular Life Sciences (2024)
-
An IL-4 signalling axis in bone marrow drives pro-tumorigenic myelopoiesis
Nature (2024)
-
Increased co-expression of PD1 and TIM3 is associated with poor prognosis and immune microenvironment heterogeneity in gallbladder cancer
Journal of Translational Medicine (2023)
-
Peripheral PD-1 and Tim-3 percentages are associated with primary sites and pathological types of peritoneal neoplasms
BMC Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.