Abstract
Hyperphosphorylation of the C-terminal domain (CTD) of the RPB1 subunit of human RNA polymerase (Pol) II is essential for transcriptional elongation and mRNA processing1,2,3. The CTD contains 52 heptapeptide repeats of the consensus sequence YSPTSPS. The highly repetitive nature and abundant possible phosphorylation sites of the CTD exert special constraints on the kinases that catalyse its hyperphosphorylation. Positive transcription elongation factor b (P-TEFb)—which consists of CDK9 and cyclin T1—is known to hyperphosphorylate the CTD and negative elongation factors to stimulate Pol II elongation1,4,5. The sequence determinant on P-TEFb that facilitates this action is currently unknown. Here we identify a histidine-rich domain in cyclin T1 that promotes the hyperphosphorylation of the CTD and stimulation of transcription by CDK9. The histidine-rich domain markedly enhances the binding of P-TEFb to the CTD and functional engagement with target genes in cells. In addition to cyclin T1, at least one other kinase—DYRK1A6—also uses a histidine-rich domain to target and hyperphosphorylate the CTD. As a low-complexity domain, the histidine-rich domain also promotes the formation of phase-separated liquid droplets in vitro, and the localization of P-TEFb to nuclear speckles that display dynamic liquid properties and are sensitive to the disruption of weak hydrophobic interactions. The CTD—which in isolation does not phase separate, despite being a low-complexity domain—is trapped within the cyclin T1 droplets, and this process is enhanced upon pre-phosphorylation by CDK7 of transcription initiation factor TFIIH1,2,3. By using multivalent interactions to create a phase-separated functional compartment, the histidine-rich domain in kinases targets the CTD into this environment to ensure hyperphosphorylation and efficient elongation of Pol II.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Harlen, K. M. & Churchman, L. S. The code and beyond: transcription regulation by the RNA polymerase II carboxy-terminal domain. Nat. Rev. Mol. Cell Biol. 18, 263–273 (2017).
Eick, D. & Geyer, M. The RNA polymerase II carboxy-terminal domain (CTD) code. Chem. Rev. 113, 8456–8490 (2013).
Zaborowska, J., Egloff, S. & Murphy, S. The pol II CTD: new twists in the tail. Nat. Struct. Mol. Biol. 23, 771–777 (2016).
Zhou, Q., Li, T. & Price, D. H. RNA polymerase II elongation control. Annu. Rev. Biochem. 81, 119–143 (2012).
Kwak, H. & Lis, J. T. Control of transcriptional elongation. Annu. Rev. Genet. 47, 483–508 (2013).
Di Vona, C. et al. Chromatin-wide profiling of DYRK1A reveals a role as a gene-specific RNA polymerase II CTD kinase. Mol. Cell 57, 506–520 (2015).
Taube, R., Lin, X., Irwin, D., Fujinaga, K. & Peterlin, B. M. Interaction between P-TEFb and the C-terminal domain of RNA polymerase II activates transcriptional elongation from sites upstream or downstream of target genes. Mol. Cell. Biol. 22, 321–331 (2002).
Darzacq, X. et al. In vivo dynamics of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 14, 796–806 (2007).
Hansen, A. S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq, X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. eLife 6, e25776 (2017).
Hansen, A. S. et al. Robust model-based analysis of single-particle tracking experiments with Spot-On. eLife 7, e33125 (2018).
Xiang, J. et al. DYRK1A regulates Hap1–Dcaf7/WDR68 binding with implication for delayed growth in Down syndrome. Proc. Natl Acad. Sci. USA 114, E1224–E1233 (2017).
Dosztányi, Z., Csizmok, V., Tompa, P. & Simon, I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21, 3433–3434 (2005).
Mitrea, D. M. & Kriwacki, R. W. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal. 14, 1 (2016).
Courchaine, E. M., Lu, A. & Neugebauer, K. M. Droplet organelles? EMBO J. 35, 1603–1612 (2016).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Lin, Y., Protter, D. S., Rosen, M. K. & Parker, R. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Ying, Y. et al. Splicing activation by Rbfox requires self-aggregation through its tyrosine-rich domain. Cell 170, 312–323.e310 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Herrmann, C. H. & Mancini, M. A. The Cdk9 and cyclin T subunits of TAK/P-TEFb localize to splicing factor-rich nuclear speckle regions. J. Cell Sci. 114, 1491–1503 (2001).
Marcello, A. et al. Recruitment of human cyclin T1 to nuclear bodies through direct interaction with the PML protein. EMBO J. 22, 2156–2166 (2003).
Salichs, E., Ledda, A., Mularoni, L., Albà, M. M. & de la Luna, S. Genome-wide analysis of histidine repeats reveals their role in the localization of human proteins to the nuclear speckles compartment. PLoS Genet. 5, e1000397 (2009).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
Mortillaro, M. J. et al. A hyperphosphorylated form of the large subunit of RNA polymerase II is associated with splicing complexes and the nuclear matrix. Proc. Natl Acad. Sci. USA 93, 8253–8257 (1996).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Acknowledgements
We thank S. McKnight, M. Geyer, J. Hurley and their colleagues for providing the various expression plasmids, and U. Schulze-Gahmen for technical help. This work was supported by the National Institutes of Health grant R01AI041757 to Q.Z. and the California Institute of Regenerative Medicine grant LA1-08013 to X.D.
Reviewer information
Nature thanks J. Lis, D. Taatjes and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
H.L., X.D. and Q.Z. conceived the studies. H.L., D.Y. and R.L. performed kinase reaction assays, cell culture, immunofluorescence staining and droplet formation experiments and analyses. A.S.H. performed and analysed the single-particle tracking experiment. S.G. performed and analysed the FRAP assay. H.L. and A.H. performed and analysed the time-lapse phase-contrast imaging experiment. H.L., A.S.H. and Q.Z. wrote the manuscript, and all authors contributed ideas and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
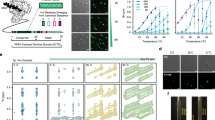
Extended Data Fig. 1 The CYCT1 HRD-dependent hyperphosphorylation of Pol II CTD52 by affinity-purified CDK9–CYCT1–Flag dimer can also be detected with anti-phospho-Ser2 antibody.
a, Examination of the affinity-purified CDK9–CYCT1–Flag dimer containing wild-type CYCT1–Flag by SDS–PAGE and silver staining. The anti-Flag affinity-purification was performed under high salt plus detergent (1 M KCl + 1% NP-40) conditions to strip away all the P-TEFb-associated factors but keep the CDK9–CYCT1 interaction intact. b, Affinity-purified CDK9–CYCT1–Flag heterodimers containing the indicated CYCT1–Flag proteins were tested in kinase reactions containing a mixture of GST-fused CTD52 and CTD9 as the substrates. The phosphorylated p-CTD52 and p-CTD9 were detected by western blotting with the anti-phospho-Ser2 antibody 3E10. Although a very similar pattern of CTD phosphorylation was detected with both anti-phospho-Ser2 and anti-phospho-Ser5 antibodies, the unphosphorylated CTD present in the ATP(–) lanes was only detected by the former antibody, making the phospho-Ser5 antibody a preferred choice for detecting CTD phosphorylation in these kinase reactions.
Extended Data Fig. 2 The CYCT1 HRD is required for efficient transcription of human immediate early genes as well as the HSP70-1 gene under both basal and heat-shock conditions.
a, Confirmation by western blotting of the knockdown of endogenous CYCT1 expression in HEK293T cells expressing the CYCT1 (also known as CCNT1)-specific shRNA (shCYCT1). b, Anti-Flag western blotting analysis of the expression of either wild-type CYCT1–Flag or CYCT1ΔHRD–Flag from the shCYCT1-resistant plasmid introduced into the knockdown cells. The α-tubulin and CDK9 levels were used as controls. c, The HRD-dependent transcription of four cellular immediate early genes. The mRNA levels of the indicated immediate early genes in the knockdown cells expressing wild-type CYCT1 or CYCT1ΔHRD–Flag (analysed in b) were examined by qRT–PCR, normalized to that of GAPDH and shown. The activity in the first column of each group was set to 1. Data are mean ± s.d., n = 3, and P values from two-tailed Student’s t-test. d, The CYCT1 HRD is required for optimal HSP70-1 transcription under both basal and heat-shock conditions. The HSP70 mRNA levels in the knockdown cells expressing wild-type CYCT1 or CYCT1ΔHRD were examined under heat-shock or non-heat-shock conditions, as in c. e, HeLa cells were transfected in twofold increments with the plasmid expressing either wild-type CYCT1–Flag or CYCT1ΔHRD–Flag. Anti-Flag immunoprecipitates (IP) from nuclear extracts were examined by western blotting for the proteins labelled on the left. The levels of the co-precipitated MEPCE, AFF4 and AFF1 were first quantified and then normalized to that of their corresponding CYCT1–Flag bait. The levels of MEPCE, AFF4 and AFF1 bound to low, middle and high levels of CYCT1ΔHRD–Flag were then divided by those of MEPCE, AFF4 and AFF1 bound to the corresponding levels of wild-type CYCT1–Flag and shown in lanes 5–7, with the numbers in lanes 2–4 set to 1 as a reference.
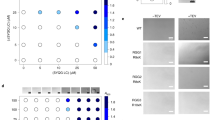
Extended Data Fig. 3 The HRD promotes the binding of CYCT1 to activated gene expression array, endogenous genes and the RNA Pol II CTD.
a, Estimates of parameters extracted from fitting of FRAP data (top) and parameters calculated from the fitted FRAP data (bottom). b, Model fit overlaid on raw FRAP data. A full description of how the reaction–diffusion FRAP model was fitted to the data is provided in Supplementary Information. c, Anti-CYCT1 western blotting analysis of the levels of endogenous CYCT1, and the stably expressed Halo–CYCT1 and Halo–CYCT1ΔHRD, in lysates of the engineered U2OS cell lines. The α-tubulin levels provided an internal control. d, Histograms of displacements at the indicated Δτ with three-state model fit overlaid. The three-state model is described in Supplementary Information. e, Table of best-fit parameters from fitting Spot-On to the raw displacements for three independent replicates (about 5–10 cells per replicate). Values in the table are mean ± s.d., n = 3.
Extended Data Fig. 4 Examination of recombinant P-TEFb and CAK for their phosphorylation of the Pol II CTD in kinase reactions and contribution of the DYRK1A HRD to DYRK1A–Pol II interaction.
a, b, Compared to CDK9 in recombinant P-TEFb, CDK7 in recombinant CAK (CDK7–CYCH–MAT1) shows decreased ability to hyperphosphorylate CTD52 in a time- and dosage-dependent manner. The indicated amounts of baculovirus-produced recombinant P-TEFb or CAK (Millipore) were added to in vitro kinase reactions that also contained GST–CTD52 as the substrate. The reactions were performed for the indicated periods of time. The products were analysed by western blotting with the phospho-Ser5 antibody. CDK7 in recombinant CAK and CDK9 in recombinant P-TEFb were also examined by western blotting. c, The deletion of the HRD causes DYRK1A to decrease interaction with RNA Pol II but not DCAF7. Nuclear extract (NE) of HeLa cells expressing the indicated proteins and anti-Flag immunoprecipitates derived from NE were analysed by western blotting with the various antibodies as labelled.
Extended Data Fig. 5 GFP–T1-IDR purified from recombinant E. coli forms phase-separated liquid droplets that are HRD-dependent and highly sensitive to elevated salt concentration and exposure to 1,6-hexanediol.
a, Alignment of CYCT1 amino acid sequences in the HRD regions among the indicated vertebrate species, with the central histidine cluster shaded in red. b, Intrinsic disorder tendency was predicted by IUPred across the entire length of CYCT1. The scores are assigned between 0 and 1, and a score above 0.5 indicates disorder. The HRD region is shaded in blue. The longest stretch of IDR is labelled as T1-IDR, and its boundaries are marked. c, d, The C-terminally Strep-tagged GFP–T1-IDR fusions (c) and N-terminally His-tagged mCherry–CTD fusion (d) were purified from recombinant E. coli BL21 cells (Supplementary Information). Ten micrograms of each of the fusion proteins was examined by SDS–PAGE followed by Coomassie blue staining. e, Protein solutions containing either wild-type GFP–T1-IDR or GFP–T1-IDRΔHRD at 6 mg ml−1 were adjusted to the indicated salt concentrations and their appearances in Eppendorf tubes are shown. f, NaCl concentrations in the wild-type GFP–T1-IDR solution were changed in the indicated sequential order and then examined under a fluorescence microscope. g, Protein solutions containing wild-type GFP–T1-IDR at 6 mg ml−1 were adjusted to 37.5 mM NaCl with or without 10% 1,6-hexanediol, and then examined under a fluorescence microscope. h, Live-cell images of a HeLa cell expressing either wild-type eGFP–CYCT1 or eGFP–CYCT1ΔHRD at levels similar to that of endogenous CYCT1.
Extended Data Fig. 6 The longest stretch of IDR in DYRK1A promotes formation of phase-separated droplets in an HRD-dependent manner.
a, Intrinsic disorder tendency was predicted by IUPred across the entire length of DYRK1A. The scores are assigned between 0 and 1, and a score above 0.5 indicates disorder. The HRD region is shaded in blue. The longest stretch of IDR is labelled as DYRK1A-IDR and its boundaries are marked. b, The indicated C-terminally Strep-tagged GFP fusions were purified from recombinant E. coli BL21 cells. Two micrograms each of the fusion proteins was examined by SDS–PAGE followed by Coomassie blue staining. c, Solutions containing 5 mg ml−1 of the indicated fusion proteins, 37.5 mM NaCl and 10% PEG8000 were trapped between coverslips and examined with a microscope under either fluorescent (488 nm) or normal white light (brightfield).
Extended Data Fig. 7 The central histidine cluster within the CYCT1 HRD is essential for promotion of phase separation by CYCT1 IDR in vitro and in cells, and for P-TEFb to hyperphosphorylate the Pol II CTD and activate HIV transcription.
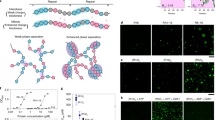
a, The nine histidines in the central histidine cluster within the CYCT1 HRD are highlighted in red and changed to alanines in the 9A mutant. b, The C-terminally Strep-tagged GFP–T1-IDR fusions containing either wild-type GFP–T1-IDR or the GFP–T1-IDR(9A) mutant sequence were purified from E. coli BL21 cells and examined by SDS–PAGE followed by Coomassie blue staining. c, CDK9–CYCT1–Flag heterodimers containing the indicated CYCT1–Flag proteins were affinity-purified from HeLa cells and tested in kinase reactions, with the reaction products analysed as in Fig. 1a. d, Plasmids expressing the Gal4 DNA binding domain fused to the indicated CYCT1 proteins were co-transfected into HeLa cell with a HIV-1 LTR–luciferase reporter construct containing the UAS for Gal4. Luciferase activities in cell extracts were measured and analysed as in Fig. 1b. e, Fixed and permeabilized HeLa cells expressing wild-type CYCT1–Flag or CYCT1(9A)–Flag were examined by indirect immunofluorescence with the mouse anti-Flag monoclonal antibody and Alexa Fluor 488-conjugated goat anti-mouse secondary antibody. DNA was counterstained using DAPI.
Extended Data Fig. 8 CYCT1 binds directly to the Pol II CTD in an HRD-dependent manner and the binding is enhanced after the CTD is phosphorylated by CAK (CDK7–CYCH–MAT1).
a, Immobilized GST–CTD was incubated with recombinant GFP–T1-IDR or GFP–T1-IDRΔHRD. The input (2.5%) and the bound proteins were analysed by western blotting. b, Binding reactions containing the purified recombinant fusion proteins indicated on the right were analysed in a 7.5 to 20% glycerol gradient containing 500 mM NaCl plus 0.5% NP-40, which was centrifuged at 55,000 r.p.m. and 4 °C for 13 h. The indicated fractions were analysed by western blotting to detect the distributions of proteins marked on the left. The entire length of the CTD could be bound by varying numbers of IDRs, resulting in the formation of a series of complexes with broad distributions in the gradient. c, mCherry–CTD was incubated with or without immobilized CAK for 6 h in kinase reactions and then analysed by SDS–PAGE and Coomassie blue staining. d, Immobilized GST–CTD was incubated with (phos.) or without (unphos.) CAK for 6 h in kinase reactions. After washing, the GST–CTD beads were incubated with GFP–T1-IDR. The indicated proteins were eluted off the beads and analysed by western blotting.
Extended Data Fig. 9 A model depicting how P-TEFb uses the CYCT1 HRD to target and recruit the Pol II CTD into a phase-separated compartment—which is formed by weak, multivalent homotypic interactions among the HRDs—to enable highly efficient phosphorylation of CTD by P-TEFb in the presence of ATP.
At 2.5% 1,6-hexanediol, the HRD-mediated phase separation but not the direct HRD–CTD binding is disrupted, making it impossible for P-TEFb to hyperphosphorylate the CTD.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1, the uncropped gel source data with size marker indications, Supplementary Methods and Supplementary References.
Supplementary Video 1
Representative video for spaSPT data.
Supplementary Video 2
Representative video for slowSPT data.
Rights and permissions
About this article
Cite this article
Lu, H., Yu, D., Hansen, A.S. et al. Phase-separation mechanism for C-terminal hyperphosphorylation of RNA polymerase II. Nature 558, 318–323 (2018). https://doi.org/10.1038/s41586-018-0174-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0174-3
This article is cited by
-
RBM22 regulates RNA polymerase II 5′ pausing, elongation rate, and termination by coordinating 7SK-P-TEFb complex and SPT5
Genome Biology (2024)
-
Crosstalk between protein post-translational modifications and phase separation
Cell Communication and Signaling (2024)
-
Tissue-specific RNA Polymerase II promoter-proximal pause release and burst kinetics in a Drosophila embryonic patterning network
Genome Biology (2024)
-
Enhancer selectivity in space and time: from enhancer–promoter interactions to promoter activation
Nature Reviews Molecular Cell Biology (2024)
-
A chaperone-like function of FUS ensures TAZ condensate dynamics and transcriptional activation
Nature Cell Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.