Abstract
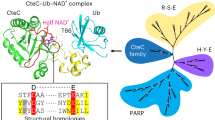
Protein ubiquitination is a multifaceted post-translational modification that controls almost every process in eukaryotic cells. Recently, the Legionella effector SdeA was reported to mediate a unique phosphoribosyl-linked ubiquitination through successive modifications of the Arg42 of ubiquitin (Ub) by its mono-ADP-ribosyltransferase (mART) and phosphodiesterase (PDE) domains. However, the mechanisms of SdeA-mediated Ub modification and phosphoribosyl-linked ubiquitination remain unknown. Here we report the structures of SdeA in its ligand-free, Ub-bound and Ub–NADH-bound states. The structures reveal that the mART and PDE domains of SdeA form a catalytic domain over its C-terminal region. Upon Ub binding, the canonical ADP-ribosyltransferase toxin turn-turn (ARTT) and phosphate-nicotinamide (PN) loops in the mART domain of SdeA undergo marked conformational changes. The Ub Arg72 might act as a ‘probe’ that interacts with the mART domain first, and then movements may occur in the side chains of Arg72 and Arg42 during the ADP-ribosylation of Ub. Our study reveals the mechanism of SdeA-mediated Ub modification and provides a framework for further investigations into the phosphoribosyl-linked ubiquitination process.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Yau, R. & Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 18, 579–586 (2016).
Hershko, A., Ciechanover, A. & Varshavsky, A. The ubiquitin system. Nat. Med. 6, 1073–1081 (2000).
Hubber, A., Kubori, T. & Nagai, H. Modulation of the ubiquitination machinery by Legionella. Curr. Top. Microbiol. Immunol. 376, 227–247 (2013).
Jeong, K. C., Sexton, J. A. & Vogel, J. P. Spatiotemporal regulation of a Legionella pneumophila T4SS substrate by the metaeffector SidJ. PLoS Pathog. 11, e1004695 (2015).
Bardill, J. P., Miller, J. L. & Vogel, J. P. IcmS-dependent translocation of SdeA into macrophages by the Legionella pneumophila type IV secretion system. Mol. Microbiol. 56, 90–103 (2005).
Horwitz, M. A. Formation of a novel phagosome by the Legionnaires’ disease bacterium (Legionella pneumophila) in human monocytes. J. Exp. Med. 158, 1319–1331 (1983).
Swanson, M. S. & Isberg, R. R. Association of Legionella pneumophila with the macrophage endoplasmic reticulum. Infect. Immun. 63, 3609–3620 (1995).
Kagan, J. C. & Roy, C. R. Legionella phagosomes intercept vesicular traffic from endoplasmic reticulum exit sites. Nat. Cell Biol. 4, 945–954 (2002).
Luo, Z. Q. & Isberg, R. R. Multiple substrates of the Legionella pneumophila Dot/Icm system identified by interbacterial protein transfer. Proc. Natl Acad. Sci. USA 101, 841–846 (2004).
Zhu, W. et al. Comprehensive identification of protein substrates of the Dot/Icm type IV transporter of Legionella pneumophila. PLoS ONE 6, e17638 (2011).
Lifshitz, Z. et al. Computational modeling and experimental validation of the Legionella and Coxiella virulence-related type-IVB secretion signal. Proc. Natl Acad. Sci. USA 110, E707–E715 (2013).
Qiu, J. & Luo, Z. Q. Legionella and Coxiella effectors: strength in diversity and activity. Nat. Rev. Microbiol. 15, 591–605 (2017).
Qiu, J. et al. Ubiquitination independent of E1 and E2 enzymes by bacterial effectors. Nature 533, 120–124 (2016).
Kotewicz, K. M. et al. A single Legionella effector catalyzes a multistep ubiquitination pathway to rearrange tubular endoplasmic reticulum for replication. Cell Host Microbe 21, 169–181 (2017).
Bhogaraju, S. et al. Phosphoribosylation of ubiquitin promotes serine ubiquitination and impairs conventional ubiquitination. Cell 167, 1636–1649.e13 (2016).
Sheedlo, M. J. et al. Structural basis of substrate recognition by a bacterial deubiquitinase important for dynamics of phagosome ubiquitination. Proc. Natl Acad. Sci. USA 112, 15090–15095 (2015).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Rao, S. T. & Rossmann, M. G. Comparison of super-secondary structures in proteins. J. Mol. Biol. 76, 241–256 (1973).
Han, S., Arvai, A. S., Clancy, S. B. & Tainer, J. A. Crystal structure and novel recognition motif of Rho ADP-ribosylating C3 exoenzyme from Clostridium botulinum: structural insights for recognition specificity and catalysis. J. Mol. Biol. 305, 95–107 (2001).
Jeong, B. R. et al. Structure function analysis of an ADP-ribosyltransferase type III effector and its RNA-binding target in plant immunity. J. Biol. Chem. 286, 43272–43281 (2011).
Dikic, I., Wakatsuki, S. & Walters, K. J. Ubiquitin-binding domains—from structures to functions. Nat. Rev. Mol. Cell Biol. 10, 659–671 (2009).
Puvar, K. et al. Ubiquitin chains modified by the bacterial ligase SdeA are protected from deubiquitinase hydrolysis. Biochemistry 56, 4762–4766 (2017).
Sakurai, J., Nagahama, M., Oda, M., Tsuge, H. & Kobayashi, K. Clostridium perfringens iota-toxin: structure and function. Toxins (Basel) 1, 208–228 (2009).
Tsurumura, T. et al. Arginine ADP-ribosylation mechanism based on structural snapshots of iota-toxin and actin complex. Proc. Natl Acad. Sci. USA 110, 4267–4272 (2013).
Tsuge, H. et al. Structural basis of actin recognition and arginine ADP-ribosylation by Clostridium perfringens ι-toxin. Proc. Natl Acad. Sci. USA 105, 7399–7404 (2008).
Akturk, A. et al. Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector. Nature https://doi.org/10.1038/s41586-018-0147-6 (2018).
Kalayil, S. et al. Insights into catalysis and function of phosphoribosyl-linked serine ubiquitination. Nature https://doi.org/10.1038/s41586-018-0145-8 (2018).
Ninio, S., Zuckman-Cholon, D. M., Cambronne, E. D. & Roy, C. R. The Legionella IcmS–IcmW protein complex is important for Dot/Icm-mediated protein translocation. Mol. Microbiol. 55, 912–926 (2005).
Cambronne, E. D. & Roy, C. R. The Legionella pneumophila IcmSW complex interacts with multiple Dot/Icm effectors to facilitate type IV translocation. PLoS Pathog. 3, e188 (2007).
Kwak, M. J. et al. Architecture of the type IV coupling protein complex of Legionella pneumophila. Nat. Microbiol. 2, 17114 (2017).
Wang, Q. S. et al. The macromolecular crystallography beamline of SSRF. Nucl. Sci. Tech. 26, 010102 (2015).
Otwinowski, Z., Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Cowtan, K. D. & Zhang, K. Y. Density modification for macromolecular phase improvement. Prog. Biophys. Mol. Biol. 72, 245–270 (1999).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. PRIMUS - a Windows-PC based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Wang, J. et al. A method for helical RNA global structure determination in solution using small-angle X-ray scattering and NMR measurements. J. Mol. Biol. 393, 717–734 (2009).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Rambo, R. P. & Tainer, J. A. Accurate assessment of mass, models and resolution by small-angle scattering. Nature 496, 477–481 (2013).
Petoukhov, M. V., Konarev, P. V., Kikhney, A. G. & Svergun, D. I. ATSAS 2.1—towards automated and websupported small-angle scattering data analysis. J. Appl. Crystallogr. 40, s223–s228 (2007).
Svergun, D. I., Barberato, C. & Koch, M. H. J. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Svergun, D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 76, 2879–2886 (1999).
Volkov, V. V. & Svergun, D. I. Uniqueness of ab initio shape determination in small-angle scattering. J. Appl. Crystallogr. 36, 860–864 (2003).
Kozin, M. B. & Svergun, D. I. Automated matching of high- and low-resolution structural models. J. Appl. Crystallogr. 34, 33–41 (2001).
Bas, D. C., Rogers, D. M. & Jensen, J. H. Very fast prediction and rationalization of pK a values for protein–ligand complexes. Proteins 73, 765–783 (2008).
Li, H., Robertson, A. D. & Jensen, J. H. Very fast empirical prediction and rationalization of protein pK a values. Proteins 61, 704–721 (2005).
Banks, J. L. et al. Integrated modeling program, applied chemical theory (IMPACT). J. Comput. Chem. 26, 1752–1780 (2005).
Dodda, L. S., Cabeza de Vaca, I., Tirado-Rives, J. & Jorgensen, W. L. LigParGen web server: an automatic OPLS-AA parameter generator for organic ligands. Nucleic Acids Res. 45, W331–W336 (2017).
Jorgensen, W. L. & Tirado-Rives, J. Molecular modeling of organic and biomolecular systems using BOSS and MCPRO. J. Comput. Chem. 26, 1689–1700 (2005).
Storer, J. W., Giesen, D. J., Cramer, C. J. & Truhlar, D. G. Class IV charge models: a new semiempirical approach in quantum chemistry. J. Comput. Aided Mol. Des. 9, 87–110 (1995).
Udier-Blagović, M., Morales De Tirado, P., Pearlman, S. A. & Jorgensen, W. L. Accuracy of free energies of hydration using CM1 and CM3 atomic charges. J. Comput. Chem. 25, 1322–1332 (2004).
Dodda, L. S., Vilseck, J. Z., Tirado-Rives, J. & Jorgensen, W. L. 1.14*CM1A-LBCC: Localized Bond-Charge Corrected CM1A Charges for Condensed-Phase Simulations. J. Phys. Chem. B 121, 3864–3870 (2017).
Bowers, K. J. et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proc. of the 2006 ACM/IEEE Conference on Supercomputing 43–43 (2006).
Kräutler, V., Van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22, 501–508 (2001).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089 (1993).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Acknowledgements
We thank B. Zhou for the gift of the yeast W303 strain and the pYES2 vector; Y. Zhang for help in the yeast experiments; J. Ren, X. Zhang and R. Qiao for discussions about the mechanisms of SdeA; S. Zhang for the electron microscopy tests of the SdeA sample; C. Yan for help with the data collection process; the Tsinghua University Branch of China National Center for Protein Sciences Beijing and Y. Xue for providing facility support for NMR analysis of the protein samples; the Protein Chemistry Facility at the Center for Biomedical Analysis of Tsinghua University and W. Zhang for sample analysis; the staff at beamline BL17U1 and BL19U1 of the Shanghai Synchrotron Radiation Facility for their assistance with data collection and X. Zuo at the Advanced Photon Source (APS), Argonne National Laboratory (ANL) and the staff of BL19U2 beamline at the National Center for Protein Science Shanghai and Shanghai Synchrotron Radiation Facility for assistance during data collection. Use of the scattering beamline 12-ID-B resource at APS, ANL is allocated under the GUP-52757 to X.F. This work was supported by the National Key Research and Development Program of China (2017YFA0506500), the National Natural Science Foundation of China (31670766, 21532004, 21475005, and 21622501) and the Fundamental Research Funds for the Central Universities (buctylkxj03).
Reviewer information
Nature thanks K. Gehring and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.F. designed and supervised the project. Y.D., Y.M., Y.H. and W.W. purified the proteins, grew and optimized the crystals, and tested the diffractions of hundreds of crystals. Y.M., Y.X., Y.D., Z.L., Y.H., H.W. and N.X. performed the in vitro activity analysis, MST analysis, yeast toxicity assay and GST pull-down assays. Y.Zho. conducted the molecular dynamics simulations, and C.-Q.X. and J.L. conducted calculations to analyse the results of the molecular dynamics simulations. Y.Zha. and X.F. performed the SAXS analysis of different constructs of SdeA. M.W., H.D. and X.L. performed the mass spectrometric analysis. M.P., T.T., and L.L. contributed to experiment design and helped to supervise the project. J.W., Y.F., M.Y. and S.F. collected and analysed the crystallographic data and solved the crystal structure. Y.F. analysed the data and wrote the paper with the help of all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sequence alignment of SidE family members, spanning the regions corresponding to the SdeA fragment used in crystallization.
Residues with 100% homology, over 75% homology and over 50% homology are shaded in dark blue, pink and light blue, respectively. Secondary structural elements of SdeA are shown above the sequences. The residue ranges of the PDE, mART, pCTD and the α-helical lobe of the mART domain are marked with brackets or lines.
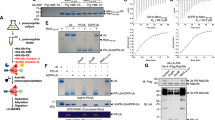
Extended Data Fig. 2 SdeA(231–1190) is an active monomer in solution.
a, SdeA, wild-type SdeA(231–1190), SdeA(231–1190)E860A/E862A or SdeA(231–1190)H277A were incubated with NAD+ and Ub in the presence or absence of His–RAB33B. Ubiquitinated His–RAB33B were analysed using tricine gels, Coomassie staining and immunoblotting with anti-His and anti-Ub antibodies. b, SdeA(231–1190) and RAB33B(15–202) were incubated with GST–Ub and NAD+, and self-ubiquitinated SdeA was detected by Coomassie staining, immunoblotting with anti-Ub antibodies, and Pro-Q diamond phosphoprotein staining. c, SdeA(231–1190), NAD+ and RAB33B were incubated with 0, 1, 2 or 5 mM NADH. The ubiquitination reactions were analysed using tricine gel and Coomassie staining. d, Analytical ultracentrifugation results showed that SdeA(231–1190) is a monomer. Analytical ultracentrifugation analysis yielded a sedimentation coefficient of 5.13 S, and a molecular mass of approximately 106 kDa. The buffer is 10 mM Tris pH 8.0, 200 mM NaCl and 5 mM DTT. e, Gel filtration profile of the SdeA(231–1190) protein and the molecular markers on Superdex-75 column (GE Healthcare) are shown. The sizes of the molecular markers are marked on top of the peaks. The samples of SdeA(231–1190) collected from the Superdex-75 column were run on SDS–PAGE gels and detected by Coomassie staining. a–e, Similar results were obtained in three independent experiments. a–c, e, Uncropped blots and gel images are shown in Supplementary Fig. 1. f, Two views of the superimposition of the structures of the two molecules in the asymmetric unit, coloured in different colours. g, Structure of the CTD region in the crystallized protein can be divided into two parts (left and right). The α helices are numbered according to their orders in the residue region from 908 to 1190. h, Topological diagram of the CTD region shown in g. The N and C termini of the pCTD domain are labelled.
Extended Data Fig. 3 Interactions between SdeA mART and SdeA PDE are essential for the activity of SdeA mART.
a, Overview of the interactions between SdeA mART and SdeA PDE. SdeA is coloured as in Fig. 1b. The major interaction region between the two domains is outlined. b, An expanded view of the region outlined in a. Interaction residues are shown in stick representation and the red dashed lines represent polar interactions. The plug loop in SdeA mART is indicated. c, The interaction between SdeA mART and SdeA PDE. SdeA mART and SdeA PDE are shown in cartoon and surface electrostatic models, respectively. d, A view of the interaction from c rotated by 180 degrees. In this view, SdeA mART and SdeA PDE are shown as surface electrostatic and cartoon models, respectively. e, Testing the ability of SdeA PDE to process ADPR-Ub into PR-Ub. SdeA(231–588), wild-type ∆NC SdeA or the H277A mutant were incubated with ADPR-Ub and RAB33B for 30 min. The samples were stained with Coomassie and Pro-Q diamond phosphoprotein stain. f, Testing the importance of domain interaction for the activity of SdeA mART. Various SdeA segments, and mixtures of SdeA(231–588) and SdeA(597–935) or SdeA(193–935)H277A, were incubated with RAB33B, NAD+ and Ub for 30 min. The samples were analysed using Coomassie staining, immunoblotting with anti-Ub antibodies and Pro-Q diamond phosphoprotein staining. e, f, Similar results were obtained in three independent experiments. Uncropped blots and gel images are shown in Supplementary Fig. 1.
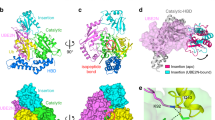
Extended Data Fig. 4 SdeA mART exhibits novel conformations of ARTT and PN loops.
a, Superimposition of SdeA mART (green) and HopU1 (pink) from Pseudomonas syringae (PDB: 3U0J). The ARTT loop is indicated. The r.m.s.d. value is indicated beside the PDB code (panels b and c are arranged in the same way). b, Superimposition of SdeA mART (green) and ADP-ribosyltransferase Vis (blue) (PDB: 4XZK). c, Superimposition of SdeA mART (green) and XopAI from Xanthomonas axonopodis pv. citri (cyan) (PDB: 4ELN). d, Superimposition of SdeA mART structure (green) and the three other structures from a–c. e, f, Mass spectra of the samples in Fig. 2f. The sample name and their molecular masses are indicated in the figures. g, h, Different fragments and different combinations of SdeA proteins were incubated with Ub, RAB33B and NAD+ at 37 °C for the indicated amounts of time. The samples were analysed using Coomassie staining and Pro-Q diamond phosphoprotein staining. i, Testing the ubiquitination ability of the catalytic core. 0.09 or 0.9 μM ∆NC SdeA or SdeA(193–935) was incubated with or without Ub and NAD+ for the indicated amounts of time. The samples were analysed using Coomassie staining, immunoblotting with anti-Ub antibodies and Pro-Q diamond phosphoprotein staining. e–i, Similar results were obtained in three independent experiments. g–i, Uncropped blots and gel images are shown in Supplementary Fig. 1.
Extended Data Fig. 5 SdeA(231–1190) binds three Ub molecules.
a, Overall structure of the SdeA(231–1190)–Ub complex. SdeA is coloured as in Fig. 1b. The three Ub molecules are coloured in magenta and labelled as Ub1–3 according to the order of their binding region in SdeA(231–1190). Q935 and S998, which are two common C termini of the clones used in this study, are shown as spheres. b, Ub binding causes prominent structural changes of SdeA. The SdeA–Ub complex structure is shown as in a, and the apo-SdeA structure is coloured in pink. The N-terminal region of SdeA pCTD which undergoes pronounced conformational changes is outlined with a circle. c, d, Expanded views of the two Ub binding sites in SdeA pCTD. The proteins are coloured as in a. Red dashed lines indicate polar interactions. e–h, Structural alignments of the Ub molecule (magenta) in the SdeA mART–Ub complex with the proximal (yellow) and distal (orange) Ubs of the K11- (e), K48- (f), K63- (g) and M1-linked (h) diubiquitins. The two R42 residues in each of the four diUbs are shown in stick representation.
Extended Data Fig. 6 Specific recognition of Ub by SdeA mART.
a, The interaction between SdeA mART and Ub. SdeA mART is shown as a surface electrostatic potential model and Ub is in magenta cartoon representation. The R42, R72 and R74 residues of Ub are shown in stick representation. b, UbR72 and UbR74 are bound in the negatively charged groove of SdeA mART. The front part of SdeA mART is cut away to reveal the inner surface. c, Superimposition of SUMO1 (PDB: 1WM3), NEDD8 (PDB: 1NDD) and Ub in the SdeA mART-Ub complex. The conserved Arg residues in Ub, SUMO1 and NEDD8 are shown in stick representation, out of which UbR42, UbR72, and UbR74 are marked. The polar interactions with UbR72 and UbR74 are shown as red dashed lines. d, The purified SUMO and NEDD8 proteins were incubated with SdeA(193–935)H277A and NAD+ under the conditions stated in the ‘Top-down LC–MS analysis of modified Ub and Ub-like proteins’ section of the Methods. Mass spectra of the samples are also shown. The sample names and their molecular masses are indicated in the figures. Similar results were obtained in three independent experiments.
Extended Data Fig. 7 Molecular dynamics simulations indicate the movements of the side chains of UbR42 and UbR72.
a–d, Wild-type ∆NC SdeA and indicated mutants were incubated with Ub, RAB33B and NAD+ at 37 °C for the indicated amounts of time. The samples were analysed using Coomassie staining and Pro-Q diamond phosphoprotein staining. Similar results were obtained in three independent experiments. Uncropped blots and gel images are shown in Supplementary Fig. 1. e, The structure of the SdeA mART–Ub–NADH complex. SdeA mART is shown as an electrostatic surface potential model. White, blue and red indicate neutral, positive and negative surfaces, respectively. Shown in green mesh is the 2Fo−Fc electron density map contoured at 1σ around the NADH molecule. f, Galactose-inducible pYES2 plasmids containing wild-type ∆NC SdeA or the mutants were transformed into yeast W303 strain. Five microlitres of cells in three tenfold serial dilutions were spotted on both glucose- and galactose-containing plates lacking uracil for two days before image acquisition. g, Purified ADPR-Ub proteins were treated with or without wild-type ∆NC SdeA. The samples were analysed using Coomassie staining and Pro-Q diamond phosphoprotein staining. h, Purified ADPR-Ub protein was subjected to top-down LC–MS analysis. The results indicated 100% ADPR-Ub. i, Wild-type ∆NC SdeA or other mutants were incubated with RAB33B and the prepared ADPR-Ub verified in g and h. The samples were analysed using SDS–PAGE, with Coomassie staining and Pro-Q diamond phosphoprotein staining. f–i, Similar results were obtained in three independent experiments. g, i, Uncropped blots and gel images are shown in Supplementary Fig. 1. j, k, The time series for the r.m.s.d. of the non-hydrogen atoms of the protein–ligand complex (j) and the ligand (k) in the SdeA mART–Ub–intermediate and SdeA mART–Ub–NAD+ systems during molecular dynamics simulations. These two plots indicate that both systems have reached equilibrium during the 200-ns simulations. l, m, The time series for the shortest distance between the NH1/2 atom of UbR72 and C1D of the ligand (l) and the distance between the NH1/2 atom of UbR42 and C1D of the ligand (m) in the two systems during molecular dynamics simulations.
Extended Data Fig. 8 The overall shape of SdeA and the function of SdeA CTD.
a, Superimposition of various Ub structures (PDB codes: 5M93 (orange), 1UBQ (pink), 5CRA (chain C, cyan), 3ZLZ (chain B, yellow) and 4BOZ (chain B, grey)) onto the SdeA mART–Ub–NADH structure with R42 residues of all the Ub molecules shown in stick representation. SdeA mART and Ub from the SdeA mART–Ub–NADH complex are shown in green and magenta, respectively. b, In vitro GST pull-down assays to detect the interactions of SdeA with IcmS or its complexes. GST-fused SdeA protein was incubated with IcmS, the IcmS–IcmW complex, the IcmS–IcmW–DotLc (residues 656–783 of DotL) ternary complex or the IcmS–IcmW–DotLc–LvgA quaternary complex. The protein samples bound to glutathione resins were washed three times and analysed by SDS–PAGE and Coomassie blue staining. IcmS/W represents IcmS + IcmW. The band marked with an asterisk represents the degraded GST tag. Similar results were obtained in three independent experiments. Uncropped blots and gel images are shown in Supplementary Fig. 1. c, Experimental PDDFs (pair distance distribution function) for SdeA(231–1190), SdeA(1–1499), SdeA(1–1190) and SdeA(1092–1496). d, Overlay of the experimental scattering profiles (exp) from the four samples in the SAXS analysis with the back-calculated scattering profile of the crystal structure of SdeA(231–1190) (cal). e, Fitting the crystal structure of SdeA(231–1190) into the SAXS envelope of SdeA(231–1190). Two perpendicular views are shown. f, Superimposition of the SAXS envelopes of SdeA(231–1190) (coloured as in e) and SdeA(1–1190) (light magenta) with the crystal structure of SdeA(231–1190) fitted. g, SAXS envelopes of SdeA(1092–1496). h, Superimposition of the SAXS envelopes of SdeA(1–1190) (light magenta), SdeA(1092–1496) (cyan) and SdeA(1–1499) (wheat) with the crystal structure of SdeA(231–1190) fitted.
Supplementary information
Supplementary Figure 1
This file contains the uncropped scans with size marker indications.
Rights and permissions
About this article
Cite this article
Dong, Y., Mu, Y., Xie, Y. et al. Structural basis of ubiquitin modification by the Legionella effector SdeA. Nature 557, 674–678 (2018). https://doi.org/10.1038/s41586-018-0146-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0146-7
This article is cited by
-
Legionella metaeffector MavL reverses ubiquitin ADP-ribosylation via a conserved arginine-specific macrodomain
Nature Communications (2024)
-
Molecular basis of threonine ADP-ribosylation of ubiquitin by bacterial ARTs
Nature Chemical Biology (2024)
-
Serine-ubiquitination regulates Golgi morphology and the secretory pathway upon Legionella infection
Cell Death & Differentiation (2021)
-
Structural basis for protein glutamylation by the Legionella pseudokinase SidJ
Nature Communications (2021)
-
Structural insights into the mechanism and inhibition of transglutaminase-induced ubiquitination by the Legionella effector MavC
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.