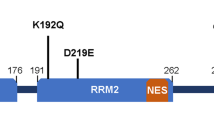
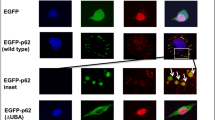
Abstract
Cytoplasmic aggregation of TAR DNA-binding protein 43 (TDP43; also known as TARDBP or TDP-43) is a key pathological feature of several neurodegenerative diseases, including amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). TDP43 typically resides in the nucleus but can shuttle between the nucleus and the cytoplasm to exert its multiple functions, which include regulation of the splicing, trafficking and stabilization of RNA. Cytoplasmic mislocalization and nuclear loss of TDP43 have both been associated with ALS and FTD, suggesting that calibrated levels and correct localization of TDP43 — achieved through an autoregulatory loop and tightly controlled nucleocytoplasmic transport — safeguard its normal function. Furthermore, TDP43 can undergo phase transitions, including its dispersion into liquid droplets and its accumulation into irreversible cytoplasmic aggregates. Thus, autoregulation, nucleocytoplasmic transport and phase transition are all part of an intrinsic control system regulating the physiological levels and localization of TDP43, and together are essential for the cellular homeostasis that is affected in neurodegenerative disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ou, S. H., Wu, F., Harrich, D., García-Martínez, L. F. & Gaynor, R. B. Cloning and characterization of a novel cellular protein, TDP-43, that binds to human immunodeficiency virus type 1 TAR DNA sequence motifs. J. Virol. 69, 3584–3596 (1995).
Buratti, E. et al. Nuclear factor TDP-43 and SR proteins promote in vitro and in vivo CFTR exon 9 skipping. EMBO J. 20, 1774–1784 (2001). This paper was the first to describe splicing regulation as a major function of TDP43 activity.
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Arai, T. et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem. Biophys. Res. Commun. 351, 602–611 (2006). Together with Neumann et al. (2006), this paper identifies TDP43 as the main component of proteinaceous inclusions in ALS and FTD, and describes nuclear depletion and cytoplasmic accumulation as major hallmarks of disease.
Heyburn, L. & Moussa, C. E.-H. TDP-43 in the spectrum of MND-FTLD pathologies. Mol. Cell. Neurosci. 83, 46–54 (2017).
Robberecht, W. & Philips, T. The changing scene of amyotrophic lateral sclerosis. Nat. Rev. Neurosci. 14, 248–264 (2013).
Ling, S.-C., Polymenidou, M. & Cleveland, D. W. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79, 416–438 (2013).
Cook, C., Zhang, Y.-J., Xu, Y., Dickson, D. W. & Petrucelli, L. TDP-43 in neurodegenerative disorders. Expert Opin. Biol. Ther. 8, 969–978 (2008).
Arai, T. et al. Phosphorylated TDP-43 in Alzheimer’s disease and dementia with Lewy bodies. Acta Neuropathol. 117, 125–136 (2009).
Gao, J., Wang, L., Huntley, M. L., Perry, G. & Wang, X. Pathomechanisms of TDP-43 in neurodegeneration. J. Neurochem. 146, 7–20 (2018).
McAleese, K. E. et al. TDP-43 pathology in Alzheimer’s disease, dementia with Lewy bodies and ageing. Brain Pathol. 27, 472–479 (2017).
James, B. D. et al. TDP-43 stage, mixed pathologies, and clinical Alzheimer’s-type dementia. Brain 139, 2983–2993 (2016).
Amador-Ortiz, C. et al. TDP-43 immunoreactivity in hippocampal sclerosis and Alzheimer’s disease. Ann. Neurol. 61, 435–445 (2007).
Nakashima-Yasuda, H. et al. Co-morbidity of TDP-43 proteinopathy in Lewy body related diseases. Acta Neuropathol. 114, 221–229 (2007).
Higashi, S. et al. Concurrence of TDP-43, tau and α-synuclein pathology in brains of Alzheimer’s disease and dementia with Lewy bodies. Brain Res. 1184, 284–294 (2007).
Schwab, C., Arai, T., Hasegawa, M., Yu, S. & McGeer, P. L. Colocalization of transactivation-responsive DNA-binding protein 43 and huntingtin in inclusions of Huntington disease. J. Neuropathol. Exp. Neurol. 67, 1159–1165 (2008).
Uryu, K. et al. Concomitant TAR-DNA-binding protein 43 pathology is present in Alzheimer disease and corticobasal degeneration but not in other tauopathies. J. Neuropathol. Exp. Neurol. 67, 555–564 (2008).
Takeda, T. Possible concurrence of TDP-43, tau and other proteins in amyotrophic lateral sclerosis/frontotemporal lobar degeneration. Neuropathology 38, 72–81 (2018).
Buratti, E. & Baralle, F. E. Characterization and functional implications of the RNA binding properties of nuclear factor TDP-43, a novel splicing regulator of CFTR exon 9. J. Biol. Chem. 276, 36337–36343 (2001).
Wang, H. Y., Wang, I. F., Bose, J. & Shen, C. K. J. Structural diversity and functional implications of the eukaryotic TDP gene family. Genomics 83, 130–139 (2004).
Ayala, Y. M. et al. Human, Drosophila, and C. elegans TDP43: nucleic acid binding properties and splicing regulatory function. J. Mol. Biol. 348, 575–588 (2005).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Lee, E. B., Lee, V. M.-Y. & Trojanowski, J. Q. Gains or losses: molecular mechanisms of TDP43-mediated neurodegeneration. Nat. Rev. Neurosci. 13, 38–50 (2012).
Tollervey, J. R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Buratti, E. et al. TDP-43 binds heterogeneous nuclear ribonucleoprotein A/B through its C-terminal tail: an important region for the inhibition of cystic fibrosis transmembrane conductance regulator exon 9 splicing. J. Biol. Chem. 280, 37572–37584 (2005).
Freibaum, B. D., Chitta, R. K., High, A. A. & Taylor, J. P. Global analysis of TDP-43 interacting proteins reveals strong association with RNA splicing and translation machinery. J. Proteome Res. 9, 1104–1120 (2010).
Budini, M. et al. Role of selected mutations in the Q/N rich region of TDP-43 in EGFP-12xQ/N-induced aggregate formation. Brain Res. 1462, 139–150 (2012).
Budini, M. et al. Cellular model of TAR DNA-binding protein 43 (TDP-43) aggregation based on its C-terminal Gln/Asn-rich region. J. Biol. Chem. 287, 7512–7525 (2012).
Cushman, M., Johnson, B. S., King, O. D., Gitler, A. D. & Shorter, J. Prion-like disorders: blurring the divide between transmissibility and infectivity. J. Cell Sci. 123, 1191–1201 (2010).
Fuentealba, R. A. et al. Interaction with polyglutamine aggregates reveals a Q/N-rich domain in TDP-43. J. Biol. Chem. 285, 26304–26314 (2010).
Jiang, L.-L. et al. Two mutations G335D and Q343R within the amyloidogenic core region of TDP-43 influence its aggregation and inclusion formation. Sci. Rep. 6, 23928 (2016).
Schmidt, H. B., Barreau, A. & Rohatgi, R. Phase separation-deficient TDP43 remains functional in splicing. Nat. Commun. 10, 4890 (2019).
Mompeán, M. et al. Structural characterization of the minimal segment of TDP-43 competent for aggregation. Arch. Biochem. Biophys. 545, 53–62 (2014).
Conicella, A. E., Zerze, G. H., Mittal, J. & Fawzi, N. L. ALS mutations disrupt phase separation mediated by α-helical structure in the TDP-43 low-complexity C-terminal domain. Structure 24, 1537–1549 (2016).
Conicella, A. E. et al. TDP-43 α-helical structure tunes liquid–liquid phase separation and function. Proc. Natl Acad. Sci. USA 117, 5883–5894 (2020).
Bolognesi, B. et al. The mutational landscape of a prion-like domain. Nat. Commun. 10, 4162 (2019).
Hughes, M. P. et al. Atomic structures of low-complexity protein segments reveal kinked β sheets that assemble networks. Science 359, 698–701 (2018).
Guenther, E. L. et al. Atomic structures of TDP-43 LCD segments and insights into reversible or pathogenic aggregation. Nat. Struct. Mol. Biol. 25, 463–471 (2018).
Guenther, E. L. et al. Atomic-level evidence for packing and positional amyloid polymorphism by segment from TDP-43 RRM2. Nat. Struct. Mol. Biol. 25, 311–319 (2018).
Sawaya, M. R. et al. Atomic structures of amyloid cross-β spines reveal varied steric zippers. Nature 447, 453–457 (2007).
Kim, H. J. et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495, 467–473 (2013).
Cao, Q., Boyer, D. R., Sawaya, M. R., Ge, P. & Eisenberg, D. S. Cryo-EM structures of four polymorphic TDP-43 amyloid cores. Nat. Struct. Mol. Biol. 26, 619–627 (2019).
Afroz, T. et al. Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation. Nat. Commun. 8, 45 (2017).
Mercado, P. A., Ayala, Y. M., Romano, M., Buratti, E. & Baralle, F. E. Depletion of TDP 43 overrides the need for exonic and intronic splicing enhancers in the human apoA-II gene. Nucleic Acids Res. 33, 6000–6010 (2005).
Gu, J. et al. Transactive response DNA-binding protein 43 (TDP-43) regulates alternative splicing of tau exon 10: implications for the pathogenesis of tauopathies. J. Biol. Chem. 292, 10600–10612 (2017).
Goedert, M., Spillantini, M. G., Jakes, R., Rutherford, D. & Crowther, R. A. Multiple isoforms of human microtubule-associated protein tau: sequences and localization in neurofibrillary tangles of Alzheimer’s disease. Neuron 3, 519–526 (1989).
Mulot, S. F. C., Hughes, K., Woodgett, J. R., Anderton, B. H. & Hanger, D. P. PHF-tau from Alzheimer’s brain comprises four species on SDS-PAGE which can be mimicked by in vitro phosphorylation of human brain tau by glycogen synthase kinase-3β. FEBS Lett. 349, 359–364 (1994).
Liu, W.-K. et al. Relationship of the extended tau haplotype to tau biochemistry and neuropathology in progressive supranuclear palsy. Ann. Neurol. 50, 494–502 (2001).
Takanashi, M., Mori, H., Arima, K., Mizuno, Y. & Hattori, N. Expression patterns of tau mRNA isoforms correlate with susceptible lesions in progressive supranuclear palsy and corticobasal degeneration. Mol. Brain Res. 104, 210–219 (2002).
Umeda, Y. et al. Alterations in human tau transcripts correlate with those of neurofilament in sporadic tauopathies. Neurosci. Lett. 359, 151–154 (2004).
Becker, L. A. et al. Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature 544, 367–371 (2017).
Ling, S. C. et al. ALS-associated mutations in TDP-43 increase its stability and promote TDP-43 complexes with FUS/TLS. Proc. Natl Acad. Sci. USA 107, 13318–13323 (2010).
Appocher, C. et al. Major hnRNP proteins act as general TDP-43 functional modifiers both in Drosophila and human neuronal cells. Nucleic Acids Res. 45, 8026–8045 (2017).
Prudencio, M. et al. Misregulation of human sortilin splicing leads to the generation of a nonfunctional progranulin receptor. Proc. Natl Acad. Sci. USA 109, 21510–21515 (2012).
Shiga, A. et al. Alteration of POLDIP3 splicing associated with loss of function of TDP-43 in tissues affected with ALS. PLoS ONE 7, e43120 (2012).
Deshaies, J.-E. et al. TDP-43 regulates the alternative splicing of hnRNP A1 to yield an aggregation-prone variant in amyotrophic lateral sclerosis. Brain 141, 1320–1333 (2018).
D’Ambrogio, A. et al. Functional mapping of the interaction between TDP-43 and hnRNP A2 in vivo. Nucleic Acids Res. 37, 4116–4126 (2009).
Xiao, S. et al. RNA targets of TDP-43 identified by UV-CLIP are deregulated in ALS. Mol. Cell. Neurosci. 47, 167–180 (2011).
Bhardwaj, A., Myers, M. P., Buratti, E. & Baralle, F. E. Characterizing TDP-43 interaction with its RNA targets. Nucleic Acids Res. 41, 5062–5074 (2013).
Fan, Z., Chen, X. & Chen, R. Transcriptome-wide analysis of TDP-43 binding small RNAs identifies miR-NID1 (miR-8485), a novel miRNA that represses NRXN1 expression. Genomics 103, 76–82 (2014).
Kapeli, K. et al. Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses. Nat. Commun. 7, 12143 (2016).
Fratta, P. et al. Mice with endogenous TDP-43 mutations exhibit gain of splicing function and characteristics of amyotrophic lateral sclerosis. EMBO J. 37, e98684 (2018).
Ling, J. P., Pletnikova, O., Troncoso, J. C. & Wong, P. C. TDP-43 repression of nonconserved cryptic exons is compromised in ALS-FTD. Science 349, 650–655 (2015).
Tan, Q. et al. Extensive cryptic splicing upon loss of RBM17 and TDP43 in neurodegeneration models. Hum. Mol. Genet. 25, 5083–5093 (2016).
Rot, G. et al. High-resolution RNA maps suggest common principles of splicing and polyadenylation regulation by TDP-43. Cell Rep. 19, 1056–1067 (2017).
Melamed, Z. et al. Premature polyadenylation-mediated loss of stathmin-2 is a hallmark of TDP-43-dependent neurodegeneration. Nat. Neurosci. 22, 180–190 (2019).
Klim, J. R. et al. ALS-implicated protein TDP-43 sustains levels of STMN2, a mediator of motor neuron growth and repair. Nat. Neurosci. 22, 167–179 (2019).
Prudencio, M. et al. Truncated stathmin-2 is a marker of TDP-43 pathology in frontotemporal dementia. J. Clin. Invest. 130, 6080–6092 (2020). Together with Melamed et al. (2019) and Klim et al. (2019), this paper identifies and describes TDP43-mediated regulation of stathmin-2 by TDP43 and its disease-related deregulation as a defining feature of ALS and FTD.
White, M. A. et al. TDP-43 gains function due to perturbed autoregulation in a Tardbp knock-in mouse model of ALS-FTD. Nat. Neurosci. 21, 552–563 (2018). This study introduces a single amino acid change in mouse TDP43 mimicking ALS-related mutant Q331K that affects the 3D structure and function of its low-complexity domain, thereby uncovering a direct relation to defective autoregulation.
Johnson, B. S., McCaffery, J. M., Lindquist, S. & Gitler, A. D. A yeast TDP-43 proteinopathy model: exploring the molecular determinants of TDP-43 aggregation and cellular toxicity. Proc. Natl Acad. Sci. USA 105, 6439–6444 (2008).
Li, Y. et al. A Drosophila model for TDP-43 proteinopathy. Proc. Natl Acad. Sci. USA 107, 3169–3174 (2010).
Diaper, D. C. et al. Loss and gain of Drosophila TDP-43 impair synaptic efficacy and motor control leading to age-related neurodegeneration by loss-of-function phenotypes. Hum. Mol. Genet. 22, 1539–1557 (2013).
Schmid, B. et al. Loss of ALS-associated TDP-43 in zebrafish causes muscle degeneration, vascular dysfunction, and reduced motor neuron axon outgrowth. Proc. Natl Acad. Sci. USA 110, 4986–4991 (2013).
Wils, H. et al. TDP-43 transgenic mice develop spastic paralysis and neuronal inclusions characteristic of ALS and frontotemporal lobar degeneration. Proc. Natl Acad. Sci. USA 107, 3858–3863 (2010).
Wegorzewska, I., Bell, S., Cairns, N. J., Miller, T. M. & Baloh, R. H. TDP-43 mutant transgenic mice develop features of ALS and frontotemporal lobar degeneration. Proc. Natl Acad. Sci. USA 106, 18809–18814 (2009).
Swarup, V. et al. Pathological hallmarks of amyotrophic lateral sclerosis/frontotemporal lobar degeneration in transgenic mice produced with TDP-43 genomic fragments. Brain 134, 2610–2626 (2011).
Dayton, R. D. et al. Selective forelimb impairment in rats expressing a pathological TDP-43 25 kDa C-terminal fragment to mimic amyotrophic lateral sclerosis. Mol. Ther. 21, 1324–1334 (2013).
Uchida, A. et al. Non-human primate model of amyotrophic lateral sclerosis with cytoplasmic mislocalization of TDP-43. Brain 135, 833–846 (2012).
Ayala, Y. M. et al. TDP-43 regulates its mRNA levels through a negative feedback loop. EMBO J. 30, 277–288 (2011). This study was the first to report that autoregulation of TDP43 is a mechanism to regulate its expression level and localization.
Avendaño-Vázquez, S. E. et al. Autoregulation of TDP-43 mRNA levels involves interplay between transcription, splicing, and alternative polyA site selection. Genes Dev. 26, 1679–1684 (2012).
Koyama, A. et al. Increased cytoplasmic TARDBP mRNA in affected spinal motor neurons in ALS caused by abnormal autoregulation of TDP-43. Nucleic Acids Res. 44, 5820–5836 (2016).
Bembich, S. et al. Predominance of spliceosomal complex formation over polyadenylation site selection in TDP-43 autoregulation. Nucleic Acids Res. 42, 3362–3371 (2014).
Pons, M. et al. Splicing factors act as genetic modulators of TDP-43 production in a new autoregulatory TDP-43 Drosophila model. Hum. Mol. Genet. 26, 3396–3408 (2017).
D’Alton, S., Altshuler, M. & Lewis, J. Studies of alternative isoforms provide insight into TDP-43 autoregulation and pathogenesis. RNA 21, 1419–1432 (2015).
Sugai, A. et al. Non-genetically modified models exhibit TARDBP mRNA increase due to perturbed TDP-43 autoregulation. Neurobiol. Dis. 130, 104534 (2019).
Flores, B. N. et al. An intramolecular salt bridge linking TDP43 RNA binding, protein stability, and TDP43-dependent neurodegeneration. Cell Rep. 27, 1133–1150.e8 (2019).
Weskamp, K. et al. Shortened TDP43 isoforms upregulated by neuronal hyperactivity drive TDP43 pathology in ALS. J. Clin. Invest. 130, 1139–1155 (2020).
Gitcho, M. A. et al. TARDBP 3′-UTR variant in autopsy-confirmed frontotemporal lobar degeneration with TDP-43 proteinopathy. Acta Neuropathol. 118, 633–645 (2009).
Lee, S. M. et al. TDP-43 cytoplasmic inclusion formation is disrupted in C9orf72-associated amyotrophic lateral sclerosis/frontotemporal lobar degeneration. Brain Commun. 1, fcz014 (2019).
Chook, Y. M. & Blobel, G. Karyopherins and nuclear import. Curr. Opin. Struct. Biol. 11, 703–715 (2001).
Prpar Mihevc, S. et al. Nuclear trafficking in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. Brain 140, 13–26 (2017).
Nishimura, A. L. et al. Nuclear import impairment causes cytoplasmic trans-activation response DNA-binding protein accumulation and is associated with frontotemporal lobar degeneration. Brain 133, 1763–1771 (2010). This study was the first to show direct interactions between TDP43 and karyopherins, alterations of which characterize ALS and FTD.
Winton, M. J. et al. Disturbance of nuclear and cytoplasmic TAR DNA-binding protein (TDP-43) induces disease-like redistribution, sequestration, and aggregate formation. J. Biol. Chem. 283, 13302–13309 (2008).
Ederle, H. et al. Nuclear egress of TDP-43 and FUS occurs independently of Exportin-1/CRM1. Sci. Rep. 8, 7084 (2018).
Murphy, R. & Wente, S. R. An RNA-export mediator with an essential nuclear export signal. Nature 383, 357–360 (1996).
Ederle, H. & Dormann, D. TDP-43 and FUS en route from the nucleus to the cytoplasm. FEBS Lett. 591, 1489–1507 (2017).
Folkmann, A. W., Dawson, T. R. & Wente, S. R. Insights into mRNA export-linked molecular mechanisms of human disease through a Gle1 structure–function analysis. Adv. Biol. Regul. 54, 74–91 (2014).
Kaneb, H. M. et al. Deleterious mutations in the essential mRNA metabolism factor, hGle1, in amyotrophic lateral sclerosis. Hum. Mol. Genet. 24, 1363–1373 (2015).
Aditi, Glass, L., Dawson, T. R. & Wente, S. R. An amyotrophic lateral sclerosis-linked mutation in GLE1 alters the cellular pool of human Gle1 functional isoforms. Adv. Biol. Regul. 62, 25–36 (2016).
Berson, A. et al. Drosophila Ref1/ALYREF regulates transcription and toxicity associated with ALS/FTD disease etiologies. Acta Neuropathol. Commun. 7, 65 (2019).
Pinarbasi, E. S. et al. Active nuclear import and passive nuclear export are the primary determinants of TDP-43 localization. Sci. Rep. 8, 7083 (2018).
Archbold, H. C. et al. TDP43 nuclear export and neurodegeneration in models of amyotrophic lateral sclerosis and frontotemporal dementia. Sci. Rep. 8, 4606 (2018).
Wickramasinghe, V. O. & Laskey, R. A. Control of mammalian gene expression by selective mRNA export. Nat. Rev. Mol. Cell Biol. 16, 431–442 (2015).
Igaz, L. M. et al. Dysregulation of the ALS-associated gene TDP-43 leads to neuronal death and degeneration in mice. J. Clin. Invest. 121, 726–738 (2011).
Walker, A. K. et al. Functional recovery in new mouse models of ALS/FTLD after clearance of pathological cytoplasmic TDP-43. Acta Neuropathol. 130, 643–660 (2015). This study shows that the reversible expression of human TARDBP carrying a deletion of the TDP43 NLS in mice results in accumulation of cytoplasmic aggregates and loss of nuclear TDP43, leading to neurodegeneration and motor dysfunction. The subsequent removal of cytoplasmic TDP43 and return of nuclear TDP43 lead to neuron preservation, muscle reinnervation and functional recovery.
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Scaber, J. & Talbot, K. What is the role of TDP-43 in C9orf72-related amyotrophic lateral sclerosis and frontemporal dementia? Brain 139, 3057–3059 (2016).
Mori, K. et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339, 1335–1338 (2013).
Zhang, K. et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature 525, 56–61 (2015).
Freibaum, B. D. et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature 525, 129–133 (2015).
Jovičić, A. et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat. Neurosci. 18, 1226–1229 (2015).
Lee, K. H. et al. C9orf72 dipeptide repeats impair the assembly, dynamics, and function of membrane-less organelles. Cell 167, 774–788 (2016).
Boeynaems, S., Bogaert, E., Van Damme, P. & Van Den Bosch, L. Inside out: the role of nucleocytoplasmic transport in ALS and FTLD. Acta Neuropathol. 132, 159–173 (2016).
Boeynaems, S. et al. Drosophila screen connects nuclear transport genes to DPR pathology in c9ALS/FTD. Sci. Rep. 6, 20877 (2016).
Chou, C.-C. et al. TDP-43 pathology disrupts nuclear pore complexes and nucleocytoplasmic transport in ALS/FTD. Nat. Neurosci. 21, 228–239 (2018).
Solomon, D. A. et al. A feedback loop between dipeptide-repeat protein, TDP-43 and karyopherin-α mediates C9orf72-related neurodegeneration. Brain 141, 2908–2924 (2018).
Babinchak, W. M. et al. The role of liquid–liquid phase separation in aggregation of the TDP-43 low-complexity domain. J. Biol. Chem. 294, 6306–6317 (2019).
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Alberti, S. & Hyman, A. A. Are aberrant phase transitions a driver of cellular aging? BioEssays 38, 959–968 (2016).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, 4382 (2017).
French, R. L. et al. Detection of TAR DNA-binding protein 43 (TDP-43) oligomers as initial intermediate species during aggregate formation. J. Biol. Chem. 294, 6696–6709 (2019).
Guo, L. et al. Nuclear-import receptors reverse aberrant phase transitions of RNA-binding proteins with prion-like domains. Cell 173, 677–692 (2018).
Gasset-Rosa, F. et al. Cytoplasmic TDP-43 de-mixing independent of stress granules drives inhibition of nuclear import, loss of nuclear TDP-43, and cell death. Neuron 102, 339–357 (2019).
Mann, J. R. et al. RNA binding antagonizes neurotoxic phase transitions of TDP-43. Neuron 102, 321–338.e8 (2019).
Gopal, P. P., Nirschl, J. J., Klinman, E. & Holzbaur, E. L. F. Amyotrophic lateral sclerosis-linked mutations increase the viscosity of liquid-like TDP-43 RNP granules in neurons. Proc. Natl Acad. Sci. USA 114, E2466–E2475 (2017).
Wang, A. et al. A single N-terminal phosphomimic disrupts TDP-43 polymerization, phase separation, and RNA splicing. EMBO J. 37, e97452 (2018).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Li, H. R., Chiang, W. C., Chou, P. C., Wang, W. J. & Huang, J. R. TAR DNA-binding protein 43 (TDP-43) liquid–liquid phase separation is mediated by just a few aromatic residues. J. Biol. Chem. 293, 6090–6098 (2018).
Jucker, M. & Walker, L. C. Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 501, 45–51 (2013).
Brettschneider, J. et al. Stages of pTDP-43 pathology in amyotrophic lateral sclerosis. Ann. Neurol. 74, 20–38 (2013).
Brettschneider, J. et al. Sequential distribution of pTDP-43 pathology in behavioral variant frontotemporal dementia (bvFTD). Acta Neuropathol. 127, 423–439 (2014).
Kawakami, I., Arai, T. & Hasegawa, M. The basis of clinicopathological heterogeneity in TDP-43 proteinopathy. Acta Neuropathol. 138, 751–770 (2019).
Hofweber, M. et al. Phase separation of FUS is suppressed by its nuclear import receptor and arginine methylation. Cell 173, 706–719.e13 (2018).
Woerner, A. C. et al. Cytoplasmic protein aggregates interfere with nucleocytoplasmic transport of protein and RNA. Science 351, 173–176 (2016). This study shows, using cell culture models, that cytoplasmic accumulation of TDP43 fragments causes the sequestration and mislocalization of proteins containing disordered and low-complexity domains. These include multiple factors involved in the nuclear import and export machinery, leading to impaired nucleocytoplasmic transport.
Kosyna, F. K. & Depping, R. Controlling the gatekeeper: therapeutic targeting of nuclear transport. Cells 7, 221 (2018).
Haines, J. D. et al. Nuclear export inhibitors avert progression in preclinical models of inflammatory demyelination. Nat. Neurosci. 18, 511–520 (2015).
Hasegawa, M. et al. Phosphorylated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Ann. Neurol. 64, 60–70 (2008).
Neumann, M. et al. Phosphorylation of S409/410 of TDP-43 is a consistent feature in all sporadic and familial forms of TDP-43 proteinopathies. Acta Neuropathol. 117, 137–149 (2009).
Acknowledgements
The authors thank somersault18:24 for valuable assistance with figure assembly and design. P.T. thanks M. Kalthoff for helpful discussions during the preparation of the manuscript. Research by P.T. and L.V.D.B. is supported by VIB, KU Leuven (C1 and Opening the Future Fund), the Fund for Scientific Research Flanders (FWO-Vlaanderen), the Thierry Latran Foundation, the Association Belge contre les Maladies neuro-Musculaires (ABMM), the Muscular Dystrophy Association (MDA), the Association Française contre les Myopathies (AFM), Target ALS, the ALS Liga België (A Cure for ALS) and the ALS Association (ALSA). P.T. is a PhD fellow of FWO-Vlaanderen (11C7621N). F.H. acknowledges support from the UK Medical Research Council (G0701498; MR/L010666/1), the MND Association (Hirth/Nov15/914-793; Hirth/Oct13/6202; Hirth/Mar12/6085; Hirth/Oct07/6233) and Alzheimer’s Research UK (Hirth/ARUK/2012).
Author information
Authors and Affiliations
Contributions
P.T. and F.H. wrote the manuscript and all authors made substantial contributions to the discussion of the content and reviewed and edited the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Neuroscience thanks E. Buratti, C. Lagier-Tourenne and J. Ule (who co-reviewed with M. Hallegger) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Heterogeneous nuclear ribonucleoprotein
-
(hnRNP). An RNA-binding protein involved in several aspects of nucleic acid metabolism such as transcriptional and translational regulation, alternative splicing and mRNA stabilization.
- Splicing
-
A form of processing of a newly synthesized precursor mRNA transcript during which introns are removed and exons are joined together, thereby transforming it into a mature mRNA.
- Proteinaceous inclusions
-
Aggregates of one or more proteins found in the nucleus and/or the cytoplasm, a main characteristic of many neurodegenerative diseases.
- Aggregates
-
Accumulations of misfolded and/or intrinsically disordered proteins that form amyloid-like fibrils and insoluble depositions in the nucleus and/or the cytoplasm and are highly associated with neurodegeneration and pathology.
- Low-complexity domain
-
An intrinsically disordered protein sequence that is unable to form orderly 3D structures, such as α-helices or β-strands, and is thus prone to intermolecular interactions and involved in phase separation and the formation of pathological aggregates.
- Stress granules
-
Membraneless organelles formed upon cellular stress by liquid demixing and accumulation of RNA and proteins.
- Phase separation
-
The creation of two different phases from a homogeneous mixture, which can occur by liquid demixing of soluble monomeric proteins and includes their dispersion into liquid droplets and accumulation into irreversible cytoplasmic aggregates.
- Polymorphs
-
Proteins whose sequences enable them to adopt different structural conformations.
- Cellular stress
-
A consequence of environmental or chemical stressors, such as extreme temperatures, alterations in pH or exposure to toxins, that threatens the internal state and, ultimately, the survival of a cell.
- Cryptic exons
-
Coding sequences within a precursor mRNA that are normally not included in the mature RNA but can occur in an alternative form of the mRNA.
- Nonsense-mediated decay
-
A surveillance and degradation mechanism in eukaryotes, mainly eliminating mRNAs containing premature stop codons.
- Alternative polyadenylation
-
A process of gene regulation leading to the production of mRNA isoforms in which the formation of poly(A) tails is initiated at different positions in the precursor mRNA.
- RNA exosome
-
A ring-structured multi-protein complex found in the cytoplasm, nucleus or nucleolus, responsible for degrading different kinds of RNA molecules.
- Molecular chaperones
-
Proteins that assist in the correct and timely folding or unfolding of other proteins or the assembly and disassembly of macromolecular complexes.
Rights and permissions
About this article
Cite this article
Tziortzouda, P., Van Den Bosch, L. & Hirth, F. Triad of TDP43 control in neurodegeneration: autoregulation, localization and aggregation. Nat Rev Neurosci 22, 197–208 (2021). https://doi.org/10.1038/s41583-021-00431-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41583-021-00431-1
This article is cited by
-
Dysregulation of stress granule dynamics by DCTN1 deficiency exacerbates TDP-43 pathology in Drosophila models of ALS/FTD
Acta Neuropathologica Communications (2024)
-
PP2A and GSK3 act as modifiers of FUS-ALS by modulating mitochondrial transport
Acta Neuropathologica (2024)
-
Inhibition of the MEK/ERK pathway suppresses immune overactivation and mitigates TDP-43 toxicity in a Drosophila model of ALS
Immunity & Ageing (2023)
-
Pathological insights from amyotrophic lateral sclerosis animal models: comparisons, limitations, and challenges
Translational Neurodegeneration (2023)
-
Preclinical investigation in FAAH inhibition as a neuroprotective therapy for frontotemporal dementia using TDP-43 transgenic male mice
Journal of Neuroinflammation (2023)