Abstract
Enhancers for embryonic stem (ES) cell-expressed genes and lineage-determining factors are characterized by conventional marks of enhancer activation in ES cells1,2,3, but it remains unclear whether enhancers destined to regulate cell-type-restricted transcription units might also have distinct signatures in ES cells. Here we show that cell-type-restricted enhancers are ‘premarked’ and activated as transcription units by the binding of one or two ES cell transcription factors, although they do not exhibit traditional enhancer epigenetic marks in ES cells, thus uncovering the initial temporal origins of cell-type-restricted enhancers. This premarking is required for future cell-type-restricted enhancer activity in the differentiated cells, with the strength of the ES cell signature being functionally important for the subsequent robustness of cell-type-restricted enhancer activation. We have experimentally validated this model in macrophage-restricted enhancers and neural precursor cell (NPC)-restricted enhancers using ES cell-derived macrophages or NPCs, edited to contain specific ES cell transcription factor motif deletions. DNA hydroxyl-methylation of enhancers in ES cells, determined by ES cell transcription factors, may serve as a potential molecular memory for subsequent enhancer activation in mature macrophages. These findings suggest that the massive repertoire of cell-type-restricted enhancers are essentially hierarchically and obligatorily premarked by binding of a defining ES cell transcription factor in ES cells, dictating the robustness of enhancer activation in mature cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Miguel-Escalada, I., Pasquali, L. & Ferrer, J. Transcriptional enhancers: functional insights and role in human disease. Curr. Opin. Genet. Dev. 33, 71–76 (2015).
Ong, C.-T. & Corces, V. G. Enhancer function: new insights into the regulation of tissue-specific gene expression. Nat. Rev. Genet. 12, 283–293 (2011).
Kaikkonen, M. U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Huang, H.-T. et al. A network of epigenetic regulators guides developmental haematopoiesis in vivo. Nat. Cell Biol 15, 1516–1525 (2013).
Epelman, S., Lavine, K. J. & Randolph, G. J. Origin and functions of tissue macrophages. Immunity 41, 21–35 (2014).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Schmidt, D. et al. A CTCF-independent role for cohesin in tissue-specific transcription. Genome Res. 20, 578–588 (2010).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
van de Werken, H. J. G. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat. Methods 9, 969–972 (2012).
Li, W. et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498, 516–520 (2013).
Ying, Q.-L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Kumar, R. M. et al. Deconstructing transcriptional heterogeneity in pluripotent stem cells. Nature 516, 56–61 (2014).
Zhuang, L. et al. Pure populations of murine macrophages from cultured embryonic stem cells. Application to studies of chemotaxis and apoptotic cell clearance. J. Immunol. Methods 385, 1–14 (2012).
Ying, Q.-L., Stavridis, M., Griffiths, D., Li, M. & Smith, A. Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nat. Biotechnol. 21, 183–186 (2003).
Wang, A. et al. Epigenetic priming of enhancers predicts developmental competence of hESC-derived endodermal lineage intermediates. Cell Stem Cell 16, 386–399 (2015).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Will, B. et al. Minimal PU.1 reduction induces a preleukemic state and promotes development of acute myeloid leukemia. Nat. Med. 21, 1172–1181 (2015).
Lichtinger, M. et al. RUNX1 reshapes the epigenetic landscape at the onset of haematopoiesis. EMBO J. 31, 4318–4333 (2012).
Gagliardi, A. et al. A direct physical interaction between Nanog and Sox2 regulates embryonic stem cell self-renewal. EMBO J 32, 2231–2247 (2013).
Han, D. et al. A highly sensitive and robust method for genome-wide 5hmC profiling of rare cell populations. Mol. Cell 63, 711–719 (2016).
Lara-Astiaso, D. et al. Immunogenetics. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
Kwak, H., Fuda, N. J., Core, L. J. & Lis, J. T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Thu, K. L. et al. Methylated DNA immunoprecipitation. J. Vis. Exp. 23, e935 (2009).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Acknowledgements
We thank J. Hightower for assistance with figure preparation, and H. Taylor for help with animal-based cell culture. M.G.R. is an investigator with the Howard Hughes Medical Institute. This work was supported by grants from NIH to M.G.R. (5R01NS093066, DK018477, DK039949, NS093066, and GM104459).
Reviewer information
Nature thanks H. Stunnenberg and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
H.S.K. and M.G.R. conceived the project. H.S.K. performed most of the experiments, with particular contributions from W.M. (CRISPR–Cas9), Y.T. (PRO-cap) and E.D. (4C-seq). D.M. and Y.T. performed most of the bioinformatics analyses. T.S. performed the DNA methylation experiment. K.O. prepared samples for deep sequencing assays. Q.M. analysed 4C-seq. Additional experiments or methods and discussion were contributed by E.D., D.S.-K. and M.F. H.S.K. and M.G.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Features of macrophage-restricted enhancers in ES cells.
a, PU.1+H3K4me2+ macrophage enhancer profiles in mouse ES cells. b, Normalized tag counts of ATAC-seq and DNase-seq in 18,405 random regions versus 18,405 macrophage enhancers versus 28,450 ES cell enhancers. c, Percentage of macrophage enhancers or random regions bound by ENOS in a −1 kb/+ 1 kb window. d, ESRRB binding in random regions versus macrophage enhancers versus ES cell enhancers in ES cells. e, ENOS binding profiles in 6,775 macrophage enhancers. f, UCSC genome browser screen shots show ENOS binding with chromatin openness but no enhancer marks, such as H3K4me2 and H3K27Ac, in macrophage-restricted enhancers in ES cells. The Irg1 gene locus has two different macrophage-restricted enhancers bound by ES cell transcription factors. The blue bar indicates macrophage-restricted enhancers. In the box plots, line shows median, and box shows 25th and 75th percentiles. P values calculated using Welch’s two-sided t-test.
Extended Data Fig. 2 ES cell transcription factors bind to heart-, kidney- and N2A cell-restricted enhancers in ES cells.
a, Profiles of cell-type-restricted enhancers in heart, kidney, and N2A neuronal cells, with percentages of enhancers that are unmarked, poised and active in ES cells. b, Target percentage of ENOS-bound heart-, kidney- and N2A cell-restricted enhancers in ES cells. c, Heat map of ESRRB, NANOG, OCT4 and SOX2 binding within a −1 kb/+ 1 kb window centred on each transcription factor in 17,561 heart-, 14,242 kidney- and 12,027 N2A cell-restricted enhancers in ES cells. d, Heat map of ATAC-seq within a −1 kb/+ 1 kb window centred on each transcription factor in heart-, kidney- and N2A cell-restricted enhancers in ES cells. Data from published sources are listed in Supplementary Table 1.
Extended Data Fig. 3 Premarked macrophage-restricted enhancers do not interact with cognate target coding genes of macrophages in ES cells.
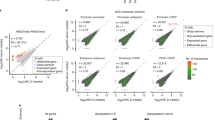
a, b, 4C–seq in ES cells and macrophages. Primer sets for 4 C are listed in Supplementary Table 2. a, 4C–seq analysis of the Il1a gene locus from the enhancer viewpoint in KLA-treated peritoneal macrophages or ES cells. Black arrows represent interaction frequency based on the trunc mean in a 400-kb window. The statistic of trunc mean calculated standard mean but with a truncated high and low value12. Red box shows position of macrophage putative enhancer 5′ of Il1a. b, 4C–seq analysis of the Tnfaip3 gene locus from the enhancer viewpoint in KLA-treated Raw 264.7 macrophages or in ES cells. Black arrows indicate interactions based on linear mean in 1-Mb window. Red box indicates position of putative Tnfaip3 enhancer and blue circle indicates Tnfaip3 promoter. c, The 634 differentially expressed genes (fold-change ≥ 4, over 20 tags in macrophage) among the proximal macrophage-expressed genes of 6,775 macrophage-restricted enhancers were visualized by heat map in ES cells and macrophages. Colour indicates normalized tag counts with log2 transform. d, Gene ontology (GO) term analysis of 634 differentially expressed genes.
Extended Data Fig. 4 Features of 6,775 macrophage enhancers in ES cells are not affected by ES cell culture medium (2i or serum).
a, b, Heat map of H3K4me2 and H3K27Ac (a) and ATAC-seq (b) in 6,775 premarked macrophage enhancers and 28,450 ES cell enhancers in 2i- or serum-cultured ES cells. Centred on ENOS and visualized in a −1 kb/+ 1 kb window.
Extended Data Fig. 5 ENOS binding correlates with future enhancer activity.
a–d, Comparison of the approximately 10% ‘most’ marked (high) and ‘least’ marked (low) enhancers selected on the basis of ENOS binding, and the level of H3K4me1 and H3K27Ac on these enhancers in mature tissues for spleen (a), lung (b), cortex (c) and bone marrow (d). In the box plots, line shows median and box shows 25th and 75th percentiles. Red dots indicate each value of H3K4me1 or H3K27Ac. P values calculated using Welch’s two-sided t-test. Data from published sources are listed in Supplementary Table 1.
Extended Data Fig. 6 Premarking in ES cells is required for robust future macrophage enhancer activation.
a, Photomicrographs of macrophages differentiated from ES cells documented by F4/80 and CD11B staining. Scale bar, 50 µM. b, Screenshot of the Tlr1 locus. The blue box corresponds to the CRISPR–Cas9 target region. c, d, Putative Tlr1 enhancer mutant sequence diagram and DNA sequence documentation of several of the homozygous mutation clones, used for analysis. e, Screenshot of the Tnfaip3 locus. The blue box corresponds to the CRISPR–Cas9 target region. f, g, Putative Tnfaip3 enhancer mutant sequence diagram and DNA sequence documentation of several of the homozygous mutation clones, used for analysis. h, Tlr1 eRNA transcription was tested in wild-type and mutant clonal cells (#26, #45) of the Tnfaip3 enhancer. One representative experiment is plotted (n = 2 biological repeats). i, UCSC genome browser shot of RNA-seq in Tlr1 locus in normal ES cells and clonal mutant ES cells (wild-type, #10, #14). Primer sets are listed in Supplementary Table 2.
Extended Data Fig. 7 Premarking in ES cells is required for robust future macrophage enhancer activation.
a, b, Putative Prdx5 enhancer mutant sequence diagram and DNA sequence documentation of several of the homozygous mutation clones, used for analysis. c, Three mutant clones (B11, F1, G11) of the putative Prdx5 enhancer decreased level of Prdx5 eRNA transcription in ESDMs (right). Each dot indicates one biological experiment (n = 3 biological repeats from two pooled different experiments; n = 2 biological repeats from two pooled different experiments for B11). ESRRB binding in ES cells is a representative experiment (left) (n = 2 biological repeats). d, Nod2 eRNA transcription was tested in wild-type and mutant (B11, F1, G11) clonal cells of the Prdx5 enhancer. Data from one representative experiment are plotted (n = 2 biological repeats). e, f, Putative Nod2 enhancer mutant sequence diagram and DNA sequence documentation of several of the homozygous mutation clones, used for analysis. g, Two mutant clones (D2, D12) of the putative Nod2 enhancer decreased level of Nod2 eRNA transcription in ESDMs (right). Each dot indicates one biological experiment (n = 3 biological repeats from two pooled different experiments; n = 2 biological repeats from two pooled different experiments for D12). ESRRB binding in ES cells is a representative experiment (left) (n = 2 biological repeats). In the box plots, line shows median and box shows 25th and 75th percentiles. P values calculated using Welch’s two-sided t-test. Primer sets are listed in Supplementary Table 2.
Extended Data Fig. 8 Premarking in ES cells is required for robust future NPC enhancer activation.
a, The putative Nek1 enhancer, referred to as N4; mutant sequence diagram and DNA sequence documentation of several mutation clones (N4C3, N4C16), used for analysis. b, Two mutant clones (N4C3, N4C16) of the N4 enhancer decreased level of N4 eRNA transcription in NPCs (right). Each dot indicates one biological experiment. A representative experiment for ESRRB binding in ES cells is presented (left) (n = 3 biological repeats for N4C3, n = 2 biological repeats for N4C16). c, N8 eRNA transcription was tested in N4 mutant clones (N4C3, N4C16) to show specificity of N4 mutant cells on inhibition of N4 eRNA transcription. A representative experiment is presented (n = 2 biological repeats). d, The putative Ankrd1 enhancer, referred to as N8; mutant sequence diagram and DNA sequence documentation of several of the mutation clones (N8C6, N8C12), used for analysis. e, Two mutant clones (N8C6, N8C12) of the N8 enhancer decreased level of N8 eRNA transcription in NPCs (right). Each dot indicates one biological experiment. ESRRB binding in ES cells is a representative experiment showing inhibition of ESRRB binding in ESRRB-deleted N8 locus (left) (n = 3 biological repeats). f, N4 eRNA transcription was examined to prove the specificity of N8 mutant cells on inhibition of N8 eRNA transcription. Results of one representative experiment are presented (n = 3 biological repeats). g, NPC differentiation was confirmed on day 6 after differentiation by testing expression of ES cell-expressing genes and NPC-expressing genes. A representative experiment is presented (n = 2 biological repeats). In the box plots, line shows median and box shows 25th and 75th percentiles. P values calculated using Welch’s two-sided t-test. Primer sets listed in Supplementary Table 2.
Extended Data Fig. 9 Binding of ES cell transcription factors in poised enhancer.
a, Profile of 214 macrophage enhancers with poised chromatin signatures in ES cells. b, ES cell transcription factors (OCT4, SOX2, NANOG, ESRRB, E2F1, TCFCP2L1, ZFX and KLF4) were analysed in 129 macrophage enhancers to test binding of ES cell transcription factors. These 129 macrophage enhancers were chosen according to the following criteria: high H3K27me3 in ES cells and high H3K27Ac (over 20 tags) in macrophages. Data from published sources are listed in Supplementary Table 1. c, Schematic diagram of Pu.1 enhancers. d, Genome browser screenshot of the poised Pu.1 enhancers. Blue arrow indicates ESRRB binding region and red arrow indicates OCT4 binding region. Red boxes indicate four clustered putative Pu.1 enhancers.
Extended Data Fig. 10 Establishment and maintenance of 5-hmC on premarked enhancers and proposed model.
a, Esrrb, Oct4 and Cebp level measured at four different time points through differentiation process from ES cells to macrophages. b, 5-hmC level in 6,775 premarked enhancers in ES cells expressing scrambled (siCtrl) or Esrrb-targeted (siEsrrb) siRNA. In the box plots, line shows median and box shows 25th and 75th percentiles. P values calculated using Welch’s two-sided t-test. c, Schematic of the haematopoietic differentiation stages24. d, H3K4me1 and H3K27Ac in 6,775 premarked enhancers were analysed during haematopoietic differentiation (ES cells, mesoderm, LT-HSC, ST-HSC, MPP, CMP, GMP and macrophages), and presented in heat map within a −3 kb/+ 3 kb window centred on PU.1. Data from published sources are listed in Supplementary Table 1. e, Model of ‘premarked’ lineage-determining and terminal-differentiation enhancer indicating that poised enhancers bind several ES cell transcription factors, whereas late-activated cell-specific enhancers are premarked by binding of a single ES cell transcription factor, causing chromatin opening, transcription of a non-coding RNA, and appearance of a 5-hmC mark in the area of the enhancer that may provide the molecular memory for what ultimately will be the PU.1–C/EBPα core from which eRNAs will be transcribed.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-2
Rights and permissions
About this article
Cite this article
Kim, H.S., Tan, Y., Ma, W. et al. Pluripotency factors functionally premark cell-type-restricted enhancers in ES cells. Nature 556, 510–514 (2018). https://doi.org/10.1038/s41586-018-0048-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0048-8
This article is cited by
-
FACT regulates pluripotency through proximal and distal regulation of gene expression in murine embryonic stem cells
BMC Biology (2023)
-
Bcl6 drives stem-like memory macrophages differentiation to foster tumor progression
Cellular and Molecular Life Sciences (2023)
-
Transcription imparts architecture, function and logic to enhancer units
Nature Genetics (2020)
-
IFI16 promotes human embryonic stem cell trilineage specification through interaction with p53
npj Regenerative Medicine (2020)
-
Nascent RNA analyses: tracking transcription and its regulation
Nature Reviews Genetics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.