Abstract
The ability of living systems to adapt to changing conditions originates from their capacity to change their molecular constitution. This is achieved by multiple mechanisms that modulate the quantitative composition and the diversity of the molecular inventory. Molecular diversification is particularly pronounced on the proteome level, at which multiple proteoforms derived from the same gene can in turn combinatorially form different protein complexes, thus expanding the repertoire of functional modules in the cell. The study of molecular and modular diversity and their involvement in responses to changing conditions has only recently become possible through the development of new ‘omics’-based screening technologies. This Review explores our current knowledge of the mechanisms regulating functional diversification along the axis of gene expression, with a focus on the proteome and interactome. We explore the interdependence between different molecular levels and how this contributes to functional diversity. Finally, we highlight several recent techniques for studying molecular diversity, with specific focus on mass spectrometry-based analysis of the proteome and its organization into functional modules, and examine future directions for this rapidly growing field.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
17 April 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41580-020-0249-5
References
Beadle, G. W. & Tatum, E. L. Genetic control of biochemical reactions in neurospora. Proc. Natl Acad. Sci. USA 27, 499–506 (1941).
Collins, F. S., Lander, E. S., Rogers, J. & Waterson, R. H. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Carter, H., Hofree, M. & Ideker, T. Genotype to phenotype via network analysis. Curr. Opin. Genet. Dev. 23, 611–621 (2013).
Rauscher, B., Valentini, E., Hardeland, U. & Boutros, M. Phenotype databases for genetic screens in human cells. J. Biotechnol. 261, 63–69 (2017).
Amberger, J., Bocchini, C. & Hamosh, A. A new face and new challenges for Online Mendelian Inheritance in Man (OMIM®). Hum. Mutat. 32, 564–567 (2011).
Arkin, A. P. & Schaffer, D. V. Network news: innovations in 21st century systems biology. Cell 144, 844–849 (2011).
Hartwell, L. H., Hopfield, J. J., Leibler, S. & Murray, A. W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Li, J. J., Bickel, P. J. & Biggin, M. D. System wide analyses have underestimated protein abundances and the importance of transcription in mammals. PeerJ 2, e270 (2014).
Jovanovic, M. et al. Dynamic profiling of the protein life cycle in response to pathogens. Science 347, 1259038 (2015).
Marguerat, S. et al. Quantitative analysis of fission yeast transcriptomes and proteomes in proliferating and quiescent cells. Cell 151, 671–683 (2012).
Thompson, D., Regev, A. & Roy, S. Comparative analysis of gene regulatory networks: from network reconstruction to evolution. Annu. Rev. Cell Dev. Biol. 31, 399–428 (2015).
Bernadskaya, Y. & Christiaen, L. Transcriptional control of developmental cell behaviors. Annu. Rev. Cell Dev. Biol. 32, 77–101 (2016).
Kilchert, C., Wittmann, S. & Vasiljeva, L. The regulation and functions of the nuclear RNA exosome complex. Nat. Rev. Mol. Cell Biol. 17, 227–239 (2016).
Magnuson, B., Bedi, K. & Ljungman, M. Genome stability versus transcript diversity. DNA Repair 44, 81–86 (2016). This review discusses sources of error from the genome to the proteome and how errors can contribute to molecular diversity and evolution.
Schneider, C., Kudla, G., Wlotzka, W., Tuck, A. & Tollervey, D. Transcriptome-wide analysis of exosome targets. Mol. Cell 48, 422–433 (2012).
Popp, M. W. & Maquat, L. E. Nonsense-mediated mRNA decay and cancer. Curr. Opin. Genet. Dev. 48, 44–50 (2018).
Wong, J. J. L. et al. Orchestrated intron retention regulates normal granulocyte differentiation. Cell 154, 583–595 (2013).
Kurosaki, T., Popp, M. W. & Maquat, L. E. Quality and quantity control of gene expression by nonsense-mediated mRNA decay. Nat. Rev. Mol. Cell Biol. 20, 406–420 (2019).
Machnicka, M. A. et al. MODOMICS: a database of RNA modification pathways — 2013 update. Nucleic Acids Res. 41, D262–D267 (2012).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Ranjan, N. & Leidel, S. A. The epitranscriptome in translation regulation: mRNA and tRNA modifications as the two sides of the same coin? FEBS Lett. 593, 1483–1493 (2019).
Zaccara, S., Ries, R. J. & Jaffrey, S. R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 20, 608–624 (2019).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Bentley, D. L. Coupling mRNA processing with transcription in time and space. Nat. Rev. Genet. 15, 163–175 (2014). This comprehensive review discusses the mechanisms and interdependencies of mRNA transcription and processing.
de Klerk, E. & ‘t Hoen, P. A. C. Alternative mRNA transcription, processing, and translation: insights from RNA sequencing. Trends Genet. 31, 128–139 (2015).
Garieri, M. et al. The effect of genetic variation on promoter usage and enhancer activity. Nat. Commun. 8, 1358 (2017).
Reyes, A. & Huber, W. Alternative start and termination sites of transcription drive most transcript isoform differences across human tissues. Nucleic Acids Res. 46, 582–592 (2018).
Proudfoot, N. J., Furger, A. & Dye, M. J. Integrating mRNA processing with transcription. Cell 108, 501–512 (2002).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Tress, M. L. et al. The implications of alternative splicing in the ENCODE protein complement. Proc. Natl Acad. Sci. USA 104, 5495–5500 (2007).
Shi, Y. Mechanistic insights into precursor messenger RNA splicing by the spliceosome. Nat. Rev. Mol. Cell Biol. 18, 655–670 (2017).
Lee, Y. & Rio, D. C. Mechanisms and regulation of alternative pre-mRNA splicing. Annu. Rev. Biochem. 84, 291–323 (2015).
Baralle, F. E. & Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Biol. 18, 437–451 (2017).
Costa, V., Aprile, M., Esposito, R. & Ciccodicola, A. RNA-seq and human complex diseases: recent accomplishments and future perspectives. Eur. J. Hum. Genet. 21, 134–142 (2013).
Pistoni, M., Ghigna, C. & Gabellini, D. Alternative splicing and muscular dystrophy. RNA Biol. 7, 441–452 (2010).
Chaneton, B. & Gottlieb, E. Rocking cell metabolism: revised functions of the key glycolytic regulator PKM2 in cancer. Trends Biochem. Sci. 37, 309–316 (2012).
Buljan, M. et al. Tissue-specific splicing of disordered segments that embed binding motifs rewires protein interaction networks. Mol. Cell 46, 871–883 (2012).
Yablonovitch, A. L., Deng, P., Jacobson, D. & Li, J. B. The evolution and adaptation of A-to-I RNA editing. PLoS Genet. 13, e1007064 (2017).
Picardi, E., D’Erchia, A. M., Lo Giudice, C. & Pesole, G. REDIportal: a comprehensive database of A-to-I RNA editing events in humans. Nucleic Acids Res. 45, D750–D757 (2017).
Maas, S., Kawahara, Y., Tamburro, K. M. & Nishikura, K. A-to-I RNA editing and human disease. RNA Biol. 3, 1–9 (2006).
Tang, S. J. et al. Cis- and trans-regulations of pre-mRNA splicing by RNA editing enzymes influence cancer development. Nat. Commun. 11, 799 (2020).
Zipeto, M. A., Jiang, Q., Melese, E. & Jamieson, C. H. M. RNA rewriting, recoding, and rewiring in human disease. Trends Mol. Med. 21, 549–559 (2015).
Benne, R. The long and the short of it. Nature 380, 391–392 (1996).
Tress, M. L., Abascal, F. & Valencia, A. Alternative splicing may not be the key to proteome complexity. Trends Biochem. Sci. 42, 98–110 (2017).
Wan, Y. & Larson, D. R. Splicing heterogeneity: separating signal from noise. Genome Biol. 19, 86 (2018).
Liu, Y., Beyer, A. & Aebersold, R. On the dependency of cellular protein levels on mRNA abundance. Cell 165, 535–550 (2016). This study discusses the relationship between mRNA abundance and protein levels.
Dikic, I. Proteasomal and autophagic degradation systems. Annu. Rev. Biochem. 86, 193–224 (2017).
Varshavsky, A. N-degron and C-degron pathways of protein degradation. Proc. Natl Acad. Sci. USA 116, 358–366 (2019).
Harper, J. W. & Bennett, E. J. Proteome complexity and the forces that drive proteome imbalance. Nature 537, 328–338 (2016). This review discusses the regulation and balancing of protein synthesis and degradation.
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513, 382–387 (2014).
Liu, Y. et al. Multi-omic measurements of heterogeneity in HeLa cells across laboratories. Nat. Biotechnol. 37, 314–322 (2019).
Beyer, A., Hollunder, J., Nasheuer, H.-P. & Wilhelm, T. Post-transcriptional expression regulation in the yeast Saccharomyces cerevisiae on a genomic scale. Mol. Cell. Proteomics 3, 1083–1092 (2004).
Malmström, J. et al. Proteome-wide cellular protein concentrations of the human pathogen Leptospira interrogans. Nature 460, 762–765 (2009).
Milo, R. What is the total number of protein molecules per cell volume? A call to rethink some published values. BioEssays 35, 1050–1055 (2013).
Brar, G. A. Beyond the triplet code: context cues transform translation. Cell 167, 1681–1692 (2016).
Kearse, M. G. & Wilusz, J. E. Non-AUG translation: a new start for protein synthesis in eukaryotes. Genes Dev. 31, 1717–1731 (2017).
Loftfield, R. B. & Vanderjagt, D. The frequency of errors in protein biosynthesis. Biochem. J. 128, 1353–1356 (1972).
Walsh, C. T., Garneau-Tsodikova, S. & Gatto, G. J. Jr. Protein posttranslational modifications: the chemistry of proteome diversifications. Angew. Chem. Int. Ed. 44, 7342–7372 (2005).
Wolan, D. W., Zorn, J. A., Gray, D. C. & Wells, J. A. Small-molecule activators of a proenzyme. Science 326, 853–858 (2009).
Shalini, S., Dorstyn, L., Dawar, S. & Kumar, S. Old, new and emerging functions of caspases. Cell Death Differ. 22, 526–539 (2015).
Puente, X. S., Sánchez, L. M., Overall, C. M. & López-Otín, C. Human and mouse proteases: a comparative genomic approach. Nat. Rev. Genet. 4, 544–558 (2003).
Khoury, G. A., Baliban, R. C. & Floudas, C. A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the Swiss-Prot database. Sci. Rep. 1, 90 (2011).
Aebersold, R. et al. How many human proteoforms are there? Nat. Chem. Biol. 14, 206–214 (2018). This study presents the current estimation of how many proteoforms are expressed in humans.
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Blume-Jensen, P. & Hunter, T. Oncogenic kinase signalling. Nature 411, 355–365 (2001).
Wu, Z., Huang, R. & Yuan, L. Crosstalk of intracellular post-translational modifications in cancer. Arch. Biochem. Biophys. 676, 108138 (2019).
Kim, M.-S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Toby, T. K., Fornelli, L. & Kelleher, N. L. Progress in top-down proteomics and the analysis of proteoforms. Annu. Rev. Anal. Chem. 9, 499–519 (2016).
Tran, J. C. et al. Mapping intact protein isoforms in discovery mode using top-down proteomics. Nature 480, 254–258 (2011).
Anderson, L. C. et al. Identification and characterization of human proteoforms by top-down LC-21 Tesla FT-ICR mass spectrometry. J. Proteome Res. 16, 1087–1096 (2017).
Kelemen, O. et al. Function of alternative splicing. Gene 514, 1–30 (2013).
Liu, F., Lössl, P., Scheltema, R., Viner, R. & Heck, A. J. R. Optimized fragmentation schemes and data analysis strategies for proteome-wide cross-link identification. Nat. Commun. 8, 15473 (2017).
Götze, M., Iacobucci, C., Ihling, C. H. & Sinz, A. A simple cross-linking/mass spectrometry workflow for studying system-wide protein interactions. Anal. Chem. 91, 10236–10244 (2019).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Feng, Y. et al. Global analysis of protein structural changes in complex proteomes. Nat. Biotechnol. 32, 1036–1044 (2014).
Sluchanko, N. N. & Gusev, N. B. Moonlighting chaperone-like activity of the universal regulatory 14-3-3 proteins. FEBS J. 284, 1279–1295 (2017).
Pennington, K., Chan, T., Torres, M. & Andersen, J. The dynamic and stress-adaptive signaling hub of 14-3-3: emerging mechanisms of regulation and context-dependent protein–protein interactions. Oncogene 37, 5587–5604 (2018).
Huttlin, E. L. et al. The BioPlex network: a systematic exploration of the human interactome. Cell 162, 425–440 (2015).
Huttlin, E. L. et al. Architecture of the human interactome defines protein communities and disease networks. Nature 545, 505–509 (2017). This study describes the largest human PPI network generated by AP-MS to date.
Drew, K. et al. Integration of over 9,000 mass spectrometry experiments builds a global map of human protein complexes. Mol. Syst. Biol. 13, 932 (2017).
Ruepp, A. et al. CORUM: the comprehensive resource of mammalian protein complexes — 2009. Nucleic Acids Res. 38, D497–D501 (2009).
Giurgiu, M. et al. CORUM: the comprehensive resource of mammalian protein complexes — 2019. Nucleic Acids Res. 47, D559–D563 (2019).
Gaulton, A. et al. The Complex Portal — an encyclopaedia of macromolecular complexes. Nucleic Acids Res. 43, D479–D484 (2014).
Casanova, E. B. et al. Complex Portal 2018: extended content and enhanced visualization tools for macromolecular complexes. Nucleic Acids Res. 47, D550–D558 (2018).
Marsh, J. A. & Teichmann, S. A. Structure, dynamics, assembly, and evolution of protein complexes. Annu. Rev. Biochem. 84, 551–575 (2015).
Levy, E. D. & Pereira-Leal, J. B. Evolution and dynamics of protein interactions and networks. Curr. Opin. Struct. Biol. 18, 349–357 (2008).
Ellis, R. J. Molecular chaperones: assisting assembly in addition to folding. Trends Biochem. Sci. 31, 395–401 (2006).
Chen, S., Synowsky, S., Tinti, M. & MacKintosh, C. The capture of phosphoproteins by 14-3-3 proteins mediates actions of insulin. Trends Endocrinol. Metab. 22, 429–436 (2011).
Collins, B. C. et al. Quantifying protein interaction dynamics by SWATH mass spectrometry: application to the 14-3-3 system. Nat. Methods 10, 1246–1253 (2013).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Jangi, M. & Sharp, P. A. Building robust transcriptomes with master splicing factors. Cell 159, 487–498 (2014).
Großbach, J. et al. Integration of transcriptome, proteome and phosphoproteome data elucidates the genetic control of molecular networks. Preprint at https://doi.org/10.1101/703140 (2019). This recent study investigates quantitative trait loci on different molecular levels and how they mediate the effects of genomic variants in multilayered molecular networks.
Williams, E. G. et al. Systems proteomics of liver mitochondria function. Science 352, aad0189 (2016).
Wu, Y. et al. Multilayered genetic and omics dissection of mitochondrial activity in a mouse reference population. Cell 158, 1415–1430 (2014).
Picotti, P. et al. A complete mass-spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature 494, 266–270 (2013).
Hrdlickova, R., Toloue, M. & Tian, B. RNA-seq methods for transcriptome analysis. Wiley Interdiscip. Rev. RNA 8, e1364 (2017).
Levy, S. E. & Myers, R. M. Advancements in next-generation sequencing. Annu. Rev. Genomics Hum. Genet. 17, 95–115 (2016).
Reuter, J. A., Spacek, D. V. & Snyder, M. P. High-throughput sequencing technologies. Mol. Cell 58, 586–597 (2015).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Schaffer, L. V. et al. Identification and quantification of proteoforms by mass spectrometry. Proteomics 19, 1800361 (2019). This recent review discusses the current state of proteoform identification and quantification by top-down proteomics.
He, Z., Huang, T., Zhao, C. & Teng, B. in Modern Proteomics — Sample Preparation, Analysis and Practical Applications (eds Mirzaei, H. & Carrasco, M.) 237–242 (Springer, 2016).
Han, X., Aslanian, A. & Yates, J. R. Mass spectrometry for proteomics. Curr. Opin. Chem. Biol. 12, 483–490 (2008).
Bantscheff, M., Lemeer, S., Savitski, M. M. & Kuster, B. Quantitative mass spectrometry in proteomics: critical review update from 2007 to the present. Anal. Bioanal. Chem. 404, 939–965 (2012).
Gillet, L. C., Leitner, A. & Aebersold, R. Mass spectrometry applied to bottom-up proteomics: entering the high-throughput era for hypothesis testing. Annu. Rev. Anal. Chem. 9, 449–472 (2016). This in-depth review discusses current mass spectrometry techniques for bottom-up proteomics.
Bunt, G. & Wouters, F. S. FRET from single to multiplexed signaling events. Biophys. Rev. 9, 119–129 (2017).
Hu, C. D., Chinenov, Y. & Kerppola, T. K. Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol. Cell 9, 789–798 (2002).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Martell, J. D. et al. Engineered ascorbate peroxidase as a genetically encoded reporter for electron microscopy. Nat. Biotechnol. 30, 1143–1148 (2012).
Becher, I. et al. Pervasive protein thermal stability variation during the cell cycle. Cell 173, 1495–1507.e18 (2018).
Piazza, I. et al. A map of protein–metabolite interactions reveals principles of chemical communication. Cell 172, 358–372.e23 (2018).
Rolland, T. et al. A proteome-scale map of the human interactome network. Cell 159, 1212–1226 (2014).
Braun, P. et al. An experimentally derived confidence score for binary protein–protein interactions. Nat. Methods 6, 91–97 (2009).
Hein, M. Y. et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163, 712–723 (2015).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification–mass spectrometry data. Nat. Methods 10, 730–736 (2013).
Trinkle-Mulcahy, L. Recent advances in proximity-based labeling methods for interactome mapping. F1000Res. 8, 135 (2019).
Gingras, A. C., Abe, K. T. & Raught, B. Getting to know the neighborhood: using proximity-dependent biotinylation to characterize protein complexes and map organelles. Curr. Opin. Chem. Biol. 48, 44–54 (2019). This excellent review discusses the current state of proximity labelling techniques to analyse protein complexes.
Gupta, G. D. et al. A dynamic protein interaction landscape of the human centrosome–cilium interface. Cell 163, 1484–1499 (2015).
Liu, X. et al. An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations. Nat. Commun. 9, 1188 (2018).
Liu, X., Yang, W., Gao, Q. & Regnier, F. Toward chromatographic analysis of interacting protein networks. J. Chromatogr. A 1178, 24–32 (2008).
Dong, M. et al. A “tagless” strategy for identification of stable protein complexes genome-wide by multidimensional orthogonal chromatographic separation and iTRAQ reagent tracking. J. Proteome Res. 7, 1836–1849 (2008).
Kristensen, A. R., Gsponer, J. & Foster, L. J. A high-throughput approach for measuring temporal changes in the interactome. Nat. Methods 9, 907–909 (2012).
Kristensen, A. R. & Foster, L. J. in Stable Isotope Labeling by Amino Acids in Cell Culture (SILAC) (ed. Warscheid, B.) 263–270 (Humana Press, 2014).
Havugimana, P. C. et al. A census of human soluble protein complexes. Cell 150, 1068–1081 (2012).
Wan, C. et al. Panorama of ancient metazoan macromolecular complexes. Nature 525, 339–344 (2015).
Salas, D., Stacey, R. G., Akinlaja, M. & Foster, L. J. Next-generation interactomics: considerations for the use of co-elution to measure protein interaction networks. Mol. Cell. Proteomics 19, 1–10 (2020). This recent review discusses the techniques, limitations and possibilities of co-fractionation mass spectrometry approaches for PPI and protein complex mapping.
Scott, N. E., Brown, L. M., Kristensen, A. R. & Foster, L. J. Development of a computational framework for the analysis of protein correlation profiling and spatial proteomics experiments. J. Proteomics 118, 112–129 (2015).
Hu, L. Z. et al. EPIC: software toolkit for elution profile-based inference of protein complexes. Nat. Methods 16, 737–742 (2019).
Heusel, M. et al. Complex-centric proteome profiling by SEC-SWATH-MS. Mol. Syst. Biol. 15, e8438 (2019).
Heusel, M. et al. A global screen for assembly state changes of the mitotic proteome by SEC-SWATH-MS. Cell Syst. 10, 133–155.e6 (2020).
Varjosalo, M. et al. The protein interaction landscape of the human CMGC kinase group. Cell Rep. 3, 1306–1320 (2013).
Scott, N. E. et al. Interactome disassembly during apoptosis occurs independent of caspase cleavage. Mol. Syst. Biol. 13, 906 (2017).
Ni, J. Z. et al. Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay. Genes Dev. 21, 708–718 (2007).
Acknowledgements
The authors thank all members of the Aebersold laboratory who provided input to the content and presentation of the Review. The authors further thank B. Collins, L. Sieverling, J. Čuklina, C. Dörig, U. Kutay and M. Claassen for reading and providing feedback on different parts of the Review. R.A. acknowledges funding support from the SystemsX.ch projects PhosphoNetX PPM and TbX, as well as from the European Research Council (ERC-20140AdG 670821). I.B. acknowledges funding support from the Swiss National Science Foundation (31003A_166435).
Author information
Authors and Affiliations
Contributions
I.B. researched data for the review. I.B. and R.A. wrote and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks Ivan Marazzi and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Complex Portal: https://www.ebi.ac.uk/complexportal/home
CORUM: https://mips.helmholtz-muenchen.de/corum/
GENCODE database: https://www.gencodegenes.org/human/stats.html
Supplementary information
Glossary
- Interactome
-
The whole set of physical interactions between molecules in a cell, here specifically referring to protein–protein interactions.
- Proteoform
-
A protein species with a unique combination of amino acid sequence and post-translational modifications. Multiple alternative proteoforms can originate from the same gene locus.
- Isoforms
-
Alternative mRNA transcripts originating from the same gene locus.
- Epitranscriptome
-
The whole set of biochemical modifications of RNAs in a cell, with the methylation of adenosine being the most prominent modification.
- Alternative polyadenylation
-
Deviations from the standard process of polyadenylation, including the usage of different poly(A) start sites or variable length of the transcript’s 3′ untranslated region.
- Synonymous substitutions
-
DNA base substitutions in an exon of a protein-coding gene that do not cause a change in the translated protein sequence.
- N-degron pathways
-
A set of proteolytic systems that can recognize proteins containing N-degrons, thereby causing the degradation of these proteins (formerly ‘N-end rule pathways’). N-degrons are degradation signals in the amino-terminal region of a protein, which relate its in vivo half-life to the identity of its amino-terminal composition.
- C-degron pathways
-
A set of proteolytic systems that can recognize proteins containing C-degrons, thereby causing the degradation of these proteins. C-degrons are degradation signals in the carboxy-terminal region of a protein, which relate its in vivo half-life to the identity of its carboxy-terminal composition.
- SUMOylation
-
The post-translational modification of a protein by covalently attached small ubiquitin-like modifier (SUMO) proteins.
- Alternative splicing-coupled nonsense-mediated mRNA decay
-
A regulatory mechanism in which alternative splicing events introduce premature termination codons that lead to the downregulation of the transcript through the process of nonsense-mediated mRNA decay.
- Quantitative trait loci
-
DNA loci that correlate with the variation of a quantitative trait in the phenotype.
- Yeast two-hybrid screens
-
(Y2H screens). A strategy to identify protein–protein interactions in which one target protein is fused with a DNA-binding domain and a second target protein is fused with a respective transcriptional activation domain. When co-expressed in yeast, if the two target proteins interact in the nucleus, they initiate the transcription of a reporter gene.
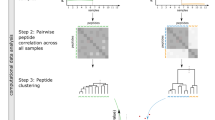
- Co-fractionation coupled to mass spectrometry
-
(CoFrac-MS). A strategy for the global mapping of protein–protein interactions and protein complexes, involving the native extraction and subsequent separation of protein complexes according to their physicochemical properties, followed by mass spectrometry analysis.
- False discovery rates
-
A metric used for the control of the error rate in experiments affected by the multiple testing problem.
- Graph partitioning
-
The process of dividing a graph (for example, a protein–protein interaction network) into smaller connected subnetworks.
Rights and permissions
About this article
Cite this article
Bludau, I., Aebersold, R. Proteomic and interactomic insights into the molecular basis of cell functional diversity. Nat Rev Mol Cell Biol 21, 327–340 (2020). https://doi.org/10.1038/s41580-020-0231-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-020-0231-2
This article is cited by
-
Spatiotemporal and direct capturing global substrates of lysine-modifying enzymes in living cells
Nature Communications (2024)
-
AI-guided pipeline for protein–protein interaction drug discovery identifies a SARS-CoV-2 inhibitor
Molecular Systems Biology (2024)
-
Integration of omics analyses into GMO risk assessment in Europe: a case study from soybean field trials
Environmental Sciences Europe (2023)
-
Analysis of RAS and drug induced homo- and heterodimerization of RAF and KSR1 proteins in living cells using split Nanoluc luciferase
Cell Communication and Signaling (2023)
-
Structural and functional complexity of HSP90 in cellular homeostasis and disease
Nature Reviews Molecular Cell Biology (2023)