Abstract
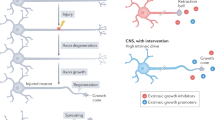
Permanent disabilities following CNS injuries result from the failure of injured axons to regenerate and rebuild functional connections with their original targets. By contrast, injury to peripheral nerves is followed by robust regeneration, which can lead to recovery of sensory and motor functions. This regenerative response requires the induction of widespread transcriptional and epigenetic changes in injured neurons. Considerable progress has been made in recent years in understanding how peripheral axon injury elicits these widespread changes through the coordinated actions of transcription factors, epigenetic modifiers and, to a lesser extent, microRNAs. Although many questions remain about the interplay between these mechanisms, these new findings provide important insights into the pivotal role of coordinated gene expression and chromatin remodelling in the neuronal response to injury.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Liu, K., Tedeschi, A., Park, K. K. & He, Z. Neuronal intrinsic mechanisms of axon regeneration. |Annu. Rev. Neurosci. 34, 131–152 (2011).
Di Giovanni, S. Molecular targets for axon regeneration: focus on the intrinsic pathways. Expert Opin. Ther. Targets 13, 1387–1398 (2009).
Schwab, M. E. & Strittmatter, S. M. Nogo limits neural plasticity and recovery from injury. Curr. Opin. Neurobiol. 27, 53–60 (2014).
Tedeschi, A. & Bradke, F. Spatial and temporal arrangement of neuronal intrinsic and extrinsic mechanisms controlling axon regeneration. Curr. Opin. Neurobiol. 42, 118–127 (2017).
Kaplan, A., Bueno, M., Hua, L. & Fournier, A. E. Maximizing functional axon repair in the injured central nervous system: lessons from neuronal development. Dev. Dyn. 247, 18–23 (2017).
Wood, M. D., Kemp, S. W., Weber, C., Borschel, G. H. & Gordon, T. Outcome measures of peripheral nerve regeneration. Ann. Anat. 193, 321–333 (2011).
Saheb-Al-Zamani, M. et al. Limited regeneration in long acellular nerve allografts is associated with increased Schwann cell senescence. Exp. Neurol. 247, 165–177 (2013).
He, Z. & Jin, Y. Intrinsic control of axon regeneration. Neuron 90, 437–451 (2016).
Quadrato, G. & Di Giovanni, S. Waking up the sleepers: shared transcriptional pathways in axonal regeneration and neurogenesis. Cell. Mol. Life Sci. 70, 993–1007 (2013).
Rishal, I. & Fainzilber, M. Axon-soma communication in neuronal injury. Nat. Rev. Neurosci. 15, 32–42 (2014).
Bradke, F., Fawcett, J. W. & Spira, M. E. Assembly of a new growth cone after axotomy: the precursor to axon regeneration. Nat. Rev. Neurosci. 13, 183–193 (2012).
Cho, Y. & Cavalli, V. HDAC signaling in neuronal development and axon regeneration. Curr. Opin. Neurobiol. 27, 118–126 (2014).
Lu, Y., Belin, S. & He, Z. Signaling regulations of neuronal regenerative ability. Curr. Opin. Neurobiol. 27, 135–142 (2014).
Byrne, A. B. & Hammarlund, M. Axon regeneration in C. elegans: worming our way to mechanisms of axon regeneration. Exp. Neurol. 287, 300–309 (2017).
Chisholm, A. D., Hutter, H., Jin, Y. & Wadsworth, W. G. The genetics of axon guidance and axon regeneration in Caenorhabditis elegans. Genetics 204, 849–882 (2016).
Hao, Y. & Collins, C. Intrinsic mechanisms for axon regeneration: insights from injured axons in Drosophila. Curr. Opin. Genet. Dev. 44, 84–91 (2017).
Rasmussen, J. P. & Sagasti, A. Learning to swim, again: axon regeneration in fish. Exp. Neurol. 287, 318–330 (2017).
Ziv, N. E. & Spira, M. E. Spatiotemporal distribution of Ca2+ following axotomy and throughout the recovery process of cultured Aplysia neurons. Eur. J. Neurosci. 5, 657–668 (1993).
Ziv, N. E. & Spira, M. E. Axotomy induces a transient and localized elevation of the free intracellular calcium concentration to the millimolar range. J. Neurophysiol. 74, 2625–2637 (1995).
Cho, Y., Sloutsky, R., Naegle, K. M. & Cavalli, V. Injury-induced HDAC5 nuclear export is essential for axon regeneration. Cell 155, 894–908 (2013). This study demonstrates that, following axon injury, a calcium wave triggers epigenetic changes through the nuclear export of HDAC5 to promote expression of pro-regenerative genes.
Ghosh-Roy, A., Wu, Z., Goncharov, A., Jin, Y. & Chisholm, A. D. Calcium and cyclic AMP promote axonal regeneration in Caenorhabditis elegans and require DLK-1 kinase. J. Neurosci. 30, 3175–3183 (2010).
Saito, A. & Cavalli, V. Signaling over distances. Mol. Cell. Proteomics 15, 382–393 (2016).
Kulbatski, I., Cook, D. J. & Tator, C. H. Calcium entry through L-type calcium channels is essential for neurite regeneration in cultured sympathetic neurons. J. Neurotrauma 21, 357–374 (2004).
Enes, J. et al. Electrical activity suppresses axon growth through Ca(v)1.2 channels in adult primary sensory neurons. Curr. Biol. 20, 1154–1164 (2010).
Sun, L. et al. Neuronal regeneration in C. elegans requires subcellular calcium release by ryanodine receptor channels and can be enhanced by optogenetic stimulation. J. Neurosci. 34, 15947–15956 (2014).
Tuck, E. & Cavalli, V. Roles of membrane trafficking in nerve repair and regeneration. Commun. Integr. Biol. 3, 209–214 (2010).
McNeil, P. L. & Kirchhausen, T. An emergency response team for membrane repair. Nat. Rev. Mol. Cell Biol. 6, 499–505 (2005).
Weng, Y. L. et al. An intrinsic epigenetic barrier for functional axon regeneration. Neuron 94, 337–346.e6 (2017).This study reveals that DNA is actively demethylated at the promoters of RAGs, which is required to promote functional axonal regeneration of peripheral sensory neurons.
Perry, R. B. & Fainzilber, M. Local translation in neuronal processes — in vivo tests of a “heretical hypothesis”. Dev. Neurobiol. 74, 210–217 (2014).
Stirling, D. P. & Stys, P. K. Mechanisms of axonal injury: internodal nanocomplexes and calcium deregulation. Trends Mol. Med. 16, 160–170 (2010).
Tedeschi, A. et al. The calcium channel subunit Alpha2delta2 suppresses axon regeneration in the adult CNS. Neuron 92, 419–434 (2016). This study shows that the calcium channel subunit VGCC-α2/δ2 (CACNA2D2) is developmentally upregulated in DRG neurons, decreasing neuronal growth capacity, and that its inhibition with the existing drug pregabalin increases regenerative capacity.
Cai, D. et al. Neuronal cyclic AMP controls the developmental loss in ability of axons to regenerate. J. Neurosci. 21, 4731–4739 (2001).
Udina, E. et al. Electrical stimulation of intact peripheral sensory axons in rats promotes outgrowth of their central projections. Exp. Neurol. 210, 238–247 (2008).
Hao, Y. et al. An evolutionarily conserved mechanism for cAMP elicited axonal regeneration involves direct activation of the dual leucine zipper kinase DLK. eLife 5, e14048 (2016). This study demonstrates that the cAMP–PKA pathway activates the DLK pathway, which is necessary for axon regeneration.
Hammarlund, M., Nix, P., Hauth, L., Jorgensen, E. M. & Bastiani, M. Axon regeneration requires a conserved MAP kinase pathway. Science 323, 802–806 (2009).
Gao, Y. et al. Activated CREB is sufficient to overcome inhibitors in myelin and promote spinal axon regeneration in vivo. Neuron 44, 609–621 (2004).
Ma, T. C. & Willis, D. E. What makes a RAG regeneration associated? Front. Mol. Neurosci. 8, 43 (2015).
Blesch, A. et al. Conditioning lesions before or after spinal cord injury recruit broad genetic mechanisms that sustain axonal regeneration: superiority to camp-mediated effects. Exp. Neurol. 235, 162–173 (2012).
Perez-Cadahia, B., Drobic, B. & Davie, J. R. Activation and function of immediate-early genes in the nervous system. Biochem. Cell Biol. 89, 61–73 (2011).
West, A. E. & Greenberg, M. E. Neuronal activity-regulated gene transcription in synapse development and cognitive function. Cold Spring Harb. Perspect. Biol. 3, a005744 (2011).
Chawla, S., Vanhoutte, P., Arnold, F. J., Huang, C. L. & Bading, H. Neuronal activity-dependent nucleocytoplasmic shuttling of HDAC4 and HDAC5. J. Neurochem. 85, 151–159 (2003).
Lonze, B. E. & Ginty, D. D. Function and regulation of CREB family transcription factors in the nervous system. Neuron 35, 605–623 (2002).
Sheng, M., Thompson, M. A. & Greenberg, M. E. CREB: a Ca(2+)-regulated transcription factor phosphorylated by calmodulin-dependent kinases. Science 252, 1427–1430 (1991).
Chrivia, J. C. et al. Phosphorylated CREB binds specifically to the nuclear protein CBP. Nature 365, 855–859 (1993).
Gaub, P. et al. The histone acetyltransferase p300 promotes intrinsic axonal regeneration. Brain 134, 2134–2148 (2011).
Tedeschi, A., Nguyen, T., Puttagunta, R., Gaub, P. & Di Giovanni, S. A p53-CBP/p300 transcription module is required for GAP-43 expression, axon outgrowth, and regeneration. Cell Death Differ 16, 543–554 (2009).
Knoll, B. Serum response factor mediated gene activity in physiological and pathological processes of neuronal motility. Front. Mol. Neurosci. 4, 49 (2011).
Stern, S. et al. The transcription factor serum response factor stimulates axon regeneration through cytoplasmic localization and cofilin interaction. J. Neurosci. 33, 18836–18848 (2013).
Xia, Z., Dudek, H., Miranti, C. K. & Greenberg, M. E. Calcium influx via the NMDA receptor induces immediate early gene transcription by a MAP kinase/ERK-dependent mechanism. J. Neurosci. 16, 5425–5436 (1996).
Chierzi, S., Ratto, G. M., Verma, P. & Fawcett, J. W. The ability of axons to regenerate their growth cones depends on axonal type and age, and is regulated by calcium, cAMP and ERK. Eur. J. Neurosci. 21, 2051–2062 (2005).
Broude, E., McAtee, M., Kelley, M. S. & Bregman, B. S. c-Jun expression in adult rat dorsal root ganglion neurons: differential response after central or peripheral axotomy. Exp. Neurol. 148, 367–377 (1997).
Perlson, E. et al. Vimentin-dependent spatial translocation of an activated MAP kinase in injured nerve. Neuron 45, 715–726 (2005).
Cavalli, V., Kujala, P., Klumperman, J. & Goldstein, L. S. Sunday Driver links axonal transport to damage signaling. J. Cell Biol. 168, 775–787 (2005).
Drerup, C. M. & Nechiporuk, A. V. JNK-interacting protein 3 mediates the retrograde transport of activated c-Jun N-terminal kinase and lysosomes. PLoS Genet. 9, e1003303 (2013).
Lindwall, C. & Kanje, M. Retrograde axonal transport of JNK signaling molecules influence injury induced nuclear changes in p-c-Jun and ATF3 in adult rat sensory neurons. Mol. Cell Neurosci. 29, 269–282 (2005).
Xiong, X. et al. Protein turnover of the Wallenda/DLK kinase regulates a retrograde response to axonal injury. J. Cell Biol. 191, 211–223 (2010).
Ben-Yaakov, K. et al. Axonal transcription factors signal retrogradely in lesioned peripheral nerve. EMBO J. 31, 1350–1363 (2012). This study shows that STAT3 is locally translated in the injured axon and retrogradely transported to the nucleus to promote sensory neuron survival.
Hanz, S. et al. Axoplasmic importins enable retrograde injury signaling in lesioned nerve. Neuron 40, 1095–1104 (2003).
O’Donovan, K. J. et al. B-RAF kinase drives developmental axon growth and promotes axon regeneration in the injured mature CNS. J. Exp. Med. 211, 801–814 (2014). This study shows that activation of BRAF synergizes with PTEN knockout to promote optic nerve regeneration.
Valakh, V., Walker, L. J., Skeath, J. B. & DiAntonio, A. Loss of the spectraplakin short stop activates the DLK injury response pathway in Drosophila. J. Neurosci. 33, 17863–17873 (2013).
Yan, D., Wu, Z., Chisholm, A. D. & Jin, Y. The DLK-1 kinase promotes mRNA stability and local translation in C. elegans synapses and axon regeneration. Cell 138, 1005–1018 (2009).
Kenney, A. M. & Kocsis, J. D. Peripheral axotomy induces long-term c-Jun amino-terminal kinase-1 activation and activator protein-1 binding activity by c-Jun and junD in adult rat dorsal root ganglia in vivo. J. Neurosci. 18, 1318–1328 (1998).
Shin, J. et al. Dual leucine zipper kinase is required for retrograde injury signaling and axonal regeneration. Neuron 74, 1015–1022 (2012).
Song, W., Cho, Y., Watt, D. & Cavalli, V. Tubulin-tyrosine ligase (TTL)-mediated increase in tyrosinated α-tubulin in injured axons is required for retrograde injury signaling and axon regeneration. J. Biol. Chem. 290, 14765–14775 (2015).
Welsbie, D. S. et al. Enhanced functional genomic screening identifies novel mediators of dual leucine zipper kinase-dependent injury signaling in neurons. Neuron 94, 1142–1154.e6 (2017). This study identifies novel members of the DLK injury signalling pathway in RGCs and demonstrates that LZK cooperates with DLK to promote RGC cell death in response to axon injury.
Watkins, T. A. et al. DLK initiates a transcriptional program that couples apoptotic and regenerative responses to axonal injury. Proc. Natl Acad. Sci. USA 110, 4039–4044 (2013).
Luo, X. et al. Enhanced transcriptional activity and mitochondrial localization of STAT3 co-induce axon regrowth in the adult central nervous system. Cell Rep. 15, 398–410 (2016).
Zigmond, R. E. gp130 cytokines are positive signals triggering changes in gene expression and axon outgrowth in peripheral neurons following injury. Front. Mol. Neurosci. 4, 62 (2011).
Zhong, J., Dietzel, I. D., Wahle, P., Kopf, M. & Heumann, R. Sensory impairments and delayed regeneration of sensory axons in interleukin-6-deficient mice. J. Neurosci. 19, 4305–4313 (1999).
Cafferty, W. B. et al. Conditioning injury-induced spinal axon regeneration fails in interleukin-6 knock-out mice. J. Neurosci. 24, 4432–4443 (2004).
Lerch, J. K. et al. Stress increases peripheral axon growth and regeneration through glucocorticoid receptor-dependent transcriptional programs. eNeuro https://doi.org/10.1523/ENEURO.0246-17.2017 (2017).
Niemi, J. P., DeFrancesco-Lisowitz, A., Cregg, J. M., Howarth, M. & Zigmond, R. E. Overexpression of the monocyte chemokine CCL2 in dorsal root ganglion neurons causes a conditioning-like increase in neurite outgrowth and does so via a STAT3 dependent mechanism. Exp. Neurol. 275, 25–37 (2016).
Muller, A., Hauk, T. G. & Fischer, D. Astrocyte-derived CNTF switches mature RGCs to a regenerative state following inflammatory stimulation. Brain 130, 3308–3320 (2007).
Leibinger, M., Andreadaki, A., Diekmann, H. & Fischer, D. Neuronal STAT3 activation is essential for CNTF- and inflammatory stimulation-induced CNS axon regeneration. Cell Death Dis. 4, e805 (2013).
Verma, N. K. et al. STAT3-stathmin interactions control microtubule dynamics in migrating T-cells. J. Biol. Chem. 284, 12349–12362 (2009).
Ng, D. C. et al. Stat3 regulates microtubules by antagonizing the depolymerization activity of stathmin. J. Cell Biol. 172, 245–257 (2006).
Selvaraj, B. T., Frank, N., Bender, F. L., Asan, E. & Sendtner, M. Local axonal function of STAT3 rescues axon degeneration in the pmn model of motoneuron disease. J. Cell Biol. 199, 437–451 (2012).
Twiss, J. L. & Merianda, T. T. Old dogs with new tricks: intra-axonal translation of nuclear proteins. Neural Regen. Res. 10, 1560–1562 (2015).
Liu, P. H., Tsai, H. Y., Chung, Y. W., Wang, Y. J. & Tseng, G. F. The proximity of the lesion to cell bodies determines the free radical risk induced in rat rubrospinal neurons subjected to axonal injury. Anat. Embryol. (Berl.) 207, 439–451 (2004).
Liu, P. H., Yang, L. H., Wang, T. Y., Wang, Y. J. & Tseng, G. F. Proximity of lesioning determines response of facial motoneurons to peripheral axotomy. J. Neurotrauma 23, 1857–1873 (2006).
Villegas-Perez, M. P., Vidal-Sanz, M., Rasminsky, M., Bray, G. M. & Aguayo, A. J. Rapid and protracted phases of retinal ganglion cell loss follow axotomy in the optic nerve of adult rats. J. Neurobiol. 24, 23–36 (1993).
Berkelaar, M., Clarke, D. B., Wang, Y. C., Bray, G. M. & Aguayo, A. J. Axotomy results in delayed death and apoptosis of retinal ganglion cells in adult rats. J. Neurosci. 14, 4368–4374 (1994).
Rishal, I. et al. A motor-driven mechanism for cell-length sensing. Cell Rep. 1, 608–616 (2012).
Brochier, C., Jones, J. I., Willis, D. E. & Langley, B. Poly(ADP-ribose) polymerase 1 is a novel target to promote axonal regeneration. Proc. Natl Acad. Sci. USA 112, 15220–15225 (2015).
Byrne, A. B. et al. Inhibiting poly(ADP-ribosylation) improves axon regeneration. eLife 5, e12734 (2016). This study demonstrates that PARylation is a conserved regulator of axon regeneration that functions downstream of DLK signalling.
Wang, X. et al. Inhibition of poly-ADP-ribosylation fails to increase axonal regeneration or improve functional recovery after adult mammalian CNS injury. eNeuro https://doi.org/10.1523/ENEURO.0270-16.2016 (2016).
Yiu, G. & He, Z. Glial inhibition of CNS axon regeneration. Nat. Rev. Neurosci. 7, 617–627 (2006).
Kosmaczewski, S. G. et al. RNA ligation in neurons by RtcB inhibits axon regeneration. Proc. Natl Acad. Sci. USA 112, 8451–8456 (2015).
Song, Y. et al. Regulation of axon regeneration by the RNA repair and splicing pathway. Nat. Neurosci. 18, 817–825 (2015).
Nix, P. et al. Axon regeneration genes identified by RNAi screening in C. elegans. J. Neurosci. 34, 629–645 (2014).
Onate, M. et al. Activation of the unfolded protein response promotes axonal regeneration after peripheral nerve injury. Sci. Rep. 6, 21709 (2016).
Berry, M., Ahmed, Z., Morgan-Warren, P., Fulton, D. & Logan, A. Prospects for mTOR-mediated functional repair after central nervous system trauma. Neurobiol. Dis. 85, 99–110 (2016).
Park, K. K. et al. Promoting axon regeneration in the adult CNS by modulation of the PTEN/mTOR pathway. Science 322, 963–966 (2008).
Abe, N., Borson, S. H., Gambello, M. J., Wang, F. & Cavalli, V. Mammalian target of rapamycin (mTOR) activation increases axonal growth capacity of injured peripheral nerves. J. Biol. Chem. 285, 28034–28043 (2010).
Liu, K. et al. PTEN deletion enhances the regenerative ability of adult corticospinal neurons. Nat. Neurosci. 13, 1075–1081 (2010).
Christie, K. J., Webber, C. A., Martinez, J. A., Singh, B. & Zochodne, D. W. PTEN inhibition to facilitate intrinsic regenerative outgrowth of adult peripheral axons. J. Neurosci. 30, 9306–9315 (2010).
Chen, W. et al. Rapamycin-resistant mTOR activity is required for sensory axon regeneration induced by a conditioning lesion. eNeuro https://doi.org/10.1523/ENEURO.0358-16.2016 (2016).
Al-Ali, H. et al. The mTOR substrate S6 kinase 1 (S6K1) is a negative regulator of axon regeneration and a potential drug target for central nervous system injury. J. Neurosci. 37, 7079–7095 (2017). This study finds that treatment with a specific S6K1 inhibitor promotes cortical spinal tract regeneration and behavioural recovery.
Duan, X. et al. Subtype-specific regeneration of retinal ganglion cells following axotomy: effects of osteopontin and mTOR signaling. Neuron 85, 1244–1256 (2015).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Leighton, L. J. et al. Experience-dependent neural plasticity, learning, and memory in the era of epitranscriptomics. Genes Brain Behav. https://doi.org/10.1111/gbb.12426 (2017).
Yoon, K. J. et al. Temporal control of mammalian cortical neurogenesis by m(6)A methylation. Cell 171, 877–889.e17 (2017).
Weng, Y. L. et al. Epitranscriptomic m(6)A regulation of axon regeneration in the adult mammalian nervous system. Neuron 97, 313–325.e6 (2018). This study uncovers an epitranscriptomic mechanism in which axon injury elevates N6-methyladenosine levels in regeneration-associated transcripts, which is essential for functional peripheral axon regeneration.
Smith, D. S. & Skene, J. H. A transcription-dependent switch controls competence of adult neurons for distinct modes of axon growth. J. Neurosci. 17, 646–658 (1997). This study establishes that successful axon regeneration requires a transcription-dependent phase.
Moore, D. L. & Goldberg, J. L. Multiple transcription factor families regulate axon growth and regeneration. Dev. Neurobiol. 71, 1186–1211 (2011).
Blackmore, M. G. Molecular control of axon growth: insights from comparative gene profiling and high-throughput screening. Int. Rev. Neurobiol. 105, 39–70 (2012).
Xu, J., Du, Y. & Deng, H. Direct lineage reprogramming: strategies, mechanisms, and applications. Cell Stem Cell 16, 119–134 (2015).
Gey, M. et al. Atf3 mutant mice show reduced axon regeneration and impaired regeneration-associated gene induction after peripheral nerve injury. Open Biol. 6, 160091 (2016).
Seijffers, R., Mills, C. D. & Woolf, C. J. ATF3 increases the intrinsic growth state of DRG neurons to enhance peripheral nerve regeneration. J. Neurosci. 27, 7911–7920 (2007).
Venkatesh, I., Simpson, M. T., Coley, D. M. & Blackmore, M. G. Epigenetic profiling reveals a developmental decrease in promoter accessibility during cortical maturation in vivo. Neuroepigenetics 8, 19–26 (2016). This study finds that, although there is an increase in H3K27me3 at RAG promoters in cortical neurons during development, manipulation of HDACs or HATs does not further promote regeneration.
Seijffers, R., Allchorne, A. J. & Woolf, C. J. The transcription factor ATF-3 promotes neurite outgrowth. Mol. Cell. Neurosci. 32, 143–154 (2006).
Lerch, J. K., Martinez-Ondaro, Y. R., Bixby, J. L. & Lemmon, V. P. cJun promotes CNS axon growth. Mol. Cell. Neurosci. 59, 97–105 (2014).
Simpson, M. T. et al. The tumor suppressor HHEX inhibits axon growth when prematurely expressed in developing central nervous system neurons. Mol. Cell. Neurosci. 68, 272–283 (2015).
Chandran, V. et al. A systems-level analysis of the peripheral nerve intrinsic axonal growth program. Neuron 89, 956–970 (2016). This study uses a multilevel bioinformatics approach to identify pharmacological compounds that enhance axon regeneration.
Raivich, G. et al. The AP-1 transcription factor c-Jun is required for efficient axonal regeneration. Neuron 43, 57–67 (2004).
Ruff, C. A. et al. Neuronal c-Jun is required for successful axonal regeneration, but the effects of phosphorylation of its N-terminus are moderate. J. Neurochem. 121, 607–618 (2012).
Zou, H., Ho, C., Wong, K. & Tessier-Lavigne, M. Axotomy-induced Smad1 activation promotes axonal growth in adult sensory neurons. J. Neurosci. 29, 7116–7123 (2009).
Saijilafu et al. PI3K-GSK3 signalling regulates mammalian axon regeneration by inducing the expression of Smad1. Nat. Commun. 4, 2690 (2013).
Finelli, M. J., Wong, J. K. & Zou, H. Epigenetic regulation of sensory axon regeneration after spinal cord injury. J. Neurosci. 33, 19664–19676 (2013). This study reveals that histone-modifying enzymes work together with SMAD1 to facilitate an increase in histone H4 acetylation and transcription of RAGs and to promote axon regeneration.
Tedeschi, A., Omura, T. & Costigan, M. CNS repair and axon regeneration: using genetic variation to determine mechanisms. Exp. Neurol. 287, 409–422 (2017).
Omura, T. et al. Robust axonal regeneration occurs in the injured CAST/Ei mouse CNS. Neuron 86, 1215–1227 (2015).
Hill, C. S. Transcriptional control by the SMADs. Cold Spring Harb. Perspect. Biol. 8, a022079 (2016).
Schwaiger, F. W. et al. Peripheral but not central axotomy induces changes in Janus kinases (JAK) and signal transducers and activators of transcription (STAT). Eur. J. Neurosci. 12, 1165–1176 (2000).
Bareyre, F. M. et al. In vivo imaging reveals a phase-specific role of STAT3 during central and peripheral nervous system axon regeneration. Proc. Natl Acad. Sci. USA 108, 6282–6287 (2011).
Lang, C., Bradley, P. M., Jacobi, A., Kerschensteiner, M. & Bareyre, F. M. STAT3 promotes corticospinal remodelling and functional recovery after spinal cord injury. EMBO Rep. 14, 931–937 (2013).
Mehta, S. T., Luo, X., Park, K. K., Bixby, J. L. & Lemmon, V. P. Hyperactivated Stat3 boosts axon regeneration in the CNS. Exp. Neurol. 280, 115–120 (2016).
Qiu, J., Cafferty, W. B., McMahon, S. B. & Thompson, S. W. Conditioning injury-induced spinal axon regeneration requires signal transducer and activator of transcription 3 activation. J. Neurosci. 25, 1645–1653 (2005).
Lingor, P. et al. ROCK inhibition and CNTF interact on intrinsic signalling pathways and differentially regulate survival and regeneration in retinal ganglion cells. Brain 131, 250–263 (2008).
Smith, P. D. et al. SOCS3 deletion promotes optic nerve regeneration in vivo. Neuron 64, 617–623 (2009).
Sun, F. et al. Sustained axon regeneration induced by co-deletion of PTEN and SOCS3. Nature 480, 372–375 (2011).
Moore, D. L. et al. KLF family members regulate intrinsic axon regeneration ability. Science 326, 298–301 (2009). This study establishes that the developmentally regulated KLF family of transcription factors regulates axon regeneration.
Blackmore, M. G. et al. Kruppel-like Factor 7 engineered for transcriptional activation promotes axon regeneration in the adult corticospinal tract. Proc. Natl Acad. Sci. USA 109, 7517–7522 (2012).
Moore, D. L., Apara, A. & Goldberg, J. L. Kruppel-like transcription factors in the nervous system: novel players in neurite outgrowth and axon regeneration. Mol. Cell. Neurosci. 47, 233–243 (2011).
Apara, A. et al. KLF9 and JNK3 interact to suppress axon regeneration in the adult CNS. J. Neurosci. 37, 9632–9644 (2017).
Blackmore, M. G. et al. High content screening of cortical neurons identifies novel regulators of axon growth. Mol. Cell. Neurosci. 44, 43–54 (2010).
Veldman, M. B., Bemben, M. A., Thompson, R. C. & Goldman, D. Gene expression analysis of zebrafish retinal ganglion cells during optic nerve regeneration identifies KLF6a and KLF7a as important regulators of axon regeneration. Dev. Biol. 312, 596–612 (2007).
Belin, S. et al. Injury-induced decline of intrinsic regenerative ability revealed by quantitative proteomics. Neuron 86, 1000–1014 (2015).
Cho, Y. et al. Activating injury-responsive genes with hypoxia enhances axon regeneration through neuronal HIF-1alpha. Neuron 88, 720–734 (2015).
Jankowski, M. P. et al. Sox11 transcription factor modulates peripheral nerve regeneration in adult mice. Brain Res. 1256, 43–54 (2009).
Jankowski, M. P., Cornuet, P. K., McIlwrath, S., Koerber, H. R. & Albers, K. M. SRY-box containing gene 11 (Sox11) transcription factor is required for neuron survival and neurite growth. Neuroscience 143, 501–514 (2006).
Jing, X., Wang, T., Huang, S., Glorioso, J. C. & Albers, K. M. The transcription factor Sox11 promotes nerve regeneration through activation of the regeneration-associated gene Sprr1a. Exp. Neurol. 233, 221–232 (2012).
Wang, Z., Reynolds, A., Kirry, A., Nienhaus, C. & Blackmore, M. G. Overexpression of Sox11 promotes corticospinal tract regeneration after spinal injury while interfering with functional recovery. J. Neurosci. 35, 3139–3145 (2015).
Norsworthy, M. W. et al. Sox11 expression promotes regeneration of some retinal ganglion cell types but kills others. Neuron 94, 1112–1120.e4 (2017). This study reveals that expression of SOX11 reactivates an axon growth programme and promotes axon regeneration in a subset of adult RGCs but kills other types of RGCs.
Di Giovanni, S. et al. The tumor suppressor protein p53 is required for neurite outgrowth and axon regeneration. EMBO J. 25, 4084–4096 (2006).
Stern, S. & Knoll, B. CNS axon regeneration inhibitors stimulate an immediate early gene response via MAP kinase-SRF signaling. Mol. Brain 7, 86 (2014).
Venkatesh, I. & Blackmore, M. G. Selecting optimal combinations of transcription factors to promote axon regeneration: why mechanisms matter. Neurosci. Lett. 652, 64–73 (2017).
Fagoe, N. D., van Heest, J. & Verhaagen, J. Spinal cord injury and the neuron-intrinsic regeneration-associated gene program. Neuromolecular Med. 16, 799–813 (2014).
Fagoe, N. D., Attwell, C. L., Kouwenhoven, D., Verhaagen, J. & Mason, M. R. Overexpression of ATF3 or the combination of ATF3, c-Jun, STAT3 and Smad1 promotes regeneration of the central axon branch of sensory neurons but without synergistic effects. Hum. Mol. Genet. 24, 6788–6800 (2015).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Takahashi, K. & Yamanaka, S. A decade of transcription factor-mediated reprogramming to pluripotency. Nat. Rev. Mol. Cell Biol. 17, 183–193 (2016).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459.e20 (2017).
Lee, T. I. & Young, R. A. Transcriptional regulation and its misregulation in disease. Cell 152, 1237–1251 (2013).
Guo, C. & Morris, S. A. Engineering cell identity: establishing new gene regulatory and chromatin landscapes. Curr. Opin. Genet. Dev. 46, 50–57 (2017).
Iwafuchi-Doi, M. & Zaret, K. S. Pioneer transcription factors in cell reprogramming. Genes Dev. 28, 2679–2692 (2014).
Zaret, K. S. & Mango, S. E. Pioneer transcription factors, chromatin dynamics, and cell fate control. Curr. Opin. Genet. Dev. 37, 76–81 (2016).
Gifford, C. A. & Meissner, A. Epigenetic obstacles encountered by transcription factors: reprogramming against all odds. Curr. Opin. Genet. Dev. 22, 409–415 (2012).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Su, Y. et al. Neuronal activity modifies the chromatin accessibility landscape in the adult brain. Nat. Neurosci. 20, 476–483 (2017).
Hu, G. et al. Single-cell RNA-seq reveals distinct injury responses in different types of DRG sensory neurons. Sci. Rep. 6, 31851 (2016). This study uses single-cell RNAseq to reveal distinctive and sustained heterogeneity of transcriptomic responses to axon injury at the single-neuron level.
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Nichterwitz, S., Benitez, J. A., Hoogstraaten, R., Deng, Q. & Hedlund, E. LCM-Seq: a method for spatial transcriptomic profiling using laser capture microdissection coupled with PolyA-based RNA sequencing. Methods Mol. Biol. 1649, 95–110 (2018).
Doyle, J. P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135, 749–762 (2008).
Yao, B. et al. Epigenetic mechanisms in neurogenesis. Nat. Rev. Neurosci. 17, 537–549 (2016).
Cholewa-Waclaw, J. et al. The role of epigenetic mechanisms in the regulation of gene expression in the nervous system. J. Neurosci. 36, 11427–11434 (2016).
Zhao, Y. & Garcia, B. A. Comprehensive catalog of currently documented histone modifications. Cold Spring Harb. Perspect. Biol. 7, a025064 (2015).
Puttagunta, R. et al. PCAF-dependent epigenetic changes promote axonal regeneration in the central nervous system. Nat. Commun. 5, 3527 (2014). This study shows that KAT2B functions downstream of ERK signalling to increase histone H3 acetylation at the promoters of key RAGs following peripheral but not central axonal injury.
Cho, Y. & Cavalli, V. HDAC5 is a novel injury-regulated tubulin deacetylase controlling axon regeneration. EMBO J. 31, 3063–3078 (2012).
Pelzel, H. R., Schlamp, C. L. & Nickells, R. W. Histone H4 deacetylation plays a critical role in early gene silencing during neuronal apoptosis. BMC Neurosci. 11, 62 (2010).
Schmitt, H. M., Pelzel, H. R., Schlamp, C. L. & Nickells, R. W. Histone deacetylase 3 (HDAC3) plays an important role in retinal ganglion cell death after acute optic nerve injury. Mol. Neurodegener. 9, 39 (2014).
Lv, L., Han, X., Sun, Y., Wang, X. & Dong, Q. Valproic acid improves locomotion in vivo after SCI and axonal growth of neurons in vitro. Exp. Neurol. 233, 783–790 (2012).
Riccio, A. Dynamic epigenetic regulation in neurons: enzymes, stimuli and signaling pathways. Nat. Neurosci. 13, 1330–1337 (2010).
Bhat, N., Park, J., Zoghbi, H. Y., Arthur, J. S. & Zaret, K. S. The chromatin modifier MSK1/2 suppresses endocrine cell fates during mouse pancreatic development. PLOS ONE 11, e0166703 (2016).
Lu, C. & Thompson, C. B. Metabolic regulation of epigenetics. Cell Metab. 16, 9–17 (2012).
Pollema-Mays, S. L., Centeno, M. V., Apkarian, A. V. & Martina, M. Expression of DNA methyltransferases in adult dorsal root ganglia is cell-type specific and up regulated in a rodent model of neuropathic pain. Front. Cell. Neurosci. 8, 217 (2014).
Zhao, J. Y. et al. DNA methyltransferase DNMT3a contributes to neuropathic pain by repressing Kcna2 in primary afferent neurons. Nat. Commun. 8, 14712 (2017).
Weng, Y. L., Joseph, J., An, R., Song, H. & Ming, G. L. Epigenetic regulation of axonal regenerative capacity. Epigenomics 8, 1429–1442 (2016).
Bachman, M. et al. 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nat. Chem. 6, 1049–1055 (2014).
Iskandar, B. J. et al. Folate regulation of axonal regeneration in the rodent central nervous system through DNA methylation. J. Clin. Invest. 120, 1603–1616 (2010).
Loh, Y. E. et al. Comprehensive mapping of 5-hydroxymethylcytosine epigenetic dynamics in axon regeneration. Epigenetics 12, 77–92 (2017).
Ghibaudi, M., Boido, M. & Vercelli, A. Functional integration of complex miRNA networks in central and peripheral lesion and axonal regeneration. Prog. Neurobiol. 158, 69–93 (2017).
Yates, L. A., Norbury, C. J. & Gilbert, R. J. The long and short of microRNA. Cell 153, 516–519 (2013).
Wu, D., Raafat, A., Pak, E., Clemens, S. & Murashov, A. K. Dicer-microRNA pathway is critical for peripheral nerve regeneration and functional recovery in vivo and regenerative axonogenesis in vitro. Exp. Neurol. 233, 555–565 (2012).
Wu, D., Raafat, M., Pak, E., Hammond, S. & Murashov, A. K. MicroRNA machinery responds to peripheral nerve lesion in an injury-regulated pattern. Neuroscience 190, 386–397 (2011).
Emde, A. & Hornstein, E. miRNAs at the interface of cellular stress and disease. EMBO J. 33, 1428–1437 (2014).
Maurel, M. & Chevet, E. Endoplasmic reticulum stress signaling: the microRNA connection. Am. J. Physiol. Cell Physiol. 304, C1117–C1126 (2013).
Ying, Z. et al. The unfolded protein response and cholesterol biosynthesis link Luman/CREB3 to regenerative axon growth in sensory neurons. J. Neurosci. 35, 14557–14570 (2015).
Hu, Y. et al. Differential effects of unfolded protein response pathways on axon injury-induced death of retinal ganglion cells. Neuron 73, 445–452 (2012).
Sun, A. X., Crabtree, G. R. & Yoo, A. S. MicroRNAs: regulators of neuronal fate. Curr. Opin. Cell Biol. 25, 215–221 (2013).
Motti, D. et al. Identification of miRNAs involved in DRG neurite outgrowth and their putative targets. FEBS Lett. 591, 2091–2105 (2017).
Strickland, I. T. et al. Axotomy-induced miR-21 promotes axon growth in adult dorsal root ganglion neurons. PLOS ONE 6, e23423 (2011).
Zhang, H. Y. et al. MicroRNAs 144, 145, and 214 are down-regulated in primary neurons responding to sciatic nerve transection. Brain Res. 1383, 62–70 (2011).
Zhou, S. et al. microRNA-222 targeting PTEN promotes neurite outgrowth from adult dorsal root ganglion neurons following sciatic nerve transection. PLOS ONE 7, e44768 (2012).
Wu, D. & Murashov, A. K. MicroRNA-431 regulates axon regeneration in mature sensory neurons by targeting the Wnt antagonist Kremen1. Front. Mol. Neurosci 6, 35 (2013).
Lisi, V. et al. Enhanced neuronal regeneration in the CAST/Ei mouse strain is linked to expression of differentiation markers after injury. Cell Rep. 20, 1136–1147 (2017).
Li, Z. & Rana, T. M. Therapeutic targeting of microRNAs: current status and future challenges. Nat. Rev. Drug Discov. 13, 622–638 (2014).
Niemi, J. P. et al. A critical role for macrophages near axotomized neuronal cell bodies in stimulating nerve regeneration. J. Neurosci. 33, 16236–16248 (2013).
Kwon, M. J. et al. Contribution of macrophages to enhanced regenerative capacity of dorsal root ganglia sensory neurons by conditioning injury. J. Neurosci. 33, 15095–15108 (2013).
Ciabrelli, F. & Cavalli, G. Chromatin-driven behavior of topologically associating domains. J. Mol. Biol. 427, 608–625 (2015).
Ali, T., Renkawitz, R. & Bartkuhn, M. Insulators and domains of gene expression. Curr. Opin. Genet. Dev. 37, 17–26 (2016).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Bonev, B. & Cavalli, G. Organization and function of the 3D genome. Nat. Rev. Genet. 17, 661–678 (2016).
Ong, C. T. & Corces, V. G. Enhancer function: new insights into the regulation of tissue-specific gene expression. Nat. Rev. Genet. 12, 283–293 (2011).
Cartoni, R. et al. The mammalian-specific protein Armcx1 regulates mitochondrial transport during axon regeneration. Neuron 92, 1294–1307 (2016).
Han, S. M., Baig, H. S. & Hammarlund, M. Mitochondria localize to injured axons to support regeneration. Neuron 92, 1308–1323 (2016).
Sainath, R. et al. Chondroitin sulfate proteoglycans negatively regulate the positioning of mitochondria and endoplasmic reticulum to distal axons. Dev. Neurobiol. 77, 1351–1370 (2017).
Seo, J., Singh, N. N., Ottesen, E. W., Lee, B. M. & Singh, R. N. A novel human-specific splice isoform alters the critical C-terminus of Survival Motor Neuron protein. Sci. Rep 6, 30778 (2016).
Li, M. et al. A human-specific AS3MT isoform and BORCS7 are molecular risk factors in the 10q24.32 schizophrenia-associated locus. Nat. Med. 22, 649–656 (2016).
Stahl, P. D. & Wainszelbaum, M. J. Human-specific genes may offer a unique window into human cell signaling. Sci Signal. 2, pe59 (2009).
Aldiri, I. et al. The dynamic epigenetic landscape of the retina during development, reprogramming, and tumorigenesis. Neuron 94, 550–568.e10 (2017).
Geoffroy, C. G., Meves, J. M. & Zheng, B. The age factor in axonal repair after spinal cord injury: a focus on neuron-intrinsic mechanisms. Neurosci. Lett. 652, 41–49 (2017).
Painter, M. W. et al. Diminished Schwann cell repair responses underlie age-associated impaired axonal regeneration. Neuron 83, 331–343 (2014).
von Schimmelmann, M. et al. Polycomb repressive complex 2 (PRC2) silences genes responsible for neurodegeneration. Nat. Neurosci. 19, 1321–1330 (2016).
Richardson, P. M. & Issa, V. M. Peripheral injury enhances central regeneration of primary sensory neurones. Nature 309, 791–793 (1984).
McQuarrie, I. G. & Grafstein, B. Axon outgrowth enhanced by a previous nerve injury. Arch. Neurol. 29, 53–55 (1973).
Neumann, S. & Woolf, C. J. Regeneration of dorsal column fibers into and beyond the lesion site following adult spinal cord injury. Neuron 23, 83–91 (1999).
Tuszynski, M. H. & Steward, O. Concepts and methods for the study of axonal regeneration in the CNS. Neuron 74, 777–791 (2012).
Ambron, R. T. & Walters, E. T. Priming events and retrograde injury signals. A new perspective on the cellular and molecular biology of nerve regeneration. Mol. Neurobiol. 13, 61–79 (1996).
Tazaki, A., Tanaka, E. M. & Fei, J. Salamander spinal cord regeneration: the ultimate positive control in vertebrate spinal cord regeneration. Dev. Biol. 432, 63–71 (2017).
Graff, J., Kim, D., Dobbin, M. M. & Tsai, L. H. Epigenetic regulation of gene expression in physiological and pathological brain processes. Physiol. Rev. 91, 603–649 (2011).
Wang, Z., Tang, B., He, Y. & Jin, P. DNA methylation dynamics in neurogenesis. Epigenomics 8, 401–414 (2016).
Guo, J. U. et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat. Neurosci. 14, 1345–1351 (2011).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why? Mol. Cell 49, 825–837 (2013).
Zhu, H., Wang, G. & Qian, J. Transcription factors as readers and effectors of DNA methylation. Nat. Rev. Genet. 17, 551–565 (2016).
Gonzalez-Sandoval, A. & Gasser, S. M. On TADs and LADs: spatial control over gene expression. Trends Genet. 32, 485–495 (2016).
Ong, C. T. & Corces, V. G. CTCF: an architectural protein bridging genome topology and function. Nat. Rev. Genet. 15, 234–246 (2014).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572.e24 (2017). This study performs an ultra-high-resolution Hi-C mapping of mouse neural differentiation and provides insights into the factors that influence the dynamics of chromatin interactions during neuronal development.
Soufi, A. & Zaret, K. S. Understanding impediments to cellular conversion to pluripotency by assessing the earliest events in ectopic transcription factor binding to the genome. Cell Cycle 12, 1487–1491 (2013).
Valentin, G. in Muller’s Archiv fur Anatomie, Physiologie und wissenschaftliche Medicin (ed. Müller, J.) 139–164. (Veit et Comp, Berlin, 1839).
Pannese, E. Number and structure of perisomatic satellite cells of spinal ganglia under normal conditions or during axon regeneration and neuronal hypertrophy. Z. Zellforsch. Mikrosk. Anat. 63, 568–592 (1964).
Pannese, E. The structure of the perineuronal sheath of satellite glial cells (SGCs) in sensory ganglia. Neuron Glia Biol. 6, 3–10 (2010).
Christie, K. et al. Intraganglionic interactions between satellite cells and adult sensory neurons. Mol. Cell. Neurosci. 67, 1–12 (2015).
Fenzi, F., Benedetti, M. D., Moretto, G. & Rizzuto, N. Glial cell and macrophage reactions in rat spinal ganglion after peripheral nerve lesions: an immunocytochemical and morphometric study. Arch. Ital. Biol. 139, 357–365 (2001).
Xie, W., Strong, J. A. & Zhang, J. M. Early blockade of injured primary sensory afferents reduces glial cell activation in two rat neuropathic pain models. Neuroscience 160, 847–857 (2009).
Zhang, H. et al. Altered functional properties of satellite glial cells in compressed spinal ganglia. Glia 57, 1588–1599 (2009).
Xu, M., Aita, M. & Chavkin, C. Partial infraorbital nerve ligation as a model of trigeminal nerve injury in the mouse: behavioral, neural, and glial reactions. J. Pain 9, 1036–1048 (2008).
Warwick, R. A. & Hanani, M. The contribution of satellite glial cells to chemotherapy-induced neuropathic pain. Eur. J. Pain 17, 571–580 (2013).
Obata, K. & Noguchi, K. MAPK activation in nociceptive neurons and pain hypersensitivity. Life Sci. 74, 2643–2653 (2004).
Mikuzuki, L. et al. Phenotypic change in trigeminal ganglion neurons associated with satellite cell activation via extracellular signal-regulated kinase phosphorylation is involved in lingual neuropathic pain. Eur. J. Neurosci. 46, 2190–2202 (2017).
Pannese, E. Perikaryal surface specializations of neurons in sensory ganglia. Int. Rev. Cytol. 220, 1–34 (2002).
Hanani, M., Huang, T. Y., Cherkas, P. S., Ledda, M. & Pannese, E. Glial cell plasticity in sensory ganglia induced by nerve damage. Neuroscience 114, 279–283 (2002).
Hanani, M. Satellite glial cells in sensory ganglia: from form to function. Brain Res. Brain Res. Rev. 48, 457–476 (2005).
Vit, J. P., Ohara, P. T., Bhargava, A., Kelley, K. & Jasmin, L. Silencing the Kir4.1 potassium channel subunit in satellite glial cells of the rat trigeminal ganglion results in pain-like behavior in the absence of nerve injury. J. Neurosci. 28, 4161–4171 (2008).
Humbertson, A. Jr., Zimmermann, E. & Leedy, M. A chronological study of mitotic activity in satellite cell hyperplasia associated with chromatolytic neurons. Z. Zellforsch. Mikrosk. Anat. 100, 507–515 (1969).
Donegan, M., Kernisant, M., Cua, C., Jasmin, L. & Ohara, P. T. Satellite glial cell proliferation in the trigeminal ganglia after chronic constriction injury of the infraorbital nerve. Glia 61, 2000–2008 (2013).
Huang, L. Y., Gu, Y. & Chen, Y. Communication between neuronal somata and satellite glial cells in sensory ganglia. Glia 61, 1571–1581 (2013).
Jessen, K. R. & Mirsky, R. The repair Schwann cell and its function in regenerating nerves. J. Physiol. 594, 3521–3531 (2016).
Cho, Y., Park, D. & Cavalli, V. Filamin A is required in injured axons for HDAC5 activity and axon regeneration. J. Biol. Chem. 290, 22759–22770 (2015).
Gaub, P. et al. HDAC inhibition promotes neuronal outgrowth and counteracts growth cone collapse through CBP/p300 and P/CAF-dependent p53 acetylation. Cell Death Differ. 17, 1392–1408 (2010).
Parikh, P. et al. Regeneration of axons in injured spinal cord by activation of bone morphogenetic protein/Smad1 signaling pathway in adult neurons. Proc. Natl Acad. Sci. USA 108, E99–E107 (2011).
Smith, R. P. et al. Transcriptional profiling of intrinsic PNS factors in the postnatal mouse. Mol. Cell. Neurosci. 46, 32–44 (2011).
Acknowledgements
The authors’ research on these topics has been generously supported by the US National Institute of Health grants NS096034, NS082446 and NS099603, the University of Missouri Spinal Cord Injury Research Program and a Philip and Sima K. Needleman Doctoral Fellowship. The authors thank H. Gabel for helpful comments and critical reading of the manuscript. The authors thank the Cavalli laboratory members for their helpful comments on the manuscript. The authors apologize to those whose studies could not be cited owing to space limitation.
Reviewer information
Nature Reviews Neuroscience thanks S. Di Giovanni, J. Twiss and the other anonymous reviewer(s), for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.M. and V.C. researched data for the article, made substantial contributions to discussions of the content, wrote the article and reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Transcription factors
-
Proteins that activate or repress the expression of genes by binding to DNA sequence motifs proximal to a gene’s transcription start site or interacting enhancer regions.
- Epigenetic modifiers
-
Proteins that post-translationally modify either DNA or histones, which affects DNA compaction and accessibility for protein binding.
- MicroRNAs
-
(miRNAs). Single-stranded RNA molecules of 20–23 nucleotides in length, generated endogenously from a single-stranded hairpin precursor, which act as post-transcriptional inhibitors in association with the RNA-induced silencing complex (RISC).
- Motor proteins
-
Proteins, such as kinesin, dynein and myosin, that use either the microtubule or the actin cytoskeleton for movement by converting chemical energy into mechanical force.
- Cofactor
-
Protein that binds with other proteins to form heteromeric complexes that alter or enhance the function of its binding partners.
- Poly(ADP-ribose)
-
A negatively charged polymer of ADP-ribose that can be added to proteins. Poly(ADP-ribose) represents a unique post-translational modification that regulates protein function.
- RNA processing
-
The process by which an RNA molecule translated from DNA undergoes modifications, including 5′ capping, 3′ polyadenylation, splicing and methylation, before the RNA is translated into a protein.
- Unfolded protein response
-
A cellular stress response that is triggered by an excess of unfolded or misfolded proteins in the endoplasmic reticulum.
- Epitranscriptomic mechanisms
-
Post-transcriptional RNA modifications that regulate mRNA half-life or translation or otherwise alter biological processes.
- Next-generation sequencing
-
High-throughput parallel sequencing of either DNA or RNA.
- Cytokines
-
Originally defined as immune system proteins, these proteins are now known to be released by most cells and are important in regulating intercellular communication, cell function and cell survival.
- Induced pluripotent stem cells
-
(iPSCs). Cells created from differentiated cell types (for example, fibroblasts) that are reprogrammed by a cocktail of transcription factors (or other approaches) back to a pluripotent state and are capable of differentiating into all three germ layers.
- Genome topology
-
The 3D DNA structure that dictates its accessibility to binding by proteins such as transcription factors and epigenetic modifiers.
Rights and permissions
About this article
Cite this article
Mahar, M., Cavalli, V. Intrinsic mechanisms of neuronal axon regeneration. Nat Rev Neurosci 19, 323–337 (2018). https://doi.org/10.1038/s41583-018-0001-8
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41583-018-0001-8
This article is cited by
-
The role of TSC1 and TSC2 proteins in neuronal axons
Molecular Psychiatry (2024)
-
The Transcription Factor Ets1 Influences Axonal Growth via Regulation of Lcn2
Molecular Neurobiology (2024)
-
Genes in Axonal Regeneration
Molecular Neurobiology (2024)
-
MST2 Acts via AKT Activity to Promote Neurite Outgrowth and Functional Recovery after Spinal Cord Injury in Mice
Molecular Neurobiology (2024)
-
M2 macrophage-derived cathepsin S promotes peripheral nerve regeneration via fibroblast–Schwann cell-signaling relay
Journal of Neuroinflammation (2023)