Abstract
Microglia and astrocytes modulate inflammation and neurodegeneration in the central nervous system (CNS)1,2,3. Microglia modulate pro-inflammatory and neurotoxic activities in astrocytes, but the mechanisms involved are not completely understood4,5. Here we report that TGFα and VEGF-B produced by microglia regulate the pathogenic activities of astrocytes in the experimental autoimmune encephalomyelitis (EAE) mouse model of multiple sclerosis. Microglia-derived TGFα acts via the ErbB1 receptor in astrocytes to limit their pathogenic activities and EAE development. Conversely, microglial VEGF-B triggers FLT-1 signalling in astrocytes and worsens EAE. VEGF-B and TGFα also participate in the microglial control of human astrocytes. Furthermore, expression of TGFα and VEGF-B in CD14+ cells correlates with the multiple sclerosis lesion stage. Finally, metabolites of dietary tryptophan produced by the commensal flora control microglial activation and TGFα and VEGF-B production, modulating the transcriptional program of astrocytes and CNS inflammation through a mechanism mediated by the aryl hydrocarbon receptor. In summary, we identified positive and negative regulators that mediate the microglial control of astrocytes. Moreover, these findings define a pathway through which microbial metabolites limit pathogenic activities of microglia and astrocytes, and suppress CNS inflammation. This pathway may guide new therapies for multiple sclerosis and other neurological disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ben Haim, L. & Rowitch, D. H. Functional diversity of astrocytes in neural circuit regulation. Nat. Rev. Neurosci. 18, 31–41 (2017).
Khakh, B. S. & Sofroniew, M. V. Diversity of astrocyte functions and phenotypes in neural circuits. Nat. Neurosci. 18, 942–952 (2015).
Sofroniew, M. V. Astrocyte barriers to neurotoxic inflammation. Nat. Rev. Neurosci. 16, 249–263 (2015).
Liddelow, S. A. & Barres, B. A. Reactive astrocytes: production, function, and therapeutic potential. Immunity 46, 957–967 (2017).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Buttgereit, A. et al. Sall1 is a transcriptional regulator defining microglia identity and function. Nat. Immunol. 17, 1397–1406 (2016).
Lee, Y. H. et al. Aryl hydrocarbon receptor mediates both proinflammatory and anti-inflammatory effects in lipopolysaccharide-activated microglia. Glia 63, 1138–1154 (2015).
Goldmann, T. et al. A new type of microglia gene targeting shows TAK1 to be pivotal in CNS autoimmune inflammation. Nat. Neurosci. 16, 1618–1626 (2013).
Croxford, A. L. et al. The cytokine GM-CSF drives the inflammatory signature of CCR2+ monocytes and licenses autoimmunity. Immunity 43, 502–514 (2015).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med. 22, 586–597 (2016).
Yeste, A. et al. Tolerogenic nanoparticles inhibit T cell-mediated autoimmunity through SOCS2. Sci. Signal. 9, ra61 (2016).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Quintana, F. J. & Sherr, D. H. Aryl hydrocarbon receptor control of adaptive immunity. Pharmacol. Rev. 65, 1148–1161 (2013).
Stockinger, B., Di Meglio, P., Gialitakis, M. & Duarte, J. H. The aryl hydrocarbon receptor: multitasking in the immune system. Annu. Rev. Immunol. 32, 403–432 (2014).
Mayo, L. et al. Regulation of astrocyte activation by glycolipids drives chronic CNS inflammation. Nat. Med. 20, 1147–1156 (2014).
Rothhammer, V. et al. Sphingosine 1-phosphate receptor modulation suppresses pathogenic astrocyte activation and chronic progressive CNS inflammation. Proc. Natl Acad. Sci. USA 114, 2012–2017 (2017).
Wheeler, M. A. & Quintana, F. J. Regulation of Astrocyte functions in multiple sclerosis. Cold Spring Harb. Perspect. Med. https://doi.org/10.1101/cshperspect.a029009 (2018).
Gutiérrez-Vázquez, C. & Quintana, F. J. Regulation of the immune response by the aryl hydrocarbon receptor. Immunity 48, 19–33 (2018).
Wikoff, W. R. et al. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl Acad. Sci. USA 106, 3698–3703 (2009).
Girolamo, F., Coppola, C., Ribatti, D. & Trojano, M. Angiogenesis in multiple sclerosis and experimental autoimmune encephalomyelitis. Acta Neuropathol. Commun. 2, 84 (2014).
Mosher, K. I. et al. Neural progenitor cells regulate microglia functions and activity. Nat. Neurosci. 15, 1485–1487 (2012).
Mor, F., Quintana, F. J. & Cohen, I. R. Angiogenesis-inflammation cross-talk: vascular endothelial growth factor is secreted by activated T cells and induces Th1 polarization. J. Immunol. 172, 4618–4623 (2004).
Gaál, E. I. et al. Comparison of vascular growth factors in the murine brain reveals placenta growth factor as prime candidate for CNS revascularization. Blood 122, 658–665 (2013).
Li, X., Kumar, A., Zhang, F., Lee, C. & Tang, Z. Complicated life, complicated VEGF-B. Trends Mol. Med. 18, 119–127 (2012).
Nag, S., Eskandarian, M. R., Davis, J. & Eubanks, J. H. Differential expression of vascular endothelial growth factor-A (VEGF-A) and VEGF-B after brain injury. J. Neuropathol. Exp. Neurol. 61, 778–788 (2002).
Junier, M. P. What role(s) for TGFα in the central nervous system? Prog. Neurobiol. 62, 443–473 (2000).
White, R. E., Yin, F. Q. & Jakeman, L. B. TGF-α increases astrocyte invasion and promotes axonal growth into the lesion following spinal cord injury in mice. Exp. Neurol. 214, 10–24 (2008).
Anderson, M. A. et al. Astrocyte scar formation aids central nervous system axon regeneration. Nature 532, 195–200 (2016).
Kigerl, K. A. et al. Gut dysbiosis impairs recovery after spinal cord injury. J. Exp. Med. 213, 2603–2620 (2016).
Mildner, A. et al. Distinct and non-redundant roles of microglia and myeloid subsets in mouse models of Alzheimer’s disease. J. Neurosci. 31, 11159–11171 (2011).
Mildner, A. et al. Microglia in the adult brain arise from Ly-6ChiCCR2+ monocytes only under defined host conditions. Nat. Neurosci. 10, 1544–1553 (2007).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protocols 7, 562–578 (2012).
Yan, Y. et al. CNS-specific therapy for ongoing EAE by silencing IL-17 pathway in astrocytes. Mol. Ther. 20, 1338–1348 (2012).
Jack, C. S. et al. TLR signaling tailors innate immune responses in human microglia and astrocytes. J. Immunol. 175, 4320–4330 (2005).
Acknowledgements
This work was supported by grants NS087867, ES02530, AI126880 and AI093903 from the National Institutes of Health, RSG-14-198-01-LIB from the American Cancer Society and RG4111A1 and JF2161-A-5 from the National Multiple Sclerosis Society to F.J.Q. F.J.Q. and J.A. received support from International Progressive Multiple Sclerosis Alliance grant PA-1604-08459. V.R. received support from an educational grant from Mallinkrodt Pharmaceuticals (A219074) and by a fellowship from the German Research Foundation (DFG RO4866 1/1). M.P. is supported by the BMBF-funded competence network of multiple sclerosis (KKNMS), the Sobek-Stiftung and the DFG (SFB 992, SFB1140, SFB/TRR167, Reinhart-Koselleck-Grant) and the Ministry of Science, Research and the Arts, Baden-Wuerttemberg (Sonderlinie ‘Neuroinflammation’). Human fetal tissue came from the Human Fetal Tissue Repository (Albert Einstein College of Medicine) and from the University of Washington Birth Defects Research Laboratory (BDRL, Project Number 5R24HD000836-51).
Reviewer information
Nature thanks S. Liddelow, M. Platten, H. Wekerle and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
V.R., D.M.B., M.C.T., C.-C.C., A.A.-F., K.A.d.L., C.G.-V., P.H., O.S., M.Bl., L.H., T.N., M.Bo., M.W., L.L.D., D.A.L. and J.I.A. performed in vitro and in vivo experiments, J.A. and M.P. provided unique reagents, discussed and/or interpreted findings, E.C.T. performed bioinformatics, V.R. and F.J.Q. wrote the manuscript and F.J.Q. designed and supervised the study and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
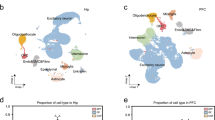
Extended Data Fig. 1 Contribution of AHR in CNS resident and infiltrating immune cells during EAE.
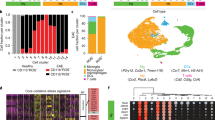
a, qPCR of indicated genes from microglia, splenic macrophages, and astrocytes from control and CX3CR1-AHR mice on day 28 after EAE induction. n = 8 independent samples per group. b, Flow cytometry analysis of AHR expression in microglia, monocytes and astrocytes from Control and CX3CR1-AHR mice 21 days after EAE induction. Thin line depicts isotype control, thick line AHR staining, and numbers indicate percentage of AHR positive cells. Representative of stainings of n = 3 mice per group. c, Spinal cord samples from naive Control and CX3CR1-AHR mice were stained for Iba-1 and DAPI and Iba-1+ microglia/mm2 were determined. n = 5 mice per group. n.s., not significant. d, TUNEL staining in Iba-1+ microglia in spinal cord sections of control and CX3CR1-AHR mice as in c. For the positive control, slides were cooked at 98 °C in citrate buffer during 60 min using a vapour cooker. Solid arrows show TUNEL positive microglia. Representative of n = 5 independent experiments. e, Number of CNS-infiltrating (top) and splenic T cells (bottom), and splenic pro-inflammatory monocytes (bottom) as determined by flow cytometry. n = 5 samples per group for CNS, n = 4 samples per group for spleen. f, Proliferation assay from splenocytes isolated on day 28 of the experiment. n = 4 biologically independent samples per group, representative of two independent experiments. g, Bone marrow chimaera were generated using wild-type mice irradiated as recipients, reconstituted with control or CX3CR1-AHR bone marrow. Recipients of bone marrow were then rested for 3 weeks and thereafter treated with weekly tamoxifen gavages (4 mg) for another 3 weeks; after a total of 6 weeks, EAE was induced and tamoxifen administration continued weekly during EAE. Left, flow cytometry analysis of AHR expression in microglia and monocytes 21 days after EAE induction. Thin line depicts isotype control, thick line denotes AHR staining, and numbers indicate the percentage of AHR-positive cells. Representative of stainings of n = 3 independent mice per group. Right, EAE clinical course in bone marrow chimaera mice. n = 4 mice per group. h, Control and CX3CR1-AHR mice were treated with oral tamoxifen weekly starting from 5 weeks of age. EAE was induced at 8 weeks under continuation of weekly tamoxifen administration. Left, intracellular FACS staining for AHR in microglia and monocytes from at day 21 of EAE. Representative of stainings of n = 3 independent mice per group. Right, clinical course of control and CX3CR1-AHR bone marrow chimaera mice. Data in a, c, e–h are mean ± s.e.m. of n = 4 mice per group. P values were determined by two-sided Student’s t-test (a, c, e) or two-way ANOVA (g, h).
Extended Data Fig. 2 Topical and molecular regulation of TGFα and VEGF-B.
a, Ingenuity pathway analysis of differentially regulated pathways in astrocytes from n = 3 control versus CX3CR1-AHR mice per group during EAE. b, Tgfa and Vegfb expression determined by qPCR in microglia from brain, cerebellum and spinal cord 21 days after EAE induction (left). n = 8 mice per group. c, Predicted NF-κB and AHR responsive sites (NREs and XREs, respectively) in Vegfb and Tgfa promoters. d, Microglia were isolated by FACS sorting from control and CX3CR1-AHR mice during EAE. Ex vivo ChIP assay of NF-κB p65 or AHR binding to predicted binding sites in the Vegfb promoter. n = 3 mice per group. Data are representative of two independent experiments. e, Reporter assay using a construct in which the Vegfb promoter controls luciferase expression (pVegfb-Luc). Luciferase activity was measured in HEK293 cells 24 h after transfection with pVegfb-Luc, pTK-Renilla, and plasmids expressing AHR or NF-κB p65. Data are representative of two independent experiments with four biological replicates. f, Ex vivo ChIP assay as in d for AHR binding to the Tgfa promoter. n = 3 mice per group. Representative of two independent experiments. g, Reporter assays using a construct in which the Tgfa promoter controls luciferase expression (pTgfa-Luc). Luciferase activity was measured in HEK293 cells 24 h after transfection with pTgfa-Luc, pTK-Renilla, and plasmids expressing AHR or control. Data are representative of two independent experiments with three biological replicates. Data in a, d–g are mean ± s.e.m. P values were determined by one-way ANOVA followed by Tukey’s post-hoc test (b, d, e, f) or two-sided Student’s t-test (g).
Extended Data Fig. 3 TGFα and VEGF-B are regulated by AHR in highly purified astrocytes and microglia.
a, b, Mouse microglia were activated with lipopolysaccharide (LPS) in the presence or absence of the AHR inhibitor CH223191. After 24 h, activation medium was removed and substituted with fresh medium after extensive washes. Then 48 h later, microglia conditioned medium (MCM) was collected and applied to cultures of primary astrocytes. a, Gene expression in microglia 24 h after activation in the presence or absence of CH223191. b, Gene expression in astrocytes after 24 h exposure to MCM. Data are representative of two independent experiments with three biological replicates. c, Representative FACS stainings for CD11b and CD45 in primary astrocyte and microglia cultures. Numbers indicate percentages in respective gate. Data are representative of three independent experiments. d, Representative FACS stainings for GFAP and GLAST in astrocyte cultures as in b. Data are representative of three independent experiments. e, f, qPCR analysis of mRNA expression in astrocyte and microglia cultures. n = 4 independent cultures. Data are representative of two independent experiments with four biological replicates. g, Effect of TGFα and VEGF-B on gene expression in primary astrocytes activated with TNF and IL-1β, determined by pPCR after 24 h. Data are representative of three independent experiments with three biological replicates. h, i, Primary mouse astrocytes were activated with TNF and IL-1β and treated with TGFα or VEGF-B. After 24 h later, culture medium was substituted by fresh medium after extensive washes. Then 48 h later, ACM was added to mouse neurons (h) and oligodendrocytes (i) in culture, and cytotoxicity was determined by quantifying lactate dehydrogenase (LDH) release after 24 h. n = 3 biological replicates. Data are representative of two independent experiments. j, CD11b+Ly6Chi monocyte migration assay performed using ACM from astrocytes activated in the presence of TGFα or VEGF-B. n = 4 biological replicates. Data are representative of two independent experiments. k, qPCR analysis of Nos2 expression in microglia co-cultured with astrocytes activated in the presence of TGFα or VEGF-B. n = 3 biological replicates. Data are representative of two independent experiments. Data in b, e–k are mean ± s.e.m. P values were determined by two-sided Student’s t-test (b, e, f) or one-way ANOVA followed by Tukey’s post-hoc test (g–k).
Extended Data Fig. 4 Phenotypical and functional effects of knockdown of microglial TGFα and VEGF-B.
a, Quantification of astrocyte numbers in spinal cord sections of knockdown mice. SOX9-positive astrocytes per mm2 were quantified in spinal cord sections of four mice per group. b, IMARIS reconstruction of GFAP+ astrocytes in spinal cord sections as in a, and quantification of dendrite length, branches, volume, terminal points, and segments of n = 4 mice per group. c, d, qPCR analysis of Tgfa and Vegfb expression in sorted CNS-infiltrating inflammatory monocytes (c) and microglia (d) from mice injected with pCD11b-shControl, pCD11b-shTgfa, and pCD11b-shVegfb 7 days after EAE induction. Representative of two independent experiments with three biological replicates. e, qPCR analysis of Erbb1 and Flt1 expression in mice injected with pGFAP-shControl, pGFAP-shErbb1, and pCD11b-shFlt1 7 days after EAE induction. Representative of two independent experiments with three biological replicates. f, Left, flow cytometry analysis of VEGF-B and TGFα expression in microglia from mice injected with pCD11b-shControl, pCD11b-shTgfa, and pCD11b-shVegfb 7 days after EAE induction. Right, quantification of VEGF-B- and TGFα-positive microglia in n = 5 mice per group. Representative of two independent experiments with five biological replicates. g, Left, flow cytometry analysis of FLT-1 and ErbB1 expression in astrocytes from mice injected with pGFAP-shControl, pGFAP-shErbb1, and pCD11b-shFlt1 7 days after EAE induction. Right, quantification of FLT-1 and ErbB1-positive microglia in n = 5 mice per group. Representative of two independent experiments with five biological replicates. h, Naive mice were injected with lysolecithin, VEGF-B, or PBS into the corpus callosum by stereotaxic injection and 6 days later, brains were analysed by myelin staining. Representative of two independent experiments with five biological replicates. Data are mean ± s.e.m. P values were determined one-way ANOVA followed by Tukey’s post-hoc test (a–h).
Extended Data Fig. 5 Directionality of TGFα and VEGF-B signalling during EAE.
a, b, EAE development in wild-type mice injected with pGFAP-shControl, pGFAP-shErbb1 and pCD11b-shFlt1 (a), or pCD11b-shControl, pCD11b-shTgfa and pCD11b-shVegfb (b). Clinical course. n = 5 mice per group. Representative of two independent experiments with n = 5 mice per group. c, Left, flow cytometry analysis of TGFα and VEGF-B expression in astrocytes as in a. Right, quantification of cytokine-positive astrocytes. Data are mean ± s.e.m. and P values were determined by one way ANOVA followed by Tukey’s post-hoc test. Representative of two independent experiments with four biological replicates. d, Left, flow cytometry analysis of FLT-1 and ErbB1 expression in microglia as in b. Right, quantification of cell-surface receptor expression of microglia. Data are mean ± s.e.m. and P values were determined by one way ANOVA followed by Tukey’s post-hoc test. Representative of two independent experiments with four biological replicates.
Extended Data Fig. 6 Regulation and transcriptional effects of TGFα and VEGF-B during EAE.
a, b, NanoString analysis of mRNA expression in astrocytes from EAE mice injected with pCD11b-shVegfb or pCD11b-shTgfa (a) and pGFAP-shFlt1 or pGFAP-shErbb1 (b; see also Fig. 2k, l). Fold change in relative expression relative to control as determined by log2(shKD/shControl). shKD, shRNA knockdown. Representative of two independent experiments with pooled RNA isolated from n = 3 mice per group. c, Principal component analysis of gene expression in astrocytes isolated as in a and b. Representative of two independent experiments with pooled RNA isolated from n = 3 mice per group. d, Ingenuity pathway analysis of significantly regulated pathways from astrocytes as in a and b. Representative of two independent experiments with pooled RNA isolated from n = 3 mice per group. e, Left, representative flow cytometry plots depicting NF-κB p65 phosphorylation in wild-type astrocytes stimulated for 15 min with vehicle (top) or TNF or IL-1β (bottom) in the presence of TGFα, VEGF-B, or their combination. Numbers indicate percentage of FITC+ cells. Bar graphs depict quantification of FITC+ cells. Data are mean ± s.e.m. and P values were determined by one way ANOVA followed by Tukey’s post-hoc test. Representative of two independent experiments with four biological replicates. f, Primary mouse astrocytes were exposed to VEGF-B or vehicle and pharmacological blocker of NF-κB activation. RNA was obtained after 18 h and subjected to qPCR analyses for the indicated genes. Data are mean ± s.e.m. and P values were determined by one way ANOVA followed by Tukey’s post-hoc test. Representative of two independent experiments with three biological replicates. g, Primary mouse astrocytes were activated with TNF or IL-1β in the presence of VEGF-B or vehicle, and a pharmacological blocker of NF-κB activation. RNA was obtained after 18 h and subjected to qPCR analyses for the indicated genes. Data are mean ± s.e.m. and P values were determined by one way ANOVA followed by Tukey’s post-hoc test. Representative of two independent experiments with n = 3 biological replicates.
Extended Data Fig. 7 Role of AHR in astrocytes and microglia during EAE.
a–c, EAE was induced in control (WT), GfapcreAhrfl/fl (GFAP-AHR), or CX3CR1-AHR EAE mice. Starting from day 7, mice were injected daily intraperitoneally with indoxyl-3-sulfate (I3S), given a tryptophan-depleted diet (TDD), or kept on a control diet. Clinical course of EAE mice under treatment conditions as indicated. Representative of two independent experiments with n = 4 mice per group. d–f, EAE was induced in wild-type mice, which were treated with lentiviruses to knockdown AHR in astrocytes (pGFAP-shAhr) or microglia (pCD11b-shAhr). A noncoding RNA was used as a control. Flow cytometry quantification of AHR expression in astrocytes and microglia by FACS. d, Representative histograms of n = 4 mice per group. Numbers indicate percentage of AHR-positive cells; thin lines denote isotype control, thick lines denote AHR staining. e, Quantification of AHR-positive astrocytes and microglia as in d. Representative of two independent experiments with four biological replicates. f, EAE mice with knock down of AHR in astrocytes, microglia, or both as in d were subjected to daily I3S injections, TDD, or control diet conditions starting on day 14 after disease induction. Clinical course of n = 4 mice per group. Representative of two independent experiments with n = 4 mice per group. g, Quantification of CNS-infiltrating pro-inflammatory monocytes as determined by FACS at day 28 of EAE. Representative of two independent experiments with three biological replicates. Data are mean ± s.e.m. P values were determined by two-way ANOVA (a, f), or one way ANOVA followed by Tukey’s post-hoc test (e, g).
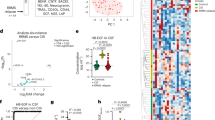
Extended Data Fig. 8 Dietary factors influence mouse and human TGFα and VEGF-B expression.
a, Ingenuity pathway analysis of NF-κB signalling comparing a TDD to a TDD plus Trp diet in control animals. Colours code for up- and downregulation of individual members in red (up) and blue (down). Normalized reads of two independent samples per group. b, mRNA expression determined by qPCR in from EAE mice as in Fig. 3a. Data are representative of two independent experiments with three replicates. c, Quantification of co-expression of AHR and CD14, VEGF-B and CD14, TGFα and CD14 in immunofluorescence stainings of human white matter brain tissue of NAWM, active, or chronic MS lesions for AHR (left), VEGF-B (middle), or TGFα (right), CD14 (green), and DAPI (blue). Data are representative of n = 12 fields from three distinct MS brains. d, Ratio of VEGF-B to TGFα intensities. Data are the ratio of mean values from Fig. 4e + s.e.m of n = 25 fields. Data in b and d are mean ± s.e.m. P values derived by one-way ANOVA followed by Tukey’s post-hoc test (b, d).
Supplementary information
Rights and permissions
About this article
Cite this article
Rothhammer, V., Borucki, D.M., Tjon, E.C. et al. Microglial control of astrocytes in response to microbial metabolites. Nature 557, 724–728 (2018). https://doi.org/10.1038/s41586-018-0119-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0119-x
This article is cited by
-
Distinct forebrain regions define a dichotomous astrocytic profile in multiple system atrophy
Acta Neuropathologica Communications (2024)
-
Bridging the gap: associations between gut microbiota and psychiatric disorders
Middle East Current Psychiatry (2024)
-
Gastrointestinal and brain barriers: unlocking gates of communication across the microbiota–gut–brain axis
Nature Reviews Gastroenterology & Hepatology (2024)
-
Microglia regulation of central nervous system myelin health and regeneration
Nature Reviews Immunology (2024)
-
Microbiota–gut–brain axis and its therapeutic applications in neurodegenerative diseases
Signal Transduction and Targeted Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.