Abstract
The NMDA (N-methyl-d-aspartate) receptor transduces the binding of glutamate and glycine, coupling it to the opening of a calcium-permeable ion channel1. Owing to the lack of high-resolution structural studies of the NMDA receptor, the mechanism by which ion-channel blockers occlude ion permeation is not well understood. Here we show that removal of the amino-terminal domains from the GluN1–GluN2B NMDA receptor yields a functional receptor and crystals with good diffraction properties, allowing us to map the binding site of the NMDA receptor blocker, MK-801. This crystal structure, together with long-timescale molecular dynamics simulations, shows how MK-801 and memantine (a drug approved for the treatment of Alzheimer’s disease) bind within the vestibule of the ion channel, promote closure of the ion channel gate and lodge between the M3-helix-bundle crossing and the M2-pore loops, physically blocking ion permeation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
30 April 2018
The Source Data file originally published with this article was missing for Figure 2. This has now been corrected.
References
Traynelis, S. F. et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol. Rev. 62, 405–496 (2010).
Paoletti, P. Molecular basis of NMDA receptor functional diversity. Eur. J. Neurosci. 33, 1351–1365 (2011).
Lee, C. H. et al. NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511, 191–197 (2014).
Karakas, E. & Furukawa, H. Crystal structure of a heteromeric NMDA receptor ion channel. Science 344, 992–997 (2014).
Mayer, M. L., Westbrook, G. L. & Guthrie, P. B. Voltage-dependent block by Mg2+ of NMDA responses in spinal cord neurones. Nature 309, 261–263 (1984).
Nowak, L., Bregestovski, P., Ascher, P., Herbet, A. & Prochiantz, A. Magnesium gates glutamate-activated channels in mouse central neurones. Nature 307, 462–465 (1984).
Johnson, J. W. & Ascher, P. Glycine potentiates the NMDA response in cultured mouse brain neurons. Nature 325, 529–531 (1987).
Bliss, T. V. P. & Collingridge, G. L. A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361, 31–39 (1993).
Paoletti, P., Bellone, C. & Zhou, Q. NMDA receptor subunit diversity: impact on receptor properties, synaptic plasticity and disease. Nat. Rev. Neurosci. 14, 383–400 (2013).
Simon, R. P., Swan, J. H., Griffiths, T. & Meldrum, B. S. Blockade of N-methyl-d-aspartate receptors may protect against ischemic damage in the brain. Science 226, 850–852 (1984).
Parsons, M. P. & Raymond, L. A. Extrasynaptic NMDA receptor involvement in central nervous system disorders. Neuron 82, 279–293 (2014).
Yuan, H., Hansen, K. B., Vance, K. M., Ogden, K. K. & Traynelis, S. F. Control of NMDA receptor function by the NR2 subunit amino-terminal domain. J. Neurosci. 29, 12045–12058 (2009).
Karakas, E., Simorowski, N. & Furukawa, H. Subunit arrangement and phenylethanolamine binding in GluN1/GluN2 NMDA receptors. Nature 475, 249–253 (2011).
Hu, N. W., Klyubin, I., Anwyl, R. & Rowan, M. J. GluN2B subunit-containing NMDA receptor antagonists prevent Aβ-mediated synaptic plasticity disruption in vivo. Proc. Natl Acad. Sci. USA 106, 20504–20509 (2009).
Yuan, H. et al. Context-dependent GluN2B-selective inhibitors of NMDA receptor function are neuroprotective with minimal side effects. Neuron 85, 1305–1318 (2015).
Parsons, C. G. et al. Comparison of the potency, kinetics and voltage-dependency of a series of uncompetitive NMDA receptor antagonists in vitro with anticonvulsive and motor impairment activity in vivo. Neuropharmacology 34, 1239–1258 (1995).
Kovacic, P. & Somanathan, R. Clinical physiology and mechanism of dizocilpine (MK-801): electron transfer, radicals, redox metabolites and bioactivity. Oxid. Med. Cell. Longev. 3, 13–22 (2010).
Reisberg, B. et al. Memantine in moderate-to-severe Alzheimer’s disease. N. Engl. J. Med. 348, 1333–1341 (2003).
Pierson, T. M. et al. GRIN2A mutation and early-onset epileptic encephalopathy: personalized therapy with memantine. Ann. Clin. Transl. Neurol. 1, 190–198 (2014).
Wollmuth, L. P. & Sobolevsky, A. I. Structure and gating of the glutamate receptor ion channel. Trends Neurosci. 27, 321–328 (2004).
Kashiwagi, K. et al. Channel blockers acting at N-methyl-d-aspartate receptors: differential effects of mutations in the vestibule and ion channel pore. Mol. Pharmacol. 61, 533–545 (2002).
Sobolevsky, A. I., Koshelev, S. G. & Khodorov, B. I. Interaction of memantine and amantadine with agonist-unbound NMDA-receptor channels in acutely isolated rat hippocampal neurons. J. Physiol. (Lond.) 512, 47–60 (1998).
Wollmuth, L. P., Kuner, T., Seeburg, P. H. & Sakmann, B. Differential contribution of the NR1- and NR2A-subunits to the selectivity filter of recombinant NMDA receptor channels. J. Physiol. (Lond.) 491, 779–797 (1996).
Wollmuth, L. P., Kuner, T. & Sakmann, B. Intracellular Mg2+ interacts with structural determinants of the narrow constriction contributed by the NR1-subunit in the NMDA receptor channel. J. Physiol. (Lond.) 506, 33–52 (1998).
Klepeis, J. L., Lindorff-Larsen, K., Dror, R. O. & Shaw, D. E. Long-timescale molecular dynamics simulations of protein structure and function. Curr. Opin. Struct. Biol. 19, 120–127 (2009).
Chang, H. R. & Kuo, C. C. The activation gate and gating mechanism of the NMDA receptor. J. Neurosci. 28, 1546–1556 (2008).
Murthy, S. E., Shogan, T., Page, J. C., Kasperek, E. M. & Popescu, G. K. Probing the activation sequence of NMDA receptors with lurcher mutations. J. Gen. Physiol. 140, 267–277 (2012).
Stern, P., Cik, M., Colquhoun, D. & Stephenson, F. A. Single channel properties of cloned NMDA receptors in a human cell line: comparison with results from Xenopus oocytes. J. Physiol. (Lond.) 476, 391–397 (1994).
Jensen, M. Ø., Jogini, V., Eastwood, M. P. & Shaw, D. E. Atomic-level simulation of current-voltage relationships in single-file ion channels. J. Gen. Physiol. 141, 619–632 (2013).
Lipton, S. A. Paradigm shift in neuroprotection by NMDA receptor blockade: memantine and beyond. Nat. Rev. Drug Discov. 5, 160–170 (2006).
Dukkipati, A., Park, H. H., Waghray, D., Fischer, S. & Garcia, K. C. BacMam system for high-level expression of recombinant soluble and membrane glycoproteins for structural studies. Protein Expr. Purif. 62, 160–170 (2008).
Baconguis, I. & Gouaux, E. Structural plasticity and dynamic selectivity of acid-sensing ion channel–spider toxin complexes. Nature 489, 400–405 (2012).
Inouye, H., Barnes, W. & Beckwith, J. Signal sequence of alkaline phosphatase of Escherichia coli. J. Bacteriol. 149, 434–439 (1982).
Reeves, P. J., Callewaert, N., Contreras, R. & Khorana, H. G. Structure and function in rhodopsin: high-level expression of rhodopsin with restricted and homogeneous N-glycosylation by a tetracycline-inducible N-acetylglucosaminyltransferase I-negative HEK293S stable mammalian cell line. Proc. Natl Acad. Sci. USA 99, 13419–13424 (2002).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012).
Armstrong, N., Jasti, J., Beich-Frandsen, M. & Gouaux, E. Measurement of conformational changes accompanying desensitization in an ionotropic glutamate receptor. Cell 127, 85–97 (2006).
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, (1948–1954 (2002).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure and homology modelling. Bioinformatics 22, 195–201 (2006).
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLOS Comput. Biol. 8, e1002708 (2012).
Hart, H. E. & Greenwald, E. B. Scintillation proximity assay (SPA)—a new method of immunoassay. Direct and inhibition mode detection with human albumin and rabbit antihuman albumin. Mol. Immunol. 16, 265–267 (1979).
Vilar, S., Cozza, G. & Moro, S. Medicinal chemistry and the molecular operating environment (MOE): application of QSAR and molecular docking to drug discovery. Curr. Top. Med. Chem. 8, 1555–1572 (2008).
Roux, B. The membrane potential and its representation by a constant electric field in computer simulations. Biophys. J. 95, 4205–4216 (2008).
Shaw, D. E. et al. Anton 2: raising the bar for performance and programmability in a special-purpose molecular dynamics supercomputer. In Proc. International Conference for High Performance Computing, Networking, Storage and Analysis 41–53 https://doi.org/10.1109/SC.2014.9 (2014).
Hackos, D. H. et al. Positive allosteric modulators of GluN2A-containing NMDARs with distinct modes of action and impacts on circuit function. Neuron 89, 983–999 (2016).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Mackerell, A. D. Jr, Feig, M. & Brooks, C. L. III Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Jensen, M. Ø. et al. Mechanism of voltage gating in potassium channels. Science 336, 229–233 (2012).
Vanommeslaeghe, K. et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690, (2010).
Yu, W., He, X., Vanommeslaeghe, K. & MacKerell, A. D. Jr. Extension of the CHARMM General Force Field to sulfonyl-containing compounds and its utility in biomolecular simulations. J. Comput. Chem. 33, 2451–2468 (2012).
Mishra, S. et al. Conformational dynamics of the nucleotide binding domains and the power stroke of a heterodimeric ABC transporter. eLife 3, e02740 (2014).
Jeschke, G., Koch, A., Jonas, U. & Godt, A. Direct conversion of EPR dipolar time evolution data to distance distributions. J. Magn. Reson. 155, 72–82 (2002).
Stein, R. A., Beth, A. H. & Hustedt, E. J. A straightforward approach to the analysis of double electron-electron resonance data. Methods Enzymol. 563, 531–567 (2015).
Acknowledgements
We thank L. Vaskalis and H. Owen for manuscript preparation, Gouaux laboratory members, S. Piana and M. Eastwood for discussions, and the Berkeley Center for Structural Biology (5.0.2) and the Advanced Photon Source (24ID-C and 24ID-E) for assistance with data collection. This work was supported by the National Institutes of Health (R01 NS038631). E.G. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
X.S. and E.G. designed the project; X.S. and C.-H.L. developed the constructs for crystallization; X.S. performed protein purification, crystallography, electrophysiology and biochemical analysis; M.Ø.J. and V.J. performed and analysed, and D.E.S. participated in the oversight of, the molecular dynamics simulations; R.A.S. and H.S.M. carried out the DEER experiments; and X.S. and E.G. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The ∆ATD NMDA receptor construct and structure.
a, Selected amino acid sequences of constructs used in these studies are compared to the wild-type sequence to highlight mutations in both subunits. Locations of mutated sites and deletions are highlighted in yellow. Insertions are shown in blue or red. The ‘A2 tail’ is derived from residues 837–847 of GluA2 AMPA receptor C terminus (NP_058957). b, Cartoon representation shows the GluN1 and GluN2B subunit constructs and modifications of the ∆ATD receptor. The locations of point mutations are highlighted with blue circles and the deletions are shown as yellow wedges. c, Superposition of the two ∆ATD NMDA receptors in the crystallographic asymmetric unit, aligned by the TMD. Black arrows show the shift between receptor 1 (light blue) and receptor 2 (magenta).
Extended Data Fig. 2 LBD dimer rearrangement and dynamics.
a, b, Top-down view from the extracellular side of the membrane, showing the LBD layer of the intact Δ2 NMDA receptor (a) with GluN1 in blue and GluN2B in yellow, and of the ∆ATD NMDA receptor (b) with GluN1 in green and GluN2B in orange. The M3–LBD linkers of GluN2B (red ribbon, Q653–S664) adopt distinct conformations in the two receptors. Shown are distances between GluN2B R739 residues (β-carbon atoms, salmon spheres), the residue selected for the DEER experiments (in Å) in both the intact and ΔATD receptors. c, Cartoon emphasizing how the ATDs participate in defining the conformation of the LBD layer and how this, in turn, keeps the GluN2B M3–D2 linker in a conformation capable of opening the channel gate. d, The Fo−Fc density (3σ, green mesh) fits loop 1 of the GluN1 subunit (blue cartoon) but not that of the GluN2B subunit (orange cartoon). e, DEER data of MTSSL-labelled GluN2B(R739C) ∆ATD (red) (sample size, n = 2) and intact NMDA receptor (blue) (sample size, n = 1). Left, peak-normalized echo decay and fits; right, probability distributions of DEER distances. The probability distributions of the DEER distances show two major peaks, one centred at 35–40 Å and a second broad peak at around 55 Å. The amplitudes of the two peaks in the ΔATD receptor are comparable, with the 55 Å peak corresponding to the ‘rearranged’ LBD layer as seen in the ΔATD crystal structure, whereas the shorter distance (approximately 35–40 Å) indicates the canonical LBD arrangement, like that observed in the intact receptor structure. The intact receptor, by contrast, shows one major narrow peak at around 40 Å, which corresponds with the predicted distance based on the intact receptor crystal structure, whereas the small peak centred around 55 Å suggests that the intact receptor may harbour a minor population with a ΔATD-like LBD arrangement.
Extended Data Fig. 3 The ∆ATD NMDA receptor channel.
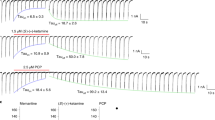
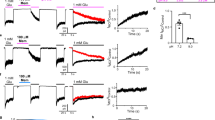
a, Inhibition of agonist-induced (300 μM glutamate and 300 μM glycine) current by 1 μM MK-801 for the ∆ATD NMDA receptor by two-electrode voltage clamp. The inhibition ratio is 0.37 ± 0.06 (mean ± s.d., n = 5). The holding potential is −60 mV. b, Superposition of the ∆2 and ∆ATD receptor TMDs shows that they adopt similar conformations. c, Side view of the ion pore with GluN2B subunits (orange ribbons) showing the van der Waals radius along the pore (magenta dots). The α-carbons of selected residues facing the pore are shown as spheres. The radius is plotted against the distance along the pore axis.
Extended Data Fig. 4 Lipid accessibility of the TMD tunnel.
a, Simulation snapshot (simulation 2) of a lipid molecule with one of its tails trapped between the M2 and M3 helices of the GluN1 subunit (chain A, green ribbons) and the M3 helix of the adjacent GluN2B subunit (light blue ribbons) viewed from within the membrane and towards the pore. Residues L612, L613, A638, I641 and V642 of GluN1 (chain A) and V637 of GluN2B (chain D) of the tunnel walls are shown as spheres with the carbon atoms in green and grey, respectively. GluN1 (chain C) and GluN2B (chain B) subunits are shown as green and light blue solid surfaces, respectively. The dark grey plane represents a cut across the lipid membrane, the remainder of which is shown as a red and white surface. b, Mean lipid occupancy (number of lipid atoms) within 3.5 Å of the tunnel walls, defined by residues L612, L613, A638, I641 and V642 of GluN1 (chains A, C) and residue V637 of the GluN2B (chains B, D) subunit. The occupancy was calculated across the closed-pore and pore-opening simulations (simulations 2 and 3) and all permeation simulations (simulations 4–17). N, all individual simulations within a given panel; n, all individual data points aggregated across all simulations. N = 16 (simulations 2–17); n ≫ 10. Error bars, s.d. of the mean calculated from all individual data points aggregated across all simulations.
Extended Data Fig. 5 Binding times of MK-801 and memantine.
a, Binding time of MK-801 in simulations 18–21. Green lines are 30-ns running medians, and red lines indicate bound and unbound states. Binding was defined as the ligand heavy-atom centre of mass being within 10 Å of the centre of mass of the α-atoms of the N612 residues of the two GluN2B subunits. The mean binding time of simulations 20 and 21 at 0 mV was 0.78 ± 0.10 µs; application of voltage in simulations 18 and 19 (−593.9 ± 3.8 and −197.9 ± 1.2 mV) did not significantly decrease the binding time. b, Binding time of memantine in simulations 24, 25, and 28–30. Simulations 26 and 27 were initiated with memantine already bound, and the binding curves from these simulations were thus omitted in the determination of the on-rates for this pore blocker. Green lines are 30-ns running medians, and red lines indicate bound and unbound states. The mean binding time in simulations 24 and 25 (at 0 mV) was 0.14 ± 0.02 µs; application of voltage in simulations 28–30 (−592.6 ± 0.3, −592.7 ± 0.3 and −196.9 ± 0.1 mV) did not significantly decrease the binding time. In simulations 29 and 30, ‘unbound’ states following binding are artefacts owing to the voltage driving memantine through the selectivity filter. N = 1 in each panel.
Extended Data Fig. 6 Blocker-induced channel closure.
a–c, The resemblance between the closed, deactivated receptor (far left) and the closed, pore-blocked receptor (far right) is shown. r.m.s.d. (red, GluN1; blue, GluN2B) of the M3-bundle-crossing region (the activation gate) relative to the closed-state Δ2 crystal structure obtained from simulations of the closed pore (simulation 2), pore opening (simulation 3), permeation (simulation 4 at −396.6 ± 2.7 mV, simulation 5 at −593.8 ± 3.8 mV, and simulation 6 (a) at −396.1 ± 2.7 mV or simulation 7 (b and c) at −415.1 ± 6.4 mV), two MK-801-binding simulations (simulations 20 (a) and 21 (b)), and one memantine-binding simulation (simulation 25 (c)). N = 1 in each panel.
Extended Data Fig. 7 Free-energy estimates of MK-801 and memantine binding.
a, Competition binding of memantine to the Δ2 receptor in the presence of 3 μM [3H]MK-801, measured by the scintillation proximity assay. The plot shows data from a representative experiment with error bars representing s.e.m. from triplicate measurements. b, Kd (circles) derived from free-energy estimates of binding of memantine and MK-801 to the open, intact, activated Δ2 receptor in which the pore has collapsed onto the ligand. The absolute experimental affinities of MK-801 (green) and memantine (red) for the Δ2 receptors are shown as squares. The free energies were calculated for four independent, ligand-bound configurations, all taken from binding simulations at zero transmembrane voltage (MK-801: simulations 20 and 21; memantine: simulations 24, 25 and 27). Each calculation consisted of 0.5 µs of simulation to solvate the ligand in water, followed by 3.0 µs of simulation of the protein–ligand complex. The average Kd values for MK-801 and memantine of around 0.08 and around 7.64 µM for the Δ2 receptor show a 100-fold difference in the affinities of these two ligands; a similar relative affinity of these two ligands has been found experimentally using the Δ2 receptor with a Kd value of around 1.1 μM for MK-801 and a Ki value of around 147.4 μM for memantine. The calculated binding free energies −9.78 ± 1.61 kcal mol−1 (MK-801) and −7.02 ± 1.24 kcal mol−1 (memantine), and thus the free-energy-derived dissociation constants, are subject to large errors, estimated as s.e.m., owing to lack of convergence from including long-range effects from lipid molecules surrounding the pore. We note that the contribution of pore-cavity collapse upon ligand binding to the binding free energy is not included in the free-energy calculations, which were performed with the pore-collapsed, intact, agonist-bound receptor. Also not included is the contribution of −ln2kBT (where kB is the Boltzmann constant and T is absolute temperature) arising from the two poses available to MK-801.
Extended Data Fig. 8 Hydrogen-bonding propensity between MK-801 and memantine, and the selectivity filter asparagine residues.
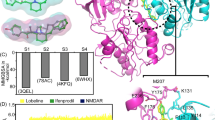
a, b, The two N-site asparagine residues N614 (GluN1) and N612 (GluN2B) of the pore-loop tips and the N + 1 asparagine residue, N613, of the GluN2B subunit, which is believed to be involved in the voltage-dependence of memantine binding. For MK-801 (a): left, data obtained at zero transmembrane voltage (simulations 20–23, N = 4); middle, −197.9 ± 1.2 mV (simulation 19, N = 1); right, −593.9 ± 3.8 mV (simulation 18, N = 1); n ≫ 10. For memantine (b): left, data obtained at zero transmembrane voltage (simulations 24–27, N = 4); middle, −196.9 ± 0.1, −205.1 ± 0.1, and −204.1 ± 0.1 mV (simulations 30–32, N = 3); right, −592.6 ± 0.3 and −592.7 ± 0.3 mV (simulations 28 and 29, N = 2); n ≫ 10. Hydrogen-bonding propensity is relatively low for N614 of GluN1, except at high voltage, suggesting that N612 and N613 of GluN2B are more important for pore-blocker binding and its voltage dependence, respectively; at non-zero transmembrane voltage, hydrogen-bonding propensity increases at the N + 1 site asparagine N613 of GluN2B. Error bars, s.d. of the mean calculated from all individual data points aggregated across all simulations.
Extended Data Fig. 9 Binding mode distributions of MK-801 and memantine.
a, MK-801 r.m.s.d. distributions (in Å; heavy atoms only) obtained from MK-801-binding simulations 21–23, with respect to all MK-801 poses obtained in binding simulation 20. Mean (µ) and standard deviation (σ) are indicated in Å; solid red lines are best fits to a normal distribution, but the distributions for simulations 21 and 23 show clear evidence of two r.m.s.d. populations, consistent with the observation that MK-801 can block the pore in two symmetry-related poses. The degree of asymmetry of the distributions observed for simulations 21 and 23 indicates non-equal occupancy of the two poses, a result of incomplete sampling. We note that in simulation 22, one of the two poses almost completely predominates. N = 1 for each panel; n ≫ 10. b, Memantine r.m.s.d. distributions (in Å; heavy atoms only), obtained from binding simulations 25–27, with respect to all poses obtained in memantine-binding simulation 24. Mean (μ) and standard deviation (σ) are indicated in Å; solid lines are best fits to a normal distribution. The relatively narrow and unimodal distributions reflect that memantine appears to predominantly block the pore in a single pose. The heavy-atom average r.m.s.d. of the main poses of memantine was 3.7 ± 0.2 Å, less than that observed for MK-801. N = 1 for each panel; n ≫ 10. c, MK-801-I poses obtained in simulations with and without selectivity filter backbone torsional corrections. Grey, the two predominant poses observed with corrections (simulation 1); cyan and orange, predominant poses identified from the initial portion (1–3 µs), before the filter deteriorated too extensively, of two different simulations without torsional corrections (pose 1, simulation 42; pose 2, simulation 47). One hundred individual poses from the initial portion (1–3 µs, uniformly separated by 0.02 µs) are shown as cyan and orange lines with the iodine atoms shown as spheres. Both poses of MK-801-I observed in our simulations with torsional backbone corrections (simulation 1) were therefore also observed, with comparable stability, in these additional simulations without these corrections. d, MK-801 poses obtained in free-binding simulations with and without filter backbone torsional corrections. Grey, the two distinct poses observed in a free binding simulation with backbone corrections (simulation 20); cyan and orange, poses in the three independent simulations without corrections in which MK-801 bound stably (simulation 50: pose identification period, 4–18 µs; simulation 51: pose identification period, 9–12 µs; simulation 53: pose identification period, 3–4 µs). MK-801 bound stably to the receptor in three (simulations 50, 51 and 53) out of five simulations performed without corrections—again in two distinct poses, as observed in our simulations with torsional corrections—and some closure of the activation gate (that is, the bundle-crossing region) was also observed in these three simulations before the filter deteriorated.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1: Molecular dynamics simulations with a detailed table legend
Video 1: Structure of ∆ATD NMDA receptor and a morph of the rearrangements of the LBD layer.
This video demonstrates the overall structure of ∆ATD NMDA receptor and a morph of the rearrangements of the LBD layer in going from the ΔATD arrangement to that of the intact, Δ2 receptor, showing how the LBD subunits swap partners
Video 2: MK-801 binding (Sim. 20) viewed along the membrane plane and toward the pore.
Only GluN1 residues 562–642 and GluN2B residues 554–634 are included in the surface cross section; asparagine residues N614 (GluN1) and N612 (GluN2B) are shown as sticks
Video 3: MK-801 binding (Sim. 20) viewed from the extracellular side of the pore and along the pore axis.
Only GluN1 residues 562–642 and GluN2B residues 554–634 are included
Video 4: Memantine binding (Sim. 25) viewed along the membrane plane and toward the pore.
Only GluN1 residues 562–650 and GluN2B residues 554–648 are included in the surface cross section; asparagine residues N614 (GluN1) and N612 (GluN2B) are shown as sticks
Video 5: Memantine binding (Sim. 25) viewed from the extracellular side of the pore and along the pore axis.
Only GluN1 residues 562–650 and GluN2B residues 554–648 are included
Rights and permissions
About this article
Cite this article
Song, X., Jensen, M.Ø., Jogini, V. et al. Mechanism of NMDA receptor channel block by MK-801 and memantine. Nature 556, 515–519 (2018). https://doi.org/10.1038/s41586-018-0039-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0039-9
This article is cited by
-
Enhanced NMDA receptor pathway and glutamate transmission in the hippocampal dentate gyrus mediate the spatial learning and memory impairment of obese rats
Pflügers Archiv - European Journal of Physiology (2024)
-
Structural mobility tunes signalling of the GluA1 AMPA glutamate receptor
Nature (2023)
-
BACE1 in PV interneuron tunes hippocampal CA1 local circuits and resets priming of fear memory extinction
Molecular Psychiatry (2023)
-
GluN2B subunit selective N-methyl-D-aspartate receptor ligands: Democratizing recent progress to assist the development of novel neurotherapeutics
Molecular Diversity (2023)
-
Cholecystokinin Signaling can Rescue Cognition and Synaptic Plasticity in the APP/PS1 Mouse Model of Alzheimer’s Disease
Molecular Neurobiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.