Abstract
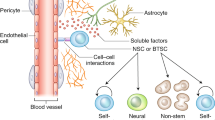
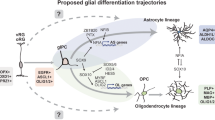
Gliomas are heterogeneous tumours derived from glial cells and remain the deadliest form of brain cancer. Although the glioma stem cell sits at the apex of the cellular hierarchy, how it produces the vast cellular constituency associated with frank glioma remains poorly defined. We explore glioma tumorigenesis through the lens of glial development, starting with the neurogenic–gliogenic switch and progressing through oligodendrocyte and astrocyte differentiation. Beginning with the factors that influence normal glial linage progression and diversity, a pattern emerges that has useful parallels in the development of glioma and may ultimately provide targetable pathways for much-needed new therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011). This review is the expanded version of the canonical primer on cancer biology.
Wainwright, E. N. & Scaffidi, P. Epigenetics and cancer stem cells: unleashing, hijacking, and restricting cellular plasticity. Trends Cancer 3, 372–386 (2017).
Takebe, N. et al. Targeting notch, hedgehog, and wnt pathways in cancer stem cells: clinical update. Nat. Rev. Clin. Oncol. 12, 445–464 (2015). This is a comprehensive review of the current pipeline therapies targeting developmental pathway components for glioma treatment.
Joruiz, S. M. & Bourdon, J. C. p53 isoforms: key regulators of the cell fate decision. Cold Spring Harb. Perspect. Med. https://doi.org/10.1101/cshperspect.a026039 (2016).
Veleva-Rotse, B. O. & Barnes, A. P. Brain patterning perturbations following PTEN loss. Front. Mol. Neurosci. 7, 35 (2014).
Molofsky, A. V. et al. Increasing p16INK4a expression decreases forebrain progenitors and neurogenesis during ageing. Nature 443, 448–452 (2006).
Sanchez-Martin, M. & Ferrando, A. The NOTCH1-MYC highway toward T cell acute lymphoblastic leukemia. Blood 129, 1124–1133 (2017).
Mok, T. S. Personalized medicine in lung cancer: what we need to know. Nat. Rev. Clin. Oncol. 8, 661–668 (2011).
Birey, F., Kokkosis, A. G. & Aguirre, A. Oligodendroglia-lineage cells in brain plasticity, homeostasis and psychiatric disorders. Curr. Opin. Neurobiol. 47, 93–103 (2017).
Molofsky, A. V. & Deneen, B. Astrocyte development: a guide for the perplexed. Glia 63, 1320–1329 (2015). This review provides a thorough and approachable description of astrocyte development.
Louis, D. N. et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 131, 803–820 (2016).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013). This study presents a characterization of adult GBM subtypes on the basis of relevant patterns of somatic mutations.
Baker, S. J., Ellison, D. W. & Gutmann, D. H. Pediatric gliomas as neurodevelopmental disorders. Glia 64, 879–895 (2016). This article presents a thorough discussion of the unique properties of paediatric gliomas, how they differ from adult gliomas and the genetic basis for their relationship to development.
Jones, C. & Baker, S. J. Unique genetic and epigenetic mechanisms driving paediatric diffuse high-grade glioma. Nat. Rev. Cancer https://doi.org/10.1038/nrc3811 (2014).
Mackay, A. et al. Integrated molecular meta-analysis of 1,000 pediatric high-grade and diffuse intrinsic pontine glioma. Cancer Cell 32, 520–537 e525 (2017). This paper provides a thorough and modern classification of paediatric glioma subtypes on the basis of patient genomic alterations.
Singh, S. K. et al. Identification of human brain tumour initiating cells. Nature 432, 396–401 (2004).
Wu, S., Wu, Y. & Capecchi, M. R. Motoneurons and oligodendrocytes are sequentially generated from neural stem cells but do not appear to share common lineage-restricted progenitors in vivo. Development 133, 581–590 (2006).
Masahira, N. et al. Olig2-positive progenitors in the embryonic spinal cord give rise not only to motoneurons and oligodendrocytes, but also to a subset of astrocytes and ependymal cells. Dev. Biol. 293, 358–369 (2006).
Ravanelli, A. M. & Appel, B. Motor neurons and oligodendrocytes arise from distinct cell lineages by progenitor recruitment. Genes Dev. 29, 2504–2515 (2015).
Mukouyama, Y. S. et al. Olig2+ neuroepithelial motoneuron progenitors are not multipotent stem cells in vivo. Proc. Natl Acad. Sci. USA 103, 1551–1556 (2006).
Stolt, C. C. et al. The Sox9 transcription factor determines glial fate choice in the developing spinal cord. Genes Dev. 17, 1677–1689 (2003).
Deneen, B. et al. The transcription factor NFIA controls the onset of gliogenesis in the developing spinal cord. Neuron 52, 953–968 (2006).
Kang, P. et al. Sox9 and NFIA coordinate a transcriptional regulatory cascade during the initiation of gliogenesis. Neuron 74, 79–94 (2012).
Glasgow, S. M. et al. Glia-specific enhancers and chromatin structure regulate NFIA expression and glioma tumorigenesis. Nat. Neurosci. 20, 1520–1528 (2017).
Hatakeyama, J. et al. Hes genes regulate size, shape and histogenesis of the nervous system by control of the timing of neural stem cell differentiation. Development 131, 5539–5550 (2004).
Park, H. C. & Appel, B. Delta-Notch signaling regulates oligodendrocyte specification. Development 130, 3747–3755 (2003).
Taylor, M. K., Yeager, K. & Morrison, S. J. Physiological notch signaling promotes gliogenesis in the developing peripheral and central nervous systems. Development 134, 2435–2447 (2007).
Namihira, M. et al. Committed neuronal precursors confer astrocytic potential on residual neural precursor cells. Dev. Cell 16, 245–255 (2009).
Suva, M. L. et al. Reconstructing and reprogramming the tumor-propagating potential of glioblastoma stem-like cells. Cell 157, 580–594 (2014).
Lan, X. et al. Fate mapping of human glioblastoma reveals an invariant stem cell hierarchy. Nature 549, 227–232 (2017). This study elegantly shows that intratumoural GBM heterogeneity arises from hierarchical fate progression of GSCs, similar to that of developmental GSCs.
Glasgow, S. M. et al. The miR-223/nuclear factor I-A axis regulates glial precursor proliferation and tumorigenesis in the CNS. J. Neurosci. 33, 13560–13568 (2013).
Song, H. R. et al. Nuclear factor IA is expressed in astrocytomas and is associated with improved survival. Neuro Oncol. 12, 122–132 (2010).
Glasgow, S. M. et al. Mutual antagonism between Sox10 and NFIA regulates diversification of glial lineages and glioma subtypes. Nat. Neurosci. 17, 1322–1329 (2014). This paper draws parallels between glial development and glioma tumorigenesis through misexpression of glial-specific transcription factors SOX10 and NFIA.
Schlierf, B. et al. Expression of SoxE and SoxD genes in human gliomas. Neuropathol. Appl. Neurobiol. 33, 621–630 (2007).
Liu, H. et al. SOX9 overexpression promotes glioma metastasis via wnt/beta-catenin signaling. Cell Biochem. Biophys. 73, 205–212 (2015).
Swartling, F. J. et al. Distinct neural stem cell populations give rise to disparate brain tumors in response to N-MYC. Cancer Cell 21, 601–613 (2012).
Teodorczyk, M. & Schmidt, M. H. Notching on cancer’s door: notch signaling in brain tumors. Front. Oncol. 4, 341 (2014).
Giachino, C. et al. A tumor suppressor function for notch signaling in forebrain tumor subtypes. Cancer Cell 28, 730–742 (2015).
Xie, Q. et al. RBPJ maintains brain tumor-initiating cells through CDK9-mediated transcriptional elongation. J. Clin. Invest. 126, 2757–2772 (2016).
Liau, B. B. et al. Adaptive chromatin remodeling drives glioblastoma stem cell plasticity and drug tolerance. Cell Stem Cell 20, 233–246 e237 (2017). This paper highlights mechanisms through which GSCs reutilize developmental pathways and epigenetic programmes to transition between proliferative states and plasticity.
Fancy, S. P., Chan, J. R., Baranzini, S. E., Franklin, R. J. & Rowitch, D. H. Myelin regeneration: a recapitulation of development? Annu. Rev. Neurosci. 34, 21–43 (2011).
Emery, B. & Lu, Q. R. Transcriptional and epigenetic regulation of oligodendrocyte development and myelination in the central nervous system. Cold Spring Harb. Perspect. Biol. 7, a020461 (2015).
Bergles, D. E. & Richardson, W. D. Oligodendrocyte development and plasticity. Cold Spring Harb. Perspect. Biol. 8, a020453 (2015).
Dimou, L. & Gallo, V. NG2-glia and their functions in the central nervous system. Glia 63, 1429–1451 (2015).
Zhu, X. et al. Age-dependent fate and lineage restriction of single NG2 cells. Development 138, 745–753 (2011).
Nishiyama, A., Boshans, L., Goncalves, C. M., Wegrzyn, J. & Patel, K. D. Lineage, fate, and fate potential of NG2-glia. Brain Res. 1638, 116–128 (2016). This paper provides an extensive mapping of the potential and fate of NG2 cells, a proposed cell of origin for glioma.
Vigano, F. & Dimou, L. The heterogeneous nature of NG2-glia. Brain Res. 1638, 129–137 (2016).
Lu, Q. R. et al. Sonic hedgehog — regulated oligodendrocyte lineage genes encoding bHLH proteins in the mammalian central nervous system. Neuron 25, 317–329 (2000).
Zhou, Q. & Anderson, D. J. The bHLH transcription factors OLIG2 and OLIG1 couple neuronal and glial subtype specification. Cell 109, 61–73 (2002).
Britsch, S. et al. The transcription factor Sox10 is a key regulator of peripheral glial development. Genes Dev. 15, 66–78 (2001).
Finzsch, M., Stolt, C. C., Lommes, P. & Wegner, M. Sox9 and Sox10 influence survival and migration of oligodendrocyte precursors in the spinal cord by regulating PDGF receptor alpha expression. Development 135, 637–646 (2008).
Raff, M. C., Lillien, L. E., Richardson, W. D., Burne, J. F. & Noble, M. D. Platelet-derived growth factor from astrocytes drives the clock that times oligodendrocyte development in culture. Nature 333, 562–565 (1988).
Fruttiger, M. et al. Defective oligodendrocyte development and severe hypomyelination in PDGF-A knockout mice. Development 126, 457–467 (1999).
Chew, L. J., Coley, W., Cheng, Y. & Gallo, V. Mechanisms of regulation of oligodendrocyte development by p38 mitogen-activated protein kinase. J. Neurosci. 30, 11011–11027 (2010).
Holland, E. C. et al. Combined activation of Ras and Akt in neural progenitors induces glioblastoma formation in mice. Nat. Genet. 25, 55–57 (2000).
Alcantara Llaguno, S. et al. Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell 15, 45–56 (2009).
Hambardzumyan, D., Parada, L. F., Holland, E. C. & Charest, A. Genetic modeling of gliomas in mice: new tools to tackle old problems. Glia 59, 1155–1168 (2011).
Liu, C. et al. Mosaic analysis with double markers reveals tumor cell of origin in glioma. Cell 146, 209–221 (2011).
Alcantara Llaguno, S. R. et al. Adult lineage-restricted CNS progenitors specify distinct glioblastoma subtypes. Cancer Cell 28, 429–440 (2015).
Persson, A. I. et al. Non-stem cell origin for oligodendroglioma. Cancer Cell 18, 669–682 (2010).
Yadavilli, S. et al. The emerging role of NG2 in pediatric diffuse intrinsic pontine glioma. Oncotarget 6, 12141–12155 (2015).
Ligon, K. L. et al. Olig2-regulated lineage-restricted pathway controls replication competence in neural stem cells and malignant glioma. Neuron 53, 503–517 (2007).
Ligon, K. L. et al. The oligodendroglial lineage marker OLIG2 is universally expressed in diffuse gliomas. J. Neuropathol. Exp. Neurol. 63, 499–509 (2004).
Mehta, S. et al. The central nervous system-restricted transcription factor Olig2 opposes p53 responses to genotoxic damage in neural progenitors and malignant glioma. Cancer Cell 19, 359–371 (2011).
Lu, F. et al. Olig2-dependent reciprocal shift in PDGF and EGF receptor signaling regulates tumor phenotype and mitotic growth in malignant glioma. Cancer Cell 29, 669–683 (2016). This paper shows excellent use of genetic tools to modulate regulators of developmental gliogenesis during glioma tumorigenesis to affect glioma subtype identity.
Zhou, J. et al. A sequentially priming phosphorylation cascade activates the gliomagenic transcription factor Olig2. Cell Rep. 18, 3167–3177 (2017).
Ferletta, M., Uhrbom, L., Olofsson, T., Ponten, F. & Westermark, B. Sox10 has a broad expression pattern in gliomas and enhances platelet-derived growth factor-B− -induced gliomagenesis. Mol. Cancer Res. 5, 891–897 (2007).
Heldin, C. H. Targeting the PDGF signaling pathway in tumor treatment. Cell Commun. Signal 11, 97 (2013).
Phillips, J. J. et al. PDGFRA amplification is common in pediatric and adult high-grade astrocytomas and identifies a poor prognostic group in IDH1 mutant glioblastoma. Brain Pathol. 23, 565–573 (2013).
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Dai, C. et al. PDGF autocrine stimulation dedifferentiates cultured astrocytes and induces oligodendrogliomas and oligoastrocytomas from neural progenitors and astrocytes in vivo. Genes Dev. 15, 1913–1925 (2001).
Mellinghoff, I. K., Schultz, N., Mischel, P. S. & Cloughesy, T. F. Will kinase inhibitors make it as glioblastoma drugs? Curr. Top. Microbiol. Immunol. 355, 135–169 (2012).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008). This paper describes a valuable database of transcriptome data from glial development that may be useful to inform glioma tumorigenesis studies.
Molofsky, A. V. et al. Expression profiling of Aldh1l1-precursors in the developing spinal cord reveals glial lineage-specific genes and direct Sox9-Nfe2l1 interactions. Glia 61, 1518–1532 (2013).
Chaboub, L. S. et al. Temporal profiling of astrocyte precursors reveals parallel roles for asef during development and after injury. J. Neurosci. 36, 11904–11917 (2016).
Tien, A. C. et al. Regulated temporal-spatial astrocyte precursor cell proliferation involves BRAF signalling in mammalian spinal cord. Development 139, 2477–2487 (2012).
Ge, W. P., Miyawaki, A., Gage, F. H., Jan, Y. N. & Jan, L. Y. Local generation of glia is a major astrocyte source in postnatal cortex. Nature 484, 376–380 (2012).
Jones, D. T. et al. Recurrent somatic alterations of FGFR1 and NTRK2 in pilocytic astrocytoma. Nat. Genet. 45, 927–932 (2013).
Tsai, H. H. et al. Regional astrocyte allocation regulates CNS synaptogenesis and repair. Science 337, 358–362 (2012).
Hochstim, C., Deneen, B., Lukaszewicz, A., Zhou, Q. & Anderson, D. J. Identification of positionally distinct astrocyte subtypes whose identities are specified by a homeodomain code. Cell 133, 510–522 (2008).
Chai, H. et al. Neural circuit-specialized astrocytes: transcriptomic, proteomic, morphological, and functional evidence. Neuron 95, 531–549 e539 (2017).
Morel, L. et al. Molecular and functional properties of regional astrocytes in the adult brain. J. Neurosci. 37, 8706–8717 (2017).
Lin, J. C. C. et al. Identification of diverse astrocyte populations and their malignant analogs. Nat. Neurosci. 20, 396–405 (2017). This study found that molecularly distinct astrocyte populations isolated from healthy brain tissue have analogous populations in glioma tumours.
das Neves, L. et al. Disruption of the murine nuclear factor I-A gene (Nfia) results in perinatal lethality, hydrocephalus, and agenesis of the corpus callosum. Proc. Natl Acad. Sci. USA 96, 11946–11951 (1999).
Sun, W. et al. SOX9 is an astrocyte-specific nuclear marker in the adult brain outside the neurogenic regions. J. Neurosci. 37, 4493–4507 (2017).
Bonni, A. et al. Regulation of gliogenesis in the central nervous system by the JAK-STAT signaling pathway. Science 278, 477–483 (1997).
Barnabe-Heider, F. et al. Evidence that embryonic neurons regulate the onset of cortical gliogenesis via cardiotrophin-1. Neuron 48, 253–265 (2005).
Martini, S. et al. A critical role for Sox9 in notch-induced astrogliogenesis and stem cell maintenance. Stem Cells 31, 741–751 (2013).
Nagao, M., Ogata, T., Sawada, Y. & Gotoh, Y. Zbtb20 promotes astrocytogenesis during neocortical development. Nat. Commun. 7, 11102 (2016).
Magnusson, J. P. et al. A latent neurogenic program in astrocytes regulated by Notch signaling in the mouse. Science 346, 237–241 (2014).
Darmanis, S. et al. Single-cell RNA-seq analysis of infiltrating neoplastic cells at the migrating front of human glioblastoma. Cell Rep. 21, 1399–1410 (2017).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
de la Iglesia, N., Puram, S. V. & Bonni, A. STAT3 regulation of glioblastoma pathogenesis. Curr. Mol. Med. 9, 580–590 (2009).
de la Iglesia, N. et al. Identification of a PTEN-regulated STAT3 brain tumor suppressor pathway. Genes Dev. 22, 449–462 (2008).
Fan, Q. W. et al. EGFR phosphorylates tumor-derived EGFRvIII driving STAT3/5 and progression in glioblastoma. Cancer Cell 24, 438–449 (2013).
Jahani-Asl, A. et al. Control of glioblastoma tumorigenesis by feed-forward cytokine signaling. Nat. Neurosci. 19, 798–806 (2016).
Jackson, C., Ruzevick, J., Amin, A. G. & Lim, M. Potential role for STAT3 inhibitors in glioblastoma. Neurosurg. Clin. N. Am. 23, 379–389 (2012).
Furtek, S. L., Backos, D. S., Matheson, C. J. & Reigan, P. Strategies and approaches of targeting STAT3 for cancer treatment. ACS Chem. Biol. 11, 308–318 (2016).
de The, H. Differentiation therapy revisited. Nat. Rev. Cancer 18, 117–127 (2018). This paper presents an interesting opinion on the ways that differentiation therapies have been used to treat certain tumours and how these treatments may be improved to treat other malignancies.
Ostrom, Q. T. et al. The epidemiology of glioma in adults: a “state of the science” review. Neuro Oncol. 16, 896–913 (2014).
Acknowledgements
The authors thank C. Gillespie for editing and reviewing this manuscript. This work was supported by grants from the US National Institutes of Health (NS071153 to B.D. and K01CA190235 and 5-T32HL092332-08 to S.M.G.) and the Cancer Prevention Research Institute of Texas (RP150334 and RP160192 to B.D.).
Reviewer information
Nature Reviews Neuroscience thanks B. Appel, W. Weiss, M. Monje and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
B.D. and D.L. researched data for the article and made a substantial contribution to the discussion of content and the writing, reviewing and editing of the manuscript before submission. S.M.G. researched data for the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Isocitrate dehydrogenase (IDH) status
-
Refers to whether a patient possesses a missense mutation in IDH1 or IDH2 that has been linked to glioma.
- Frank glioma
-
A clinically diagnosed glioma.
- Xenograft
-
A commonly used animal tumour model in which patient-derived cancer cells are transplanted under the skin or into the organ of interest in immune-deficient animals.
Rights and permissions
About this article
Cite this article
Laug, D., Glasgow, S.M. & Deneen, B. A glial blueprint for gliomagenesis. Nat Rev Neurosci 19, 393–403 (2018). https://doi.org/10.1038/s41583-018-0014-3
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41583-018-0014-3
This article is cited by
-
Two Mixed-Ligand Co(II) Complexes as Luminescent Materials and Loaded with Temozolomide-gel Particles in Nursing Against Glioma
Journal of Fluorescence (2024)
-
Stalled oligodendrocyte differentiation in IDH-mutant gliomas
Genome Medicine (2023)
-
TCAF2 drives glioma cellular migratory/invasion properties through STAT3 signaling
Molecular and Cellular Biochemistry (2023)
-
Temporal and spatial stability of the EM/PM molecular subtypes in adult diffuse glioma
Frontiers of Medicine (2023)
-
A brain precursor atlas reveals the acquisition of developmental-like states in adult cerebral tumours
Nature Communications (2022)