Abstract
The cell microenvironment, which is critical for stem cell maintenance, contains both cellular and non-cellular components, including secreted growth factors and the extracellular matrix1,2,3. Although Notch and other signalling pathways have previously been reported to regulate quiescence of stem cells4,5,6,7,8,9, the composition and source of molecules that maintain the stem cell niche remain largely unknown. Here we show that adult muscle satellite (stem) cells in mice produce extracellular matrix collagens to maintain quiescence in a cell-autonomous manner. Using chromatin immunoprecipitation followed by sequencing, we identified NOTCH1/RBPJ-bound regulatory elements adjacent to specific collagen genes, the expression of which is deregulated in Notch-mutant mice. Moreover, we show that Collagen V (COLV) produced by satellite cells is a critical component of the quiescent niche, as depletion of COLV by conditional deletion of the Col5a1 gene leads to anomalous cell cycle entry and gradual diminution of the stem cell pool. Notably, the interaction of COLV with satellite cells is mediated by the Calcitonin receptor, for which COLV acts as a surrogate local ligand. Systemic administration of a calcitonin derivative is sufficient to rescue the quiescence and self-renewal defects found in COLV-null satellite cells. This study reveals a Notch–COLV–Calcitonin receptor signalling cascade that maintains satellite cells in a quiescent state in a cell-autonomous fashion, and raises the possibility that similar reciprocal mechanisms act in diverse stem cell populations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
29 May 2018
In the originally published article, two panel labels were missing in the legend for Extended Data Fig. 1. This has now been corrected.
References
Raymond, K., Deugnier, M. A., Faraldo, M. M. & Glukhova, M. A. Adhesion within the stem cell niches. Curr. Opin. Cell Biol. 21, 623–629 (2009).
Moore, K. A. & Lemischka, I. R. Stem cells and their niches. Science 311, 1880–1885 (2006).
Watt, F. M. & Huck, W. T. Role of the extracellular matrix in regulating stem cell fate. Nat. Rev. Mol. Cell Biol. 14, 467–473 (2013).
Mourikis, P. et al. A critical requirement for notch signaling in maintenance of the quiescent skeletal muscle stem cell state. Stem Cells 30, 243–252 (2012).
Bjornson, C. R. et al. Notch signaling is necessary to maintain quiescence in adult muscle stem cells. Stem Cells 30, 232–242 (2012).
Rozo, M., Li, L. & Fan, C. M. Targeting β1-integrin signaling enhances regeneration in aged and dystrophic muscle in mice. Nat. Med. 22, 889–896 (2016).
Cheung, T. H. et al. Maintenance of muscle stem-cell quiescence by microRNA-489. Nature 482, 524–528 (2012).
Zismanov, V. et al. Phosphorylation of eIF2α is a translational control mechanism regulating muscle stem cell quiescence and self-renewal. Cell Stem Cell 18, 79–90 (2016).
Chakkalakal, J. V., Jones, K. M., Basson, M. A. & Brack, A. S. The aged niche disrupts muscle stem cell quiescence. Nature 490, 355–360 (2012).
Shen, H. et al. The Notch coactivator, MAML1, functions as a novel coactivator for MEF2C-mediated transcription and is required for normal myogenesis. Genes Dev. 20, 675–688 (2006).
Buas, M. F., Kabak, S. & Kadesch, T. The Notch effector Hey1 associates with myogenic target genes to repress myogenesis. J. Biol. Chem. 285, 1249–1258 (2010).
Castel, D. et al. Dynamic binding of RBPJ is determined by Notch signaling status. Genes Dev. 27, 1059–1071 (2013).
Vasyutina, E. et al. RBP-J (Rbpsuh) is essential to maintain muscle progenitor cells and to generate satellite cells. Proc. Natl Acad. Sci. USA 104, 4443–4448 (2007).
Sun, M. et al. Targeted deletion of collagen V in tendons and ligaments results in a classic Ehlers–Danlos syndrome joint phenotype. Am. J. Pathol. 185, 1436–1447 (2015).
Gilbert, P. M. et al. Substrate elasticity regulates skeletal muscle stem cell self-renewal in culture. Science 329, 1078–1081 (2010).
Yennek, S., Burute, M., Théry, M. & Tajbakhsh, S. Cell adhesion geometry regulates non-random DNA segregation and asymmetric cell fates in mouse skeletal muscle stem cells. Cell Reports 7, 961–970 (2014).
Leitinger, B. Transmembrane collagen receptors. Annu. Rev. Cell Dev. Biol. 27, 265–290 (2011).
Vogel, W., Gish, G. D., Alves, F. & Pawson, T. The discoidin domain receptor tyrosine kinases are activated by collagen. Mol. Cell 1, 13–23 (1997).
Paavola, K. J., Sidik, H., Zuchero, J. B., Eckart, M. & Talbot, W. S. Type IV collagen is an activating ligand for the adhesion G protein-coupled receptor GPR126. Sci. Signal. 7, ra76 (2014).
Luo, R. et al. G protein-coupled receptor 56 and collagen III, a receptor–ligand pair, regulates cortical development and lamination. Proc. Natl Acad. Sci. USA 108, 12925–12930 (2011).
Yamaguchi, M. et al. Calcitonin receptor signaling inhibits muscle stem cells from escaping the quiescent state and the niche. Cell Reports 13, 302–314 (2015).
Evans, B. N., Rosenblatt, M. I., Mnayer, L. O., Oliver, K. R. & Dickerson, I. M. CGRP-RCP, a novel protein required for signal transduction at calcitonin gene-related peptide and adrenomedullin receptors. J. Biol. Chem. 275, 31438–31443 (2000).
Rocheteau, P., Gayraud-Morel, B., Siegl-Cachedenier, I., Blasco, M. A. & Tajbakhsh, S. A subpopulation of adult skeletal muscle stem cells retains all template DNA strands after cell division. Cell 148, 112–125 (2012).
Mourikis, P. & Tajbakhsh, S. Distinct contextual roles for Notch signalling in skeletal muscle stem cells. BMC Dev. Biol. 14, 2 (2014).
Machado, L. et al. In situ fixation redefines quiescence and early activation of skeletal muscle stem cells. Cell Reports 21, 1982–1993 (2017).
Mourikis, P., Gopalakrishnan, S., Sambasivan, R. & Tajbakhsh, S. Cell-autonomous Notch activity maintains the temporal specification potential of skeletal muscle stem cells. Development 139, 4536–4548 (2012).
Haldar, M., Karan, G., Tvrdik, P. & Capecchi, M. R. Two cell lineages, myf5 and myf5-independent, participate in mouse skeletal myogenesis. Dev. Cell 14, 437–445 (2008).
Murphy, M. M., Lawson, J. A., Mathew, S. J., Hutcheson, D. A. & Kardon, G. Satellite cells, connective tissue fibroblasts and their interactions are crucial for muscle regeneration. Development 138, 3625–3637 (2011).
Murtaugh, L. C., Stanger, B. Z., Kwan, K. M. & Melton, D. A. Notch signaling controls multiple steps of pancreatic differentiation. Proc. Natl Acad. Sci. USA 100, 14920–14925 (2003).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Han, H. et al. Inducible gene knockout of transcription factor recombination signal binding protein-J reveals its essential role in T versus B lineage decision. Int. Immunol. 14, 637–645 (2002).
Sun, M. et al. Collagen V is a dominant regulator of collagen fibrillogenesis: dysfunctional regulation of structure and function in a corneal-stroma-specific Col5a1-null mouse model. J. Cell Sci. 124, 4096–4105 (2011).
Sambasivan, R. et al. Distinct regulatory cascades govern extraocular and pharyngeal arch muscle progenitor cell fates. Dev. Cell 16, 810–821 (2009).
Hicks, C. et al. A secreted Delta1-Fc fusion protein functions both as an activator and inhibitor of Notch1 signaling. J. Neurosci. Res. 68, 655–667 (2002).
Vasconcelos, F. F. et al. MyT1 counteracts the neural progenitor program to promote vertebrate neurogenesis. Cell Reports 17, 469–483 (2016).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔC t method. Methods 25, 402–408 (2001).
Gao, M. et al. Discovery and optimization of 3-(2-(Pyrazolo[1,5-a]pyrimidin-6-yl)ethynyl)benzamides as novel selective and orally bioavailable discoidin domain receptor 1 (DDR1) inhibitors. J. Med. Chem. 56, 3281–3295 (2013).
Shinin, V., Gayraud-Morel, B., Gomes, D. & Tajbakhsh, S. Asymmetric division and cosegregation of template DNA strands in adult muscle satellite cells. Nat. Cell Biol. 8, 677–687 (2006).
Yoshida, N., Yoshida, S., Koishi, K., Masuda, K. & Nabeshima, Y. Cell heterogeneity upon myogenic differentiation: down-regulation of MyoD and Myf-5 generates ‘reserve cells’. J. Cell Sci. 111, 769–779 (1998).
Morita, S., Kojima, T. & Kitamura, T. Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther. 7, 1063–1066 (2000).
Acknowledgements
We thank H. Stunnenberg for the ChIP–seq and RNA sequencing data; D. Castro for the RBPJ ChIP protocol; D. Greenspan for the anti-a3-COLV antibody and Col5a3-knockout muscle samples; C. Moali for the SPR assay; F. Auradé and the Protein Core Facility, Institut Curie, for the production of CalcR proteins; K. Ding for the 7rh DDR1 inhibitor; F. Ruggiero for suggesting the on-cell enzyme-linking immunosorbent assay experiment; and the Cytometry platforms of Institut Pasteur and IMRB, Inserm U955, Creteil. F.R. was funded by the Association Française contre les Myopathies via TRANSLAMUSCLE (PROJECT 19507), Agence Nationale pour la Recherche grant Satnet (ANR-15-CE13-0011-01) and RHU CARMMA (ANR-15-RHUS-0003). S.T. was funded by Institut Pasteur, Centre National pour la Recherche Scientific and the Agence Nationale de la Recherche (Laboratoire d’Excellence Revive, Investissement d’Avenir; ANR-10-LABX- 73) and the European Research Council (Advanced Research Grant 332893). M.B.B. was funded by the Doctoral School grant and Fondation pour la Recherche Médicale.
Reviewer information
Nature thanks I. Conboy, G. Kardon and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.B.B., S.T. and P.M. proposed the concept, designed experiments and wrote the manuscript, F.R. oversaw revisions, and S.T. funded most of the study. P.M. and D.C. conducted initial experiments on enhancer analysis. D.C. and L.M. performed and analysed ChIP experiments. M.B.B. performed the remaining experiments and, together with P.M., analysed the data. S.F. and D.E.B. provided mouse models.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of NICD/RBPJ-bound enhancers and response to activation of Notch signalling.
a, Gene expression microarray data show that satellite cells express a specific subset of collagen types, which include the fibrillar COLI (Col1a1 and Col1a2), COLIII (Col3a1, possibly as (α1(III))3 homodimer) and COLV (Col5a1, Col5a2 and Col5a3) and the non-fibrillar COLIV (Col4a1 and Col4a2), COLVI (Col6a1 and Col6a2) and COLXV (Col15a1, possibly as (α1(XV))3 homodimer). Data are shown as a heat map of normalized collagen transcripts expressed at different developmental time points (E12.5, E17.5 and post-natal day (P)8; Tg-Pax7-nGFP, Gene Expression Omnibus (GEO) accession number GSE52192), quiescent and post-injury (t = 60 h after BaCl2 injury). b, ChIP–seq tracks indicating NICD/RBPJ-occupied enhancers, associated with mouse Col5a1, Col5a3, Col6a1 and Col6a2 loci. H3K4me1, H3K27ac, p300 and NICD are shown. Orange rectangles indicate RBPJ binding positions and asterisks indicate the enhancers used for transcriptional activity assays in c. c, Core sequences of the selected NICD/RBPJ-bound enhancers (asterisked orange rectangle in Fig. 1a and in b). The RBPJ consensus binding motif is highlighted in yellow. d, Transcriptional response of isolated enhancers to activation of Notch signalling in C2C12 cells. Firefly luciferase signal was measured in cells with doxycycline-inducible expressed human Notch1–GFP (NICD) and GFP control cells treated with (2S)-N-[(3,5-difluorophenyl)acetyl]-l-alanyl-2-phenyl]glycine 1,1-dimethylethyl ester (DAPT) and were normalized to internal control (pCMV-Renilla). Data are expressed as relative luminescence units (n = 3 independent experiments). Data are mean ± s.d.; two-sided paired t-test. e, Expression measurements, based on RNA sequencing, of collagen genes in myogenic C2C12 cells, with active (treated with Delta-like 1) or inhibited (treated with DAPT) Notch signalling for 6 or 24 h (data available at GEO, accession number GSE37184). Data are shown as Delta-like 1-to-DAPT ratios of average reads per kilobase of exon model per million mapped reads (RPKMs). Genes with low expression (RPKM < 2) were eliminated. HeyL and Hey1 transcripts indicate Notch pathway activation. Red line designates no change (ratio = 1).
Extended Data Fig. 2 Notch signalling regulates Col5 and Col6 expression in vivo.
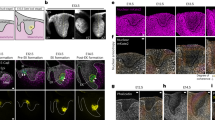
a, Satellite cells isolated by FACS at day 10 after tamoxifen injections, from resting tibialis anterior muscle from control (Tg:Pax7-CT2;Rbpj+/−;R26mTmG/+) and Rbpj-null (Tg:Pax7-CT2;Rbpjflox/−;R26mTmG/+) mice immunostained for RBPJ. b, RT–qPCR of collagen genes in Rbpj cKO and control satellite cells. Hey1 used as control for Notch signalling (n = 3 mice per genotype). c, Induction of collagen genes in E17.5 control (Myf5Cre/+;R26mTmG/+) and Myf5Cre-NICD (Myf5Cre/+;R26stop-NICD-nGFP/+) cells isolated by FACS. RT–qPCR was normalized to Gapdh, n = 3 fetuses per genotype. HeyL reports Notch activity. d, FACS plots showing fractionation of GFP+ cells from E17.5 Tg:Pax7-nGFP fetuses into Pax7high (20% of population), Pax7mid (40%), and Pax7low (20%). The intensity of the GFP signal reflects the activity of the Pax7 promoter. e, Transcript levels of GFP+ cells isolated by FACS show a tight correlation between lineage progression, Notch signalling activity and collagen gene expression (n = 3 fetuses per genotype). f, Specificity of α3-COLV antibody assessed by immunostaining of tibialis anterior muscle transverse section from wild-type and Col5a3 cKO P14 postnatal pups (n = 3 mice per genotype). g, Time course of gene expression performed by RT–qPCR on freshly isolated satellite cells (Quiescent), 48 h or 60 h after cardiotoxin injury of tibialis anterior muscle (48 hours post injury (hpi), 60 hpi), and isolated single myofibres from extensor digitorum longus muscle of Tg:Pax7-nGFP mice. Col5a1 and Col5a3 were strongly downregulated in activated and differentiated cells. Quiescence (Pax7, Calcr) and differentiation (Myog) markers are indicated. Col4a2, a major component of the basement membrane, is expressed mainly by myofibres (n = 3 mice per condition). Data are mean ± s.d.; one-sided unpaired t-test. Scale bars, 50 μm.
Extended Data Fig. 3 COLV delays proliferation and differentiation of satellite cells.
a, Experimental scheme: isolated Tg:Pax7-nGFP satellite cells cultured overnight (o/n) before collagen treatment. b, Myosin heavy chain (MyHC) and EdU staining of satellite cells treated with COLI or COLV. Fusion index: 82%, 86% and 33% for HOAc solvent, COLI and COLV, respectively (n = 3 mice, ≥250 cells, 2 wells per condition). c, Percentage of EdU+ primary myogenic cells after ten days of culture with indicated collagens. EdU: 2.6%, 1.3% and 18.2% for COLI, COLVI and COLV, respectively (n = 3 mice, ≥250 cells, 2 wells per condition). d, Experimental scheme for control and cKO mice. Satellite cells were plated overnight before collagen treatment. e, GFP and MyHC immunostaining of Rbpj cKO satellite cells (n = 3 mice per condition) incubated 60 h in presence of COLI or COLV, or with HOAc control (n = 3 mice, ≥200 cells, 2 wells per condition). f, Percentage of EdU+ cells (2 h pulse) of Rbpj-null primary myogenic cells, after ten days of culture with HOAc or indicated collagens. EdU: 1.0% and 7.6% for COLI and COLV, respectively (n = 3 mice, ≥150 cells, 2 wells per condition). g, RT–qPCR on GFP+ Rbpj-null satellite cells isolated by FACS and cultured for 72 h in the presence of COLI or COLV. Results are normalized to Tbp. Data are mean ± s.d.; two-sided paired t-test; P value: two-sided unpaired t-test. Scale bars, 50 μm.
Extended Data Fig. 4 COLV—and specifically α3-COLV—is critical for satellite cell self-renewal.
a, RT–qPCR of Col5a1 in control (Ctr; Tg:Pax7-CT2;Col5a1+/+;R26mTmG), heterozygous (Het; Tg:Pax7-CT2;Col5a1flox/+;R26mTmG) and conditional knockout (cKO; Tg:Pax7CT2;Col5a1flox/flox;R26mTmG) mice two weeks after tamoxifen diet (n = 3 mice per genotype). b, Transcript levels of the different Col5 mRNA chains in C2C12 after transfection of either control scramble, Col5a1 or Col5a3 siRNA, showing the specificity of each siRNA for its given targeted mRNA. Data are normalized to Tbp gene expression (n = 3 independent assays). c, Col5a1 and Col5a3 siRNA transfection of Tg:Pax7-nGFP isolated single myofibres cultured for 72 h and immunostained for GFP and MYOD. Resident satellite cells enter the myogenic program and form clusters composed of proliferating (PAX7+MYOD+MYOG−), differentiated (PAX7−MYOG+) and self-renewed (PAX7+MYOD−) cells within 72 h. Quantification of PAX7+MYOD−, PAX7+MYOD+ and PAX7−MYOD+ populations 72 h after transfection. Scramble siRNA was used as negative control (n ≥ 15 fibres counted from 3 mice). Data are mean ± s.d.; a, two-sided unpaired t-test; b, c, two-sided paired t-test. Scale bar, 50 μm.
Extended Data Fig. 5 Screening for COLV receptor candidates identifies CALCR.
a, Screening for the COLV receptor: satellite cells from Tg:Pax7-nGFP mice were incubated for ten days with COLV and candidate receptors were targeted with respective inhibitors: 7rh for DDR1 (sub-panels C, D), the broad-spectrum Integrin-binding competitor RGDS peptide (sub-panels E, F), Obtustatin for Integrin α1β1 (sub-panels G, H), TC-I 15 for Integrin α2β1 (sub-panels I, J). DMSO solvent was used as a control for TC-I 15 and 7rh (sub-panels A, B). Satellite cell differentiation was assayed by MyHC immunostaining. b, EdU (2 h pulse) and CALCR staining of GFP+ C2C12 cells isolated by FACS and transduced with Calcr-GFP or mock-GFP retrovirus and cultured for 24 h with COLI (top) or COLV (bottom). Quantification of EdU+ Calcr-transduced C2C12 cells or mock-GFP cells treated for 24h with COLV or with the controls, COLI and HOAc (n = 5 independent experiments, ≥250 cells counted, 2 wells per condition). There was no significant difference between HOAc and COLI treated samples (data not shown). c, Experimental scheme of tamoxifen administration to control (Ctr) (Calcr+/+) and cKO (Calcrflox/flox) mice. FACS plot of satellite cells from Pax7CreERT2/+;Calcrflox/flox;R26stop-YFP and Pax7CreERT2/+;Calcr+/+;R26stop-YFP mice. Cells sorted based on YFP expression. d, Control and Calcr cKO satellite cells isolated by FACS, fixed immediately after sorting and immunostained for CALCR to confirm the absence of CALCR protein from recombined cells. For control (upper panel), two fields from the same culture dish are shown, separated by a white line. Asterisk shows a non-recombined, CALCR+ cell in the cKO sample (lower panel). e, Quantification of PAX7+, Myogenin+ and EdU+ cells in Calcr-depleted satellite cells (Pax7CT2/+;Calcrflox/flox;R26stop-YFP) isolated by FACS and treated for 32 h or 72 h with COLI or COLV (n = 3 mice, ≥250 cells counted, 2 wells per condition). f, Quantification of total PAX7+ (GFP), Myogenin+ and EdU+ myogenic cells isolated by FACS from Tg:Pax7-nGFP mice three days after cardiotoxin injury of tibialis anterior muscle, and incubated for 72 h in presence of COLI or COLV, or HOAc as a control, in the culture medium (n = 3 mice, ≥200 cells counted). g, CALCR protein in freshly isolated satellite cells, or satellite cells cultured for 12 h, from Tg:Pax7-nGFP mice, demonstrating that CALCR protein is still present when satellite cells are treated with different collagens (see Extended Data Fig. 2). h, Induction of Calcr transcript expression by RT–qPCR of Tg:Pax7-nGFP satellite cells isolated by FACS and cultured for 72 h in the presence of COLI or COLV. Results are normalized to Tbp (n = 3 mice). i, Immunostainings for CALCR protein of Tg:Pax7-nGFP satellite cells cultured for 72 h in presence of COLI or COLV (n = 3 mice, ≥50 cells, 2 wells per condition). Data are mean ± s.d.; b, two-sided unpaired t-test; c–i, two-sided paired t-test. Scale bars, 25 μm (g), 50 μm (a, b, d, i).
Extended Data Fig. 6 CALCR ligand elcatonin can substitute the depletion of the surrogate ligand COLV.
a, Intracellular levels of cAMP in Calcr-transduced C2C12 cells treated with COLV for up to 480 min (n = 4 independent assays). b, Rescue of loss of COLV by elcatonin in an ex vivo self-renewal reserve-cell model, where PAX7+ non-proliferative cells return to quiescence (see Methods). MyHC and PAX7 staining of control (Ctr: Tg:Pax7-CT2;Col5a1+/+;R26mTmG) and Col5a1-null (Tg:Pax7-CT2;Col5a1flox/flox;R26mTmG) cells, non-treated (NT) or treated with elcatonin. No GFP+EdU+ cells (12 h pulse) could be detected under any of the conditions, indicating GFP+ cells are quiescent (data not shown). c, Quantification of percentage of reserve cells (PAX7+ per total nuclei) (n = 3 mice per genotype and condition, ≥350 cells counted). Elcat, elcatonin. Data are mean ± s.d.; two-sided paired t-test; #, P value calculated by two-sided unpaired t-test. Scale bar, 50 μm.
Extended Data Fig. 7 Schematic of Notch–COLV–CALCR axis in satellite cells.
A Notch–COLV–CALCR signalling cascade actively maintains satellite cell quiescence. Satellite cells are in direct contact with the plasma membrane of the myofibre (black outline) and an overlying basement membrane (blue line). Activation of the Notch receptor is achieved by a ligand (probably DLL1 or DLL4) present on the muscle fibre. Induction of Col5a1 and Col5a3 (and also Col6a1 and Col6a2) genes occurs via distal regulatory elements (grey box). Satellite-cell-produced COLV is deposited under the basement membrane and acts as a surrogate ligand of the plasma membrane receptor CALCR, also expressed by the satellite cells, thereby propagating a cell-autonomous signalling system in the local niche. In the absence of COLV (deletion of Col5a1) the quiescent niche is disturbed, CALCR signalling is abrogated, and satellite cells spontaneously differentiate and fuse to myofibres, leading to exhaustion of the muscle stem cell pool.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3
Rights and permissions
About this article
Cite this article
Baghdadi, M.B., Castel, D., Machado, L. et al. Reciprocal signalling by Notch–Collagen V–CALCR retains muscle stem cells in their niche. Nature 557, 714–718 (2018). https://doi.org/10.1038/s41586-018-0144-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0144-9
This article is cited by
-
Neurofibromin 1 controls metabolic balance and Notch-dependent quiescence of murine juvenile myogenic progenitors
Nature Communications (2024)
-
Post-transcriptional regulation of myogenic transcription factors during muscle development and pathogenesis
Journal of Muscle Research and Cell Motility (2024)
-
Multi-omics analysis of sarcospan overexpression in mdx skeletal muscle reveals compensatory remodeling of cytoskeleton-matrix interactions that promote mechanotransduction pathways
Skeletal Muscle (2023)
-
Extracellular matrix-induced signaling pathways in mesenchymal stem/stromal cells
Cell Communication and Signaling (2023)
-
Sox11 is enriched in myogenic progenitors but dispensable for development and regeneration of the skeletal muscle
Skeletal Muscle (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.