Abstract
In vertebrate hearts, the ventricular trabecular myocardium develops as a sponge-like network of cardiomyocytes that is critical for contraction and conduction, ventricular septation, papillary muscle formation and wall thickening through the process of compaction1. Defective trabeculation leads to embryonic lethality2,3,4 or non-compaction cardiomyopathy (NCC)5. There are divergent views on when and how trabeculation is initiated in different species. In zebrafish, trabecular cardiomyocytes extrude from compact myocardium6, whereas in chicks, chamber wall thickening occurs before overt trabeculation7. In mice, the onset of trabeculation has not been described, but is proposed to begin at embryonic day 9.0, when cardiomyocytes form radially oriented ribs2. Endocardium–myocardium communication is essential for trabeculation, and numerous signalling pathways have been identified, including Notch2,8 and Neuregulin (NRG)4. Late disruption of the Notch pathway causes NCC5. Whereas it has been shown that mutations in the extracellular matrix (ECM) genes Has2 and Vcan prevent the formation of trabeculae in mice9,10 and the matrix metalloprotease ADAMTS1 promotes trabecular termination3, the pathways involved in ECM dynamics and the molecular regulation of trabeculation during its early phases remain unexplored. Here we present a model of trabeculation in mice that integrates dynamic endocardial and myocardial cell behaviours and ECM remodelling, and reveal new epistatic relationships between the involved signalling pathways. NOTCH1 signalling promotes ECM degradation during the formation of endocardial projections that are critical for individualization of trabecular units, whereas NRG1 promotes myocardial ECM synthesis, which is necessary for trabecular rearrangement and growth. These systems interconnect through NRG1 control of Vegfa, but act antagonistically to establish trabecular architecture. These insights enabled the prediction of persistent ECM and cardiomyocyte growth in a mouse NCC model, providing new insights into the pathophysiology of congenital heart disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Moorman, A. F. & Christoffels, V. M. Cardiac chamber formation: development, genes, and evolution. Physiol. Rev. 83, 1223–1267 (2003).
Grego-Bessa, J. et al. Notch signaling is essential for ventricular chamber development. Dev. Cell 12, 415–429 (2007).
Stankunas, K. et al. Endocardial Brg1 represses ADAMTS1 to maintain the microenvironment for myocardial morphogenesis. Dev. Cell 14, 298–311 (2008).
Lai, D. et al. Neuregulin 1 sustains the gene regulatory network in both trabecular and nontrabecular myocardium. Circ. Res. 107, 715–727 (2010).
Luxán, G. et al. Mutations in the NOTCH pathway regulator MIB1 cause left ventricular noncompaction cardiomyopathy. Nat. Med. 19, 193–201 (2013).
Staudt, D. W. et al. High-resolution imaging of cardiomyocyte behavior reveals two distinct steps in ventricular trabeculation. Development 141, 585–593 (2014).
Icardo, J. M. & Fernandez-Terán, A. Morphologic study of ventricular trabeculation in the embryonic chick heart. Acta Anat. 130, 264–274 (1987).
D’Amato, G. et al. Sequential Notch activation regulates ventricular chamber development. Nat. Cell Biol. 18, 7–20 (2016).
Camenisch, T. D. et al. Disruption of hyaluronan synthase-2 abrogates normal cardiac morphogenesis and hyaluronan-mediated transformation of epithelium to mesenchyme. J. Clin. Invest. 106, 349–360 (2000).
Hatano, S. et al. Versican/PG-M is essential for ventricular septal formation subsequent to cardiac atrioventricular cushion development. Glycobiology 22, 1268–1277 (2012).
Zhou, Z. et al. The cerebral cavernous malformation pathway controls cardiac development via regulation of endocardial MEKK3 signaling and KLF expression. Dev. Cell 32, 168–180 (2015).
Gessler, M. et al. Mouse gridlock: no aortic coarctation or deficiency, but fatal cardiac defects in Hey2 −/− mice. Curr. Biol. 12, 1601–1604 (2002).
VanDusen, N. J. et al. Hand2 is an essential regulator for two Notch-dependent functions within the embryonic endocardium. Cell Rep. 9, 2071–2083 (2014).
Ribatti, D. & Crivellato, E. “Sprouting angiogenesis”, a reappraisal. Dev. Biol. 372, 157–165 (2012).
Iivanainen, E. et al. Intra- and extracellular signaling by endothelial neuregulin-1. Exp. Cell Res. 313, 2896–2909 (2007).
Ferrara, N. et al. Heterozygous embryonic lethality induced by targeted inactivation of the VEGF gene. Nature 380, 439–442 (1996).
Bourke, L. M. et al. Loss of rearranged L-Myc fusion (RLF) results in defects in heart development in the mouse. Differentiation 94, 8–20 (2017).
Britsch, S. et al. The ErbB2 and ErbB3 receptors and their ligand, neuregulin-1, are essential for development of the sympathetic nervous system. Genes Dev. 12, 1825–1836 (1998).
Daxinger, L. et al. An ENU mutagenesis screen identifies novel and known genes involved in epigenetic processes in the mouse. Genome Biol. 14, R96 (2013).
Davy, A. & Soriano, P. Ephrin-B2 forward signaling regulates somite patterning and neural crest cell development. Dev. Biol. 304, 182–193 (2007).
Chen, H. et al. BMP10 is essential for maintaining cardiac growth during murine cardiogenesis. Development 131, 2219–2231 (2004).
Koni, P. A. et al. Conditional vascular cell adhesion molecule 1 deletion in mice: impaired lymphocyte migration to bone marrow. J. Exp. Med. 193, 741–754 (2001).
Gustafsson, E., Brakebusch, C., Hietanen, K. & Fässler, R. Tie-1-directed expression of Cre recombinase in endothelial cells of embryoid bodies and transgenic mice. J. Cell Sci. 114, 671–676 (2001).
Wu, B. et al. Endocardial cells form the coronary arteries by angiogenesis through myocardial–endocardial VEGF signaling. Cell 151, 1083–1096 (2012).
Radtke, F. et al. Deficient T cell fate specification in mice with an induced inactivation of Notch1. Immunity 10, 547–558 (1999).
Han, H. et al. Inducible gene knockout of transcription factor recombination signal binding protein-J reveals its essential role in T versus B lineage decision. Int. Immunol. 14, 637–645 (2002).
Haigh, J. J. et al. Cortical and retinal defects caused by dosage-dependent reductions in VEGF-A paracrine signaling. Dev. Biol. 262, 225–241 (2003).
Watanabe, Y. et al. Activation of Notch1 signaling in cardiogenic mesoderm induces abnormal heart morphogenesis in mouse. Development 133, 1625–1634 (2006).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Shalaby, F. et al. Failure of blood-island formation and vasculogenesis in Flk-1-deficient mice. Nature 376, 62–66 (1995).
Kanzler, B., Kuschert, S. J., Liu, Y. H. & Mallo, M. Hoxa-2 restricts the chondrogenic domain and inhibits bone formation during development of the branchial area. Development 125, 2587–2597 (1998).
Del Monte, G., Grego-Bessa, J., González-Rajal, A., Bolós, V. & De La Pompa, J. L. Monitoring Notch1 activity in development: evidence for a feedback regulatory loop. Dev. Dyn. 236, 2594–2614 (2007).
del Monte, G. et al. Differential Notch signaling in the epicardium is required for cardiac inflow development and coronary vessel morphogenesis. Circ. Res. 108, 824–836 (2011).
Soufan, A. T. et al. Three-dimensional reconstruction of gene expression patterns during cardiac development. Physiol. Genomics 13, 187–195 (2003).
de Boer, B. A. et al. The interactive presentation of 3D information obtained from reconstructed datasets and 3D placement of single histological sections with the 3D portable document format. Development 138, 159–167 (2011).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Nishiyama, T. et al. Functional analysis of an established mouse vascular endothelial cell line. J. Vasc. Res. 44, 138–148 (2007).
Maier, M. M. & Gessler, M. Comparative analysis of the human and mouse Hey1 promoter: Hey genes are new Notch target genes. Biochem. Biophys. Res. Commun. 275, 652–660 (2000).
Sturm, K. & Tam, P. P. Isolation and culture of whole postimplantation embryos and germ layer derivatives. Methods Enzymol. 225, 164–190 (1993).
Baldwin, H. S. & Solursh, M. Degradation of hyaluronic acid does not prevent looping of the mammalian heart in situ. Dev. Biol. 136, 555–559 (1989).
Budi, E. H., Patterson, L. B. & Parichy, D. M. Embryonic requirements for ErbB signaling in neural crest development and adult pigment pattern formation. Development 135, 2603–2614 (2008).
Acknowledgements
We thank D. Y. Stainier, J. S. Rasouli, A. V. Cherian, J. M. Polo, F. J. Rossello, G. Chapman, V. Sardesai, B. Shewale, J. O’Rourke, S. Tyler, L. Madigan, A. Ahmad, C. Onie and M. Tondl for their contributions, and J. Moreau and E. Forte for critical review of the manuscript. This work was funded by ARC (DP160104858, DP140101067), NHMRC (APP1118576; 1074386; 573732; 573705; 1032851, CDF 1049980), Foundation Leducq (15 CVD 03) and Perpetual Trust (FR2012/0435). J.L.d.l.P. received grants SAF2016-78370-R and CB16/11/00399 (CIBER CV) from MEIC.
Author information
Authors and Affiliations
Contributions
G.d.M.-N. was involved in the conceptualization, methodology, investigation, validation, analysis, writing, project administration, visualization and supervision of this project; M.R. carried out bioinformatics analyses; A.A.S.A. provided experimental support; B.W., B.Z., A.A., G.D., E.T., L.M.B., S.K.H., M.E.P., R.H.A., H.C., W.S. and J.L.d.l.P. provided resources. R.P.H. provided supervision, acquired funding and was involved in writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Analysis of trabeculation by 3D morphological time course reconstructions and electron microscopy.
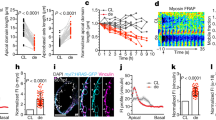
Related to Fig. 1. a–g, Time-course of left ventricle 3D reconstructions (see Supplementary Figs. 1–7) of wild-type mice (a–c, e, f), Tie2creNotch1fl/fl mice at E8.5 (d) and Nrg1tm/tm mice at E9.0 (g). In all panels: endocardium (green), myocardium (red, transparent red in a–g (left)), ECM (blue, transparent blue in the right panels of a, c, d). Lateral view of the whole heart (a–g, left) or endocardium only (a–g, second left) showing that the endocardial behaviours described in the study occur exclusively in the ventricular outer curvature (white arrows) and not in the inner curvature (arrowheads), and the progressive ECM reduction seen as the distance between the endocardium and the ventricular outer layer as trabeculation proceeds (black arrows). This distance is greater in Tie2creNotch1fl/fl (d) and smaller in Nrg1tm/tm (g) hearts. a–g, Second from the right, view towards the endocardium from the outer curvature. a–d, Second from the right, note the formation of endocardial ridges between the leading touchdowns (white arrows) generating endocardial domes (white asterisks). d, Second from the right, incomplete touchdown formation and lack of endocardial domes in Tie2creNotch1fl/fl hearts. e–g, Second from the right, outer curvature view showing the contact points between endocardium and myocardium (green, white arrow) without ECM, and ECM bubbles (blue, black arrow) in wild-type (e, f), and the increased endocardial contacts and decreased ECM in Nrg1tm/tm (g) hearts. a–g, Right, luminal view of whole heart (a) or bisected hearts (b–g) showing ventricular chamber segmentation by endocardial domes/ECM bubbles (asterisks) in wild-type hearts (a–c, e, f) and its defects in Tie2creNotch1fl/fl (d) and Nrg1tm/tm (g) hearts. h–j, Electron microscopy images of wild-type ventricles at E9.0 showing endocardium (endoc) and compact myocardium interaction at the trabecular base. h, Desmosomes (arrow). i, Endocardial cell in close apposition to a cardiomyocyte (CM). j, Cellular projections originating from both cell types. k–m, Area quantifications on wild-type left ventricle showing trabecular myocardium area normalized to chamber perimeter (k), total ECM area (l) and ECM area of apical and basal trabecular regions (m) normalized to trabecular myocardium. n–p, ISH of Hopx (n), Gja5 (o) and Bmp10 (p) in heart sections of wild-type embryos at E8.0. Marker expression (arrow), no marker expression (arrowhead). q, r, Quantification of touchdown numbers in Nrg1tm/tm (q) and Tie2creNotch1fl/fl (r) compared to wild-type embryos at E8.5 and E8.5–E9.0. a–r, n = 3 independent embryos per genotype, except for k, l, n = 4 E8.0 (8–9 ps) and E8.5 (13–15 ps). Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. k, 1, P = 0.035; 2, P = 0.025; 3, P = 0.046 4, P = 0.023; 5, P = 0.037; 6, P = 0.041. l, 1, P = 0.046; 2, P = 0.046; 3, P = 0.0003; 4, P = 0.013; 5, P = 0.0025; 6, P = 0.026; 7, P = 0.005. m, 1, P = 0.038; 2, P = 0.013; 3, P = 0.0048; 4, P = 0.041; 5, P = 0.012; 6, P = 0.0024. r, 1, P = 0.018; 2, P = 0.0048. Scale bars, as indicated (h–j) and 20 μm in n–p.
Extended Data Fig. 2 Time course expression analysis of ECM synthesis markers.
Related to Figs. 1, 2. a–m, Detailed time-course analysis of marker expression using ISH and immunofluorescence in embryonic left ventricle at E8.0, E8.5, E9.0, E9.5, E10.5, E11.5, E13.5 and E16.5 in wild-type embryos (a, c, e, g–m) as well as in Nrg1tm/tm mutants (b, d, f) relative to somite-matched wild-type embryos at E8.0, E8.5, E9.0 and E9.5. a, b, e, f, Analysis of Has2 (a, b) and Vcan (e, f) by ISH in wild-type (a, e) and Nrg1tm/tm embryos (b, f). c, d, Immunofluorescence analysis of HABP in wild-type (c) and Nrg1tm/tm (d) embryos. g–m, Immunofluorescence analysis of CD44 (g), fibronectin (h), perlecan (i), aggrecan (j), laminin (k), collagen type I (l) and collagen type IV (m) in wild-type embryos. Perlecan showed a similar pattern of enrichment in trabecular myocardium and endocardium as HABP and fibronectin, although expression persisted after termination. Aggrecan was expressed in both compact and trabecular myocardium and only became restricted to trabeculae after termination. Laminin and collagen types I and IV did not show restriction. In all panels showing gene/protein expression: marker expression (arrow), reduced marker expression (arrowhead), increased marker expression (thick arrow). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue). a–m, n = 3 independent embryos were used per genotype. Scale bars, 20 μm (E8.0), 25 μm (E8.5), 30 μm (E9.0, E9.5), 50 μm (E10.5), 60 μm (E11.5), 120 μm (E13.5) and 200 μm (E16.5).
Extended Data Fig. 3 Time course expression analysis of ECM degradation, NOTCH, NRG1 and VEGFA pathway markers.
Related to Figs. 1–3. a–m, Detailed time-course analysis of marker expression using ISH and immunofluorescence in embryonic left ventricle at E8.0, E8.5, E9.0, E9.5, E10.5, E11.5, E13.5 and E16.5 in wild-type embryos (a–e, g, i, k–m) as well as in Tie2creNotch1fl/fl (f, h) and Nrg1tm/tm (j) embryos relative to somite-matched wild-type embryos at E8.0, E8.5, E9.0 and E9.5. Data are organized according to respective ECM degradation, Notch, neuregulin-1 (NRG1) and VEGFA pathways. a–d, Analysis of ECM degradation markers Adamts1 (a), Mmp2 (b) and Hyal2 (c) by ISH and neoversican (d) by immunofluorescence. e–h, NRG1 pathway analysis of Nrg1 by RNA-scope (e, f) and pERBB2 (g, h) by immunofluorescence in wild-type (e, g) and Tie2creNotch1fl/fl (f, h) embryos. i–k, Notch pathway analysis of N1ICD (i, j) in wild-type (i) and Nrg1tm/tm (j) embryos, and DLL4 in wild-type embryos (k) by immunofluorescence. l, m, VEGFA pathway analysis of Vegfa by ISH (l) and VEGFR2 by immunofluorescence (m). In all panels showing gene/protein expression: marker expression (arrow), reduced marker expression (arrowhead), increased marker expression (thick arrow). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue). a–m, n = 3 independent embryos were used for each genotype. Scale bars, 20 μm (E8.0), 25 μm (E8.5), 30 μm (E9.0, E9.5), 50 μm (E10.5), 60 μm (E11.5), 120 μm (E13.5) and 200 μm (E16.5).
Extended Data Fig. 4 Morphological and molecular analysis of mutants with trabeculation defects.
Related to Figs. 1–3. Histological and marker analysis of left ventricle in mutant embryos and somite-matched wild-type control embryos at indicated stages. a–d, Analysis of wild-type and Erbb2−/− mutants showing histology (a), normalized trabecular and ECM area quantifications (b), immunofluorescence for N1ICD (c) and fold change in number of N1ICD+ cells (d). e, f, Immunofluorescence analysis of pERBB2 in Erbb2−/− (e (right)) and Nrg1tm/tm (f (right)) mutant embryos compared to wild-type (e, f). g–j, Analysis of wild-type and Tie1creVEGFR2fl/fl mutants showing histology (g), normalized tissue area quantifications (h) and immunofluorescence of VEGFR2 (i) and pAKT (j). k–p, Analysis of wild-type and mutant histology and normalized tissue area quantifications for the following strains: Tie2creRbpjfl/fl (k, l), Efnb2gfp/gfp (m, n) and Bmp10−/− (o, p). q–u, Time-course analysis of histology using Alcian blue staining of Nrg1tm/tm (q–u (middle)) and Tie2creNotch1fl/fl (q–u (right)) embryos relative to somite-matched wild-type (q–u (left)) at indicated stages. In all H&E panels, trabecular myocardium (black arrows), ECM (white arrows). In all panels showing molecular expression: marker expression (arrows), reduced marker expression (arrowheads). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue).The cardiac lumen is indicated by ‘l’. Trabecular myocardium (black arrows), ECM (white arrows). a–u, n = 3 independent embryos per genotype. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. b, 1, P = 0.018; 2, P = 0.047. d, 1, P = 0.034. h, 1, P = 0.04; 2, P = 0.033. l, 1, P = 0.035. Scale bars, 20 μm (q), 25 μm (r), 30 μm (a, c, e–g, i–k, m, o, s) and 35 μm (t, u).
Extended Data Fig. 5 Marker analysis in Nrg1tm/tm and Tie2creNotch1fl/fl mutant strains.
Related to Fig. 2. a–i, Analysis of markers in left ventricle sections by ISH, immunofluorescence and qPCR in Nrg1tm/tm (a–i (second left)), Tie2-Notch1fl/fl (a–i (right)) mutant embryos and somite-matched wild-type embryo (a–h (left, second from the right)) at indicated optimal time points. Data are organized around markers for chamber myocardium (Mest, Nppa), trabecular myocardium (Bmp10, Gja5, Hopx) and compact myocardium (Hey2, Mycn) as indicated. ISH (a–h); qPCR (i). i, Panels show qPCR analysis of the genes described in a–h in Nrg1tm/tm mutants at E9.0 and E10.0 (i (left column)) and in Tie2creNotch1fl/fl mutants at E8.5 and E9.5 (i (right column)), the latter time points in each case being beyond the phenotypic onset stages for each mutant. In all panels showing gene/protein expression: marker expression (arrow), reduced marker expression (arrowhead), increased marker expression (thick arrow). a–h, n = 3 independent embryos per genotype. i, n = 3 biological replicates each evaluated for three experimental replicates. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. i, Top left, 1, P = 0.013; bottom left, 1, P = 0.0015; 2, P = 0.018; 3, P = 0.002; 4, P = 0.0027; 5, P = 0.048; 6, P = 0.0044; bottom right, 1, P = 0.04; 2, P = 0.0098; 3, P = 0.047; 4, P = 0.012; 5, P = 0.022; 6, P = 0.0029; 7, P = 0.02. Scale bars, 30 μm (a–h).
Extended Data Fig. 6 Analysis of ECM synthesis and degradation markers, and proliferation, in Nrg1tm/tm and Tie2creNotch1fl/fl mutants.
Related to Fig. 2. a–k, m, o–q, Analysis of ECM synthesis (a–h) and degradation (i–k) markers, BRG1 (m), Notch pathway (o–p) and VEGFA pathway (q) in left ventricle of Nrg1tm/tm mutants at E9.0 (a–q (second left)) and Tie2creNotch1fl/fl mutants at E8.5 (a–q (right)) relative to wild-type embryos (a–q (left, second from the right)) showing immunofluorescence analysis of HABP (a), CD44 (b), fibronectin (c), perlecan (d), aggrecan (e), laminin (f), collagen type I (g), collagen type IV (h), BRG1 (m), DLL4 (o) and VEGFR2 (q); ISH for Mmp2 (i), Adamts5 (j), Hyal2 (k) and Efnb2 (p); and qPCR (l). n, Percentage of BRG1 positive cells compared to the total number of cells in the left ventricle endocardium and myocardium of Nrg1tm/tm(n (left)) and Tie2creNotch1fl/fl (n (right)) mutants. r–u, Proliferation analysis by Ki-67 immunofluorescence of Nrg1tm/tm mutants at E9.0 (r (second left)) and E9.5–E10.0 (t (second left)); Tie2creNotch1fl/fl mutants at E8.5 (r (right)) and E9.5 (t (right)), relative to wild-type embryos (r, t, (left, second from the right); and the related quantifications of total myocardium proliferation and the proportion of trabecular cardiomyocytes compared to the total cardiomyocytes (s, u). Gene/protein expression panels: expression (arrow), reduced expression (arrowhead). In all immunofluorescence: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue). a–u, n = 3 independent embryos per genotype. l, n = 3 biological replicates each evaluated for three experimental replicates. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. l, Left, 1, P = 0.0066; 2, P = 0.0038; 3, P = 0.017, 4, P = 0.035; 5, P = 0.036; right, 1, P = 0.0039; 2, P = 0.039; 3, P = 0.00053; 4, P = 0.0074; 5, P = 0.022. n, 1, P = 0.006. s, Left, 1, P = 0.029; right, 1, P = 0.012. u, Left, 1, P = 0.045; 2, P = 0.017; right, 1, P = 0.04; 2, P = 0.0031. Scale bars, 55 μm (a–k, m, o–r) and 40 μm (t).
Extended Data Fig. 7 Manipulation of ECM dynamics in RBEC assays and marker analysis of Tie2creNotch1GOF mutant embryos.
Related to Fig. 2. a–d, H&E analysis of wild-type and Tie2creNotch1fl/fl embryos treated with 20 turbidity reducing units (TRU) of hyaluronidase (HYAL) or PBS as control (a, b) or wild-type and Nrg1tm/tm embryos treated with 20 μM GM6001 or DMSO (c, d) in RBECs for 24 h, and accompanying ECM and trabecular area quantifications normalized to chamber perimeter (b, d). e–g, H&E of Tie2creNotchGOF embryos (e, f (right)) compared to wild-type embryos (e, f (left)) at E8.5 and E10.0 with morphological quantification of ECM and trabecular areas normalized to chamber perimeter (g). h–q, Molecular analysis of Tie2creNotchGOF embryos relative to wild-type embryos at E8.5 (h–l) and E9.5 (m–q): ISH of Adamts1 (h, m) and Has2 (l, p); immunofluorescence analysis of neoversican (i, n) and pERBB2 (k); RNA-scope of Nrg1 (o); qPCR analysis of the respective markers (j, q). Gene/protein expression panels: expression (arrow), reduced expression (arrowhead), increased expression (thick arrow). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue). e–i, k–p, n = 3 independent embryos were used; a–d, n = 5, except in d, n = 6 wild-type embryos treated with GM6001. j, q, n = 3 biological replicates each evaluated for three experimental replicates. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. b, 1, P = 1.99 × 10−6; 2, P = 0.025; 3, P = 9.28 × 10−6; 4, P = 0.00085; 5 = 2.83 × 10−5; 6, P = 0.00067; 7, P = 0.026; 8, P = 0.00014. d, 1, P = 0.024; 2, P = 0.0024; 3, P = 0.018; 4, P = 0.000021; 5, P = 0.016; 6, P = 0.022; 7, P = 0.028. g, Top, 1, P = 0.018; 2, P = 0.017; bottom, 1, P = 0.0071; 2, P = 0.03. j, 1, P = 0.03; 2, P = 0.041. q, 1, P = 0.034; 2, P = 0.00026; 3, P = 0.025; 4 = 0.038. Scale bars, 55 μm (a, c), 30 μm (e, h–l) and 40 μm (m–p).
Extended Data Fig. 8 Genetic and chemical mutant rescue assays.
Related to Figs. 2, 3. a–d, Analysis of wild-type, Nrg1tm/tm, Tie2creRbpjfl/fl and Tie2creRbpjfl/flNrg1tm/tm mutant embryos showing histology (a), normalized ECM and trabecular area quantifications (b), immunofluorescence of N1ICD (c) and fold change in the number of N1ICD+ cells (d). e–g, Analysis of wild-type, Nrg1tm/tm, Tie2creNotch1fl/fl and Tie2creNotch1fl/flNrg1tm/tm mutant embryos showing histology (e), normalized ECM and trabecular area quantifications (f) and immunofluorescence of N1ICD (g). Note the persistent N1ICD staining in the Tie2creRbpjfl/flNrg1tm/tm mutant due to mosaic Cre-mediated deletion of Notch1. Here, N1ICD appears to nucleate touchdowns. Cre-mediated deletion was complete in Tie2creNotch1fl/flNrg1tm/tm mutants at E9.0, although mosaicism in Cre-recombination at an earlier time point may allow N1ICD restriction and nucleation of touchdowns. h, i, H&E of Nrg1tm/tm and wild-type embryos treated with 25 nM NRG1 or PBS as control for 24 h (h) with associated area quantifications (i). j–l, Immunofluorescence of N1ICD (j) and DLL4 (k) with quantification the number of N1ICD+ cells (l). m–q, Analysis of wild-type and Tie2creNotch1fl/fl embryos treated with 10 μM SU5416 or DMSO control for 24 h, showing histology (m), ECM and trabecular area quantifications (n) and immunofluorescence in SU5416- and control-treated wild-type embryos of N1ICD (o) and DLL4 (p), with fold change in the number of N1ICD+ cells (q). In H&E panels: trabecular myocardium (black arrows), ECM (white arrows). In all panels showing molecular expression: expression (arrows), reduced expression (arrowheads), increased expression (thick arrows). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue). a–f, j–l, o–q, n = 3 independent embryos were used; h, i, m, n, n = 5, except in h, i, n = 6 wild-type embryos treated with NRG1. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. b, 1, P = 0.013; 2, P = 0.0003; 3, P = 0.012; 4, P = 0.009; 5, P = 0.009; 6, P = 0.029; 7, P = 0.033. d, 1, P = 0.024; 2, P = 0.03. f, 1, P = 0.039; 2, P = 0.023; 3, P = 0.034; 4, P = 0.012; 5, P = 0.009; 6, P = 0.045; 7, P = 0.034. i, 1, P = 0.0014; 2, P = 0.0006; 3, P = 0.03; 4, P = 0.0004; 5, P = 0.0046; 6, P = 0.022. l, 1, P = 0.02; 2, P = 0.0042. n, 1, P = 0.0025; 2, P = 0.024; 3, P = 0.016; 4, P = 0.0033; 5, P = 0.00015; 6, P = 0.0017. q, 1, P = 0.022. Scale bars, 50 μm (a, c, e, g, h, j, k, m, o, p).
Extended Data Fig. 9 Analysis of deletion efficiency of the R26R-LacZ Cre-reporter using Tie2cre and Nfatc1cre driver lines and quantification analysis of late NOTCH1 loss-of-function and gain-of-function mutants.
Related to Figs. 1–3. a–h, Time-course analysis of staining of β-galactosidase in whole-mount embryos and representative sections from Tie2creR26R-LacZ and Nfatc1creR26R-LacZ embryos at indicated stages. i–l, Immunofluorescence analysis of N1ICD in Tie2creNotch1fl/fl embryos at 8 ps and 14 ps, and in Nfatc1creNotch1fl/fl embryos at 14 ps and 20 ps, compared to wild-type embryos, as indicated. m–p, Quantification of normalized ECM and trabecular myocardium areas measuring the total, apical or basal trabecular regions in Nfatc1creNotch1fl/fl (m, n) and Nfatc1creNotch1GOF (o, p) mutant embryos. In all panels showing gene/protein expression: marker (arrow), reduced expression (arrowhead), increased expression (thick arrow). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium stained for SMA (green dulled mask); nuclei (DAPI, blue). a–p, n = 3 independent embryos were used. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. m, Top, 1, P = 0.0057; 2, P = 0.021; middle, 1, P = 0.048; 2, P = 0.045; 3, P = 0.046; bottom, 1, P = 0.018; 2, P = 0.041. n, Top, 1, P = 0.0099; 2, P = 0.043; 3, P = 0.025; 4, P = 0.012; middle, 1, P = 0.03; bottom, 1, P = 0.036; 2, P = 0.030. o, Top, 1, P = 0.032; 2, P = 0.035; middle, 1, P = 0.025; 2, P = 0.0048; bottom, 1, P = 0.042. p, Top, 1, P = 0.014; middle, 1, P = 0.049; bottom, 1, P = 0.048; 2, P = 0.034. Scale bars, 250 μm (a, b (left)), 50 μm (a, b (right)), 340 μm (c, d (left)), 65 μm (c, d (right)), 360 μm (e, f (left)), 70 μm (e, f (right)), 430 μm (g, h (left)), 80 μm (g, h (right), i–k) and 100 μm (l).
Extended Data Fig. 10 Marker analysis in Nfatc1creNotch1fl/fl, Nfatc1creNotch1GOF and RlfD28/D28 mutant strains.
Related to Fig. 3. a–p, Analysis of markers in left ventricle sections of Nfatc1creNotch1fl/fl (a–p (second left)) and Nfatc1creNotch1GOF (a–p (right)) mutant embryos and somite-matched wild-type embryos (a–p (left, second from the right)). Data are organized around markers for chamber myocardium (Mest, Nppa), trabecular myocardium (Bmp10, Gja5, Hopx), compact myocardium (Hey2, Mycn), ECM synthesis (CD44, Vcan), ECM degradation (Adamts1), NRG1 pathway (Nrg1) and VEGFA pathway (Vegfa, VEGFR2, pAKT), and proliferation (Ki-67) as indicated. ISH (a–h, j, l); RNA-scope (k); immunofluorescence (i, m, n, p). o, qPCR analysis of the genes described in a–n in Nfatc1creNotch1fl/fl mutants at E10.0 (left) and Nfatc1creNotch1GOF mutants at E10.5 (right). q, Quantification of myocardial proliferation as the ratio of the number of Ki-67 positive cardiomyocytes versus the total number of cardiomyocytes and the proportion of trabecular cardiomyocytes out of the total number of cardiomyocytes in left ventricle from Nfatc1creNotch1fl/fl mutants at E10.0 (left two panels) and Nfatc1creNotch1GOF mutants at E10.5 (right two panels). r–v, Analysis of wild-type and RlfD28/D28 mutants showing right ventricle (RV) histology at E14.5 (r), normalized ECM area quantification in both ventricles (s), immunofluorescence of neoversican in the right ventricle (t) and pERBB2 in both ventricles (v), and ISH of Has2 in the right ventricle (u) at E13.5. In all panels showing gene/protein expression: expression (arrow), reduced expression (arrowhead), increased expression (thick arrow). In all immunofluorescence images: markers associated with endocardium or ECM (red); markers associated with myocardium (orange); myocardium is stained for SMA (green dulled mask); nuclei (DAPI, blue). a–n, p–q, t–v, n = 3 independent embryos were used; s, n = 4 E11.5 and wild-type E14.5, n = 5 E13.5, n = 6 RlfD28/D28 E14.5. o, n = 3 biological replicates each evaluated for three experimental replicates. Quantitative data are shown as mean ± s.e.m. Two-sided Student’s t-tests (without corrections for multiple comparisons) were used. Significant comparisons are indicated by numbers as follows. o, Left, 1, P = 0.047; 2, P = 0.04; 3, P = 0.0029; 4, P = 0.044; 5, P = 0.041; 6, P = 0.017; right, 1, P = 0.0037; 2, P = 0.025; 3, P = 0.036; 4, P = 0.017; 5, P = 0.00057; 6, P = 0.017. s, 1, P = 0.045; 2, P = 0.026; 3, P = 0.042; 4 = 0.0033. Scale bars, 50 μm (a–n, p, unless otherwise indicated), 60 μm (h–n, right two panels) 85 μm (t–v) and 150 μm (r).
Supplementary information
Supplementary Information
This file contains full descriptions for Supplementary Figures 1-7.
Supplementary Figure 1
This file contains a 3D model of a whole E8.0 WT heart.
Supplementary Figure 2
This file contains a 3D model of WT E8.0-E8.5 left ventricle
Supplementary Figure 3
This file contains a 3D model of WT E8.5 left ventricle
Supplementary Figure 4
This file contains a 3D model of WT E9.0 left ventricle
Supplementary Figure 5
This file contains a 3D model of WT E9.0-E9.5 left ventricle
Supplementary Figure 6
This file contains a 3D model of WT VS Tie2-cre;Notch1fl/fl E8.5 left ventricle
Supplementary Figure 7
This file contains a 3D model of WT VS Nrg1tm/tm E9.0 left ventricle
Rights and permissions
About this article
Cite this article
del Monte-Nieto, G., Ramialison, M., Adam, A.A.S. et al. Control of cardiac jelly dynamics by NOTCH1 and NRG1 defines the building plan for trabeculation. Nature 557, 439–445 (2018). https://doi.org/10.1038/s41586-018-0110-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0110-6
This article is cited by
-
A novel gene-trap line reveals the dynamic patterns and essential roles of cysteine and glycine-rich protein 3 in zebrafish heart development and regeneration
Cellular and Molecular Life Sciences (2024)
-
Endothelial deletion of PTBP1 disrupts ventricular chamber development
Nature Communications (2023)
-
miR-455-5p promotes pathological cardiac remodeling via suppression of PRMT1-mediated Notch signaling pathway
Cellular and Molecular Life Sciences (2023)
-
Myocardial Biomechanics and the Consequent Differentially Expressed Genes of the Left Atrial Ligation Chick Embryonic Model of Hypoplastic Left Heart Syndrome
Annals of Biomedical Engineering (2023)
-
MiRNA sequencing of Embryonic Myogenesis in Chengkou Mountain Chicken
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.