Abstract
During our daily life, we depend on memories of past experiences to plan future behaviour. These memories are represented by the activity of specific neuronal groups or ‘engrams’1,2. Neuronal engrams are assembled during learning by synaptic modification, and engram reactivation represents the memorized experience1. Engrams of conscious memories are initially stored in the hippocampus for several days and then transferred to cortical areas2. In the dentate gyrus of the hippocampus, granule cells transform rich inputs from the entorhinal cortex into a sparse output, which is forwarded to the highly interconnected pyramidal cell network in hippocampal area CA33. This process is thought to support pattern separation4 (but see refs. 5,6). CA3 pyramidal neurons project to CA1, the hippocampal output region. Consistent with the idea of transient memory storage in the hippocampus, engrams in CA1 and CA2 do not stabilize over time7,8,9,10. Nevertheless, reactivation of engrams in the dentate gyrus can induce recall of artificial memories even after weeks2. Reconciliation of this apparent paradox will require recordings from dentate gyrus granule cells throughout learning, which has so far not been performed for more than a single day6,11,12. Here, we use chronic two-photon calcium imaging in head-fixed mice performing a multiple-day spatial memory task in a virtual environment to record neuronal activity in all major hippocampal subfields. Whereas pyramidal neurons in CA1–CA3 show precise and highly context-specific, but continuously changing, representations of the learned spatial sceneries in our behavioural paradigm, granule cells in the dentate gyrus have a spatial code that is stable over many days, with low place- or context-specificity. Our results suggest that synaptic weights along the hippocampal trisynaptic loop are constantly reassigned to support the formation of dynamic representations in downstream hippocampal areas based on a stable code provided by the dentate gyrus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ramirez, S. et al. Creating a false memory in the hippocampus. Science 341, 387–391 (2013).
Kitamura, T. et al. Engrams and circuits crucial for systems consolidation of a memory. Science 356, 73–78 (2017).
Pernía-Andrade, A. J. & Jonas, P. Theta-gamma-modulated synaptic currents in hippocampal granule cells in vivo define a mechanism for network oscillations. Neuron 81, 140–152 (2014).
Chawla, M. K. et al. Sparse, environmentally selective expression of Arc RNA in the upper blade of the rodent fascia dentata by brief spatial experience. Hippocampus 15, 579–586 (2005).
Nakashiba, T. et al. Young dentate granule cells mediate pattern separation, whereas old granule cells facilitate pattern completion. Cell 149, 188–201 (2012).
Danielson, N. B. et al. Distinct contribution of adult-born hippocampal granule cells to context encoding. Neuron 90, 101–112 (2016).
Rubin, A., Geva, N., Sheintuch, L. & Ziv, Y. Hippocampal ensemble dynamics timestamp events in long-term memory. eLife 4, e12247 (2015).
Mankin, E. A. et al. Neuronal code for extended time in the hippocampus. Proc. Natl Acad. Sci. USA 109, 19462–19467 (2012).
Kentros, C. G., Agnihotri, N. T., Streater, S., Hawkins, R. D. & Kandel, E. R. Increased attention to spatial context increases both place field stability and spatial memory. Neuron 42, 283–295 (2004).
Mankin, E. A., Diehl, G. W., Sparks, F. T., Leutgeb, S. & Leutgeb, J. K. Hippocampal CA2 activity patterns change over time to a larger extent than between spatial contexts. Neuron 85, 190–201 (2015).
GoodSmith, D. et al. Spatial representations of granule cells and mossy cells of the dentate gyrus. Neuron 93, 677–690.e5 (2017).
Senzai, Y. & Buzsáki, G. Physiological properties and behavioral correlates of hippocampal granule cells and mossy cells. Neuron 93, 691–704.e5 (2017).
Sheffield, M. E. J., Adoff, M. D. & Dombeck, D. A. Increased prevalence of calcium transients across the dendritic arbor during place field formation. Neuron 96, 490–504.e5 (2017).
Rajasethupathy, P. et al. Projections from neocortex mediate top-down control of memory retrieval. Nature 526, 653–659 (2015).
Nitz, D. & McNaughton, B. Differential modulation of CA1 and dentate gyrus interneurons during exploration of novel environments. J. Neurophysiol. 91, 863–872 (2004).
Leutgeb, J. K., Leutgeb, S., Moser, M.-B. & Moser, E. I. Pattern separation in the dentate gyrus and CA3 of the hippocampus. Science 315, 961–966 (2007).
Diamantaki, M., Frey, M., Berens, P., Preston-Ferrer, P. & Burgalossi, A. Sparse activity of identified dentate granule cells during spatial exploration. eLife 5, e20252 (2016).
Burgalossi, A., von Heimendahl, M. & Brecht, M. Deep layer neurons in the rat medial entorhinal cortex fire sparsely irrespective of spatial novelty. Front. Neural Circuits 8, 74 (2014).
Stefanelli, T., Bertollini, C., Lüscher, C., Muller, D. & Mendez, P. Hippocampal somatostatin interneurons control the size of neuronal memory ensembles. Neuron 89, 1074–1085 (2016).
Lee, S.-H. et al. Parvalbumin-positive basket cells differentiate among hippocampal pyramidal cells. Neuron 82, 1129–1144 (2014).
Cai, D. J. et al. A shared neural ensemble links distinct contextual memories encoded close in time. Nature 534, 115–118 (2016).
Driscoll, L. N., Pettit, N. L., Minderer, M., Chettih, S. N. & Harvey, C. D. Dynamic reorganization of neuronal activity patterns in parietal cortex. Cell 170, 986–999.e16 (2017).
Tsao, A. et al. Integrating time in the entorhinal cortex. Society for Neuroscience 084.21.2017 (2017).
Bittner, K. C., Milstein, A. D., Grienberger, C., Romani, S. & Magee, J. C. Behavioral time scale synaptic plasticity underlies CA1 place fields. Science 357, 1033–1036 (2017).
Peters, A. J., Chen, S. X. & Komiyama, T. Emergence of reproducible spatiotemporal activity during motor learning. Nature 510, 263–267 (2014).
Brandalise, F. & Gerber, U. Mossy fiber-evoked subthreshold responses induce timing-dependent plasticity at hippocampal CA3 recurrent synapses. Proc. Natl Acad. Sci. USA 111, 4303–4308 (2014).
Roy, D. S. et al. Memory retrieval by activating engram cells in mouse models of early Alzheimer’s disease. Nature 531, 508–512 (2016).
Bernier, B. E. et al. Dentate gyrus contributes to retrieval as well as encoding: evidence from context fear conditioning, recall, and extinction. J. Neurosci. 37, 6359–6371 (2017).
van Groen, T., Miettinen, P. & Kadish, I. The entorhinal cortex of the mouse: organization of the projection to the hippocampal formation. Hippocampus 13, 133–149 (2003).
Kheirbek, M. A. et al. Differential control of learning and anxiety along the dorsoventral axis of the dentate gyrus. Neuron 77, 955–968 (2013).
Dombeck, D. A., Harvey, C. D., Tian, L., Looger, L. L. & Tank, D. W. Functional imaging of hippocampal place cells at cellular resolution during virtual navigation. Nat. Neurosci. 13, 1433–1440 (2010).
Malvache, A., Reichinnek, S., Villette, V., Haimerl, C. & Cossart, R. Awake hippocampal reactivations project onto orthogonal neuronal assemblies. Science 353, 1280–1283 (2016).
Schmidt-Hieber, C. & Häusser, M. Cellular mechanisms of spatial navigation in the medial entorhinal cortex. Nat. Neurosci. 16, 325–331 (2013).
Aghajan, Z. M. et al. Impaired spatial selectivity and intact phase precession in two-dimensional virtual reality. Nat. Neurosci. 18, 121–128 (2015).
Sheffield, M. E. J. & Dombeck, D. A. Calcium transient prevalence across the dendritic arbour predicts place field properties. Nature 517, 200–204 (2015).
Franklin, K. B. J. & Paxinos, G. The Mouse Brain in Stereotaxic Coordinates (Elsevier, Amsterdam, 2008).
Kaifosh, P., Zaremba, J. D., Danielson, N. B. & Losonczy, A. SIMA: Python software for analysis of dynamic fluorescence imaging data. Front. Neuroinform. 8, 80 (2014).
Dombeck, D. A., Khabbaz, A. N., Collman, F., Adelman, T. L. & Tank, D. W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Hosp, J. A. et al. Morpho-physiological criteria divide dentate gyrus interneurons into classes. Hippocampus 24, 189–203 (2014).
Hainmüller, T., Krieglstein, K., Kulik, A. & Bartos, M. Joint CP-AMPA and group I mGlu receptor activation is required for synaptic plasticity in dentate gyrus fast-spiking interneurons. Proc. Natl Acad. Sci. USA 111, 13211–13216 (2014).
Bartos, M., Vida, I. & Jonas, P. Synaptic mechanisms of synchronized gamma oscillations in inhibitory interneuron networks. Nat. Rev. Neurosci. 8, 45–56 (2007).
Varga, C., Golshani, P. & Soltesz, I. Frequency-invariant temporal ordering of interneuronal discharges during hippocampal oscillations in awake mice. Proc. Natl Acad. Sci. USA 109, E2726–E2734 (2012).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Tukker, J. J. et al. Distinct dendritic arborization and in vivo firing patterns of parvalbumin-expressing basket cells in the hippocampal area CA3. J. Neurosci. 33, 6809–6825 (2013).
Arriaga, M. & Han, E. B. Dedicated hippocampal inhibitory networks for locomotion and immobility. J. Neurosci. 37, 9222–9238 (2017).
Garcia-Junco-Clemente, P. et al. An inhibitory pull-push circuit in frontal cortex. Nat. Neurosci. 20, 389–392 (2017).
Skaggs, W. E., McNaughton, B. L., Gothard, K. M. & Markus, E. J. An information-theoretic approach to deciphering the hippocampal code. In Advances in Neural Information Processing Systems (NIPS) 1030–1037 (1993).
Danielson, N. B. et al. Sublayer-specific coding dynamics during spatial navigation and learning in hippocampal area CA1. Neuron 91, 652–665 (2016).
Acknowledgements
We thank H.-J. Weber, C. Paun and C. Schmidt-Hieber for advice and help with setting up the virtual environment system; K. Winterhalter and K. Semmler for technical support; and J. Sauer, M. Strueber and M. Eyre for comments on earlier versions of the manuscript. This work was funded by the German Research Foundation (FOR2143, M.B.) and ERC-AdG 787450 (M.B.). This work was supported in part by the Excellence Initiative of the German Research Foundation (GSC-4, Spemann Graduate School; T.H.).
Reviewer information
Nature thanks M. Brecht and S. Leutgeb for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
T.H. and M.B. conceived the study, designed the experiments and wrote the manuscript. T.H. performed experiments and analysed data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Virtual environment behavioural paradigm for head-fixed mice.
Related to Fig. 1. a, Mean number of licks per spatial bin (10-cm wide) for one example mouse on day 1 (left) and day 2 (right) on the familiar (top, grey) and novel (bottom, blue) linear tracks. Blue shaded areas indicate reward zones. b, Mean lick rate per bin as a function of distance from the next reward location for the familiar (black traces) and novel (blue traces) contexts. Shaded areas denote s.e.m., n = 15 experiments. c, Mean lick rate over the entire familiar (left) or novel (right) track on day 1 (left) or day 2 (right). Grey lines denote individual experiments (n = 15) and black circles with error bars show mean ± s.e.m. d, As in c but for mean lick rate in the reward zones only. e, Reward-related licking plotted for the experiments continued over 5 days. In this subset of the data (n = 5 experiments) no significant difference in licking between contexts was observed on any day (repeated-measures one-way ANOVA), although there was a trend towards lower lick rates in the novel context. f, Lick rate increase in the reward zone (as compared to licking on the remaining track). An increase in the fraction of reward-related licks was observed only between day 1 and 2 (n = 5 experiments). g, Screenshots of the familiar context and the four different novel context sceneries. One novel context was randomly selected for each experiment from this set and maintained through all days. If more than one experiment was performed using a given animal, a different novel context was chosen for each of the experiments. h, Intact spatial memory was probed in the experimental mice and GFP-injected controls in a Barnes maze learning paradigm (see Methods). Blue traces show a mouse’s trajectory in the last of four sessions on the first (top) and third (bottom) days of the experiment. i, Mean path length per session for implanted mice used for imaging experiments (n = 8 mice; green) and GFP-injected controls (n = 8 mice; grey) on days 1–3 (repeated-measures one-way ANOVA). There was no significant difference in path length between groups (day 1: P = 0.96, day 2: P = 0.806, day 3: P = 0.915, two-sided t-test). c, d, f, Two-sided paired t-test. *P < 0.05, **P < 0.01, ***P < 0.001, n.s. not significant. Error bars denote s.e.m. throughout. For exact P values see Supplementary Table 1.
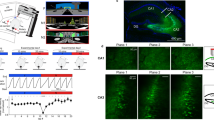
Extended Data Fig. 2 Imaging configuration and CA3 imaging locations.
a, Schematic of the transcortical window implantation. A stainless steel cannula (3 mm diameter) with a circular coverslip attached to the bottom is implanted into the brain and rests on the external capsule on top of the hippocampus (see Methods). b, Illustrative fluorescence time series of an active GC, including the time-averaged GCaMP fluorescence (AVG, left). c, Photograph of a mouse in the virtual environment setup. Depicted is a mouse in the CA2/3 recording group with tilted objective for lateral access view (see Methods). d, Anatomical drawing of the dorsal hippocampus indicating the imaging planes for CA3 recordings. Depicted is the location of the middle plane from a three-plane imaging volume of 40–80 µm. Coloured numbers indicate individual mice from which the data are derived; in cases for which more than one experiment was performed in a mouse, letters denote the locations of the respective experiments. e, Box plot of place field stability values over days from familiar-context place cells in the individual recordings shown in d. White numbers denote the number of place cells from each experiment that are represented by the boxes. Boxes, 25th to 75th percentiles; white bars, median; whiskers, 99% range.
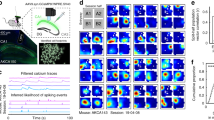
Extended Data Fig. 3 Properties of place coding in the familiar and novel virtual contexts.
Related to Fig. 2. a, Cumulative distribution of mean calcium activity, based on the area under the calcium curve, for all GCs (red), CA2/3 PYRs (blue) and CA1 PYRs (black) recorded in the familiar context. b, As in a, but for the calcium transient rate. c, Cumulative distribution of place field widths for all day 1 familiar-context place cells in the respective hippocampal areas. d, Experimental schematic: place field consistency was measured as the correlation of the average activity on the first and second blocks of five runs in the same context (familiar: F–F′ or novel: N–N′). Place field discrimination was assessed as the correlation of activity maps for different contexts (F–N). e, Activity over distance for cells with a place field on the novel track. Cells were sorted according to their peak activity in novel-track runs and are plotted separately for the first (left) and second (middle) blocks of runs on the novel track and for all runs on the familiar track (right). For the same plots with familiar-context place cells, see Fig. 2c. f, Number of cells with significant place fields (see Methods) in each experiment. Thin lines denote individual experiments (DG, n = 12; CA2/3, n = 7; CA1, n = 15 experiments), thick lines with error bars show mean ± s.e.m. (one-way repeated measures ANOVA). g, Median activity-map correlations between the familiar and novel contexts for experiments with at least 20 place cells on day 1 (circles; DG, n = 6; CA2/3, n = 4; CA1, n = 11 experiments) and their mean ± s.e.m. (ANOVA, Holm–Sidak). h, Pearson’s correlation of place-related activity for cells that had a place field in the familiar context (left) or novel context (right) on day 1. Left box of each pair shows correlation of activity maps for trial blocks on the same track (‘consistency’) and right bar shows correlations with trials on the other track (‘discrimination’) in the same session. Correlations within the same context were always significantly higher than between contexts, except for GCs that had newly acquired a place field on the novel track. i, Trial-to-trial reliability of place cell responses. Place-related firing was more reliable in the familiar than in the novel context and higher in GCs than in PYRs in the familiar context. Differences between areas were not significant for the novel context. j, Each figure shows the run-by-run calcium activity (colour coded) over distance from one individual place cell for one session in the familiar context. Rows denote individual runs. h, i, Boxes, 25th to 75th percentiles; white bars, median; whiskers, 99% range. a–c, h, i, One-way ANOVA on ranks, Dunn’s test. *P < 0.05, **P < 0.01, ***P < 0.001; n.s., not significant. For exact P values see Supplementary Table 1.
Extended Data Fig. 4 Task-related activity of PV-expressing interneurons in CA1 and DG.
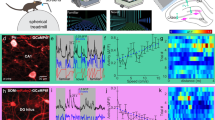
a, Illustrative, time-averaged fluorescence image of GCaMP6f (pseudocolour white) and tdT (red) expressed in PVIs. Insets below show each fluorescence channel separately for the area indicated with the dotted line. b, Calcium trace of a representative PVI in CA1 (black) and distance on the virtual linear track (blue) over time. This PVI is active particularly at times when the animal moves fast. c, Calcium activity (colour coded) of the representative cell shown in b as a function of linear track distance for multiple runs (rows). Graph at the bottom shows the average of activity over distance for this cell in the familiar context. Shaded areas denote s.d. The same analysis was performed for 78 CA1 and 19 DG PVIs. d, As in c, but activity was plotted as a function of running speed. e, f, Mean activity maps over distance for all recorded CA1 (e) or DG (f) PVIs in the familiar and novel contexts. g, h, Same as in e, f, but mean activity was plotted over running speed. Activity in most DG PVIs is suppressed in the novel context. i, Activity of PVIs in the DG on the familiar (red) and novel track (light red) over running speed. Shaded areas denote s.d. R denotes Pearson’s R. j, Mean calcium activity during familiar-track running plotted against novel-track activity for PVIs in the DG. Inset, bars denote mean ± s.e.m. of running-related activity for novel and familiar tracks (two-sided signed rank-sum test). k, l, As in i, j but for CA1 PVIs. ***P < 0.001; n.s., not significant. For exact P values see Supplementary Table 1.
Extended Data Fig. 5 Differential stability and context discrimination of CA1 and DG are not due to interindividual differences.
a, Experimental schematic. Left, fluorescence image of GCaMP6f (white) and tdT in PVIs (red) in a post mortem coronal brain section. Dotted line indicates position of the imaging window. In this mouse, recordings were made from CA1 PYRs and DG GCs in separate, sequential experiments (exemplary image planes are shown in middle images). For each experiment, the same familiar and a different novel context (right) were used. Thereby, the coding properties of PYRs and GCs in the same animal could be compared. b, Calcium activity over distance for CA1 PYRs (top) and DG GCs (bottom) with place fields on the familiar track sorted for their peak activity on day 1 in the familiar context. Activity of the same cells with the same sorting on day 2 in the familiar context (middle) and on day 1 in the novel context (right). c, Mean cellular activity map correlations (colour coded: Pearson’s R) over two days and contexts as indicated on the x-axis. Data sampled only for place cells recorded in the selected mouse. Each row shows mean correlation values for cells that had a place field on the day and track indicated on the y-axis (n denoted as white numbers). d, Left, correlations of activity maps in the familiar context between days plotted for cells that had a place field in the familiar context. Cells were sampled only from measurements in this particular mouse. Right, activity map correlations between the familiar and novel contexts on day 1 for all cells that had a place field in the familiar context on that day. Stability over time and activity map similarity between contexts are significantly higher for GCs than for CA1 PYRs in the same mouse. e–g, As in b–d, but calculated separately on cells from another mouse. h, Activity map correlations between days 1 and 2 were calculated for familiar-context place cells and medians (circles) are displayed for each animal that had a minimum of 20 such place cells (DG, n = 4; CA2/3, n = 4; CA1, n = 9 mice). The means ± s.e.m. of these per-animal medians (bars) were compared statistically. i, As in h, but for activity map correlations of familiar-context place cells on day 1 between the familiar and novel contexts (DG, n = 3; CA2/3, n = 3; CA1, n = 9 mice). Higher temporal stability and higher inter-context similarity are a feature of GCs that is consistently observed in different mice. h, i, One-way ANOVA with Holm–Sidak test. Error bars denote s.e.m. d, g, Boxes, 25th to 75th percentiles; white bars, median; whiskers, 99% range. Two-sided rank-sum test. *P < 0.05, **P < 0.01, ***P < 0.001; n.s., not significant. For exact P values see Supplementary Table 1.
Extended Data Fig. 6 Spatial firing of DG GCs requires external reference cues.
a, Screenshot from the simplified virtual linear track devoid of any visual reference cues, except for patterned walls to provide a visual percept of self-motion. b, Cumulative distribution of activity levels for all GCs imaged in the standard familiar and novel contexts, as well as the simplified version shown in a. c, Cumulative distribution of spatial information values in all cells with a minimum activity of 0.01 transients per second in the familiar, novel and self-motion-only based paradigms. **P < 0.01, ***P < 0.001; n.s., not significant (one-way ANOVA on ranks with Dunn’s test). For exact P values see Supplementary Table 1.
Extended Data Fig. 7 Place field remapping between days.
Related to Fig. 3. a, Mean cellular activity map correlations (Pearson’s R) over two days and contexts as indicated on the x-axis. Each row shows mean correlation values for cells (white numbers denote n) that had a place field on the day and track indicated on the y-axis. b, Median day-to-day correlation (‘stability’) of familiar-context place cell activity for all experiments in which at least 20 cells had a place field in the familiar context on either day (circles; DG, n = 6; CA2/3, n = 6; CA1, n = 12 experiments). Bars denote mean ± s.e.m. of these per-experiment values (one-way ANOVA with Holm-Sidak test). c, For all cells that had a place field in the familiar context on both days of the experiment, the centres of mass for the activity in these place fields was determined. The graph shows the cumulative distribution of the distances between these centres (shift) between days 1 and 2, which gives a measure of the relocation of place fields between days. d, Trial-by-trial correlation of place cell activity (reliability) in the familiar and novel contexts on days 1 and 2. N = 354, 367, 189, 226, 242, 364, 252, 285, 1,009, 1,044, 660 and 1,074 place cells per group (left to right). Place cell reliability for the novel context place cells increases in CA1 and CA2/3 between the first and second days. Boxes, 25th to 75th percentiles; white bars, median; whiskers, 99% range. c, d, One-way ANOVA on ranks with Dunn’s test. *P < 0.05, **P < 0.01, ***P < 0.001; n.s., not significant. For exact P values see Supplementary Table 1.
Extended Data Fig. 8 CA1 place coding is degraded in the absence of visual references, but does not scale with environmental complexity.
a, Screenshots of the three different virtual linear tracks. Left, the ‘poor’ track had patterned walls, but no other cues; middle, the ‘normal’ track with visual references; right, the ‘rich’ multisensory environment with many visual objects, sound, odour and tactile cues (Supplementary Video 6). b, Calcium activity over distance for CA1 PYRs with place fields on the tracks depicted above, sorted for their peak activity on day 1 (right) and, with the same sorting, on day 2 (left). Higher activity levels and day-to-day stability can be observed in the ‘normal’ and ‘rich’ environments. c, Number of cells with significant place fields (see Methods) on the first recording day per experiment. Thin lines denote individual experiments (n = 7), thick lines with error bars the means ± s.e.m. (repeated measures one-way ANOVA). d, Calcium activity levels (AUC) of the place cells detected in the three settings. e, As in d, but for spatial information. f, Activity map correlations between days for all cells that had a place field on the corresponding track. d–f, One-way ANOVA on ranks with Dunn’s test. Boxes, 25th to 75th percentiles; white bars, median; whiskers, 99% range. ***P < 0.001; n.s., not significant. For exact P values see Supplementary Table 1.
Extended Data Fig. 9 Superficial and deep layer CA1 pyramidal cells differ in their task-related coding properties.
a, Experimental schematic. Cells with somata close to the border of the stratum oriens (deep CA1 PYRs) and those close to the border of the stratum radiatum (superficial CA1 PYRs) were identified in different z-planes and separated for analysis. Illustrative pictures to the right show time-averaged fluorescence of GCaMP6f (white) and td-Tomato (red) in PVIs. b, Calcium activity over distance for deep CA1 PYRs with a place field on the familiar track. Cells were sorted according to their peak activity on the familiar track (left) and are plotted in the same order for runs on day 1 on the novel track (middle) and for the runs on the familiar track on the second day (right). c, As in b, but for novel-context place cells on the familiar context (middle) and novel context on the second day (right). d, e, As in b, c, but for superficial PYRs. f, Calcium activity rate for superficial and deep CA1 PYRs in the familiar and novel contexts. Activity rates increased significantly in the novel context in superficial but not in deep PYRs (two-sided signed rank-sum test). g, Trial-to-trial correlations of place cell activity compared between layers for familiar-context (left) and novel-context (right) runs. h, Activity map correlations between contexts for cells with a place field in the familiar context on day 1. i, Cumulative distribution of place cell activity map correlations between days 1 and 2 during familiar context runs. g–i, Two-sided rank-sum test. *P < 0.05, **P < 0.01; n.s., not significant. Boxes, 25th to 75th percentiles; white bars, median; whiskers, 99% range. For exact P values see Supplementary Table 1.
Extended Data Fig. 10 Differential stability of hippocampal place fields over extended time spans.
Related to Fig. 4. a, The same place cells were imaged over multiple days. Pictures show an illustrative example of the time-averaged fluorescence of GCaMP6f (pseudocolour white) expressed pan-neuronally and tdT (red) expressed in PVIs for the same field of view in CA1 on five subsequent days. b, Activity maps for cells that had a place field on the novel track on any of the five days. Cells are sorted by their activity peaks on day three (grey shading). c, Activity map correlations as function of days passed for cells with place field on the familiar (dark colours) or novel track (light colours). Dotted lines show corresponding levels of random correlations generated by shuffling cell IDs. Dark and light coloured asterisks underneath the dotted lines indicate significant differences of the actual versus random correlations for familiar- and novel-context place cells, respectively. Black asterisks between traces indicate significant differences between the mean correlation values of novel- and familiar-context place cells. *P < 0.05, **P < 0.01, ***P < 0.001. Two-sided rank-sum test for each time-difference (days) with Bonferroni correction. Error bars denote s.e.m. For exact P values and N numbers in c see Supplementary Table 1.
Supplementary information
Supplementary Table 1
Tabulated summary of all statistics throughout the manuscript.
Video 1 A head fixed mouse performing goal-oriented learning in the virtual environment.
The mouse runs on a spherical treadmill, rotation of the treadmill is translated into movement through the virtual world.
Video 2 Fluorescence stack through the hippocampus of an anaesthetized mouse.
GCaMP6f (white) is expressed in all neurons, tdTomato (red) only in parvalbumin expressing interneurons. The stack starts in stratum oriens of CA1 and focusses down into the hippocampus until the lower blade of the DG granule cell layer is reached.
Video 3 In vivo acquired, motion corrected video of calcium signals from GCs.
Depicted is one of three simultaneously acquired imaging planes acquired at a z-distance of ~25 µm and all cutting through the granule cell layer at different depths. Similar video data were obtained in 12 independent experiments from 6 mice. Supplementary videos 3-5 were convolved with a Gaussian kernel in the time domain for displaying purposes.
Video 4 In vivo acquired, motion corrected video of calcium signals from pyramidal cells in CA3.
Recorded in the CA3 pyramidal cell layer. Similar video data were obtained in 7 independent experiments from 5 mice. Statements of supplementary video 3 apply, accordingly.
Video 5 In vivo acquired, motion corrected video of calcium signals from pyramidal cells in CA1.
Recorded in the CA1 pyramidal cell layer. Similar video data were obtained in 15 independent experiments from 11 mice. Statements of supplementary video 3 apply, accordingly.
Video 6 Mouse exploring the rich multisensory virtual paradigm.
To the left of the mouse, a carousel with tangible objects is installed on a stepper motor and moves according to the mouse’s position on the virtual track. A high-pitched tone is displayed repeatedly as an auditory contextual cue.
Rights and permissions
About this article
Cite this article
Hainmueller, T., Bartos, M. Parallel emergence of stable and dynamic memory engrams in the hippocampus. Nature 558, 292–296 (2018). https://doi.org/10.1038/s41586-018-0191-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0191-2
This article is cited by
-
Subfield-specific interneuron circuits govern the hippocampal response to novelty in male mice
Nature Communications (2024)
-
A persistent prefrontal reference frame across time and task rules
Nature Communications (2024)
-
Dentate gyrus is needed for memory retrieval
Molecular Psychiatry (2024)
-
A consistent map in the medial entorhinal cortex supports spatial memory
Nature Communications (2024)
-
Hippocampal subfield plasticity is associated with improved spatial memory
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.