Abstract
Schizophrenia is a severe psychiatric disorder with considerable morbidity and mortality. Although the past two decades have seen limited improvement in the treatment of schizophrenia, research into the genetic causes of this condition has made important advances that offer new insights into the aetiology of schizophrenia. This Review summarizes the evidence for a polygenic architecture of schizophrenia that involves a large number of risk alleles across the whole range of population frequencies. These genetic risk loci implicate biological processes related to neurodevelopment, neuronal excitability, synaptic function and the immune system in the pathogenesis of schizophrenia. Mathematical models also suggest a substantial overlap between schizophrenia and psychiatric, behavioural and cognitive traits, a situation that has implications for understanding its clinical epidemiology, psychiatric nosology and pathobiology. Looking ahead, further genetic discoveries are expected to lead to clinically relevant predictive approaches for identifying high-risk individuals, improved diagnostic accuracy, increased yield from drug development programmes and improved stratification strategies to address the heterogeneous disease course and treatment responses observed among affected patients.
Key points
-
Schizophrenia is characterized by ‘positive’ psychotic symptoms (including hallucinations and delusions) and ‘negative’ symptoms (including blunted affect, apathy and social impairment); this disorder is associated with considerable morbidity and mortality.
-
In the past decade, important advances have been made in our understanding of the genetics of schizophrenia.
-
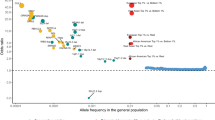
The polygenic architecture of schizophrenia is accounted for by thousands of common genetic variants with small effect sizes and a few rare variants with large effect sizes.
-
These genetic risk variants implicate dysregulation of biological processes linked to neurodevelopment, neuronal excitability, synaptic function and the immune system in schizophrenia.
-
Genetic risk factors associated with schizophrenia transcend diagnostic boundaries and form a continuum with normal psychosocial traits, which challenges current psychiatric nosology.
-
Although increasingly larger sample sizes will accelerate the discovery of genetic variants, novel statistical methodologies could also improve the efficiency of analyses, render discoveries clinically relevant and facilitate precision medicine approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Charlson, F. J. et al. Global epidemiology and burden of schizophrenia: findings from the Global Burden of Disease study 2016. Schizophr. Bull. 44, 1195–1203 (2018).
Whiteford, H. A. et al. Global burden of disease attributable to mental and substance use disorders: findings from the Global Burden of Disease Study 2010. Lancet 382, 1575–1586 (2013).
Laursen, T. M., Nordentoft, M. & Mortensen, P. B. Excess early mortality in schizophrenia. Annu. Rev. Clin. Psychol. 10, 425–448 (2014).
Fusar-Poli, P. et al. Cognitive functioning in prodromal psychosis: a meta-analysis. Arch. Gen. Psychiatry 69, 562–571 (2012).
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders (DSM-5). (APA, 2013).
World Health Organization. International classification of diseases for mortality and morbidity statistics (11th revision). https://icd.who.int/browse11/l-m/en. (WHO, 2018).
Lieberman, J. A. et al. Effectiveness of antipsychotic drugs in patients with chronic schizophrenia. N. Engl. J. Med. 353, 1209–1223 (2005).
Kahn, R. S. & Keefe, R. S. Schizophrenia is a cognitive illness: time for a change in focus. JAMA Psychiatry 70, 1107–1112 (2013).
Thornicroft, G. et al. Global pattern of experienced and anticipated discrimination against people with schizophrenia: a cross-sectional survey. Lancet 373, 408–415 (2009).
Gustavsson, A. et al. Cost of disorders of the brain in Europe 2010. Eur. Neuropsychopharmacol. 21, 718–779 (2011).
Demyttenaere, K. et al. Prevalence, severity, and unmet need for treatment of mental disorders in the World Health Organization World Mental Health Surveys. JAMA 291, 2581–2590 (2004).
Gottesman, I. I. & Wolfgram, D. L. Schizophrenia Genesis: The Origins of Madness. (Freeman, 1991).
Howes, O. D. & Kapur, S. The dopamine hypothesis of schizophrenia: version III — the final common pathway. Schizophr. Bull. 35, 549–562 (2009).
Moghaddam, B. & Javitt, D. From revolution to evolution: the glutamate hypothesis of schizophrenia and its implication for treatment. Neuropsychopharmacology 37, 4–15 (2012).
Nakazawa, K. et al. GABAergic interneuron origin of schizophrenia pathophysiology. Neuropharmacology 62, 1574–1583 (2012).
Lieberman, J. A. & First, M. B. Psychotic disorders. N. Engl. J. Med. 379, 270–280 (2018).
Carlsson, A. & Lindqvist, M. Effect of chlorpromazine or haloperidol on formation of 3methoxytyramine and normetanephrine in mouse brain. Acta Pharmacol. Toxicol. 20, 140–144 (1963).
Carlsson, A., Lindqvist, M. & Magnusson, T. 3,4-Dihydroxyphenylalanine and 5-hydroxytryptophan as reserpine antagonists. Nature 180, 1200 (1957).
Howes, O., McCutcheon, R. & Stone, J. Glutamate and dopamine in schizophrenia: an update for the 21st century. J. Psychopharmacol. 29, 97–115 (2015).
Javitt, D. C. & Zukin, S. R. Recent advances in the phencyclidine model of schizophrenia. Am. J. Psychiatry 148, 1301–1308 (1991).
Krystal, J. H. et al. Subanesthetic effects of the noncompetitive NMDA antagonist, ketamine, in humans. Psychotomimetic, perceptual, cognitive, and neuroendocrine responses. Arch. Gen. Psychiatry 51, 199–214 (1994).
Olney, J. W. & Farber, N. B. Glutamate receptor dysfunction and schizophrenia. Arch. Gen. Psychiatry 52, 998–1007 (1995).
Dalmau, J. et al. An update on anti-NMDA receptor encephalitis for neurologists and psychiatrists: mechanisms and models. Lancet Neurol. 18, 1045–1057 (2019).
Akbarian, S. et al. Gene expression for glutamic acid decarboxylase is reduced without loss of neurons in prefrontal cortex of schizophrenics. Arch. Gen. Psychiatry 52, 258–266 (1995).
Lewis, D. A., Hashimoto, T. & Volk, D. W. Cortical inhibitory neurons and schizophrenia. Nat. Rev. Neurosci. 6, 312–324 (2005).
Gonzalez-Burgos, G., Cho, R. Y. & Lewis, D. A. Alterations in cortical network oscillations and parvalbumin neurons in schizophrenia. Biol. Psychiatry 77, 1031–1040 (2015).
Paulus, M. P. & Thompson, W. K. The challenges and opportunities of small effects: the new normal in academic psychiatry. JAMA Psychiatry 76, 353–354 (2019).
Murray, R. M. & Lewis, S. W. Is schizophrenia a neurodevelopmental disorder? Br. Med. J. 295, 681–682 (1987).
Weinberger, D. R. Implications of normal brain development for the pathogenesis of schizophrenia. Arch. Gen. Psychiatry 44, 660–669 (1987).
Howes, O. D. & Murray, R. M. Schizophrenia: an integrated sociodevelopmental-cognitive model. Lancet 383, 1677–1687 (2014).
Insel, T. R. Rethinking schizophrenia. Nature 468, 187–193 (2010).
Glantz, L. A. & Lewis, D. A. Decreased dendritic spine density on prefrontal cortical pyramidal neurons in schizophrenia. Arch. Gen. Psychiatry 57, 65–73 (2000).
Glausier, J. R. & Lewis, D. A. Dendritic spine pathology in schizophrenia. Neuroscience 251, 90–107 (2013).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science 362, eaat8127 (2018).
Stevens, B. et al. The classical complement cascade mediates CNS synapse elimination. Cell 131, 1164–1178 (2007).
Salter, M. W. & Stevens, B. Microglia emerge as central players in brain disease. Nat. Med. 23, 1018–1027 (2017).
Miller, B. J., Buckley, P., Seabolt, W., Mellor, A. & Kirkpatrick, B. Meta-analysis of cytokine alterations in schizophrenia: clinical status and antipsychotic effects. Biol. Psychiatry 70, 663–671 (2011).
Pillinger, T. et al. A meta-analysis of immune parameters, variability, and assessment of modal distribution in psychosis and test of the immune subgroup hypothesis. Schizophr. Bull. 45, 1120–1133 (2019).
Haijma, S. V. et al. Brain volumes in schizophrenia: a meta-analysis in over 18 000 subjects. Schizophr. Bull. 39, 1129–1138 (2013).
van Erp, T. G. M. et al. Cortical brain abnormalities in 4474 individuals with schizophrenia and 5098 control subjects via the Enhancing Neuro Imaging Genetics through Meta Analysis (ENIGMA) Consortium. Biol. Psychiatry 84, 644–654 (2018).
van Erp, T. G. et al. Subcortical brain volume abnormalities in 2028 individuals with schizophrenia and 2540 healthy controls via the ENIGMA consortium. Mol. Psychiatry 21, 547–553 (2016).
Alnaes, D. et al. Brain heterogeneity in schizophrenia and its association with polygenic risk. JAMA Psychiatry 76, 739–748 (2019).
Fusar-Poli, P. et al. Progressive brain changes in schizophrenia related to antipsychotic treatment? A meta-analysis of longitudinal MRI studies. Neurosci. Biobehav. Rev. 37, 1680–1691 (2013).
Cannon, T. D. et al. Progressive reduction in cortical thickness as psychosis develops: a multisite longitudinal neuroimaging study of youth at elevated clinical risk. Biol. Psychiatry 77, 147–157 (2015).
Ho, B. C., Andreasen, N. C., Ziebell, S., Pierson, R. & Magnotta, V. Long-term antipsychotic treatment and brain volumes: a longitudinal study of first-episode schizophrenia. Arch. Gen. Psychiatry 68, 128–137 (2011).
Brown, A. S. & Derkits, E. J. Prenatal infection and schizophrenia: a review of epidemiologic and translational studies. Am. J. Psychiatry 167, 261–280 (2010).
Cannon, M., Jones, P. B. & Murray, R. M. Obstetric complications and schizophrenia: historical and meta-analytic review. Am. J. Psychiatry 159, 1080–1092 (2002).
Khandaker, G. M., Zimbron, J., Dalman, C., Lewis, G. & Jones, P. B. Childhood infection and adult schizophrenia: a meta-analysis of population-based studies. Schizophr. Res. 139, 161–168 (2012).
van Os, J., Kenis, G. & Rutten, B. P. The environment and schizophrenia. Nature 468, 203–212 (2010).
Cantor-Graae, E. & Selten, J. P. Schizophrenia and migration: a meta-analysis and review. Am. J. Psychiatry 162, 12–24 (2005).
Kraepelin, E., Barclay, R. M. & Robertson, G. M. Dementia præcox and paraphrenia. (E. & S. Livingstone, 1919).
Sullivan, P. F., Kendler, K. S. & Neale, M. C. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch. Gen. Psychiatry 60, 1187–1192 (2003).
Hilker, R. et al. Heritability of schizophrenia and schizophrenia spectrum based on the Nationwide Danish twin register. Biol. Psychiatry 83, 492–498 (2018).
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
Hall, J. M. et al. Linkage of early-onset familial breast cancer to chromosome 17q21. Science 250, 1684–1689 (1990).
Risch, N. & Merikangas, K. The future of genetic studies of complex human diseases. Science 273, 1516–1517 (1996).
Ng, M. Y. et al. Meta-analysis of 32 genome-wide linkage studies of schizophrenia. Mol. Psychiatry 14, 774–785 (2009).
Lewis, C. M. et al. Genome scan meta-analysis of schizophrenia and bipolar disorder, part II: Schizophrenia. Am. J. Hum. Genet. 73, 34–48 (2003).
Corder, E. H. et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 261, 921–923 (1993).
Gatt, J. M., Burton, K. L., Williams, L. M. & Schofield, P. R. Specific and common genes implicated across major mental disorders: a review of meta-analysis studies. J. Psychiatr. Res. 60, 1–13 (2015).
Psychiatric GWAS Consortium Steering Committee. A framework for interpreting genome-wide association studies of psychiatric disorders. Mol. Psychiatry 14, 10–17 (2009).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Li, Z. et al. Genome-wide association analysis identifies 30 new susceptibility loci for schizophrenia. Nat. Genet. 49, 1576–1583 (2017).
Pardinas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Lam, M. et al. Comparative genetic architectures of schizophrenia in East Asian and European populations. Nat. Genet. 51, 1670–1678 (2019).
Manolio, T. A. et al. New models of collaboration in genome-wide association studies: the Genetic Association Information Network. Nat. Genet. 39, 1045–1051 (2007).
O’Donovan, M. C. et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat. Genet. 40, 1053–1055 (2008).
International Schizophrenia, C. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Lee, S. H. et al. Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat. Genet. 44, 247–250 (2012).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Frei, O. et al. Bivariate causal mixture model quantifies polygenic overlap between complex traits beyond genetic correlation. Nat. Commun. 10, 2417 (2019).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Lee, S. H., Wray, N. R., Goddard, M. E. & Visscher, P. M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Speed, D. & Balding, D. J. SumHer better estimates the SNP heritability of complex traits from summary statistics. Nat. Genet. 51, 277–284 (2019).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Devor, A. et al. Genetic evidence for role of integration of fast and slow neurotransmission in schizophrenia. Mol. Psychiatry 22, 792–801 (2017).
Lips, E. S. et al. Functional gene group analysis identifies synaptic gene groups as risk factor for schizophrenia. Mol. Psychiatry 17, 996–1006 (2012).
Wang, Y. et al. Leveraging genomic annotations and pleiotropic enrichment for improved replication rates in schizophrenia GWAS. PLoS Genet. 12, e1005803 (2016).
Darnell, J. C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Verkerk, A. J. et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65, 905–914 (1991).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Wray, N. R., Wijmenga, C., Sullivan, P. F., Yang, J. & Visscher, P. M. Common disease is more complex than implied by the core gene omnigenic model. Cell 173, 1573–1580 (2018).
Schrode, N. et al. Synergistic effects of common schizophrenia risk variants. Nat. Genet. 51, 1475–1485 (2019).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Schork, A. J. et al. All SNPs are not created equal: genome-wide association studies reveal a consistent pattern of enrichment among functionally annotated SNPs. PLoS Genet. 9, e1003449 (2013).
Roussos, P. et al. A role for noncoding variation in schizophrenia. Cell Rep. 9, 1417–1429 (2014).
Flint, J. & Ideker, T. The great hairball gambit. PLoS Genet. 15, e1008519 (2019).
Maki-Marttunen, T. et al. Biophysical psychiatry – how computational neuroscience can help to understand the complex mechanisms of mental disorders. Front. Psychiatry 10, 534 (2019).
Visscher, P. M. et al. 10 years of GWAS discovery: biology, function, and translation. Am. J. Hum. Genet. 101, 5–22 (2017).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Cross-Disorder Group of the Psychiatric Genomics Consortium. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Brainstorm Consortium et al. Analysis of shared heritability in common disorders of the brain. Science 360, eaap8757 (2018).
Bipolar Disorder and Schizophrenia Working Group of the Psychiatric Genomics Consortium. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715.e16 (2018).
Andreassen, O. A. et al. Improved detection of common variants associated with schizophrenia by leveraging pleiotropy with cardiovascular-disease risk factors. Am. J. Hum. Genet. 92, 197–209 (2013).
Pasman, J. A. et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal influence of schizophrenia. Nat. Neurosci. 21, 1161–1170 (2018).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919 (2018).
Smeland, O. B. et al. Identification of genetic loci jointly influencing schizophrenia risk and the cognitive traits of verbal-numerical reasoning, reaction time, and general cognitive function. JAMA Psychiatry 74, 1065–1075 (2017).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Smeland, O. B. et al. Genome-wide analysis reveals extensive genetic overlap between schizophrenia, bipolar disorder, and intelligence. Mol. Psychiatry 25, 844–853 (2020).
Le Hellard, S. et al. Identification of gene loci that overlap between schizophrenia and educational attainment. Schizophr. Bull. 43, 654–664 (2017).
Smeland, O. B. et al. Genetic overlap between schizophrenia and volumes of hippocampus, putamen, and intracranial volume indicates shared molecular genetic mechanisms. Schizophr. Bull. 44, 854–864 (2018).
Lee, P. H. et al. Partitioning heritability analysis reveals a shared genetic basis of brain anatomy and schizophrenia. Mol. Psychiatry 21, 1680–1689 (2016).
Ohi, K. et al. Genetic correlations between subcortical brain volumes and psychiatric disorders. Br. J. Psychiatry 216, 280–283 (2020).
Power, R. A. et al. Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat. Neurosci. 18, 953–955 (2015).
Lo, M. T. et al. Genome-wide analyses for personality traits identify six genomic loci and show correlations with psychiatric disorders. Nat. Genet. 49, 152–156 (2017).
Smeland, O. B. et al. Identification of genetic loci shared between schizophrenia and the Big Five personality traits. Sci. Rep. 7, 2222 (2017).
Solovieff, N., Cotsapas, C., Lee, P. H., Purcell, S. M. & Smoller, J. W. Pleiotropy in complex traits: challenges and strategies. Nat. Rev. Genet. 14, 483–495 (2013).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Lee, S. H., Yang, J., Goddard, M. E., Visscher, P. M. & Wray, N. R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism-derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542 (2012).
Stahl, E. A. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat. Genet. 51, 793–803 (2019).
Ikeda, M. et al. Re-evaluating classical body type theories: genetic correlation between psychiatric disorders and body mass index. Psychol. Med. 48, 1745–1748 (2018).
Bahrami, S. et al. Shared genetic loci between body mass index and major psychiatric disorders: a genome-wide association study. JAMA Psychiatry 77, 503–512 (2020).
Green, M. F. Cognitive impairment and functional outcome in schizophrenia and bipolar disorder. J. Clin. Psychiatry 67(Suppl. 9), 3–8 (2006).
Hill, W. D. et al. Age-dependent pleiotropy between general cognitive function and major psychiatric disorders. Biol. Psychiatry 80, 266–273 (2016).
Palmer, B. W. et al. Is it possible to be schizophrenic yet neuropsychologically normal? Neuropsychology 11, 437–446 (1997).
Hibar, D. P. et al. Common genetic variants influence human subcortical brain structures. Nature 520, 224–229 (2015).
Franke, B. et al. Genetic influences on schizophrenia and subcortical brain volumes: large-scale proof of concept. Nat. Neurosci. 19, 420–431 (2016).
Holland, D. et al. Beyond SNP heritability: polygenicity and discoverability of phenotypes estimated with a univariate gaussian mixture model. PLoS Genet. https://doi.org/10.1371/journal.pgen.1008612 (2019).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Power, R. A. et al. Fecundity of patients with schizophrenia, autism, bipolar disorder, depression, anorexia nervosa, or substance abuse vs their unaffected siblings. JAMA Psychiatry 70, 22–30 (2013).
Keller, M. C. & Miller, G. Resolving the paradox of common, harmful, heritable mental disorders: which evolutionary genetic models work best? Behav. Brain Sci. 29, 385–404 (2006).
Jarvik, L. F. & Deckard, B. S. The Odyssean personality. A survival advantage for carriers of genes predisposing to schizophrenia? Neuropsychobiology 3, 179–191 (1977).
Mullins, N. et al. Reproductive fitness and genetic risk of psychiatric disorders in the general population. Nat. Commun. 8, 15833 (2017).
Escott-Price, V. et al. The relationship between common variant schizophrenia liability and number of offspring in the UK Biobank. Am. J. Psychiatry 176, 661–666 (2019).
Crow, T. J. Is schizophrenia the price that Homo sapiens pays for language? Schizophr. Res. 28, 127–141 (1997).
Srinivasan, S. et al. Genetic markers of human evolution are enriched in schizophrenia. Biol. Psychiatry 80, 284–292 (2016).
Xu, K., Schadt, E. E., Pollard, K. S., Roussos, P. & Dudley, J. T. Genomic and network patterns of schizophrenia genetic variation in human evolutionary accelerated regions. Mol. Biol. Evol. 32, 1148–1160 (2015).
Charlesworth, B. The effects of deleterious mutations on evolution at linked sites. Genetics 190, 5–22 (2012).
McClellan, J. M., Susser, E. & King, M. C. Schizophrenia: a common disease caused by multiple rare alleles. Br. J. Psychiatry 190, 194–199 (2007).
Marshall, C. R. et al. Contribution of copy number variants to schizophrenia from a genome-wide study of 41,321 subjects. Nat. Genet. 49, 27–35 (2017).
Singh, T. et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat. Neurosci. 19, 571–577 (2016).
Steinberg, S. et al. Truncating mutations in RBM12 are associated with psychosis. Nat. Genet. 49, 1251–1254 (2017).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Genovese, G. et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat. Neurosci. 19, 1433–1441 (2016).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Gandal, M. J., Leppa, V., Won, H., Parikshak, N. N. & Geschwind, D. H. The road to precision psychiatry: translating genetics into disease mechanisms. Nat. Neurosci. 19, 1397–1407 (2016).
Singh, T. et al. The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability. Nat. Genet. 49, 1167–1173 (2017).
Sullivan, P. F. & Geschwind, D. H. Defining the genetic, genomic, cellular, and diagnostic architectures of psychiatric disorders. Cell 177, 162–183 (2019).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Stefansson, H. et al. Large recurrent microdeletions associated with schizophrenia. Nature 455, 232–236 (2008).
Murphy, K. C., Jones, L. A. & Owen, M. J. High rates of schizophrenia in adults with velo-cardio-facial syndrome. Arch. Gen. Psychiatry 56, 940–945 (1999).
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Huckins, L. M. et al. Gene expression imputation across multiple brain regions provides insights into schizophrenia risk. Nat. Genet. 51, 659–674 (2019).
Mancuso, N. et al. Integrating Gene Expression with Summary Association Statistics to Identify Genes Associated with 30 Complex Traits. Am. J. Hum. Genet. 100, 473–487 (2017).
Gamazon, E. R., Zwinderman, A. H., Cox, N. J., Denys, D. & Derks, E. M. Multi-tissue transcriptome analyses identify genetic mechanisms underlying neuropsychiatric traits. Nat. Genet. 51, 933–940 (2019).
Wainberg, M. et al. Opportunities and challenges for transcriptome-wide association studies. Nat. Genet. 51, 592–599 (2019).
Li, M. et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362, eaat7615 (2018).
de la Torre-Ubieta, L. et al. The dynamic landscape of open chromatin during human cortical neurogenesis. Cell 172, 289–304 (2018).
Skene, N. G. et al. Genetic identification of brain cell types underlying schizophrenia. Nat. Genet. 50, 825–833 (2018).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Sullivan, P. F. et al. Psychiatric genomics: an update and an agenda. Am. J. Psychiatry 175, 15–27 (2018).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Ursini, G. et al. Convergence of placenta biology and genetic risk for schizophrenia. Nat. Med. 24, 792–801 (2018).
Casey, B. J. et al. The Adolescent Brain Cognitive Development (ABCD) study: Imaging acquisition across 21 sites. Dev. Cogn. Neurosci. 32, 43–54 (2018).
Magnus, P. et al. Cohort profile: the Norwegian Mother and Child Cohort Study (MoBa). Int. J. Epidemiol. 35, 1146–1150 (2006).
Hess, J. L. et al. A polygenic resilience score moderates the genetic risk for schizophrenia. Mol. Psychiatry https://doi.org/10.1038/s41380-019-0463-8 (2019).
Smoller, J. W. et al. Psychiatric genetics and the structure of psychopathology. Mol. Psychiatry 24, 409–420 (2019).
Allardyce, J. et al. Association between schizophrenia-related polygenic liability and the occurrence and level of mood-incongruent psychotic symptoms in bipolar disorder. JAMA Psychiatry 75, 28–35 (2018).
Insel, T. et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am. J. Psychiatry 167, 748–751 (2010).
Richards, A. L. et al. The relationship between polygenic risk scores and cognition in schizophrenia. Schizophr. Bull. 46, 336–344 (2020).
Frank, J. et al. Identification of increased genetic risk scores for schizophrenia in treatment-resistant patients. Mol. Psychiatry 20, 913 (2015).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
van der Meer, D. et al. Making the MOSTest of imaging genetics. Biol. Psychiatry 87 (Suppl.), S304–S305
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in ~700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Zhang, Y., Qi, G., Park, J. H. & Chatterjee, N. Estimation of complex effect-size distributions using summary-level statistics from genome-wide association studies across 32 complex traits. Nat. Genet. 50, 1318–1326 (2018).
Zhu, X. & Stephens, M. Bayesian large-scale multiple regression with summary statistics from genome-wide association studies. Ann. Appl. Stat. 11, 1561–1592 (2017).
Zeng, J. et al. Signatures of negative selection in the genetic architecture of human complex traits. Nat. Genet. 50, 746–753 (2018).
Smeland, O. B. et al. The emerging pattern of shared polygenic architecture of psychiatric disorders, conceptual and methodological challenges. Psychiatr. Genet. 29, 152–159 (2019).
Smeland, O. B. et al. Discovery of shared genomic loci using the conditional false discovery rate approach. Hum. Genet. 139, 85–94 (2020).
Schork, A. J., Wang, Y., Thompson, W. K., Dale, A. M. & Andreassen, O. A. New statistical approaches exploit the polygenic architecture of schizophrenia-implications for the underlying neurobiology. Curr. Opin. Neurobiol. 36, 89–98 (2016).
Andreassen, O. A. et al. Genetic pleiotropy between multiple sclerosis and schizophrenia but not bipolar disorder: differential involvement of immune-related gene loci. Mol. Psychiatry 20, 207–214 (2015).
McLaughlin, R. L. et al. Genetic correlation between amyotrophic lateral sclerosis and schizophrenia. Nat. Commun. 8, 14774 (2017).
Smeland, O. B. et al. Genome-wide association analysis of Parkinson’s disease and schizophrenia reveals shared genetic architecture and identifies novel risk loci. Biol. Psychiatry https://doi.org/10.1016/j.biopsych.2020.01.026 (2020).
Acknowledgements
The authors’ research is supported by National Institutes of Health (NIH) grants NS057198 and EB00790 and NIH National Institute on Drug Abuse (NIDA)/National Cancer Institute (NCI) grant U24DA041123 to A.M.D; and Research Council of Norway grants 229129, 213837, 248778, 223273 and 249711, South-East Norway Regional Health Authority grant 2017-112, and funding from K.G. Jebsen Stiftelsen (SKGJ) to O.A.A. The authors also thank N. Karadag for preparation of the original artwork for figure 2.
Author information
Authors and Affiliations
Contributions
O.A.A., O.B.S., O.F. and A.M.D. researched data for the article, contributed substantially to discussions of its content and participated in review or editing of the manuscript before submission. In addition, O.A.A. and O.B.S. wrote the initial draft.
Corresponding author
Ethics declarations
Competing interests
O.A.A. declares that he has received a speaker’s honorarium from Lundbeck and is a consultant for HealthLytix. A.M.D. declares that he is a founder of and holds equity interest in CorTechs Labs, that he is a member of the scientific advisory boards of CorTechs Labs and HealthLytix, and receives research funding from General Electric Healthcare. The terms of these arrangements have been reviewed and approved by the University of California San Diego in accordance with its conflict of interest policies. The other authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Conditional false discovery rate (FDR) software: https://github.com/precimed/pleiofdr
MiXeR software: https://github.com/precimed/mixer.
Glossary
- Polygenic risk score
-
(PRS). An estimate of overall genetic propensity to develop a given phenotype, derived from the sum of a given individual’s risk alleles weighted by their effect sizes.
- SNP-based heritability
-
The fraction of phenotypic variation attributable to common genetic variants detected in genome-wide association studies. Heritability of continuous traits (such as height) is estimated by comparison with the observed range (observed scale), whereas heritability of binary traits (such as schizophrenia) is estimated as a propensity score that takes into account population prevalence (liability scale).
- Linkage disequilibrium
-
The tendency for genes, alleles or other genetic markers to be non-randomly inherited in association with each other owing to physical proximity to one another on the same chromosome.
- Genetic pleiotropy
-
A genetic variant that affects more than one phenotype.
- Protein isoform
-
Many human genes encode multiple protein variants generated by alternative promoters, alternative mRNA splicing or post-translational modification.
Rights and permissions
About this article
Cite this article
Smeland, O.B., Frei, O., Dale, A.M. et al. The polygenic architecture of schizophrenia — rethinking pathogenesis and nosology. Nat Rev Neurol 16, 366–379 (2020). https://doi.org/10.1038/s41582-020-0364-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41582-020-0364-0
This article is cited by
-
Establishment and analysis of a novel diagnostic model for systemic juvenile idiopathic arthritis based on machine learning
Pediatric Rheumatology (2024)
-
Correlation analysis and gender differences of cognitive function based on mini-mental state examination (MMSE) and suicidal tendency in patients with schizophrenia
BMC Psychiatry (2024)
-
Genetics of human brain development
Nature Reviews Genetics (2024)
-
Shared genetic architecture between irritable bowel syndrome and psychiatric disorders reveals molecular pathways of the gut-brain axis
Genome Medicine (2023)
-
Progressive trajectories of schizophrenia across symptoms, genes, and the brain
BMC Medicine (2023)