Abstract
Despite intense interest in discovering drugs that cause G-protein-coupled receptors (GPCRs) to selectively stimulate or block arrestin signalling, the structural mechanism of receptor-mediated arrestin activation remains unclear1,2. Here we reveal this mechanism through extensive atomic-level simulations of arrestin. We find that the receptor’s transmembrane core and cytoplasmic tail—which bind distinct surfaces on arrestin—can each independently stimulate arrestin activation. We confirm this unanticipated role of the receptor core, and the allosteric coupling between these distant surfaces of arrestin, using site-directed fluorescence spectroscopy. The effect of the receptor core on arrestin conformation is mediated primarily by interactions of the intracellular loops of the receptor with the arrestin body, rather than the marked finger-loop rearrangement that is observed upon receptor binding. In the absence of a receptor, arrestin frequently adopts active conformations when its own C-terminal tail is disengaged, which may explain why certain arrestins remain active long after receptor dissociation. Our results, which suggest that diverse receptor binding modes can activate arrestin, provide a structural foundation for the design of functionally selective (‘biased’) GPCR-targeted ligands with desired effects on arrestin signalling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Violin, J. D., Crombie, A. L., Soergel, D. G. & Lark, M. W. Biased ligands at G-protein-coupled receptors: promise and progress. Trends Pharmacol. Sci. 35, 308–316 (2014).
DeWire, S. M. & Violin, J. D. Biased ligands for better cardiovascular drugs: dissecting G-protein-coupled receptor pharmacology. Circ. Res. 109, 205–216 (2011).
Palczewski, K., Buczyłko, J., Imami, N. R., McDowell, J. H. & Hargrave, P. A. Role of the carboxyl-terminal region of arrestin in binding to phosphorylated rhodopsin. J. Biol. Chem. 266, 15334–15339 (1991).
Gurevich, V. V. & Benovic, J. L. Visual arrestin interaction with rhodopsin. Sequential multisite binding ensures strict selectivity toward light-activated phosphorylated rhodopsin. J. Biol. Chem. 268, 11628–11638 (1993).
Gurevich, V. V. & Gurevich, E. V. The structural basis of arrestin-mediated regulation of G-protein-coupled receptors. Pharmacol. Ther. 110, 465–502 (2006).
Kang, Y. et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523, 561–567 (2015).
Zhou, X. E. et al. Identification of phosphorylation codes for arrestin recruitment by G protein-coupled receptors. Cell 170, 457–469 (2017).
Nobles, K. N., Guan, Z., Xiao, K., Oas, T. G. & Lefkowitz, R. J. The active conformation of beta-arrestin1: direct evidence for the phosphate sensor in the N-domain and conformational differences in the active states of beta-arrestins1 and -2. J. Biol. Chem. 282, 21370–21381 (2007).
Shukla, A. K. et al. Structure of active β-arrestin-1 bound to a G-protein-coupled receptor phosphopeptide. Nature 497, 137–141 (2013).
Kumari, P. et al. Functional competence of a partially engaged GPCR-β-arrestin complex. Nat. Commun. 7, 13416 (2016).
Kumari, P. et al. Core engagement with β-arrestin is dispensable for agonist-induced vasopressin receptor endocytosis and ERK activation. Mol. Biol. Cell 28, 1003–1010 (2017).
Thomsen, A. R. B. et al. GPCR–G protein–β-arrestin super-complex mediates sustained G protein signaling. Cell 166, 907–919 (2016).
Cahill, T. J. III et al. Distinct conformations of GPCR–β-arrestin complexes mediate desensitization, signaling, and endocytosis. Proc. Natl Acad. Sci. USA 114, 2562–2567 (2017).
Chen, Q. et al. Structural basis of arrestin-3 activation and signaling. Nat. Commun. 8, 1427 (2017).
Granzin, J. et al. Crystal structure of p44, a constitutively active splice variant of visual arrestin. J. Mol. Biol. 416, 611–618 (2012).
Kim, Y. J. et al. Crystal structure of pre-activated arrestin p44. Nature 497, 142–146 (2013).
Sommer, M. E., Farrens, D. L., McDowell, J. H., Weber, L. A. & Smith, W. C. Dynamics of arrestin-rhodopsin interactions: loop movement is involved in arrestin activation and receptor binding. J. Biol. Chem. 282, 25560–25568 (2007).
Tobin, A. B. G-protein-coupled receptor phosphorylation: where, when and by whom. Br. J. Pharmacol. 153 (Suppl. 1), S167–S176 (2008).
Hirsch, J. A., Schubert, C., Gurevich, V. V. & Sigler, P. B. The 2.8 Å crystal structure of visual arrestin: a model for arrestin’s regulation. Cell 97, 257–269 (1999).
Sommer, M. E., Smith, W. C. & Farrens, D. L. Dynamics of arrestin–rhodopsin interactions: arrestin and retinal release are directly linked events. J. Biol. Chem. 280, 6861–6871 (2005).
Sensoy, O., Moreira, I. S. & Morra, G. Understanding the differential selectivity of arrestins toward the phosphorylation state of the receptor. ACS Chem. Neurosci. 7, 1212–1224 (2016).
Jung, S. R., Kushmerick, C., Seo, J. B., Koh, D. S. & Hille, B. Muscarinic receptor regulates extracellular signal regulated kinase by two modes of arrestin binding. Proc. Natl Acad. Sci. USA 114, E5579–E5588 (2017).
Jala, V. R., Shao, W. H. & Haribabu, B. Phosphorylation-independent β-arrestin translocation and internalization of leukotriene B4 receptors. J. Biol. Chem. 280, 4880–4887 (2005).
Yang, F. et al. Phospho-selective mechanisms of arrestin conformations and functions revealed by unnatural amino acid incorporation and (19)F-NMR. Nat. Commun. 6, 8202 (2015).
Eichel, K. et al. Catalytic activation of β-arrestin by GPCRs. Nature https://doi.org/10.1038/s41586-018-0079-1 (2018).
Lee, M. H. et al. The conformational signature of β-arrestin2 predicts its trafficking and signalling functions. Nature 531, 665–668 (2016).
Tohgo, A. et al. The stability of the G protein-coupled receptor-beta-arrestin interaction determines the mechanism and functional consequence of ERK activation. J. Biol. Chem. 278, 6258–6267 (2003).
Eichel, K., Jullié, D. & von Zastrow, M. β-Arrestin drives MAP kinase signalling from clathrin-coated structures after GPCR dissociation. Nat. Cell Biol. 18, 303–310 (2016).
Nuber, S. et al. β-Arrestin biosensors reveal a rapid, receptor-dependent activation/deactivation cycle. Nature 531, 661–664 (2016).
Goodman, O. B. Jr et al. β-arrestin acts as a clathrin adaptor in endocytosis of the β2-adrenergic receptor. Nature 383, 447–450 (1996).
Laporte, S. A., Miller, W. E., Kim, K. M. & Caron, M. G. β-Arrestin/AP-2 interaction in G protein-coupled receptor internalization: identification of a β-arrestin binging site in β2-adaptin. J. Biol. Chem. 277, 9247–9254 (2002).
Pulvermüller, A. et al. Functional differences in the interaction of arrestin and its splice variant, p44, with rhodopsin. Biochemistry 36, 9253–9260 (1997).
Dror, R. O. et al. Structural basis for nucleotide exchange in heterotrimeric G proteins. Science 348, 1361–1365 (2015).
Han, M., Gurevich, V. V., Vishnivetskiy, S. A., Sigler, P. B. & Schubert, C. Crystal structure of β-arrestin at 1.9 Å: possible mechanism of receptor binding and membrane translocation. Structure 9, 869–880 (2001).
Mahalingam, M., Martínez-Mayorga, K., Brown, M. F. & Vogel, R. Two protonation switches control rhodopsin activation in membranes. Proc. Natl Acad. Sci. USA 105, 17795–17800 (2008).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for position of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Betz, R. M. Dabble. https://doi.org/10.5281/zenodo.836914 (2018).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Best, R. B., Mittal, J., Feig, M. & MacKerell, A. D. Jr. Inclusion of many-body effects in the additive CHARMM protein CMAP potential results in enhanced cooperativity of α-helix and β-hairpin formation. Biophys. J. 103, 1045–1051 (2012).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Vanommeslaeghe, K., Raman, E. P. & MacKerell, A. D. Jr. Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model. 52, 3155–3168 (2012).
Vanommeslaeghe, K. & MacKerell, A. D. Jr. Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model. 52, 3144–3154 (2012).
Salomon-Ferrer, R., Götz, A. W., Poole, D., Le Grand, S. & Walker, R. C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald. J. Chem. Theory Comput. 9, 3878–3888 (2013).
Case, D. A. et al. AMBER 2017 (University of California, San Francisco, 2017).
Hopkins, C. W., Le Grand, S., Walker, R. C. & Roitberg, A. E. Long-time-step molecular dynamics through hydrogen mass repartitioning. J. Chem. Theory Comput. 11, 1864–1874 (2015).
Shaw, D. E. et al. Millisecond-scale molecular dynamics simulations on Anton. Proc. ACM/IEEE Conf. Supercomputing (SC09) (2009).
Eastman, P. et al. OpenMM 4: A reusable, extensible, hardware independent library for high performance molecular simulation. J. Chem. Theory Comput. 9, 461–469 (2013).
Eastman, P. & Pande, V. S. Efficient nonbonded interactions for molecular dynamics on a graphics processing unit. J. Comput. Chem. 31, 1268–1272, https://doi.org/10.1002/jcc.21413 (2010).
Tribello, G. A., Bonomi, M., Branduardi, D., Camilloni, C. & Bussi, G. PLUMED2: New feathers for an old bird. Comp. Phys. Comm. 186, 604–613 (2014).
Cheng, X., Wang, H., Grant, B., Sine, S. M. & McCammon, J. A. Targeted molecular dynamics study of C-loop closure and channel gating in nicotinic receptors. PLOS Comput. Biol. 2, e134 (2006).
Roe, D. R. & Cheatham, T. E. III PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 27–38 (1996).
Sommer, M. E., Hofmann, K. P. & Heck, M. Distinct loops in arrestin differentially regulate ligand binding within the GPCR opsin. Nat. Commun. 3, 995 (2012).
Garwin, G. G. & Saari, J. C. High-performance liquid chromatography analysis of visual cycle retinoids. Methods Enzymol. 316, 313–324 (2000).
Schleicher, A., Kühn, H. & Hofmann, K. P. Kinetics, binding constant, and activation energy of the 48-kDa protein-rhodopsin complex by extra-metarhodopsin II. Biochemistry 28, 1770–1775 (1989).
McDowell, J. H., Nawrocki, J. P. & Hargrave, P. A. Isolation of isoelectric species of phosphorylated rhodopsin. Methods Enzymol. 315, 70–76 (2000).
Lally, C. C., Bauer, B., Selent, J. & Sommer, M. E. C-edge loops of arrestin function as a membrane anchor. Nat. Commun. 8, 14258 (2017).
Sommer, M. E., Hofmann, K. P. & Heck, M. Arrestin-rhodopsin binding stoichiometry in isolated rod outer segment membranes depends on the percentage of activated receptors. J. Biol. Chem. 286, 7359–7369 (2011).
Vogel, R. & Siebert, F. Conformations of the active and inactive states of opsin. J. Biol. Chem. 276, 38487–38493 (2001).
Mansoor, S. E., McHaourab, H. S. & Farrens, D. L. Mapping proximity within proteins using fluorescence spectroscopy. A study of T4 lysozyme showing that tryptophan residues quench bimane fluorescence. Biochemistry 41, 2475–2484 (2002).
Hanson, S. M. et al. A model for the solution structure of the rod arrestin tetramer. Structure 16, 924–934 (2008).
Mukherjee, S. et al. Aspartic acid 564 in the third cytoplasmic loop of the luteinizing hormone/choriogonadotropin receptor is crucial for phosphorylation-independent interaction with arrestin2. J. Biol. Chem. 277, 17916–17927 (2002).
Isberg, V. et al. Generic GPCR residue numbers—aligning topology maps while minding the gaps. Trends Pharmacol. Sci. 36, 22–31 (2015).
Piana, S., Lindorff-Larsen, K. & Shaw, D. E. How robust are protein folding simulations with respect to force field parameterization? Biophys. J. 100, L47–L49 (2011).
Vishnivetskiy, S. A. et al. How does arrestin respond to the phosphorylated state of rhodopsin? J. Biol. Chem. 274, 11451–11454 (1999).
Bouvier, M. et al. Removal of phosphorylation sites from the β2-adrenergic receptor delays onset of agonist-promoted desensitization. Nature 333, 370–373 (1988).
Ohguro, H., Palczewski, K., Walsh, K. A. & Johnson, R. S. Topographic study of arrestin using differential chemical modifications and hydrogen/deuterium exchange. Protein Sci. 3, 2428–2434 (1994).
Gurevich, V. V. & Benovic, J. L. Mechanism of phosphorylation-recognition by visual arrestin and the transition of arrestin into a high affinity binding state. Mol. Pharmacol. 51, 161–169 (1997).
Vishnivetskiy, S. A. et al. An additional phosphate-binding element in arrestin molecule. Implications for the mechanism of arrestin activation. J. Biol. Chem. 275, 41049–41057 (2000).
Richardson, M. D. et al. Human substance P receptor lacking the C-terminal domain remains competent to desensitize and internalize. J. Neurochem. 84, 854–863 (2003).
Hanson, S. M. & Gurevich, V. V. The differential engagement of arrestin surface charges by the various functional forms of the receptor. J. Biol. Chem. 281, 3458–3462 (2006).
Shukla, A. K. et al. Distinct conformational changes in β-arrestin report biased agonism at seven-transmembrane receptors. Proc. Natl Acad. Sci. USA 105, 9988–9993 (2008).
Gimenez, L. E. et al. Role of receptor-attached phosphates in binding of visual and non-visual arrestins to G protein-coupled receptors. J. Biol. Chem. 287, 9028–9040 (2012).
Vishnivetskiy, S. A., Baameur, F., Findley, K. R. & Gurevich, V. V. Critical role of the central 139-loop in stability and binding selectivity of arrestin-1. J. Biol. Chem. 288, 11741–11750 (2013).
Acknowledgements
We thank P. Eastman, M. Sultan, R. Betz and C. Brinton for assistance with simulation setup and analysis, and K. Eichel, M. Masureel, A. Venkatakrishnan, B. Kobilka and M. von Zastrow for valuable discussions. This work was supported by National Institutes of Health (NIH) grant R01GM127359 to R.O.D., the Deutsche Forschungsgemeinschaft (SO1037/1-2 to M.E.S.), the Berlin Institute of Health (Delbrück Fellowship BIH_PRO_314 to M.E.S.), an NIH postdoctoral fellowship to S.A.H. (T15-LM007033-33), a Stanford Terman Faculty Fellowship to R.O.D., and NSF Graduate Research Fellowships to N.R.L. and R.J.L.T. A few preliminary simulations were performed on an Anton machine at the Pittsburgh Supercomputing Center, donated by D.E. Shaw Research and supported by NIH grant R01GM116961.
Reviewer information
Nature thanks C. Hoffmann and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
N.R.L., M.E.S. and R.O.D. designed the research. N.R.L and J.K.W. performed and analysed simulations, with assistance from R.J.L.T., J.E.O., S.A.H. and R.O.D. B.B. prepared arrestin mutants. M.E.S. performed and analysed fluorescence spectroscopy experiments. H.E.X. provided structural information and insights. N.R.L, M.E.S. and R.O.D. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Interdomain twist angles and global projection metric values for six arrestin-1 simulation conditions.
Dashed lines indicate the interdomain twist angles in the inactive (0°) and active (18°) state crystal structures and the projection metric values in the inactive (0.0 Å) and active (8.15 Å) state structures. Thick traces indicate the moving average smoothed over a 50-ns window, and thin traces represent unsmoothed data. For each simulation, a pair of plots is shown, one immediately above the other. The top plot (dark colours) shows the interdomain twist angle. The bottom plot shows the projection metric, an alternative means of capturing global conformational change. In one simulation (red box, lower right corner), the RP tail became unbound from arrestin, which resulted in inactive twist angles. All other simulations in that condition maintained stable binding to the RP tail.
Extended Data Fig. 2 Global conformational behaviour of arrestin-1 in simulation.
a, The r.m.s.d. from the inactive structure for representative simulations of arrestin-1 starting in its inactive state with: the C tail removed (green), arrestin-1 with the C tail present (grey), or arrestin-1 bound to full-length rhodopsin (blue). The simulation of arrestin-1 with its C tail removed transitions to active conformations and achieves r.m.s.d. values that match those of rhodopsin-bound active-state simulations. r.m.s.d. is computed on arrestin C-domain β-strands after alignment on the N domain. b, Mean r.m.s.d. from active and inactive structures across all six independent simulations for each condition, calculated after removing the first 500 ns of each simulation. c, We used PCA to compare the conformational states visited under the various arrestin-1 simulation conditions (see Methods; n = 8100 simulation frames as input). Each principal component corresponds to a mode of motion or variance in Cartesian coordinate space. The star on the left in each plot corresponds to the position of the active-state crystal structure, and the star on the right corresponds to the inactive-state structure. Simulations of the two crystallographic conditions separate clearly along the first principal component (PC1) and along the third principal component (PC3) but not along the second principal component (PC2). Simulations starting from the inactive state or active state with the arrestin C tail removed and no receptor present explore similar ranges of PC1 and PC2 coefficients and have some overlap in the range of PC3 coefficients. Simulations with either the receptor core or RP tail bound closely overlap with simulations performed in the presence of the full-length receptor. The x axis is shifted to the right in the first plot in each row relative to the second and third plots in order to show the full range of values of PC1 coefficients. d, Images that show the motion of arrestin-1 along each principal component. e, Variance explained by each principal component. The cumulative distribution function (CDF) shows the variance explained by all principal components up to and including a given one.
Extended Data Fig. 3 Conformation of arrestin favoured by binding of receptor core.
a, b, In simulations started from an inactive conformation of arrestin bound to the receptor core alone (a, see Methods), arrestin preferred to adopt active interdomain twist angles (cyan histogram) (b). By contrast, simulations of arrestin initiated from the same conformation without the receptor (grey histogram) were less likely to spontaneously adopt interdomain twist angles matching those seen in the active-state structure. c, Traces for all simulations, 20 per condition. The difference between the grey and cyan histograms increases with simulation time and would be likely to increase further with additional simulation time, but the differences in the mean interdomain twist angle achieved between the two conditions are already highly significant (P = 3 × 10−5).
Extended Data Fig. 4 Simulation traces from targeted and restrained molecular dynamics simulations.
a, We started simulations of arrestin-1 from the inactive state and pulled the finger loop towards its active, helical conformation (grey trace). The finger loop quickly reached its active state (top), but this failed to induce active interdomain twist angles (bottom) on timescales of hundreds of nanoseconds. b, Similarly, in simulations of arrestin-1 started from the active state (without a receptor present), restraining the finger loop conformation to its active conformation did not prevent arrestin-1 from visiting inactive interdomain twist angles. c, d, By contrast, pulling the gate loop to its active state to mimic RP-tail binding (c), or pulling the IL2-binding crevice apart to mimic receptor core binding (d), consistently induced active interdomain twist angles in arrestin (P = 4 × 10−6, P = 0.002, respectively, compared to unbiased simulations). All traces are smoothed using an averaging window of 20 ns. In all cases we attempted to apply external forces to mimic binding of various structural elements of the receptor. To mimic the effect of binding at the receptor core interface, we aligned and pulled on the same residues within the C loop and middle loop contacted by the receptor. The broad range of interdomain twist angles may reflect the fact that the restraints do not perfectly mimic the effect of the receptor core. Nonetheless, other simulations, including unbiased simulations starting from the inactive state, suggest that separation of the interdomain crevice or the presence of the receptor core favour active interdomain twist angles (Extended Data Figs. 3, 6), providing independent support for the proposed effect of core binding on arrestin activation. Six independent simulations were performed for each condition.
Extended Data Fig. 5 Gate loop motion may be restrained in the inactive state by an ionic interaction with the finger loop.
In certain inactive-state crystal structures of arrestin-1 (for example, PDB entry 1CF1, chain D) and arrestin-2 (for example, PDB entry 1G4M, chain A), a lysine in the gate loop (K298 in arrestin-1, K292 or K294 in arrestin-2) forms an ionic interaction with a carboxylic acid in the finger loop (D71 in arrestin-1, E66 in arrestin-2). Simulations initiated from these structures with the C tail removed exhibited less frequent transitions of the gate loop to fully active conformations than simulations initiated from crystal structures in which this ionic interaction between the gate loop and the figure loop was not formed (for example, PDB entry 1CF1, chain A). Thus a particular finger loop conformation might mildly increase the stability of the inactive-state gate loop conformation. In simulations, we observed additional sets of ionic interactions between gate loop lysines and either D67 on the finger loop or D135 on the middle loop (according to arrestin-2 numbering), which also appeared to prevent motion of the gate loop towards the active state. Certain finger loop conformations might also favour the inactive state through interactions with the C tail of arrestin64.
Extended Data Fig. 6 Conformational changes at IL2 and IL3 interfaces correlate with interdomain twist angle.
a, The C loop contains residues S251 and D253, which interact with Y67 in the N domain in the inactive state of arrestin (cyan, left). In the rhodopsin-bound crystal structure, this network of residues separates when IL2 binds in the central crevice between the N and C domains (purple, right). b, c, We measured separation between the Y67–S251 side-chain hydroxyl oxygens (b) and between the C143–D253 Cα atoms (c). Conformational changes at the IL2 interface correlate with interdomain twist angles. This is particularly noticeable in simulations starting from the inactive state but with the arrestin C tail removed (green), where increased interdomain twist angles correlated with disruption of the Y67–S251 interaction (R2 = 0.35) and with increased separation distance between the two domains, as measured through the C143–D253 Cα distance (R2 = 0.36) (six independent simulations). Plots and correlations refer to trajectories downsampled every 10 ns, with no frames removed at the beginning of simulation. One caveat is that in simulations started from active state without the arrestin C tail, the interdomain crevice frequently collapsed at the beginning of simulation, so that even when arrestin visited more active interdomain twist angles, the crevice did not re-open. It is possible that these simulations reached a local energy minimum not typically visited in the equivalent simulations started from the inactive conformation. d, Conformational changes at the IL3 interface correlate with interdomain twist angles. Compared to the inactive state (blue) of arrestin, in the active state (purple), the back loop, located in the arrestin C domain (residues 311–320), extends away from the arrestin body (motion indicated by the black arrow). In this conformation, the back loop contacts the third intracellular loop in rhodopsin via an ionic interaction between R318 (arrestin) and E239 (rhodopsin). e, The position of the back loop correlates with the interdomain twist angle for simulations of arrestin with its C tail removed, starting from either the inactive (green) or active (purple) state (R2 = 0.50 and R2 = 0.58, respectively; six independent simulations). Back loop position is measured by projecting the coordinates of the back loop onto the vector connecting the crystallographic inactive- and active-state back loop structures (see Methods). f Similarly, in simulations of arrestin bound to the receptor core only, movement away from active interdomain twist angles weakly correlated with disruption of the R318–E239 interaction (R2 = −0.14; six independent simulations). Our simulations therefore indicate that interaction between arrestin and receptor at IL3 may control the interdomain twist angle. We speculate that this occurs because the back loop is coupled to the C loop via a set of β-strands. Thus, the receptor is likely to also modulate interdomain twisting by extending the shape of the back loop. When the back loop moves towards its active conformation, its motion appears to couple to the C domain through β-sheet formation with the C loop. Indeed, previous studies have indicated that acidic residues on IL3 might facilitate arrestin engagement. For example, an acidic residue on IL3 of the human luteinizing hormone receptor is critical for binding to arrestin-2 and arrestin-3, albeit to different extents for each65. Our simulations support the idea that binding via the IL3 interface could help to trigger arrestin activation. Arrestins 1 to 4 share a conserved basic residue at position 313 (bovine arrestin-1 numbering). A qualitative examination of GPCR sequences reveals that several receptors, including the M2 muscarinic receptor, melatonin receptors, β2AR, A2AR, NTS1R, apelin receptor and H1R, all contain acidic residues at the 5x73–5x75 positions (GPCRdb numbering66), which extend into IL3 and may facilitate arrestin activation in the absence of RP-tail phosphorylation.
Extended Data Fig. 7 Gate loop conformation and behaviour of the R175–D296 polar core interaction across arrestin-1 conditions.
a, Gate loop conformation (grey) is measured by projecting the coordinates of the gate loop onto the vector connecting the crystallographic inactive and active gate loop structures (see Methods). In simulations where arrestin-1 maintains a stable active interdomain twist, such as in simulations performed in the presence of the RP tail (right column), the R175–D296 interaction occasionally reforms transiently (blue traces), although the separation of the R175–D296 Cα atoms (red traces) continues to resemble the distance seen in active-state crystal structures. The CHARMM36 force field used here might slightly overstabilize this ionic interaction, increasing the propensity for R175–D296 to reform in these simulations67. b, Conformations of the polar core and gate loop are tightly coupled. Crystal structures of arrestin-1 bound to its C tail (PDB entry 1CF1) and of arrestin-1 bound to the RP tail (PDB entry 5W0P) reveal distinct arrangements of the residues in the polar core (D30, R175, D296, D303 and R382) and a surrounding polar network. As described in the main text, the position of D296 is tightly coupled to interdomain twist. In the active state (right), binding of the RP tail has two effects: first, a phosphorylated serine, S338, engages the gate loop through a direct interaction with K300. In doing so, D296 shifts away from its inactive-state position towards the C domain. In doing so, the interaction between D296 and R175 breaks, disturbing the ionic network of the polar core. Second, S338 also engages R29, which stabilizes the rearrangement of residues in and around the polar core. D30 now engages R175 through an ionic interaction, and D296 is free to interact with other residues, including S308. c, Our simulations reveal how the position of D296 is coupled to interdomain twisting. After the gate loop undergoes a conformational change from its inactive conformation to an active conformation in simulations started from the inactive state with arrestin C tail removed, D296 can shift between its inactive and active positions. In snapshots such as the one shown (simulation 8, right), shifting of D296 towards its active position moves a small β-strand (G292–D296), which is connected to a large β-strand (N271–L280) in the C domain. These observations explain the fact that the R175E and D296R mutations—which would force D296 towards its active position by ionic repulsion—cause phosphorylation-independent arrestin activity, whereas the combination of the two mutations, which would maintain the polar core salt bridge between positions 175 and 296, does not5,68.
Extended Data Fig. 8 Centrifugal pull-down analysis of fluorescently labelled arrestin mutants.
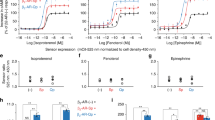
a, Arrestin mutants were mixed with rod outer segment membranes containing rhodopsin (Rho), phosphorylated rhodopsin (RhoP), opsin (Ops) or phosphorylated opsin (OpsP). Rhodopsin samples were illuminated (> 495 nm, 15 s) to obtain activated rhodopsin (Rho*) and phosphorylated activated rhodopsin (Rho*P), and then all samples were centrifuged at 20,800g for 10 min. The supernatant was removed, and the pellets were solubilized in loading buffer. Samples were subjected to SDS–PAGE, and gels were stained with Coomassie blue. MW, molecular weight marker kDa. Arrestin migrates slower than rhodopsin or opsin (arrestin (A) and receptor (R) bands are indicated by arrows). As controls, samples of arrestin in buffer alone (NS, nonspecific pull-down in isotonic or low salt buffer) or rhodopsin alone (bkd, background) were centrifuged alongside the other samples. The total amount of arrestin present in each assay (2.25 μg) is indicated in the lanes marked ‘Arr’. Arrestin ‘cysless’ corresponds to the background construct for all fluorescently labelled arrestin mutants (C63A, C128S, C143A, W194F) and is functionally equivalent to native wild-type bovine arrestin-161. Representative gels, cropped to show desired lanes, are shown. Experimental conditions: 1 μM arrestin, 10 μM receptor, 50 μl sample volume; 50 mM HEPES, 130 mM NaCl pH 7 (isotonic buffer) for samples containing rhodopsin, 50 mM HEPES pH 8.5 (low-salt buffer) for samples containing opsin, 20 °C. b, Arrestin bands were quantified by densitometry using the program GelQuant.NET v.1.8.2. Each band is expressed as the fraction of total arrestin that was present in each experiment, and bars represent averages from n = 2 (arr cysless), n = 5 (Rho* and Rho*P, L173F), and n = 4 (all other conditions) independent experiments ± s.e.m. Essentially no background density from ROS was present at the molecular-weight range of arrestin. All mutants showed some amount of nonspecific pull-down in the different buffer conditions. Note that this nonspecific pull-down is subtracted from the pull-down data reported in the main text. The fluorescent NBD-labelled arrestin mutants bound to the different receptor variants at similar levels as the cysless arrestin control (Ops < Rho* < OpsP < Rho*P).
Extended Data Fig. 9 Arrestin-2 undergoes similar fluctuations in simulation as arrestin-1, suggesting a potential common activation mechanism.
Simulations initiated from the inactive conformation but with the arrestin C tail removed reached active conformations, and simulations initiated from the active conformation but with the co-crystallized RP tail removed reached inactive conformations. a, Interdomain twist angle as a function of time for simulations of arrestin-2 performed under four conditions: active arrestin-2 bound to the V2 vasopressin receptor C-terminal phosphopeptide (PDB entry 4JQI) with the crystallographic Fab30 fragment removed; active arrestin-2 with the V2R RP tail removed; and inactive arrestin-2 with its crystallographic C tail present or absent (PDB entry 1G4M). In these simulations, arrestin-2 appears to favour more inactive-like conformations than those seen in the majority of our arrestin-1 simulations, but this may be due to the specific choice of crystal structure from which the simulations were initiated; see Extended Data Fig. 5. Dashed lines represent the inactive and active state interdomain twist angles for the arrestin-2 crystal structures. b, Snapshot of an active-like rotational state observed in simulations started from an inactive-state structure with the C tail removed (simulation 63, dark red), overlaid on the active-state structure (purple). c, Simulation snapshot from a simulation started from the inactive state with the C tail removed, in which the gate loop moves into an intermediate state (simulation 62, dark red). The absence of a structure of a receptor-bound β-arrestin leaves open the possibility that receptors might bind β-arrestins differently from arrestin-1, and even if the binding mode is similar, the activation mechanism might be different.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-2
Rights and permissions
About this article
Cite this article
Latorraca, N.R., Wang, J.K., Bauer, B. et al. Molecular mechanism of GPCR-mediated arrestin activation. Nature 557, 452–456 (2018). https://doi.org/10.1038/s41586-018-0077-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0077-3
This article is cited by
-
Tail engagement of arrestin at the glucagon receptor
Nature (2023)
-
Molecular mechanism of biased signaling at the kappa opioid receptor
Nature Communications (2023)
-
Screening of small molecule compounds targeting hnRNPA2 protein
Medicinal Chemistry Research (2023)
-
Allosteric modulation of GPCR-induced β-arrestin trafficking and signaling by a synthetic intrabody
Nature Communications (2022)
-
β-arrestin1 and 2 exhibit distinct phosphorylation-dependent conformations when coupling to the same GPCR in living cells
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.