Abstract
Evidence accumulated over the past decade shows that long non-coding RNAs (lncRNAs) are widely expressed and have key roles in gene regulation. Recent studies have begun to unravel how the biogenesis of lncRNAs is distinct from that of mRNAs and is linked with their specific subcellular localizations and functions. Depending on their localization and their specific interactions with DNA, RNA and proteins, lncRNAs can modulate chromatin function, regulate the assembly and function of membraneless nuclear bodies, alter the stability and translation of cytoplasmic mRNAs and interfere with signalling pathways. Many of these functions ultimately affect gene expression in diverse biological and physiopathological contexts, such as in neuronal disorders, immune responses and cancer. Tissue-specific and condition-specific expression patterns suggest that lncRNAs are potential biomarkers and provide a rationale to target them clinically. In this Review, we discuss the mechanisms of lncRNA biogenesis, localization and functions in transcriptional, post-transcriptional and other modes of gene regulation, and their potential therapeutic applications.
Similar content being viewed by others
Introduction
Genomes are extensively transcribed and give rise to thousands of long non-coding RNAs (lncRNAs), which are defined as RNAs longer than 200 nucleotides that are not translated into functional proteins. This broad definition encompasses a large and highly heterogeneous collection of transcripts that differ in their biogenesis and genomic origin. Statistics from Human GENCODE suggest that the human genome contains more than 16,000 lncRNA genes, but other estimates exceed 100,000 human lncRNAs1,2. These mainly include lncRNAs transcribed by RNA polymerase II (Pol II), but also by other RNA polymerases; and lncRNAs from intergenic regions (lincRNAs) as well as sense or antisense transcripts that overlap with other genes. The resulting lncRNAs are often capped by 7-methyl guanosine (m7G) at their 5′ ends, polyadenylated at their 3′ ends and spliced similarly to mRNAs (Fig. 1a). It is worthwhile noting that enhancer and promoter regions are also transcribed into enhancer RNAs (eRNAs) and promoter upstream transcripts, respectively3.
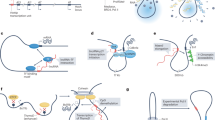
a | Biogenesis of long non-coding RNAs (lncRNAs). Unlike mRNAs, many RNA polymerase II (Pol II)-transcribed lncRNAs are inefficiently processed4,5,113 and are retained in the nucleus6,10,12,13,16 (mechanisms of lncRNA nuclear retention are shown in parts b–e), whereas others are spliced and exported to the cytoplasm. The lncRNAs (and mRNAs) that contain one or only few exons are exported to the cytoplasm by nuclear RNA export factor 1 (NXF1)23. b | Some lncRNAs are transcribed by dysregulated Pol II, remain on chromatin and, subsequently, are degraded by the nuclear exosome12. c | Numerous lncRNAs with a certain U1 small nuclear RNA (U1 snRNA) binding motif can recruit the U1 small nuclear ribonucleoprotein (U1 snRNP) and through it associate with Pol II at various loci13. d | In many lncRNAs, the sequence between the 3′ splice site and the branch point is longer and contains a shorter polypyrimidine tract (PPT) than in mRNAs10,17, which results in inefficient splicing. e | Sequence motifs in cis and factors in trans coordinately contribute to nuclear localization of lncRNAs. A nuclear retention element (NRE) U1 snRNA-binding site and C-rich motifs can recruit U1 snRNP19 and heterogeneous nuclear ribonucleoprotein K (hnRNPK)20,21, respectively, to enhance lncRNA nuclear localization. Other, differentially expressed RNA-binding proteins (RBPs), such as peptidylprolyl isomerase E (PPIE)6, inhibit splicing of groups of lncRNAs, resulting in their nuclear retention. f | In the cytoplasm, lncRNAs usually interact with diverse RBPs. g | Many lncRNAs in the cytoplasm are associated with ribosomes through ‘pseudo’ 5′ untranslated regions (UTRs); ribosome-associated lncRNAs tend to have short half-lives owing to unknown mechanisms24. h | Several lncRNAs are sorted into mitochondria by unknown mechanisms26,27. For example, the RNA component of mitochondrial RNA-processing endoribonuclease (RMRP) is recruited to mitochondria and is stabilized by binding G-rich RNA sequence-binding factor 1 (GRSF1)28. i | Some lncRNAs are also found in other organelles, such as exosomes29, probably by forming lncRNA–RBP complexes30,31. m7G, 7-methyl guanosine 5′ cap; (A)n, poly(A) 3′ tail.
The number of functional lncRNAs is still debated. Although evidence is still lacking to support the functionality of most lncRNAs, thereby rendering them transcription by-products, it is well documented that a growing number of lncRNAs have important cellular functions. The expression of a considerable number of lncRNAs is regulated and some have roles in different mechanisms of gene regulation. Several lncRNAs control the expression of nearby genes by affecting their transcription, and also affect other facets of chromatin biology, such as DNA replication or the response to DNA damage and repair. Other lncRNAs function away from their loci; their functions can be of a structural and/or regulatory nature and involve different stages of mRNA life, including splicing, turnover and translation, as well as signalling pathways. Consequently, lncRNAs affect several cellular functions that are of great physiological relevance, and alteration of their expression is inherent to numerous diseases. The specific expression patterns of these functional lncRNAs have the potential of being used as optimal disease biomarkers, and strategies are under development for their therapeutic targeting.
In this Review, we discuss emerging themes in lncRNA biology, including recent understanding of their biogenesis and their regulatory functions in cis and in trans at the transcriptional and post-transcriptional levels. We then discuss the pathological consequences of lncRNA dysregulation in neuronal disorders, haematopoiesis, immune responses and cancer. Finally, we discuss how the existing knowledge of lncRNAs allows the development of lncRNA-based therapeutic targeting.
Biogenesis of lncRNAs
Most lncRNA species are transcribed by Pol II. As such, many have 5′-end m7G caps and 3′-end poly(A) tails, and are presumed to be transcribed and processed similarly to mRNAs. However, recent studies have begun to reveal distinct transcription, processing, export and turnover of lncRNAs, which are closely linked with their cellular fates and functions.
Transcription and processing of lncRNAs
Compared with mRNAs, a greater proportion of lncRNAs are localized in the nucleus4,5,6, raising the fundamental question of what drives their differential localization. Dissection of the global features of lncRNAs and mRNAs suggests that lncRNA genes are less evolutionarily conserved, contain fewer exons and are less abundantly expressed6,7,8. Early studies indicated that lncRNA genes likely contain fewer exons than mRNAs6,7,8. The recently developed RNA capture long seq enabled better annotation of the full length of lncRNAs, including their 5′ ends9,10, revealing little length difference with mRNAs, although lncRNAs contain fewer and longer exons. Single-cell sequencing found that some lncRNAs can be abundantly expressed in the human neocortex11.
Whereas the low expression of lncRNAs is likely related to the presence of repressive histone modifications at their gene promoters9,10, their mode of transcription may partially explain some of their other distinctive features. The phosphorylation status of the Pol II carboxy-terminal domain corresponds with different transcription stages, and a significant fraction of lncRNAs are transcribed by phosphorylation-dysregulated Pol II12. Such lncRNAs appear to be weakly co-transcriptionally spliced and transcription termination at these genes is independent of polyadenylation signals, leading to temporal accumulation of lncRNAs on chromatin, followed by their rapid degradation by the RNA exosome12 (Fig. 1b). These findings provide insights into why lncRNAs are frequently nuclear, and suggest that functional lncRNAs must escape this nuclear surveillance process to accumulate at high levels in specific cell types. However, chromatin-tethered lncRNAs may not always be targeted by the nuclear surveillance process. Some chromatin-localized lncRNAs contain high levels of U1 small nuclear RNA binding sites, which recruit the U1 small nuclear ribonucleoprotein (U1 snRNP) to transcriptionally engaged Pol II, resulting in the tethering of numerous non-coding RNAs to chromatin13 (Fig. 1c). Accumulation of certain lncRNAs on chromatin can occur when the function of the Pol II-associated elongation factor SPT6 is abolished14,15. The loss of SPT6 generates redistribution of histone H3 trimethylated at Lys36 (H3K36me3; a mark of active transcription) from protein-coding genes to lncRNA genes, thereby increasing their transcription. Concomitantly, SPT6 loss impairs recruitment to chromatin of the transcription termination Integrator complex, leading to accumulation of long non-coding transcripts on chromatin in the form of DNA damage-associated R-loops15.
Overall, lncRNAs are spliced less efficiently than mRNAs6,10,16. They have weaker internal splicing signals and longer distances between the 3′ splice site and the branch point10,17, which correlate with augmented nuclear retention6,10,16 (Fig. 1d). Other factors, such as differential expression of certain splicing regulators, also contribute to the accumulation of lncRNAs in the nucleus. For example, in mouse embryonic stem cells (mESCs), the highly expressed splicing inhibitor peptidylprolyl isomerase E suppresses splicing of a subset of lncRNAs, leading to significant nuclear accumulation of many lncRNAs in mESCs6 (Fig. 1e). Alternative polyadenylation signals within lncRNAs may also modulate their subcellular localization. For example, the CCAT1 lncRNA gene (full names of all lncRNAs are provided in the footnote of Table 1) produces two isoforms: the long isoform (CCAT1-L) is nuclear and contains an internal polyadenylation site corresponding with the 3′ end of the short isoform (CCAT1-S), which is cytoplasmic18.
In addition to these general features of lncRNA transcription and processing, lncRNAs often contain embedded sequence motifs that can recruit certain nuclear factors, which promote the nuclear localization and function of the lncRNA (Fig. 1e). For example, the lncRNA maternally expressed gene 3 (MEG3) contains a 356-nucleotide nuclear retention element that associates with U1 snRNP, which in turn retains MEG3 in the nucleus19. Repeat elements also likely have roles in driving lncRNA nuclear retention. Recent studies using the high-throughput massively parallel RNA assays (MPRNA) have uncovered a C-rich sequence derived from Alu repeats that can promote the nuclear retention of lncRNAs through their association with the nuclear matrix protein heterogeneous nuclear ribonucleoprotein K (hnRNPK)20,21 (Fig. 1e). Other repeats can also guide lncRNA nuclear localization. For instance, the lncRNA functional intergenic repeating RNA element (FIRRE) contains many unique repeats, ranging in length from 67 to 804 bp, termed repeating RNA domains (RRDs), which establish FIRRE chromatin localization by interacting with hnRNPU22.
In summary, the nuclear localization and fate of lncRNAs are coordinately regulated at multiple layers, from transcription and processing to nuclear export through multiple sequence motifs in cis and factors in trans. In addition to being tethered to chromatin, some nuclear retained functional lncRNAs are specifically localized to membraneless nuclear domains (see below). Although the most representative lncRNAs of this type are processed by unusual biogenesis pathways (reviewed in ref.3), the molecular mechanism that traps such lncRNAs in specific nuclear domains still remains largely unknown. Nonetheless, given the diverse formats, sizes and functions of lncRNAs (Table 1), more work is warranted to dissect the distinctions and commonalities of mechanisms that control different nuclear localization patterns of lncRNAs.
Export of lncRNAs to the cytosol
A large fraction of lncRNAs are exported to the cytosol; these lncRNAs presumably share the same processing and export pathways with mRNAs. Indeed, a recent study revealed that long and A/U-rich transcripts with one or only few exons are dependent on the nuclear RNA export factor 1 (NXF1) pathway for export23. As lncRNAs tend to have fewer exons compared with mRNAs10, they preferentially exploit this export pathway. Upon arrival in the cytoplasm, lncRNAs likely undergo specific sorting processes that assign different lncRNAs to specific organelles or are distributed in the cytoplasm and associate with diverse RNA-binding proteins (RBPs) (Fig. 1f). It is estimated that half the pools of 70% of cytoplasmic lncRNAs are found in polysome fractions24. Certain cis elements contribute to the localization of lncRNAs with ribosomes, such as long ‘pseudo’ 5′ untranslated regions, so called because they precede ‘pseudo-open reading frames’ in the lncRNAs24,25 (Fig. 1g). The degradation of ribosome-associated lncRNAs may be triggered by a translation-dependent mechanism24. Whether the ribosome-associated lncRNAs are engaged by ribosomes for translation, have roles in translation or inertly reside in ribosomes is unknown.
Analyses of human mitochondrial transcriptomes revealed that lncRNAs exported from the nucleus can be sorted into mitochondria26,27. The RNA component of mitochondrial RNA-processing endoribonuclease (RMRP) is associated with the RBP HuR in the nucleus and exported to the cytosol by exportin 1. As soon as RMRP arrives at mitochondria, it is bound and stabilized by G-rich RNA sequence-binding factor 1 (GRSF1), thereby allowing its accumulation at the mitochondrial matrix28 (Fig. 1h). RNA sequencing of human blood exosomes29 revealed that they include many lncRNAs. It is still unknown how lncRNAs are sorted into exosomes, but the mechanism likely involves the binding of specific sequence motifs by RBPs30,31 (Fig. 1i). Considering the growing list of cytoplasmic lncRNAs with important roles in modulating mRNA stability, translation and signalling pathways (see below), it will be important to examine how each functional lncRNA is being escorted to its site of function. Our current understanding of this aspect of lncRNA biology is still very limited.
Gene regulation by lncRNAs
Gene expression is regulated by lncRNAs at multiple levels. By interacting with DNA, RNA and proteins, lncRNAs can modulate chromatin structure and function and the transcription of neighbouring and distant genes, and affect RNA splicing, stability and translation. Furthermore, lncRNAs are involved in the formation and regulation of organelles and nuclear condensates.
Chromatin regulation
The detection of RNA–chromatin association in a genome-wide fashion32,33,34,35, combined with chromatin conformation capture techniques, has unveiled complex lncRNA regulation of chromatin architecture and gene expression36,37. Although these lncRNA-mediated regulatory mechanisms should be explored individually, RNA has inherent regulatory potential. The negative charge of RNA can neutralize the positively charged histone tails, leading to chromatin de-compaction38, so RNA-mediated opening and closing of chromatin might function as a rapid switch of gene expression. Mechanistically, both cis-acting and trans-acting nuclear lncRNAs establish interactions with DNA to alter the chromatin environment, sometimes indirectly by virtue of their affinity for proteins that can associate with both RNA and DNA, and in other cases by binding DNA in a sequence-specific manner.
Protein–lncRNA localization and function on chromatin
Numerous lncRNAs localize on chromatin, where they can interact with proteins, facilitating or inhibiting their binding and activity at targeted DNA regions (Fig. 2a,b). Moreover, protein-assisted long-range chromatin interactions, such as CCCTC-binding factor (CTCF)-mediated chromatin interactions, can also act as facilitators of direct lncRNA transcriptional effects on target genes18,39,40. Although the binding of lncRNA to chromatin factors has raised considerable interest, caution is advised when evaluating such interactions, and rigorous methodologies should be applied in these studies. Moreover, the expression levels of a given lncRNA in relation to the factors it interacts with can define the extent of the effects that lncRNAs exert on the targeted chromatin41 (Supplementary Box 1).
a | Long non-coding RNAs (lncRNAs) can interact with chromatin modifiers and recruit them to target-gene promoters in order to activate or suppress their transcription in cis45,91,92, or in trans at distant, often multiple, loci46. For example, HOXA transcript at the distal tip (HOTTIP)61,62 acts in cis at the 5′ genes of the HOXA gene cluster, with which it interacts through chromatin looping. HOTTIP interacts with WD repeat-containing protein 5 (WDR5), thereby targeting the complex WDR5–myeloid/lymphoid or mixed-lineage leukaemia (MLL) to the promoters of the HOXA genes and promoting histone H3 Lys4 trimethylation (H3K4me3). b | lncRNAs can act as decoys of specific chromatin modifiers by sequestering them from the promoters of target genes. For example, p53-regulated and embryonic stem cell-specific lncRNA (lncPRESS1)63 supports the pluripotency of human embryonic stem cells by sequestering the histone deacetylase sirtuin 6 (SIRT6) from the promoters of numerous pluripotency genes. In this manner, lncPRESS keeps the active-gene H3 acetylated at Lys56 (H3K56ac) and Lys9 (H3K9ac) modifications as its target genes, thereby preventing the switch to activation of differentiation genes. During p53-mediated differentiation or following depletion of lncPRESS1, SIRT6 localizes to the chromatin and ensures the maintenance of pluripotency. c | lncRNAs can interact with DNA and co-transcriptionally form RNA–DNA hybrids such as R-loops, which are recognized by chromatin modifiers that activate or inhibit target-gene transcription43,77 or by transcription factors78. The lncRNA TCF21 antisense RNA inducing demethylation (TARID)79 forms an R-loop upstream of the promoter of its target gene transcription factor 21 (TCF21). The R-loop is recognized by growth arrest and DNA damage inducible-α (GADD45A), which drives the demethylation of the TCF21 promoter DNA by interacting with thymine–DNA glycosylase (TDG) and ten–eleven translocation 1 (TET1). R-loops can also form in trans, with similar possible outcomes. For example, auxin-regulated promoter loop (APOLO)80 is responsible for the activation of auxin responsive genes in Arabidopsis thaliana. APOLO and auxin target genes are normally silenced by H3K27me3 and the presence of chromatin loops maintained by the Polycomb factor like heterochromatin protein 1 (LHP1). Following transcriptional activation of APOLO in response to auxin, the lncRNA recognizes specific motifs at the promoters of its target genes, where it binds and generates R-loops that act as decoys of LHP1, thereby allowing target-gene expression80. Pol II, RNA polymerase II.
Polycomb repressive complex 2 (PRC2) binding and spreading across targeted chromatin has been particularly described to be facilitated by several lncRNAs, in some cases through well-characterized sequence elements42,43,44. This type of interaction can occur in cis and in trans, as is the case of the lncRNA ANRIL, which mediates PRC1 and PRC2 recruitment to the promoters of its neighbouring CDKN2A and CDKN2B genes, thereby controlling their expression and regulating cell senescence45. Furthermore, ANRIL can also act in trans through Alu sequences, which drive ANRIL recruitment of PRC1 and PRC2 proteins to distant targets46. Although PRC2 requires RNA to efficiently bind to chromatin47, the role of lncRNAs in PRC2 chromatin targeting is still under debate given the low specificity of RNA binding by PRC2 (ref.48). One example of many is the controversy over the trans-acting lncRNA HOX transcript antisense RNA (HOTAIR) in recruiting chromatin-modifying complexes to repress the distal HOXD genes, which has been described in detail elsewhere49,50,51.
Other factors are likely involved in regulating lncRNA-mediated PRC targeting. For example, extensive studies performed in mice have shown that hnRNPK and other chromatin-associated factors interact with X-inactive specific transcript (Xist) and other lncRNAs at imprinted genomic loci, such as Kcnq1ot1 and antisense of IGF2R non-protein coding RNA (Airn) to promote the spread of Polycomb complexes across different chromatin domains41,52,53,54,55,56,57. Transcription factors, such as the ubiquitously expressed YY1, are also able to target lncRNA-bound chromatin modifiers and other nascent RNAs to specific genomic loci46,52,58,59.
In addition to gene-silencing factors, lncRNAs can recruit chromatin modifiers that promote gene activation60,61. The lncRNA HOTTIP is one of several lncRNAs that regulate the HOXA gene cluster (see below) — it binds several HOXA genes at the 5′ region of the cluster through chromatin looping, and its expression contributes to the maintenance of chromatin organization in this region. HOTTIP drives the WDR5–myeloid/lymphoid or mixed-lineage leukaemia (MLL; also known as KMT2A) histone methyltransferase complex to gene promoters, thereby facilitating gene expression through H3K4me3 and acting as an important regulator of mouse haematopoietic stem cells61,62 (Fig. 2a). Finally, instead of recruiting chromatin modifiers, lncRNAs may function as decoys. The p53-regulated lncPRESS1 is a pluripotency-associated lncRNA acting as decoy for the deacetylase sirtuin 6, which represses several pluripotency genes and, thus, promotes differentiation. In human ESCs (hESCs), lncPRESS interacts with sirtuin 6 and sequesters it from chromatin, thereby maintaining the transcription-permissive H3 acetylated at Lys56 (H3K56ac) and H3K9ac at pluripotency-related genes63 (Fig. 2b).
Direct interactions between lncRNAs and DNA
An essential feature of lncRNAs is their potential to generate hybrid structures with DNA to influence chromatin accessibility. Such interactions can take the form of triple helices (triplexes) or R-loops. The actual prevalence of both of these types of structures is still unknown owing to the difficulty of detecting them in vivo. Nevertheless, the formation of triplexes and R-loops is probably widespread and essential for the regulatory activity of many lncRNAs.
RNA–DNA–DNA triplexes have been proposed as an example of non-coding RNA–DNA interplay in mediating gene silencing64,65,66 or activation66,67,68. The potential to form triplexes relies primarily on the RNA sequence69,70,71. Recently, TrIP-seq (targeted RNA immunoprecipitation sequencing) has been developed to study triplex-forming sequences in vivo72. An example of triplex-mediated gene regulation links the function of a lncRNA with an eRNA in the activation of a neighbouring proto-oncogene, sphingosine kinase 1 (SPHK1). In response to cell proliferation signals, the lncRNA KHPS1 (SPHK1 gene antisense) forms a triple helix upstream of the SPHK1 enhancer, which helps recruit chromatin modifiers that activate the transcription of eRNA-SPHK1 and promote the expression of SPHK1 (refs71,73). Remarkably, the role of the triplex in driving gene regulation was further shown by exchanging the KHPS1 triplex-forming region with the MEG3 triplex-forming region, causing KHPS1 to switch its specificity to the MEG3 target gene71.
A more extensively studied mode of lncRNA interaction with chromatin occurs at R-loops, which have long been considered a threat to genome stability. However, the transient nature of R-loops makes them ideal regulatory hubs, and recent findings argue for their re-evaluation as regulators of gene expression74,75,76 and as coordinators of DNA repair (Box 1). Several lncRNAs regulate gene expression in the context of R-loops, with the aid of proteins that recognize these structures, causing a wide spectrum of outcomes77,78 (Fig. 2a). In mESCs, the lncRNA TARID generates an R-loop at the CpG-rich promoter of the gene TCF21, which is transcribed in the opposite direction. GADD45A recognizes and binds to the R-loop at the TCF21 promoter, recruiting the DNA demethylating factor TET1, leading to transcriptional activation of TCF21 (ref.79) (Fig. 2c). Although many R-loop-forming lncRNAs act in cis, these R-loops can also be produced in trans to regulate the expression of protein-coding genes. For example, the lncRNA APOLO is able to form R-loops in trans in Arabidopsis thaliana as part of a widespread regulation of auxin responsive genes80 (Fig. 2c).
Transcription regulation
The relative position between a lncRNA and its neighbouring genes is a key determinant of their regulatory relationship. As widespread antisense and bidirectional lncRNA transcription was found to be evolutionarily conserved81, the non-random genomic distribution of lncRNAs could represent an evolutionary adaptation of genes to regulating their own expression in a context-specific manner. For instance, the genomic arrangement of divergent lncRNAs is key for gene regulation in cis82. This regulation can be mediated by two main, non-mutually exclusive mechanisms: the lncRNA transcript can regulate neighbouring loci, and/or the act of transcription or splicing of the lncRNA can generate a chromatin state or steric impediment that influences the expression of nearby genes. Thus, interpretation of several orthogonal loss-of-function and gain-of-function experiments is required to discern these possible modes of lncRNA functionality83.
Gene silencing by lncRNAs
The best-known mechanisms of gene repression mediated by lncRNAs are related to gene-dosage compensation. The main representative of this functionality is the lncRNA XIST, which is responsible for X chromosome inactivation in cells of female mammals. During embryonic development, XIST molecules spread over one of the two X chromosomes and cause the silencing of a large proportion of its genes84. XIST is able to silence large chromosomal regions even when it is ectopically expressed from a different chromosome85. A complex interplay of protein interactors contributes to XIST-mediated gene silencing55,56,57,58. In addition, a study in mESCs has revealed that the rapid coating of the X chromosome by Xist depends on the ability of the lncRNA to exploit the 3D chromatin organization, which allows it to spread from sites that are spatially proximal to its locus to distant loci, while it modifies the target chromatin structure through its interactions with chromatin modifiers86. This confers XIST a role in shaping the 3D architecture of the inactivated X chromosome, a process that, once initiated, has been shown to persist even in the absence of XIST, thereby defining the role of the lncRNA as an initiator of epigenetic memory, which is maintained during the later phases of X chromosome inactivation by the protein complexes recruited by XIST to chromatin56,87,88.
At other loci, cis-acting lncRNAs can promote an inactive chromatin state by directly or indirectly interacting with chromatin in proximity to their site of transcription. For instance, the R-loop formed in cis by the lncRNA ANRASSF1 directs PRCs to their targets to regulate gene expression43,89,90. In A. thaliana, the lncRNA COOLAIR, which is environmentally induced by cold at the FLOWERING locus, lingers at its site of transcription and coats the locus to promote PRC2-dependent H3K27me3 (refs91,92).
lncRNAs can suppress gene expression by interfering with the transcription machinery, which leads to alteration of the recruitment of transcription factors or Pol II at the inhibited promoter53, alteration of histone modifications53,93 and reduction of chromatin accessibility94,95. An example of this group of regulators is the mouse imprinted Airn lncRNA, which determines the onset of allele-specific expression during mESC differentiation53. Airn transcription from the paternal allele causes the displacement of Pol II from the overlapping Igf2r promoter, resulting in transcription pausing and gene silencing53,96,97 (Fig. 3a). Another mechanism by which a lncRNA can regulate widespread transcriptional interference is represented by the conserved lncRNA CHD2 adjacent, suppressive regulatory RNA (Chaserr)95, which is located upstream of the chromatin remodeller Chd2 gene. Chaserr depletion increased accessibility at the Chd2 promoter, as well as at several other promoters, which are all regulated by CHD2. The allele specificity of Chaserr towards Chd2 in Chaserr-mutated mouse models confirmed Chaserr functions strictly in cis. Interestingly, CHD2 binds nascent RNAs, including Chaserr, and promotes their expression. The reciprocal regulation of CHD2 and Chaserr represents a regulatory feedback loop, in which CHD2 regulates its own expression using Chaserr as a sensor of CHD2 levels95.
a | Long non-coding RNAs (lncRNAs) can inhibit gene expression in a transcript-dependent and/or in a transcription-dependent (that is, transcript-independent) manner. In mouse extra-embryonal tissues, antisense of IGF2R non-protein coding RNA (Airn) functions in trans as it is guided through a specific 3D chromosome conformation (not shown) to the promoters of two distal imprinted target genes, solute carrier family 22 member 2 (Slc22a2) and Slc22a3. Once there, Airn recruits Polycomb repressive complex 2 (PRC2), which catalyses histone H3 Lys27 trimethylation (H3K27me3) and gene silencing. Airn also functions in cis, on its overlapping protein-coding gene insulin-like growth factor 2 receptor (Igf2r). Airn transcription causes steric hindrance for RNA polymerase II (Pol II) at the transcription start site of Igfr2r, which is followed by promoter methylation (not shown) and Igfr2r silencing53,96,97. b | lncRNAs and enhancer RNAs (eRNAs) can promote the expression of protein coding genes (PCGs) that are in close proximity to their enhancers through preformed chromatin loops (for example, the eRNA P53BER (p53-bound enhancer region)266 and the enhancer-associated lncRNA (elncRNA) SWINGN (SWI/SNF interacting GAS6 enhancer non-coding RNA)113), thereby allowing recruitment of chromatin-activating complexes to the promoters of the PCGs. c | An important feature of some eRNAs and elncRNAs is their ability to regulate distant genes by directly promoting chromatin looping through the recruitment of looping factors18,36,139,267. For example, following oestrogen receptor (ER) transcription activation, the NRIP1 enhancer (eNRIP) is bi-directionally transcribed into an eRNA, which recruits cohesin to form short-range (solid line) and long-range (dashed line) chromatin loops, thereby promoting contact between the NRIP1 enhancer and the promoters of NRIP1 and trefoil factor 1 (TFF1), two of the several genes activated in response to ER activation267. d | lncRNAs can activate gene expression in a transcript-independent manner. Transcription of Bend4-regulating effects not dependent on the RNA (Bendr) is sufficient to activate enhancer elements (e) embedded in its locus, which promotes the formation of an active chromatin state (marked by H3K4me3) at the promoter of the proximal gene BEN domain containing protein 4 (Bend4)116. e | Example of a complex regulatory unit formed by the lncRNAs Upperhand (Uph) and Handsdown (Hdn) in regulating the PCG heart and neural crest derivatives expressed 2 (Hand2). An enhancer embedded in Uph activates the transcription of the proximal Hand2 gene when the lncRNA gene is transcribed, without requiring chromatin reorganization118. By contrast, chromatin looping is necessary for Hdn function, as it puts its promoter in spatial proximity with Hand2-activating enhancers. When Hdn transcription is activated, the Hand2 enhancers become unavailable for Hand2 promoter activation, thereby inhibiting its expression. Removal of Hdn or reduction of its transcription leads to increased expression of Hand2. CTCF, CCCTC-binding factor; NRIP1, nuclear receptor interacting protein 1; TF, transcription factor.
lncRNAs transcribed at enhancers
Active enhancers can be transcribed into two major types of non-coding RNAs: eRNAs and enhancer-associated lncRNAs (elncRNAs). The main distinction between the two groups of transcripts is based on their features: eRNAs are relatively short, bidirectional capped transcripts, which are generally unspliced, non-polyadenylated and unstable98,99. By contrast, elncRNAs are mostly unidirectional, polyadenylated and spliced. The distinction between the two transcript types is not always clear-cut and they can be confused in the literature. Although the correlation between enhancer activity and eRNA expression is well established, whether the eRNA transcripts per se are functional is still under debate. Nevertheless, some eRNAs have been functionally linked with gene expression. In addition to functioning through pre-existing chromatin conformations (Fig. 3b), some eRNAs can facilitate or directly drive chromatin looping by interacting with scaffold proteins such as the Mediator or the structural maintenance of chromosomes complex cohesin. These interactions generate regulatory contacts between enhancers and promoters that can be located several megabases apart100,101,102 (Fig. 3c).
Some enhancer loci produce elncRNAs, the expression of which is related to that of their enhancer elements103,104,105,106. Notably, elncRNA splicing has been positively associated with the activity of their associated enhancers and correlated with the abundance of the neighbouring protein-coding genes107,108. Furthermore, elncRNAs can modulate chromatin structure and topology in cooperation with chromatin-regulating proteins40,109. Gene-activating mechanisms described for eRNA function can also define the functions of elncRNAs (Fig. 3b,c).
Gene activation by elncRNAs often results in complex phenotypes related to human diseases110,111,112. The lncRNA SWINGN is located at the boundary of a topologically associating domain that includes its target gene GAS6 (ref.113). SWINGN promotes the interaction between SWI/SNF chromatin remodelling complexes and the transcription start site of GAS6, but also with additional distant loci involved in malignant phenotypes, accounting for its pro-oncogenic role113. In addition, some lncRNAs are able to promote the formation of genomic domains comprising many inter-loci interactions, as shown for the lncRNAs ESR1 locus enhancing and activating non-coding RNAs (ELEANORs)114. Together with other similarly acting lncRNAs, these transcripts are examples of how transcription can regulate the formation of genomic compartments to drive gene expression36,112.
As described above, it should be considered that lncRNAs can activate gene expression in a transcript-independent fashion, adding complexity to the interpretation of their gene regulatory functions. For example, functional DNA elements embedded in lncRNA loci can activate the expression of neighbouring genes115,116,117. The lncRNA Bend4-regulating effects not dependent on the RNA (Bendr) regulates its neighbouring gene BEND4 in cis through the presence of enhancer elements in Bendr that are activated by its transcription. Deletion of the Bendr promoter, but not insertion of a premature poly(A) in the first exon of Bendr, suppressed occupancy of the BEND4 promoter by Pol II116 (Fig. 3d). Other lncRNAs have been identified with similar roles in activation of proximal enhancers116,117,118.
Regulatory networks involving cis-acting lncRNAs
It is becoming increasingly clear that regulation in cis by lncRNAs is not only determined by one-on-one effects of a lncRNA on a neighbouring gene. lncRNAs are part of complex regulatory units, in which the expression of a protein-coding gene may be regulated by the coordinated activity of two or more lncRNAs and of transcript-dependent and transcript-independent mechanisms. Several of these units act on essential developmental genes or on loci with important functions in maintaining the equilibrium between normal and hyperproliferative processes.
The heart and neural crest derivatives expressed 2 (Hand2) gene encodes a transcription factor essential for heart development, in which dosage imbalance can cause serious malformations119. Two lncRNA loci are found in the vicinity of Hand2, which regulate its expression through different mechanisms (Fig. 3e). Deletion in mice of either of these lncRNA genes leads to embryonic lethality. One of these lncRNAs, Upperhand is transcribed from a bidirectional promoter, divergently from the Hand2 promoter. A study that analysed the effects of Upperhand deletion on Hand2, by intercrossing Hand2 and Upperhand knockout heterozygous mice, revealed that Upperhand controls Hand2 transcription in cis118. Furthermore, the insertion of a polyadenylation signal downstream of the Upperhand transcription start site (thereby abolishing transcription) affected Hand2 expression, whereas Hand2 expression was unaffected by depletion of the mature Upperhand transcript, elegantly demonstrating that Upperhand controls Hand2 in cis in a transcription-dependent but transcript-independent fashion118. A separate study reports partially conflicting data on the outcomes of Upperhand deletion, obtained from three different knockout mouse models, in which the effects on Hand2 expression are more subtle120. However, in both studies, alteration of Upperhand expression resulted in strong cardiac abnormalities linked with Upperhand-mediated regulation of Hand2. Additional studies will be required to fully unveil the complex interaction between Hand2 and the lncRNAs regulating it (Fig. 3e).
The lncRNA Handsdown is located several kilobases downstream of Hand2, and inhibits Hand2 expression within a preformed chromatin loop mediated by CTCF121. The mechanism behind this regulation involves looping-mediated interaction between Handsdown promoter and regulatory elements upstream of the Hand2 gene in mouse embryonic cardiomyocytes, which thus become unavailable for Hand2 activation (Fig. 3e). Upperhand and Handsdown exemplify how lncRNAs can act in association to finely tune the expression of essential genes.
Another regulatory possibility is that the functions of the transcript and of the locus of a lncRNA are uncoupled and promote opposite outcomes. The locus of the lncRNA HOXA upstream non-coding transcript (Haunt) contains enhancers that activate the expression of HOXA genes. By contrast, the Haunt transcript acts as a decoy for the enhancers embedded in its own locus, thereby inhibiting the expression of HOXA genes122. These opposing outcomes have been implicated in preventing aberrant HOXA expression.
In conclusion, several interdependent factors emerge as crucial regulators of lncRNA function: the relative position of the lncRNA and target gene, the formation of co-transcriptional RNA–DNA and RNA–protein interactions, and whether the regulatory effect is mediated by the lncRNA transcript or by its transcription. The cell-specific co-occurrence of these factors determines the regulatory potential of individual lncRNAs.
Roles in scaffolding and condensates
Nuclear condensates are membraneless RNA–protein compartments involved in many cellular processes123. By virtue of their scaffolding or regulatory activities, several abundant lncRNAs are essential for the assembly and function of different nuclear condensates.
The lncRNA nuclear paraspeckle assembly transcript 1 (NEAT1) underlies the complex organization and functions of paraspeckles124,125 (Fig. 4a). The NEAT1 gene produces two isoforms that share a common 5′ end but have different 3′ ends: NEAT1 short, which has a poly(A) tail produced from an upstream polyadenylation signal; and NEAT1 long, which has a stable 3′ U–A·U triple-helix structure that is cleaved by RNase P126,127,128. NEAT1 long, but not NEAT1 short, is essential for the assembly of paraspeckles126,127,128. The middle region (8–16.6 kb) of NEAT1 long contains two subdomains (12–13 kb and 15.4–16.6 kb) responsible for recruiting the paraspeckle core proteins NONO and SFPQ to initiate the assembly of paraspeckles through liquid–liquid phase separation129 (Fig. 4a). How exactly NEAT1 long is assembled into the core, spherical shape of paraspeckles remains unclear. Future dissection of the key structural modules of NEAT1 long should be helpful in gaining mechanistic insights into NEAT1-scaffolded condensates. As an interesting note, however, global RNA structure mapping showed that NEAT1 long might not contain long-range intramolecular interactions and structure130,131
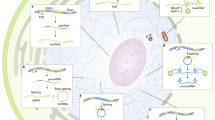
a | The long non-coding RNA (lncRNA) nuclear paraspeckle assembly transcript 1 (NEAT1) is essential for the formation of paraspeckles125. NEAT1 sequesters numerous paraspeckle proteins to form a highly organized core–shell (dark and light purple, respectively) spheroidal nuclear body124. The middle region of NEAT1 is localized in the centre of paraspeckles and the 3′ and 5′ regions are localized in the periphery124. Different paraspeckle proteins are embedded by NEAT1 into the spheroidal structure in the core region (non-POU domain containing octamer binding (NONO), fused in sarcoma (FUS) and splicing factor, proline- and glutamine-rich (SFPQ)) or the shell region (RNA binding motif protein 14 (RBM14))124. b | The lncRNA metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) is localized at the periphery of nuclear speckles126,132 and is involved in the regulation of pre-mRNA splicing116,134,135. At the periphery, MALAT1 interacts with the U1 small nuclear RNA (U1 snRNA)139, whereas proteins such as SON DNA and RNA binding protein (SON) and splicing component, 35 kDa (SC35) are localized at the centre of nuclear speckles138. c | 5′ small nucleolar RNA-capped and 3′-polyadenylated lncRNAs (SPAs)141 and small nucleolar RNA-related lncRNAs (sno-lncRNAs)140 accumulate at their transcription sites and interact with several splicing factors such as RNA binding protein fox-1 homologue 2 (RBFOX2), TAR DNA-binding protein 43 (TDP43) and heterogeneous nuclear ribonucleoprotein M (hnRNPM) to form a microscopically visible nuclear body that is involved in the regulation of alternative splicing141. d | The perinucleolar compartment contains the lncRNA pyrimidine-rich non-coding transcript (PNCTR), which sequesters pyrimidine tract-binding protein 1 (PTBP1) and, thus, suppresses PTPBP1-mediated pre-mRNA splicing elsewhere in the nucleoplasm142. e | Functional intergenic RNA repeat element (Firre) is transcribed from the mouse X chromosome and interacts with the nuclear matrix factor hnRNPU to tether chromosome (Chr) X, 2, 9, 15 and 17 into a nuclear domain151,153. The size of each type of nuclear body is indicated in parts a–d268.
The lncRNA metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) is perhaps the most abundant lncRNA in most cultured cells. It is specifically localized in nuclear speckles126,132, has important roles in pre-mRNA splicing and in transcription133,134,135, and is involved in cancer progression and metastasis136,137. Although MALAT1 interacts with many proteins, depletion of MALAT1 does not affect the formation of nuclear speckles but, rather, causes defects in their composition138. Each nuclear speckle is a multilayered compartment, in which nuclear speckle proteins such as the splicing factors SON and SC35 (also known as SRSF2) are localized at the centre and MALAT1 at the periphery138 (Fig. 4b). How this unique organization of MALAT1 facilitates the formation and function of nuclear speckles remains to be investigated. Unlike NEAT1, MALAT1 forms many long-range structures131, which are perhaps involved in its multivalent interactions with different RBPs and pre-mRNAs.
Application of the recently developed RNA in situ conformation sequencing (RIC-seq) revealed that MALAT1 functions as an RNA hub for many highly expressed RNAs. For example, a high-confidence NEAT1–RNA interaction analysis suggested that the 5′ region of NEAT1 interacts with MALAT1 in trans139. RIC-seq also revealed multiple interaction sites between U1 small nuclear RNA and MALAT1 (ref.139), which were also found using psoralen analysis of RNA interactions and structures131 (Fig. 4b). Given the peripheral localization of MALAT1 in nuclear speckles138, it will be interesting to understand how the RNA hub function of MALAT1 is achieved at the surface of nuclear speckles. These studies reveal a complicated regulatory network, which could be uncovered by further dissecting the structural modules of MALAT1 and its functions of scaffolding different RBPs.
The scaffolding nature of lncRNA-mediated gene regulation is also illustrated by small nucleolar RNA-related lncRNAs and 5′ small nucleolar RNA-capped and 3′ polyadenylated lncRNAs (SPAs), which are lncRNA species produced from the Prader–Willi syndrome (PWS; a neurodevelopmental disorder) minimal deletion of chromosome 15q11–13. Whereas induced pluripotent stem cells derived from individuals with PWS conspicuously lack these lncRNAs, they are abundantly expressed and accumulate in cis in normal hESCs, and sequester more than 1% of each tested splicing factor, including RBFOX2, TDP43 and hnRNPM140,141 (Fig. 4c). Importantly, hESCs lacking PWS-associated lncRNAs exhibited altered patterns of alternative splicing and protein binding to pre-mRNAs associated with neuronal functions141. Similarly, gene regulation by multivalent interactions between lncRNAs and RBPs was also found for PNCTR in the perinucleolar compartment142 (Fig. 4d). PNCTR is a short, tandem repeat-enriched RNA generated from the ribosomal DNA intergenic spacer; it is highly expressed in cancer cells and required for lung cancer cell survival. This lncRNA contains hundreds of polypyrimidine tract-binding protein 1 (PTBP1) binding motifs, and therefore sequesters PTBP1 to the perinucleolar compartment and suppresses its splicing activity elsewhere in the nucleoplasm142. Together, these studies show that multivalent binding between lncRNAs and RBPs is an effective mechanism of regulating disease-specific alternative splicing.
Nuclear stress bodies are another type of nuclear condensate. Their formation requires heat shock transcription factor 1 and transcription of the heterogenous lncRNAs highly repetitive satellite III (HSATIII) in conditions of heat and chemical stresses143. HSATIII lncRNAs accumulate at their transcription sites, sequester scaffold attachment factor B144, Serine and arginine-rich (SR) proteins145,146 and transcription factors147, and assemble them into nuclear stress bodies. HSATIII lncRNAs were proposed to promote intron retention at hundreds of mRNAs by modulating the phosphorylation of SR proteins148. Similar lncRNA-induced stress bodies were also found for intergenic spacer RNAs under heat shock and other stresses149,150.
In addition to functioning as scaffolds of proteins and RNAs at nuclear condensates, lncRNAs can bring different chromosomes into proximity in nuclear domains. FIRRE consists of numerous RNA variants transcribed from the X chromosome; it interacts with the nuclear matrix factor hnRNPU to maintain a nuclear domain through its scaffolding function151 (Fig. 4e). In mice, gene-expression changes caused by Firre deletion can be partially rescued by expressing transgenic Firre RNA, indicating that it has functions in trans152. Indeed, Firre is localized in proximity to its locus on the X chromosome, as well as on mouse chromosomes 2, 9, 15 and 17, and functions in trans as a chromosome scaffold lncRNA151,153 (Fig. 4e). It remains to be determined whether such lncRNA-anchored inter-chromosomal structures are phase separated.
Roles in post-transcriptional regulation
In addition to their roles in transcription regulation and nuclear organization, lncRNAs control several other aspects of gene expression, and some lncRNAs are even translated into functional peptides154. Nevertheless, as bona fide non-coding RNAs, lncRNAs mainly act through their ability to establish interactions with proteins and nucleic acids (Fig. 5). Here, we highlight a few of the many different modes of lncRNA function as post-transcriptional, translational and post-translational regulators.
A | trans-Acting long non-coding RNAs (lncRNAs) interact with RNA-binding proteins (RBPs) through sequence motifs or by forming unique structural motifs. Aa | Pyrimidine-rich non-coding transcript (PNCTR) sequesters pyrimidine tract-binding protein 1 (PTBP1) to the perinucleolar compartment (PNC) and, thus, suppresses PTBP1-mediated mRNA splicing elsewhere in the nucleoplasm142. Ab | In the cytosol, non-coding RNA activated by DNA damage (NORAD) sequesters Pumilio (PUM) RBPs, which repress the stability and translation of mRNAs to which they bind156,157,269. Ac | Human FOXD3 antisense transcript 1 (FAST) forms several structural modules that bind the E3 ligase β-transducin repeats-containing protein (β-TrCP), thereby blocking the degradation of its substrate β-catenin (β-cat), leading to activation of WNT signalling in human embryonic stem cells6. B | trans-Acting lncRNAs directly interact with RNAs through base pairing. Ba | Terminal differentiation-induced ncRNA (TINCR)160 or half-STAU1-binding site RNAs (1/2-sbsRNAs)161 promote or suppress mRNA stability, respectively, by forming intermolecular duplexes that bind Staufen homologue 1 (STAU1), the key protein of Staufen-mediated mRNA decay160,161,162. Bb | The SINEB2 repeat of mouse antisense to ubiquitin carboxyterminal hydrolase L1 (AS-Uchl1) complementarily binds the Uchl1 mRNA and promotes polysome association with Uchl1 and translation164. C | Some abundant lncRNAs affect gene expression by functioning as competitive endogenous RNAs (ce-RNAs)165,166. For example, lncRNA-PNUTS is generated by alternative splicing of the PNUTS pre-mRNA by heterogeneous nuclear ribonucleoprotein E1 (hnRNPE1)169. lncRNA-PNUTS contains seven miR-205 binding sites, which reduce the availability of miR-205 to bind and suppress the zinc finger E-box-binding homeobox 1 (ZEB1) and ZEB2 mRNAs169. GSK3, glycogen synthase kinase 3; miR, microRNA; P, phosphate group; PNUTS, phosphatase 1 nuclear targeting subunit; PRE, Pumilio response element.
Modes of direct lncRNA–protein interactions
lncRNAs are involved in post-transcriptional regulation by sequestering proteins through their binding to RNA sequence motifs or structures to form specific lncRNA–protein complexes (lncRNPs), resulting in altered mRNA splicing and turnover, and, in certain biological contexts, in the modulation of signalling pathways (Fig. 5A). Abundant lncRNAs, such as the aforementioned small nucleolar RNA-related lncRNAs140 and SPAs140,141 in the PWS region (Fig. 4c) and PNCTR142 (Fig. 4d), contain clusters of motifs that sequester different splicing factors — including UGCAU and GCAUG motifs, which are bound by RBFOX2 (ref.140), UG-rich sequences bound by TDP43 (refs140,141) and YUCUYY and YYUCUY motifs bound by PTBP1142 — thereby suppressing the splicing of pre-mRNAs containing the same motifs140,141,142 (Fig. 5A). Other mechanisms of lncRNA-mediated splicing regulation involve lncRNA modulating post-translational modifications of splicing factors, splicing repression through the formation of RNA–RNA hybrids with target pre-mRNAs and fine-tuning of target-gene splicing through chromatin remodelling (reviewed in ref.155). In the cytosol, non-coding RNA activated by DNA damage (NORAD) is highly expressed following DNA damage and maintains genomic stability by sequestering Pumilio proteins156,157. Pumilio proteins bind to a specific motif in mRNA 3′ untranslated regions and facilitates mRNA decay through deadenylation and decapping158. Each NORAD molecule contains 15 Pumilio binding motifs and, consequently, the ~500–1,000 copies of NORAD expressed in a single HCT116 cell can sequester ~7,500–15,000 Pumilio protein molecules, thereby sequestering most Pumilio from target mRNAs involved in maintaining genomic stability156,157 (Fig. 5A). It should be noted, however, that although NORAD offers a sufficient number of Pumilio binding sites to sequester the Pumilio proteins, the number may be a small portion of the total number of Pumilio binding sites offered by all cellular transcripts.
In addition to binding to sequence motifs, lncRNAs can fold into structures that interact with proteins involved in key signalling pathways. For example, FAST is transcribed from the antisense strand of the FOXD3 gene6, is highly expressed in hESCs and is required for the maintenance of hESC pluripotency. FAST depletion resulted in hESC differentiation owing to impaired WNT signalling. Each FAST molecule forms five stem–loops, which provide a multivalent platform for interacting with and blocking the E3 ubiquitin ligase β-TrCP from binding to phosphorylated β-catenin and mediating its degradation. FAST therefore enables β-catenin translocation into the nucleus to activate the transcription of WNT-dependent pluripotency genes6 (Fig. 5A). Other lncRNAs block sites of post-translational modification; for example, NF-κB interacting lncRNA (NKILA) forms two distinct hairpins, hairpin A (nucleotides 322–359) and hairpin B (nucleotides 395–418), which both bind to p65. Hairpin B can stabilize the association between NKILA and the NF-κB transcription complex and with the kinase IκB to modulate T cell activation-induced cell death by inhibiting NF-κB activity159. The molecular basis of how such non-canonical RBPs proteins interact with lncRNAs remains to be explored. Nonetheless, the stoichiometric relationship between this group of lncRNAs and their interacting proteins should be carefully evaluated (Supplementary Box 1).
Pairing with other RNAs to recruit protein complexes
Some lncRNAs can directly base pair with other RNAs and subsequently recruit proteins involved in mRNA degradation. For example, Staufen-mediated mRNA decay is carried out by the double-stranded RNA-binding protein Staufen homologue 1 (STAU1), which binds 3′ untranslated regions of mRNAs undergoing translation160,161,162. lncRNAs containing Alu retroelements in human161 or other short interspersed elements (SINEs) in mouse162 can promote Staufen-mediated mRNA decay of mRNAs bearing partial or full complementarity with these repeats by recruiting STAU1. By contrast, the lncRNA TINCR, which is highly expressed during and required for epidermal differentiation, contains several 25-nucleotide motifs that base pair with complementary sequences in differentiation mRNAs; TINCR also recruits STAU1, and the TINCR–STAU1 complex stabilizes the differentiation mRNAs160 (Fig. 5B). Of note, a recent study indicated that TINCR may code a peptide163.
In another example, base pairing in trans appears to be crucial for loading mRNAs on active polyribosomes (Fig. 5B). Antisense to ubiquitin carboxy-terminal hydrolase L1 (AS-Uchl1) is a nuclear lncRNA containing a SINEB2 repeat, which is involved in brain function and neurodegenerative diseases in mice164. Upon activation of stress signalling pathways, for example following mTORC1 inhibition by rapamycin, AS-Uchl1 shuttles from the nucleus to the cytoplasm, where its SINEB2 element undergoes base pairing with the 5′ end of Uchl1 to enhance the translation of the mRNA164.
trans-Acting lncRNAs are emerging as important post-transcriptional regulators. Future studies are warranted not only to better dissect the molecular basis of individual lncRNA–protein interactions by identifying the functional modules of the lncRNAs but also to reveal mechanistic commonalities among different lncRNPs.
Sponging microRNAs
Some abundant lncRNAs bearing microRNA (miRNA)-complementary sites can regulate gene expression as competitive endogenous RNAs or ‘sponges’ of miRNAs, thereby reducing miRNA availability to target mRNAs165,166 (Fig. 5C). The stoichiometric relationship between a potential competitive endogenous lncRNA and miRNAs is important for achieving a measurable effect on target-mRNA expression167,168 (Supplementary Box 1). In tumours, the PNUTS lncRNA is generated by alternative splicing of the PNUTS pre-mRNA, which is mediated through the binding of hnRNPE1 (ref.169) (Fig. 5C). The resulting lncRNA-PNUTS contains seven binding sites for miR-205, a well-established suppressor of the transcription repressors ZEB1 and ZEB2 and a factor required for epithelial cell maintenance. The sequestering of miR-205 by lncRNA-PNUTS results in the upregulation of ZEB1 and ZEB2, and consequently the promotion of epithelial–mesenchymal transition and breast cancer cell migration and invasion169.
Regulating functions of organelles
Interestingly, numerous lncRNAs are localized to specific organelles, such as exosomes and mitochondria (Fig. 1h,i). Because exosomes are regularly released into the extracellular environment, exosome-localized lncRNAs can be secreted and end up in recipient cells, where such lncRNAs are found to be involved in epigenetic regulation, cell-type reprogramming and genomic instability (reviewed in ref.170). Mitochondria-localized lncRNAs can be encoded by both nuclear DNA and mitochondrial DNA, and are often associated with mitochondrial metabolism, apoptosis and the crosstalk of mitochondria with nuclei171. The nuclear-encoded lncRNA survival associated mitochondrial melanoma specific oncogenic non-coding RNA (SAMMSON) controls mitochondrial homeostasis172, mitochondrial 16S ribosomal RNA maturation and expression of mitochondria-encoded polypeptides173. The three abundant mitochondria-encoded lncRNAs lncND5, lncND6 and lncCyt b form intermolecular duplexes with mRNAs and regulate their stability and expression26. Discovery of other organelle-specific lncRNAs will likely provide additional mechanistic insight into the connection between lncRNA regulation and organelle homeostasis.
Physiopathological roles of lncRNAs
The various gene regulatory activities of lncRNAs affect different aspects of physiology, from cell differentiation, growth and responses to diverse stresses and stimuli, to key roles in the nervous, muscular174,175, cardiovascular176, adipose177, haematopoietic and immune178 systems and their associated pathologies. Here, we highlight some aspects and examples of the physiological roles of lncRNAs; we refer the reader to other Reviews for additional information174,175,176,177,178,179.
Neuronal differentiation and disorders
The development of the central nervous system is a particularly intricate process that requires precise spatio-temporal gene regulation. The mammalian brain is a transcriptionally highly complex organ that expresses approximately 40% of mammalian lncRNAs180. Cell culture and mouse models have implicated lncRNAs in neuronal differentiation181 and regeneration after injury182,183. These lncRNAs are often related to protein-coding genes with specific roles in neurogenesis. For example, the lncRNA Silc1 and the transcription factor SOX11 are exquisitely co-expressed in cells of mouse dorsal root ganglia and co-induced following nerve injury. During response to injury, the cis-acting Silc1 is necessary for activation of the SOX11 transcriptional programme and nerve regeneration. The mechanism behind Silc1 interaction with the Sox11 locus to promote its activation is not well understood, but is known to be allele-specific183. In line with their roles in neuronal differentiation, the deregulation of some lncRNAs has been associated with Parkinson disease, Huntington disease, lateral amyotrophic sclerosis or Alzheimer disease184. For example, BACE1-AS, antisense of the gene encoding β-site amyloid precursor protein cleaving enzyme 1 (BACE1; also known as β-secretase 1), promotes BACE1 mRNA stability leading to increase in the levels of neurotoxic amyloid plates in the brain of individuals with Alzheimer disease185. BACE1-AS can be detected in the plasma of these individuals, and thereby serves as a potential disease biomarker186.
Haematopoiesis and immune responses
The extensively investigated roles of lncRNAs in haematopoietic cell differentiation underscore the coordinated activity of differentiation-driving transcription factors and lncRNAs187. Thus, lncRNAs are decisive in activating or suppressing the expression of genes encoding inflammatory molecules178. Interestingly, the induction of key immunity genes may depend on the expression of their regulating lncRNAs prior to the inflammatory stimulus, representing a necessary step for immune gene priming in trained immunity.
One of these immune gene-priming lncRNAs, named UMLILO, was characterized in monocytes, where it functions in cis on the promoters of several chemokine genes located within the same topologically associating domain, thereby facilitating the deposition of H3K4me3 by the WDR5–MLL1 complex following priming treatment60. Several other immunomodulatory lncRNAs are involved in chromatin regulation. lincRNA erythroid prosurvival (lincRNA-EPS), which is expressed in erythrocytes, macrophages and dendritic cells188, and lnc13, which is expressed in macrophages189, repress transcription of immunity genes. lnc13 has been related to inflammatory disease, as SNPs that affect its expression lead to higher levels of lnc13-regulated genes and predispose to coeliac disease189.
Besides those involved in adaptive immunity, mammalian lncRNAs are also related to the control of innate immunity in response to viral infection, which relies on the interferon response as one of its main axes. A signature of lncRNAs is induced by viral infection, including SARS-associated coronavirus, influenza virus, herpes simplex virus 1 and hepatitis C virus190,191, and a significant subset of these lncRNAs is upregulated in response to interferon. The interferon-induced lncRNA negative regulator of interferon response (NRIR) is a negative regulator of several antiviral genes, thereby favouring hepatitis B virus replication192. Similarly, eosinophil granule ontogeny transcript (EGOT), which in liver cells is strongly upregulated by interferon-α and by influenza, hepatitis C virus and Semliki Forest virus infections, inhibits a set of interferon-response genes193.
In summary, lncRNA activities are involved in responses to differentiation cues and stresses that trigger gene expression programmes, in which they exhibit highly specific regulatory functions that are required for correct differentiation and tissue homeostasis.
lncRNAs with cancer-relevant functions
The number of lncRNAs implicated in cancer initiation and progression is continuously growing (Supplementary Box 2), and can be found compiled in curated databases such as Lnc2Cancer194 or the Cancer LncRNA Census195. lncRNAs have been implicated in the acquisition of every hallmark of cancer cells, from the intrinsic capacity of proliferation and survival, through increased metabolism, to the relationship with the tumour microenvironment. Early evidence of the involvement of lncRNAs in cancer came from their transcriptional regulation by key oncogenic or tumour-suppressive transcription factors such as p53 (refs196,197), MYC198,199, the oestrogen receptor200 or signalling cascades such as the Notch pathway201. These lncRNAs contribute to the functional output of the oncogenic or tumour-suppressive responses. Some lncRNAs are activated by p53 following DNA damage. Mouse lincRNA-p21 promotes apoptosis by contributing to p53-dependent transcription repression in trans197 and to activation in cis in a transcript-independent manner of cyclin-dependent kinase inhibitor 1 (refs202,203). Human PANDA204 regulates p53-dependent apoptosis and cell cycle arrest; human DINO stabilizes p53 in the nucleus, thereby reinforcing its transcriptional activity205; GUARDIN preserves genomic integrity through two independent cytoplasmic and nuclear mechanisms206 (Fig. 6a,b). Furthermore, lncRNAs such as MEG3 participate in the p53 regulatory network without being transcriptional targets of p53. The imprinted MEG3 is downregulated in multiple cancers207 and contains an evolutionary conserved RNA structure that mediates p53 activation in trans208.
a | Long non-coding RNAs (lncRNAs) located in the same (human or mouse) genomic region of the cyclin-dependent kinase inhibitor 1A (CDKN1A) gene are direct targets and effectors of p53 following DNA damage. Long intergenic non-coding RNA p21 (lincRNA-p21) functions in trans to recruit the transcription repressor heterogeneous nuclear ribonucleoprotein K (hnRNPK) to the promoter of target genes in response to p53 activation197, or in cis, where it promotes activation of Cdkn1a in two possible ways: lincRNA-p21 can recruit hnRNPK to the promoter of Cdkn1a from its site of transcription203; and another in vivo study has revealed the presence of multiple enhancers (green rectangles) in the lincRNA-p21 locus, which are responsible for transcript-independent regulation in cis of Cdkn1a (ref.202). p21-associated ncRNA DNA damage-activated (PANDA) functions as a decoy for nuclear transcription factor Y subunit-α (NF-YA), thereby removing it from the promoters of its target genes and reducing apoptosis and cell senescence in a p53-dependent fashion261. Damage induced non-coding (DINO) interacts with p53 in the nucleus and promotes p53 tetramer stabilization (consequently reinforcing p53 signalling). Furthermore, DINO co-localizes with p53 at the promoters of several of its target genes, including CDKN1A (ref.205). b | GUARDIN (also known as long non-coding transcriptional activator of miR34a) is activated by p53 following DNA damage and contributes to genome integrity through two separate activities. Part of the GUARDIN pool is exported to the cytoplasm, where it acts as a sponge of miR-23a, thus preventing the destabilization of its main mRNA target, telomeric repeat-binding factor 2 (TRF2), which encodes a factor involved in telomere capping and stability. In the nucleus, GUARDIN functions as a scaffold that enables the interaction of breast cancer type 1 susceptibility protein homologue (BRCA1) and BRCA1 associated RING domain 1 (BARD1), which is important for the recruitment of DNA double-strand break (DSB) repair machinery206. c | MYC oncogene expression is tightly regulated by numerous non-coding RNAs, and relies on the function of several enhancers (green box labelled ‘e’) in the MYC genomic region. Among them, the super-enhancer lncRNA colon cancer associated transcript 1-long (CCAT1-L) promotes chromatin interactions between MYC enhancers and promoters through recruiting the DNA-binding protein CCCTC-binding factor (CTCF), thereby activating Myc expression18,136. Furthermore, the 5′ end of CCAT1-L interacts with hnRNPK, and both interact with the MYC promoter and with the lncRNA plasmacytoma variant translocation 1 (PVT1) to coordinate their expression270. PVT1 competes with the MYC promoter for the availability of enhancers; thus, when PVT1 is expressed, MYC levels are kept low. In the presence of PVT1-inactivating somatic mutations, which are frequent in some cancers, or when PVT1 expression is experimentally repressed using CRISPR interference (CRISPRi), MYC expression is favoured219. HR, homologous recombination; miR, microRNA; NHEJ, non-homologous DNA end joining.
In contrast to these p53-related functions, numerous lncRNAs are either regulated by198,199,209 or regulate210,211 the expression of the proto-oncogene MYC. An intricate regulatory network involving numerous non-coding genomic elements occurs around the MYC locus. MYC resides in the frequently amplified 8q24 chromosomal region, which contains several cancer-associated SNPs within enhancers that form tissue-specific, long-range chromatin interactions with the MYC gene212,213. Several lncRNAs are expressed from this region18,210,214,215,216, which also span SNPs that predispose to cancer215,216,217. For example, CCAT1-L has a role in the transcriptional regulation of MYC by promoting long-range chromatin looping18,214,215 (Fig. 6c). PVT1 is co-amplified with MYC in cancer, and in mice functions as an oncogene by stabilizing the MYC protein210,218. Interestingly, in some human cell types, the promoter of PVT1 limits MYC transcription by competing in cis for the use of specific enhancers and by acting as a DNA boundary element that regulates the expression of MYC, in a manner independent of the PVT1 lncRNA219 (Fig. 6c).
In summary, there is a large body of evidence indicating that cellular homeostasis is dependent on the action of lncRNAs. Although only a fraction of the thousands of lncRNAs expressed may function at some level in cancer cells, these still remain largely understudied. Relevant questions such as the role of lncRNAs in responses to chemotherapy and immunotherapy, their relationship with tumour prognosis and their effect on the tumour microenvironment warrant further investigation.
lncRNAs as therapeutic targets
lncRNAs with key roles in disease could be therapeutic targets. This possibility is supported by theoretical clinical advantages that represent several of their characteristics. The high tissue-specificity and regulation of specific facets of cellular networks suggests that lncRNAs are superior to proteins in terms of potential, undesired toxic effects associated with their targeting. Furthermore, the lack of translation, fast turnover and low expression levels may facilitate quicker effects with lower doses.
The most advanced attempts of therapeutic lncRNA targeting are currently based on the use of antisense oligonucleotides (ASOs). These molecules are essentially single-stranded DNA oligos that can be quickly designed based on sequence homology and RNA accessibility. Importantly, ASOs are suitable for downregulating lncRNAs that are retained in the nucleus: they bind to the target RNA through Watson–Crick base pairing and can induce RNase H-mediated co-transcriptional cleavage at the ASO binding site, leading to premature transcription termination and reduced lncRNA levels220,221. ASOs have high efficacy in cells, although there are limitations to using ASOs in the clinic, mainly because of in vivo toxicity and the lack of proper delivery systems, which hampers tissue targeting by an adequate dose of therapeutic ASOs. To improve their pharmacological properties, ASOs typically are chemically modified to enhance hybridization affinity to their target RNA, thereby increasing resistance to degradation by nucleases and reducing unspecific immunostimulatory activity. These chemical variations include the GapmeR ASOs, RNA–DNA–RNA single-stranded oligonucleotide chains in which ribonucleotides may contain a 2′-O-methoxyethyl modified sugar backbone222 or additional modifications such as locked nucleic acids and S-constrained ethyl residues223. Moreover, fused aptamers may also be used for targeted intracellular delivery of these oligo-based drugs224. Several mRNA-targeting ASOs have already been approved by the FDA and the European Medicines Agency225 or have advanced to clinical trials225, and several ASOs targeting oncogenic lncRNAs are under development and protected by patents.
Less developed is the use of small molecules for the targeting of lncRNAs. Obtaining successful molecules that bind lncRNAs with high affinity and specificity requires the identification of relevant RNA motifs with sufficient structural complexity226. This level of structural knowledge is so far available only for a limited number of lncRNAs, revealing that lncRNAs often fold into several modular domains potentially involved in different molecular interactions208,227,228,229,230,231. Blocking functional interactions between lncRNAs and proteins could be desirable from a therapeutic perspective. Alternatively, synthetic molecules that mimic the structure and binding properties of lncRNAs may work as decoys by competing with the lncRNA for protein binding, and therefore interfering with its function. All of these promising approaches will become more practical as the structural and molecular features of lncRNAs become better understood.
Finally, tools based on CRISPR–Cas systems are among the most versatile and promising for the precise modulation of lncRNAs. The different versions of CRISPR–Cas-engineered molecules allow the deletion (using CRISPR–Cas9)232, inhibition (CRISRPi)233 or activation (CRISPRa)234 of lncRNA-encoding genes, as well as the degradation of the transcripts themselves (CRISPR–Cas13)235. These technologies enable relatively fast knockout, knockdown or overexpression of lncRNAs, are already widely used for research applications at single lncRNA loci and are increasingly applied to thousands of loci for high-throughput loss-of-function and gain-of-function screening in diverse experimental settings236. However, because of their lack of functional open reading frames, in vivo targeting of lncRNAs using CRISPR–Cas is more difficult than targeting protein-coding genes. It is therefore expected that the therapeutic application of CRISPR–Cas systems at lncRNA loci will lag behind that of protein-coding genes.
Concluding remarks
In recent years we have witnessed remarkable progress in our understanding of lncRNAs, and we now have a clearer picture of the features and functional versatility of these molecules. Nevertheless, this knowledge represents only a small fraction of the landscape of their gene regulatory potential. Several aspects of lncRNA biology still require rigorous investigation, for instance, we are still far from understanding how lncRNA sequences and structural features relate to their functions, given their non-coding nature and low sequence conservation. Interestingly, a recent study has shown that lncRNAs with similar k-mer content have related functions despite their lack of linear homology237. This study implies that short sequence elements in lncRNAs mediate interactions with proteins (and/or other molecules), and thus are key determinants of lncRNA function. However, the nature and dynamics of such interactions still need to be elucidated. It is also increasingly evident that multiple features of lncRNAs can define their functionality. These features include their sequence, expression levels, processing, cellular localization, structural organization and interactions with other molecules. The integrated knowledge of all these features will hopefully increase the identification and classification of functional lncRNAs.
How lncRNAs influence complex physiological processes and the onset of diseases are questions of great relevance. Our current knowledge indicates that lncRNAs fine-tune cell specification and disease. These functions require deeper comprehension, not only to provide a complete picture of physiopathological processes but also because lncRNAs can readably be therapeutically targeted with high specificity. Given their characteristics, disease-related lncRNAs will predictably gain greater relevance in the context of personalized medicine. Progress in this area will go hand in hand with better understanding of the gene regulation modalities of lncRNAs.
Change history
08 January 2021
A Correction to this paper has been published: https://doi.org/110.1038/s41580-021-00330-4
References
Uszczynska-Ratajczak, B., Lagarde, J., Frankish, A., Guigo, R. & Johnson, R. Towards a complete map of the human long non-coding RNA transcriptome. Nat. Rev. Genet. 19, 535–548 (2018).
Fang, S. et al. NONCODEV5: a comprehensive annotation database for long non-coding RNAs. Nucleic Acids Res. 46, D308–D314 (2018).
Wu, H., Yang, L. & Chen, L. L. The diversity of long noncoding RNAs and their generation. Trends Genet. 33, 540–552 (2017).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Tian, B. & Manley, J. L. Alternative polyadenylation of mRNA precursors. Nat. Rev. Mol. Cell Biol. 18, 18–30 (2017).
Guo, C. J. et al. Distinct processing of lncRNAs contributes to non-conserved functions in stem cells. Cell 181, 621–636.e22 (2020). This paper shows that conserved lncRNAs exhibit distinct subcellular localization and function in human compared with mouse pluripotent cells and that the relatively high evolutionary plasticity of lncRNAs can support species-specific gene expression programmes.
Quinn, J. J. et al. Rapid evolutionary turnover underlies conserved lncRNA–genome interactions. Genes Dev. 30, 191–207 (2016).
Hezroni, H. et al. Principles of long noncoding RNA evolution derived from direct comparison of transcriptomes in 17 species. Cell Rep. 11, 1110–1122 (2015).
Lagarde, J. et al. High-throughput annotation of full-length long noncoding RNAs with capture long-read sequencing. Nat. Genet. 49, 1731–1740 (2017). This paper re-annotates GENCODE lncRNAs by developing RNA capture long seq for full-length lncRNAs, which provides a comprehensive lncRNA annotation resource.
Mele, M. et al. Chromatin environment, transcriptional regulation, and splicing distinguish lincRNAs and mRNAs. Genome Res. 27, 27–37 (2017).
Liu, S. & Trapnell, C. Single-cell transcriptome sequencing: recent advances and remaining challenges. F1000Res 5, 182 (2016).
Schlackow, M. et al. Distinctive patterns of transcription and RNA processing for human lincRNAs. Mol. Cell 65, 25–38 (2017). This paper applies mammalian native elongating transcript sequencing (mNET-seq), and reveals the distinct transcription patterns, chromatin tethering and degradation between mRNA and lncRNA genes.
Yin, Y. et al. U1 snRNP regulates chromatin retention of noncoding RNAs. Nature 580, 147–150 (2020).
Vos, S. M. et al. Structure of activated transcription complex Pol II–DSIF–PAF–SPT6. Nature 560, 607–612 (2018).
Nojima, T. et al. Deregulated expression of mammalian lncRNA through loss of SPT6 induces R-loop formation, replication stress, and cellular senescence. Mol. Cell 72, 970–984.e7 (2018).
Zuckerman, B. & Ulitsky, I. Predictive models of subcellular localization of long RNAs. RNA 25, 557–572 (2019).
Rosenberg, A. B., Patwardhan, R. P., Shendure, J. & Seelig, G. Learning the sequence determinants of alternative splicing from millions of random sequences. Cell 163, 698–711 (2015).
Xiang, J. F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
Azam, S. et al. Nuclear retention element recruits U1 snRNP components to restrain spliced lncRNAs in the nucleus. RNA Biol. 16, 1001–1009 (2019).
Shukla, C. J. et al. High-throughput identification of RNA nuclear enrichment sequences. EMBO J. 37, e98452 (2018).
Lubelsky, Y. & Ulitsky, I. Sequences enriched in Alu repeats drive nuclear localization of long RNAs in human cells. Nature 555, 107–111 (2018).
Hacisuleyman, E., Shukla, C. J., Weiner, C. L. & Rinn, J. L. Function and evolution of local repeats in the Firre locus. Nat. Commun. 7, 11021 (2016).
Zuckerman, B., Ron, M., Mikl, M., Segal, E. & Ulitsky, I. Gene architecture and sequence composition underpin selective dependency of nuclear export of long RNAs on NXF1 and the TREX complex. Mol. Cell 79, 251–267 (2020). By systematically depleting RNA export factors, this study reveals that export of long RNAs is modulated by gene architecture, sequence composition, RNA secondary structure, sequence motifs and post-transcriptional modifications.
Carlevaro-Fita, J., Rahim, A., Guigo, R., Vardy, L. A. & Johnson, R. Cytoplasmic long noncoding RNAs are frequently bound to and degraded at ribosomes in human cells. RNA 22, 867–882 (2016).
Zeng, C. & Hamada, M. Identifying sequence features that drive ribosomal association for lncRNA. BMC Genomics 19, 906 (2018).
Mercer, T. R. et al. The human mitochondrial transcriptome. Cell 146, 645–658 (2011).
Rackham, O. et al. Long noncoding RNAs are generated from the mitochondrial genome and regulated by nuclear-encoded proteins. RNA 17, 2085–2093 (2011).
Noh, J. H. et al. HuR and GRSF1 modulate the nuclear export and mitochondrial localization of the lncRNA RMRP. Genes Dev. 30, 1224–1239 (2016).
Li, S. et al. exoRBase: a database of circRNA, lncRNA and mRNA in human blood exosomes. Nucleic Acids Res. 46, D106–D112 (2018).
Gudenas, B. L. & Wang, L. Prediction of lncRNA subcellular localization with deep learning from sequence features. Sci. Rep. 8, 16385 (2018).
Statello, L. et al. Identification of RNA-binding proteins in exosomes capable of interacting with different types of RNA: RBP-facilitated transport of RNAs into exosomes. PLoS ONE 13, e0195969 (2018).
Chu, C., Qu, K., Zhong, F. L., Artandi, S. E. & Chang, H. Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA–chromatin interactions. Mol. Cell 44, 667–678 (2011).
Bonetti, A. et al. RADICL-seq identifies general and cell type-specific principles of genome-wide RNA–chromatin interactions. Nat. Commun. 11, 1018 (2020).
Li, X. et al. GRID-seq reveals the global RNA–chromatin interactome. Nat. Biotechnol. 35, 940–950 (2017).
Bell, J. C. et al. Chromatin-associated RNA sequencing (ChAR-seq) maps genome-wide RNA-to-DNA contacts. eLife 7, e27024 (2018).
Isoda, T. et al. Non-coding transcription instructs chromatin folding and compartmentalization to dictate enhancer–promoter communication and T cell fate. Cell 171, 103–119.e18 (2017).
Mumbach, M. R. et al. HiChIRP reveals RNA-associated chromosome conformation. Nat. Methods 16, 489–492 (2019).
Dueva, R. et al. Neutralization of the positive charges on histone tails by RNA promotes an open chromatin structure. Cell Chem. Biol. 26, 1436–1449.e5 (2019).
Yang, F. et al. The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation. Genome Biol. 16, 52 (2015).
Saldana-Meyer, R. et al. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cell 76, 412–422.e5 (2019).
Schertzer, M. D. et al. lncRNA-induced spread of polycomb controlled by genome architecture, RNA abundance, and CpG island DNA. Mol. Cell 75, 523–537.e10 (2019).
Kotzin, J. J. et al. The long non-coding RNA morrbid regulates Bim and short-lived myeloid cell lifespan. Nature 537, 239–243 (2016).
Beckedorff, F. C. et al. The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1A promoter, reducing the expression of RASSF1A and increasing cell proliferation. PLoS Genet. 9, e1003705 (2013).
Marin-Bejar, O. et al. The human lncRNA LINC-PINT inhibits tumor cell invasion through a highly conserved sequence element. Genome Biol. 18, 202 (2017).
Yap, K. L. et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell 38, 662–674 (2010).
Holdt, L. M. et al. Alu elements in ANRIL non-coding RNA at chromosome 9p21 modulate atherogenic cell functions through trans-regulation of gene networks. PLoS Genet. 9, e1003588 (2013).
Long, Y. et al. RNA is essential for PRC2 chromatin occupancy and function in human pluripotent stem cells. Nat. Genet. 52, 931–938 (2020).
Blanco, M. R. & Guttman, M. Re-evaluating the foundations of lncRNA–Polycomb function. EMBO J. 36, 964–966 (2017).
Kopp, F. & Mendell, J. T. Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393–407 (2018).
Portoso, M. et al. PRC2 is dispensable for HOTAIR-mediated transcriptional repression. EMBO J. 36, 981–994 (2017).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Lee, J. T. & Bartolomei, M. S. X-inactivation, imprinting, and long noncoding RNAs in health and disease. Cell 152, 1308–1323 (2013).
Latos, P. A. et al. Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science 338, 1469–1472 (2012).
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008).
Pintacuda, G. et al. hnRNPK recruits PCGF3/5-PRC1 to the Xist RNA B-repeat to establish Polycomb-mediated chromosomal silencing. Mol. Cell 68, 955–969.e10 (2017).
Colognori, D., Sunwoo, H., Kriz, A. J., Wang, C. Y. & Lee, J. T. Xist deletional analysis reveals an interdependency between Xist RNA and Polycomb complexes for spreading along the inactive X. Mol. Cell 74, 101–117.e10 (2019).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Jeon, Y. & Lee, J. T. YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146, 119–133 (2011).
Sigova, A. A. et al. Transcription factor trapping by RNA in gene regulatory elements. Science 350, 978–981 (2015).
Fanucchi, S. et al. Immune genes are primed for robust transcription by proximal long noncoding RNAs located in nuclear compartments. Nat. Genet. 51, 138–150 (2019).
Wang, K. C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Luo, H. et al. HOTTIP lncRNA promotes hematopoietic stem cell self-renewal leading to AML-like disease in mice. Cancer Cell 36, 645–659.e8 (2019).
Jain, A. K. et al. lncPRESS1 is a p53-regulated lncRNA that safeguards pluripotency by disrupting SIRT6-mediated de-acetylation of histone H3K56. Mol. Cell 64, 967–981 (2016).
Schmitz, K. M., Mayer, C., Postepska, A. & Grummt, I. Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Genes Dev. 24, 2264–2269 (2010).
Martianov, I., Ramadass, A., Serra Barros, A., Chow, N. & Akoulitchev, A. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445, 666–670 (2007).
O’Leary, V. B. et al. PARTICLE, a triplex-forming long ncRNA, regulates locus-specific methylation in response to low-dose irradiation. Cell Rep. 11, 474–485 (2015).
Mondal, T. et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA–DNA triplex structures. Nat. Commun. 6, 7743 (2015).
Grote, P. et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 24, 206–214 (2013).
Kuo, C. C. et al. Detection of RNA–DNA binding sites in long noncoding RNAs. Nucleic Acids Res. 47, e32 (2019).
Li, Y., Syed, J. & Sugiyama, H. RNA–DNA triplex formation by long noncoding RNAs. Cell Chem. Biol. 23, 1325–1333 (2016).
Blank-Giwojna, A., Postepska-Igielska, A. & Grummt, I. lncRNA KHPS1 activates a poised enhancer by triplex-dependent recruitment of epigenomic regulators. Cell Rep. 26, 2904–2915.e4 (2019).
Maldonado, R., Schwartz, U., Silberhorn, E. & Langst, G. Nucleosomes stabilize ssRNA–dsDNA triple helices in human cells. Mol. Cell 73, 1243–1254.e6 (2019).
Postepska-Igielska, A. et al. lncRNA Khps1 regulates expression of the proto-oncogene SPHK1 via triplex-mediated changes in chromatin structure. Mol. Cell 60, 626–636 (2015).
Niehrs, C. & Luke, B. Regulatory R-loops as facilitators of gene expression and genome stability. Nat. Rev. Mol. Cell Biol. 21, 167–178 (2020).
Tan-Wong, S. M., Dhir, S. & Proudfoot, N. J. R-loops promote antisense transcription across the mammalian genome. Mol. Cell 76, 600–616.e6 (2019).
Sun, Q., Csorba, T., Skourti-Stathaki, K., Proudfoot, N. J. & Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340, 619–621 (2013).
Gibbons, H. R. et al. Divergent lncRNA GATA3-AS1 regulates GATA3 transcription in T-helper 2 cells. Front. Immunol. 9, 2512 (2018).
Boque-Sastre, R. et al. Head-to-head antisense transcription and R-loop formation promotes transcriptional activation. Proc. Natl Acad. Sci. USA 112, 5785–5790 (2015).
Arab, K. et al. GADD45A binds R-loops and recruits TET1 to CpG island promoters. Nat. Genet. 51, 217–223 (2019).
Ariel, F. et al. R-loop mediated trans action of the APOLO long noncoding RNA. Mol. Cell 77, 1055–1065.e4 (2020).
Seila, A. C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008).
Luo, S. et al. Divergent lncRNAs regulate gene expression and lineage differentiation in pluripotent cells. Cell Stem Cell 18, 637–652 (2016).
Gil, N. & Ulitsky, I. Regulation of gene expression by cis-acting long non-coding RNAs. Nat. Rev. Genet. 21, 102–117 (2019).
Wutz, A. Gene silencing in X-chromosome inactivation: advances in understanding facultative heterochromatin formation. Nat. Rev. Genet. 12, 542–553 (2011).
Jiang, J. et al. Translating dosage compensation to trisomy 21. Nature 500, 296–300 (2013).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013). This study shows that at the onset of X-chromosome inactivation, Xist binds in a sequence-independent manner to distinct regions of the X chromosome, which are brought into 3D proximity, subsequently spreading to more distal regions that reposition in the 3D space.
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Pandya-Jones, A. et al. A protein assembly mediates Xist localization and gene silencing. Nature 587, 1–7 (2020).
Herzog, V. A. et al. A strand-specific switch in noncoding transcription switches the function of a Polycomb/Trithorax response element. Nat. Genet. 46, 973–981 (2014).
Alecki, C. et al. RNA–DNA strand exchange by the Drosophila Polycomb complex PRC2. Nat. Commun. 11, 1781 (2020).
Rosa, S., Duncan, S. & Dean, C. Mutually exclusive sense-antisense transcription at FLC facilitates environmentally induced gene repression. Nat. Commun. 7, 13031 (2016).
Csorba, T., Questa, J. I., Sun, Q. & Dean, C. Antisense COOLAIR mediates the coordinated switching of chromatin states at FLC during vernalization. Proc. Natl Acad. Sci. USA 111, 16160–16165 (2014).
Stojic, L. et al. Transcriptional silencing of long noncoding RNA GNG12-AS1 uncouples its transcriptional and product-related functions. Nat. Commun. 7, 10406 (2016).
Thebault, P. et al. Transcription regulation by the noncoding RNA SRG1 requires Spt2-dependent chromatin deposition in the wake of RNA polymerase II. Mol. Cell. Biol. 31, 1288–1300 (2011).
Rom, A. et al. Regulation of CHD2 expression by the Chaserr long noncoding RNA gene is essential for viability. Nat. Commun. 10, 5092 (2019).
Sleutels, F., Zwart, R. & Barlow, D. P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002).
Santoro, F. & Pauler, F. M. Silencing by the imprinted Airn macro lncRNA: transcription is the answer. Cell Cycle 12, 711–712 (2013).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Hon, C. C. et al. An atlas of human long non-coding RNAs with accurate 5′ ends. Nature 543, 199–204 (2017).
Kim, Y. J., Xie, P., Cao, L., Zhang, M. Q. & Kim, T. H. Global transcriptional activity dynamics reveal functional enhancer RNAs. Genome Res. 28, 1799–1811 (2018).
Jiao, W. et al. HPSE enhancer RNA promotes cancer progression through driving chromatin looping and regulating hnRNPU/p300/EGR1/HPSE axis. Oncogene 37, 2728–2745 (2018).
Li, W. et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498, 516–520 (2013).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Marques, A. C. et al. Chromatin signatures at transcriptional start sites separate two equally populated yet distinct classes of intergenic long noncoding RNAs. Genome Biol. 14, R131 (2013).
Orom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
De Santa, F. et al. A large fraction of extragenic RNA Pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010).
Gil, N. & Ulitsky, I. Production of spliced long noncoding RNAs specifies regions with increased enhancer activity. Cell Syst. 7, 537–547.e3 (2018).
Tan, J. Y., Biasini, A., Young, R. S. & Marques, A. C. Splicing of enhancer-associated lincRNAs contributes to enhancer activity. Life Sci. Alliance 3, e202000663 (2020).
Amaral, P. P. et al. Genomic positional conservation identifies topological anchor point RNAs linked to developmental loci. Genome Biol. 19, 32 (2018).
Tan, J. Y. et al. cis-Acting complex-trait-associated lincRNA expression correlates with modulation of chromosomal architecture. Cell Rep. 18, 2280–2288 (2017).
Fanucchi, S. & Mhlanga, M. M. Enhancer-derived lncRNAs regulate genome architecture: fact or fiction? Trends Genet. 33, 375–377 (2017).
van Steensel, B. & Furlong, E. E. M. The role of transcription in shaping the spatial organization of the genome. Nat. Rev. Mol. Cell Biol. 20, 327–337 (2019).
Grossi, E. et al. A lncRNA–SWI/SNF complex crosstalk controls transcriptional activation at specific promoter regions. Nat. Commun. 11, 936 (2020).
Tomita, S. et al. A cluster of noncoding RNAs activates the ESR1 locus during breast cancer adaptation. Nat. Commun. 6, 6966 (2015).
Dao, L. T. M. et al. Genome-wide characterization of mammalian promoters with distal enhancer functions. Nat. Genet. 49, 1073–1081 (2017).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016). This paper dissects the effects in cis of 12 lncRNAs, concluding that only 5 of the lncRNAs affected the expression of the neighbouring gene, in a transcript-independent manner.
Paralkar, V. R. et al. Unlinking an lncRNA from its associated cis element. Mol. Cell 62, 104–110 (2016).
Anderson, K. M. et al. Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature 539, 433–436 (2016).
Laurent, F. et al. HAND2 target gene regulatory networks control atrioventricular canal and cardiac valve development. Cell Rep. 19, 1602–1613 (2017).
Han, X. et al. The lncRNA Hand2os1/Uph locus orchestrates heart development through regulation of precise expression of Hand2. Development 146, dev176198 (2019).
Ritter, N. et al. The lncRNA locus Handsdown regulates cardiac gene programs and is essential for early mouse development. Dev. Cell 50, 644–657.e8 (2019).
Yin, Y. et al. Opposing roles for the lncRNA Haunt and its genomic locus in regulating HOXA gene activation during embryonic stem cell differentiation. Cell Stem Cell 16, 504–516 (2015).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
West, J. A. et al. Structural, super-resolution microscopy analysis of paraspeckle nuclear body organization. J. Cell Biol. 214, 817–830 (2016).
Clemson, C. M. et al. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol. Cell 33, 717–726 (2009).
Hutchinson, J. N. et al. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8, 39 (2007).
Sasaki, Y. T., Ideue, T., Sano, M., Mituyama, T. & Hirose, T. MENε/β noncoding RNAs are essential for structural integrity of nuclear paraspeckles. Proc. Natl Acad. Sci. USA 106, 2525–2530 (2009).
Sunwoo, H. et al. MEN ε/β nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res. 19, 347–359 (2009).
Yamazaki, T. et al. Functional domains of NEAT1 architectural lncRNA induce paraspeckle assembly through phase separation. Mol. Cell 70, 1038–1053 e1037 (2018). This paper shows that the middle domain of NEAT1 is key for de novo assembly of the phase-separated paraspeckles.
Lin, Y., Schmidt, B. F., Bruchez, M. P. & McManus, C. J. Structural analyses of NEAT1 lncRNAs suggest long-range RNA interactions that may contribute to paraspeckle architecture. Nucleic Acids Res. 46, 3742–3752 (2018).
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016).
Wilusz, J. E. et al. A triple helix stabilizes the 3′ ends of long noncoding RNAs that lack poly(A) tails. Genes Dev. 26, 2392–2407 (2012).
Engreitz, J. M. et al. RNA–RNA interactions enable specific targeting of noncoding RNAs to nascent pre-mRNAs and chromatin sites. Cell 159, 188–199 (2014).
Tripathi, V. et al. SRSF1 regulates the assembly of pre-mRNA processing factors in nuclear speckles. Mol. Biol. Cell 23, 3694–3706 (2012).
Yang, L. et al. ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147, 773–788 (2011).
Arun, G. et al. Differentiation of mammary tumors and reduction in metastasis upon Malat1 lncRNA loss. Genes Dev. 30, 34–51 (2016).
Malakar, P. et al. Long noncoding RNA MALAT1 promotes hepatocellular carcinoma development by SRSF1 upregulation and mTOR activation. Cancer Res. 77, 1155–1167 (2017).
Fei, J. et al. Quantitative analysis of multilayer organization of proteins and RNA in nuclear speckles at super resolution. J. Cell Sci. 130, 4180–4192 (2017).
Cai, Z. et al. RIC-seq for global in situ profiling of RNA–RNA spatial interactions. Nature 582, 432–437 (2020).
Yin, Q. F. et al. Long noncoding RNAs with snoRNA ends. Mol. Cell 48, 219–230 (2012).
Wu, H. et al. Unusual processing generates SPA lncRNAs that sequester multiple RNA binding proteins. Mol. Cell 64, 534–548 (2016).
Yap, K. et al. A short tandem repeat-enriched RNA assembles a nuclear compartment to control alternative splicing and promote cell survival. Mol. Cell 72, 525–540.e13 (2018).
Jolly, C., Usson, Y. & Morimoto, R. I. Rapid and reversible relocalization of heat shock factor 1 within seconds to nuclear stress granules. Proc. Natl Acad. Sci. USA 96, 6769–6774 (1999).
Weighardt, F. et al. A novel hnRNP protein (HAP/SAF-B) enters a subset of hnRNP complexes and relocates in nuclear granules in response to heat shock. J. Cell Sci. 112, 1465–1476 (1999).
Denegri, M. et al. Stress-induced nuclear bodies are sites of accumulation of pre-mRNA processing factors. Mol. Biol. Cell 12, 3502–3514 (2001).
Metz, A., Soret, J., Vourc’h, C., Tazi, J. & Jolly, C. A key role for stress-induced satellite III transcripts in the relocalization of splicing factors into nuclear stress granules. J. Cell Sci. 117, 4551–4558 (2004).
Jolly, C. et al. Stress-induced transcription of satellite III repeats. J. Cell Biol. 164, 25–33 (2004).
Ninomiya, K. et al. lncRNA-dependent nuclear stress bodies promote intron retention through SR protein phosphorylation. EMBO J. 39, e102729 (2020).
Audas, T. E., Jacob, M. D. & Lee, S. Immobilization of proteins in the nucleolus by ribosomal intergenic spacer noncoding RNA. Mol. Cell 45, 147–157 (2012).
Wang, M. et al. Stress-induced low complexity RNA activates physiological amyloidogenesis. Cell Rep. 24, 1713–1721.e4 (2018).
Hacisuleyman, E. et al. Topological organization of multichromosomal regions by the long intergenic noncoding RNA Firre. Nat. Struct. Mol. Biol. 21, 198–206 (2014).
Lewandowski, J. P. et al. The Firre locus produces a trans-acting RNA molecule that functions in hematopoiesis. Nat. Commun. 10, 5137 (2019).
Maass, P. G., Barutcu, A. R., Weiner, C. L. & Rinn, J. L. Inter-chromosomal contact properties in live-cell imaging and in Hi-C. Mol. Cell 69, 1039–1045.e3 (2018).
Hartford, C. C. R. & Lal, A. When long noncoding becomes protein coding. Mol. Cell. Biol. 40, e00528-19 (2020).
Romero-Barrios, N., Legascue, M. F., Benhamed, M., Ariel, F. & Crespi, M. Splicing regulation by long noncoding RNAs. Nucleic Acids Res. 46, 2169–2184 (2018).
Lee, S. et al. Noncoding RNA NORAD regulates genomic stability by sequestering PUMILIO proteins. Cell 164, 69–80 (2016).
Tichon, A., Perry, R. B., Stojic, L. & Ulitsky, I. SAM68 is required for regulation of Pumilio by the NORAD long noncoding RNA. Genes Dev. 32, 70–78 (2018).
Miller, M. A. & Olivas, W. M. Roles of Puf proteins in mRNA degradation and translation. Wiley Interdiscip. Rev. RNA 2, 471–492 (2011).
Liu, B. et al. A cytoplasmic NF-κB interacting long noncoding RNA blocks IκB phosphorylation and suppresses breast cancer metastasis. Cancer Cell 27, 370–381 (2015).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Gong, C. & Maquat, L. E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–288 (2011).
Wang, J., Gong, C. & Maquat, L. E. Control of myogenesis by rodent SINE-containing lncRNAs. Genes Dev. 27, 793–804 (2013).
Eckhart, L., Lachner, J., Tschachler, E. & Rice, R. H. TINCR is not a non-coding RNA but encodes a protein component of cornified epidermal keratinocytes. Exp. Dermatol. 29, 376–379 (2020).
Carrieri, C. et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 491, 454–457 (2012).
Cesana, M. et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 147, 358–369 (2011).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146, 353–358 (2011).
Bosson, A. D., Zamudio, J. R. & Sharp, P. A. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol. Cell 56, 347–359 (2014).
Denzler, R., Agarwal, V., Stefano, J., Bartel, D. P. & Stoffel, M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol. Cell 54, 766–776 (2014).
Grelet, S. et al. A regulated PNUTS mRNA to lncRNA splice switch mediates EMT and tumour progression. Nat. Cell Biol. 19, 1105–1115 (2017).
Fatima, F. & Nawaz, M. Vesiculated long non-coding RNAs: offshore packages deciphering trans-regulation between cells, cancer progression and resistance to therapies. Noncoding RNA 3, 10 (2017).
Zhao, Y., Sun, L., Wang, R. R., Hu, J. F. & Cui, J. The effects of mitochondria-associated long noncoding RNAs in cancer mitochondria: new players in an old arena. Crit. Rev. Oncol. Hematol. 131, 76–82 (2018).
Leucci, E. et al. Melanoma addiction to the long non-coding RNA SAMMSON. Nature 531, 518–522 (2016).
Vendramin, R. et al. SAMMSON fosters cancer cell fitness by concertedly enhancing mitochondrial and cytosolic translation. Nat. Struct. Mol. Biol. 25, 1035–1046 (2018). This study discovers that the mitochondrially localized oncogenic lncRNA SAMMSON modulates RNA–protein complex formation in distinct cellular compartments, including in mitochondria, the cytosol and the nucleus to promote cell growth.
Fatica, A. & Bozzoni, I. Long non-coding RNAs: new players in cell differentiation and development. Nat. Rev.Genet. 15, 7–21 (2014).
Sweta, S., Dudnakova, T., Sudheer, S., Baker, A. H. & Bhushan, R. Importance of long non-coding RNAs in the development and disease of skeletal muscle and cardiovascular lineages. Front. Cell Dev. Biol. 7, 228 (2019).
Hobuss, L., Bar, C. & Thum, T. Long non-coding RNAs: at the heart of cardiac dysfunction? Front. Physiol. 10, 30 (2019).
Sun, L. & Lin, J. D. Function and mechanism of long noncoding RNAs in adipocyte biology. Diabetes 68, 887–896 (2019).
Chen, Y. G., Satpathy, A. T. & Chang, H. Y. Gene regulation in the immune system by long noncoding RNAs. Nat. Immunol. 18, 962–972 (2017).
Hezroni, H., Perry, R. B. T. & Ulitsky, I. Long noncoding RNAs in development and regeneration of the neural lineage. Cold Spring Harb. Symp. Quant. Biol. 84, 165–177 (2019).
Briggs, J. A., Wolvetang, E. J., Mattick, J. S., Rinn, J. L. & Barry, G. Mechanisms of long non-coding RNAs in mammalian nervous system development, plasticity, disease, and evolution. Neuron 88, 861–877 (2015).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013).
He, D. et al. lncRNA functional networks in oligodendrocytes reveal stage-specific myelination control by an lncOL1/Suz12 complex in the CNS. Neuron 93, 362–378 (2017).
Perry, R. B., Hezroni, H., Goldrich, M. J. & Ulitsky, I. Regulation of neuroregeneration by long noncoding RNAs. Mol. Cell 72, 553–567.e5 (2018).
Wei, C. W., Luo, T., Zou, S. S. & Wu, A. S. The role of long noncoding RNAs in central nervous system and neurodegenerative diseases. Front. Behav. Neurosci. 12, 175 (2018).
Faghihi, M. A. et al. Evidence for natural antisense transcript-mediated inhibition of microRNA function. Genome Biol. 11, R56 (2010).
Feng, L. et al. Plasma long non-coding RNA BACE1 as a novel biomarker for diagnosis of Alzheimer disease. BMC Neurol. 18, 4 (2018).
Luo, M. et al. Long non-coding RNAs control hematopoietic stem cell function. Cell Stem Cell 16, 426–438 (2015).
Atianand, M. K. et al. A long noncoding RNA lincRNA-EPS acts as a transcriptional brake to restrain inflammation. Cell 165, 1672–1685 (2016).
Castellanos-Rubio, A. et al. A long noncoding RNA associated with susceptibility to celiac disease. Science 352, 91–95 (2016).
Peng, X. et al. Unique signatures of long noncoding RNA expression in response to virus infection and altered innate immune signaling. mBio 1, e00206–e00210 (2010).
Hadjicharalambous, M. R. & Lindsay, M. A. Long non-coding RNAs and the innate immune response. Noncoding RNA 5, 34 (2019).
Kambara, H. et al. Negative regulation of the interferon response by an interferon-induced long non-coding RNA. Nucleic Acids Res. 42, 10668–10680 (2014).
Carnero, E. et al. Long noncoding RNA EGOT negatively affects the antiviral response and favors HCV replication. EMBO Rep. 17, 1013–1028 (2016).
Gao, Y. et al. Lnc2Cancer v2.0: updated database of experimentally supported long non-coding RNAs in human cancers. Nucleic Acids Res. 47, D1028–D1033 (2019).
Carlevaro-Fita, J. et al. Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun. Biol. 3, 56 (2020).
Sanchez, Y. et al. Genome-wide analysis of the human p53 transcriptional network unveils a lncRNA tumour suppressor signature. Nat. Commun. 5, 5812 (2014). This study shows that following DNA damage, p53 directly activates the expression of dozens of lncRNAs in human cells, which is required for the complete tumour suppressor response of p53.
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Hart, J. R., Roberts, T. C., Weinberg, M. S., Morris, K. V. & Vogt, P. K. MYC regulates the non-coding transcriptome. Oncotarget 5, 12543–12554 (2014).
Kim, T. et al. Role of MYC-regulated long noncoding RNAs in cell cycle regulation and tumorigenesis. J. Natl Cancer Inst. 107, https://doi.org/10.1093/jnci/dju505 (2015).
Chakravarty, D. et al. The oestrogen receptor α-regulated lncRNA NEAT1 is a critical modulator of prostate cancer. Nat. Commun. 5, 5383 (2014).
Trimarchi, T. et al. Genome-wide mapping and characterization of notch-regulated long noncoding RNAs in acute leukemia. Cell 158, 593–606 (2014).
Groff, A. F. et al. In vivo characterization of Linc-p21 reveals functional cis-regulatory DNA elements. Cell Rep. 16, 2178–2186 (2016).
Dimitrova, N. et al. lincRNA-p21 activates p21 in cis to promote Polycomb target gene expression and to enforce the G1/S checkpoint. Mol. Cell 54, 777–790 (2014).
Hung, T. et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 43, 621–629 (2011).
Schmitt, A. M. et al. An inducible long noncoding RNA amplifies DNA damage signaling. Nat. Genet. 48, 1370–1376 (2016).
Hu, W. L. et al. GUARDIN is a p53-responsive long non-coding RNA that is essential for genomic stability. Nat. Cell Biol. 20, 492–502 (2018).
Zhou, Y., Zhang, X. & Klibanski, A. MEG3 noncoding RNA: a tumor suppressor. J. Mol. Endocrinol. 48, R45–R53 (2012).
Uroda, T. et al. Conserved pseudoknots in lncRNA MEG3 are essential for stimulation of the p53 pathway. Mol. Cell 75, 982–995.e9 (2019). This study identifies conserved structural elements in the lncRNA MEG3 that are required for the activation of the p53 response by the lncRNA.
Winkle, M. et al. Long noncoding RNAs as a novel component of the Myc transcriptional network. FASEB J. 29, 2338–2346 (2015).
Tseng, Y. Y. et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 512, 82–86 (2014).
Wang, Z. et al. lncRNA epigenetic landscape analysis identifies EPIC1 as an oncogenic lncRNA that interacts with MYC and promotes cell-cycle progression in cancer. Cancer Cell 33, 706–720.e9 (2018).
Jia, L. et al. Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet. 5, e1000597 (2009).
Sur, I., Tuupanen, S., Whitington, T., Aaltonen, L. A. & Taipale, J. Lessons from functional analysis of genome-wide association studies. Cancer Res. 73, 4180–4184 (2013).
Ling, H. et al. CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 23, 1446–1461 (2013).
Kim, T. et al. Long-range interaction and correlation between MYC enhancer and oncogenic long noncoding RNA CARLo-5. Proc. Natl Acad. Sci. USA 111, 4173–4178 (2014).
Prensner, J. R. et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat. Biotechnol. 29, 742–749 (2011).
Yang, L. et al. lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs. Nature 500, 598–602 (2013).
Cui, M. et al. Long non-coding RNA PVT1 and cancer. Biochem. Biophys. Res. Commun. 471, 10–14 (2016).
Cho, S. W. et al. Promoter of lncRNA gene PVT1 is a tumor-suppressor DNA boundary element. Cell 173, 1398–1412.e22 (2018). This study shows that a DNA element within the promoter of the lncRNA PVT1 regulates the expression of MYC in a lncRNA-independent manner.
Lee, J. S. & Mendell, J. T. Antisense-mediated transcript knockdown triggers premature transcription termination. Mol. Cell 77, 1044–1054.e3 (2020).
Lai, F., Damle, S. S., Ling, K. K. & Rigo, F. Directed RNase H cleavage of nascent transcripts causes transcription termination. Mol. Cell 77, 1032–1043.e4 (2020). Together with Lee and Mendell (2020), this paper shows that the RNase H-mediated cleavage of transcripts guided by ASOs leads to transcriptional termination, which should be considered when interpreting the transcription phenotypes.
Monia, B. P. et al. Evaluation of 2′-modified oligonucleotides containing 2′-deoxy gaps as antisense inhibitors of gene expression. J. Biol. Chem. 268, 14514–14522 (1993).
Seth, P. P. et al. Design, synthesis and evaluation of constrained methoxyethyl (cMOE) and constrained ethyl (cEt) nucleoside analogs. Nucleic Acids Symp. Ser. 52, 553–554 (2008).
Dassie, J. P. & Giangrande, P. H. Current progress on aptamer-targeted oligonucleotide therapeutics. Ther. Deliv. 4, 1527–1546 (2013).
Dhuri, K. et al. Antisense oligonucleotides: an emerging area in drug discovery and development. J. Clin. Med. 9, 2004 (2020).
Warner, K. D., Hajdin, C. E. & Weeks, K. M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 17, 547–558 (2018).
Hawkes, E. J. et al. COOLAIR antisense RNAs form evolutionarily conserved elaborate secondary structures. Cell Rep. 16, 3087–3096 (2016).
Novikova, I. V., Hennelly, S. P. & Sanbonmatsu, K. Y. Structural architecture of the human long non-coding RNA, steroid receptor RNA activator. Nucleic Acids Res. 40, 5034–5051 (2012).
Somarowthu, S. et al. HOTAIR forms an intricate and modular secondary structure. Mol. Cell 58, 353–361 (2015).
Perez-Pinera, P. et al. Synergistic and tunable human gene activation by combinations of synthetic transcription factors. Nat. Methods 10, 239–242 (2013).
Kim, D. N. et al. Zinc-finger protein CNBP alters the 3-D structure of lncRNA Braveheart in solution. Nat. Commun. 11, 148 (2020).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Perez-Pinera, P. et al. RNA-guided gene activation by CRISPR–Cas9-based transcription factors. Nat. Methods 10, 973–976 (2013).
Abudayyeh, O. O. et al. RNA targeting with CRISPR–Cas13. Nature 550, 280–284 (2017).
Esposito, R. et al. Hacking the cancer genome: profiling therapeutically actionable long non-coding RNAs using CRISPR–Cas9 screening. Cancer Cell 35, 545–557 (2019).
Kirk, J. M. et al. Functional classification of long non-coding RNAs by k-mer content. Nat. Genet. 50, 1474–1482 (2018). This study shows that lncRNAs can be classified into functional groups based on their content of 6-nucleotide k-mers, and demonstrates experimentally the k-mer dependence of gene repression by lncRNAs.
Arab, K. et al. Long noncoding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol. Cell 55, 604–614 (2014).
Kong, Y., Hsieh, C. H. & Alonso, L. C. ANRIL: a lncRNA at the CDKN2A/B locus with roles in cancer and metabolic disease. Front. Endocrinol. 9, 405 (2018).
Mira-Bontenbal, H. & Gribnau, J. New Xist-interacting proteins in X-chromosome inactivation. Curr. Biol. 26, R338–R342 (2016).
Duszczyk, M. M., Wutz, A., Rybin, V. & Sattler, M. The Xist RNA A-repeat comprises a novel AUCG tetraloop fold and a platform for multimerization. RNA 17, 1973–1982 (2011).
Lu, Z., Carter, A. C. & Chang, H. Y. Mechanistic insights in X-chromosome inactivation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 372, 20160356 (2017).
Chaligne, R. & Heard, E. X-chromosome inactivation in development and cancer. FEBS Lett. 588, 2514–2522 (2014).
Yildirim, E. et al. Xist RNA is a potent suppressor of hematologic cancer in mice. Cell 152, 727–742 (2013).
Chen, D. L. et al. Long noncoding RNA XIST expedites metastasis and modulates epithelial–mesenchymal transition in colorectal cancer. Cell Death Dis. 8, e3011 (2017).
Marcho, C., Bevilacqua, A., Tremblay, K. D. & Mager, J. Tissue-specific regulation of Igf2r/Airn imprinting during gastrulation. Epigenetics Chromatin 8, 10 (2015).
Mohammad, F., Mondal, T., Guseva, N., Pandey, G. K. & Kanduri, C. Kcnq1ot1 noncoding RNA mediates transcriptional gene silencing by interacting with Dnmt1. Development 137, 2493–2499 (2010).
Higashimoto, K., Soejima, H., Saito, T., Okumura, K. & Mukai, T. Imprinting disruption of the CDKN1C/KCNQ1OT1 domain: the molecular mechanisms causing Beckwith–Wiedemann syndrome and cancer. Cytogenet. Genome Res. 113, 306–312 (2006).
Liu, Z., Chen, Q. & Hann, S. S. The functions and oncogenic roles of CCAT1 in human cancer. Biomed. Pharmacother. 115, 108943 (2019).
Sharma, U. et al. Long non-coding RNA TINCR as potential biomarker and therapeutic target for cancer. Life Sci. 257, 118035 (2020).
Huang, D. et al. NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat. Immunol. 19, 1112–1125 (2018).
Meng, Q., Ren, M., Li, Y. & Song, X. lncRNA-RMRP acts as an oncogene in lung cancer. PLoS ONE 11, e0164845 (2016).
Rosenbluh, J. et al. RMRP is a non-coding RNA essential for early murine development. PLoS ONE 6, e26270 (2011).
Dong, P. et al. Long non-coding RNA NEAT1: a novel target for diagnosis and therapy in human tumors. Front. Genet. 9, 471 (2018).
Adriaens, C. et al. p53 induces formation of NEAT1 lncRNA-containing paraspeckles that modulate replication stress response and chemosensitivity. Nat. Med. 22, 861–868 (2016).
Nakagawa, S. et al. The lncRNA Neat1 is required for corpus luteum formation and the establishment of pregnancy in a subpopulation of mice. Development 141, 4618–4627 (2014).
Engreitz, J. M. et al. RNA–RNA interactions enable specific targeting of noncoding RNAs to nascent pre-mRNAs and chromatin sites. Cell 159, 188–199 (2014).
Wilusz, J. E., Freier, S. M. & Spector, D. L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 135, 919–932 (2008).
Michalik, K. M. et al. Long noncoding RNA MALAT1 regulates endothelial cell function and vessel growth. Circ. Res. 114, 1389–1397 (2014).
Shi, X. et al. lncRNA FIRRE is activated by MYC and promotes the development of diffuse large B-cell lymphoma via Wnt/β-catenin signaling pathway. Biochem. Biophys. Res. Commun. 510, 594–600 (2019).
Hung, T. et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 43, 621–629 (2011).
Munschauer, M. et al. The NORAD lncRNA assembles a topoisomerase complex critical for genome stability. Nature 561, 132–136 (2018).
Yang, Z. et al. Noncoding RNA activated by DNA damage (NORAD): biologic function and mechanisms in human cancers. Clin. Chim. Acta 489, 5–9 (2019).
Kopp, F. et al. PUMILIO hyperactivity drives premature aging of Norad-deficient mice. eLife 8, e42650 (2019).
Marchese, F. P. et al. A long noncoding RNA regulates sister chromatid cohesion. Mol. Cell 63, 397–407 (2016).
Melo, C. A. et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell 49, 524–535 (2013).
Li, W. et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498, 516–520 (2013).
Krause, H. M. New and prospective roles for lncRNAs in organelle formation and function. Trends Genet. 34, 736–745 (2018).
Wang, X., McLachlan, J., Zamore, P. D. & Hall, T. M. Modular recognition of RNA by a human Pumilio-homology domain. Cell 110, 501–512 (2002).
Cai, Z. et al. RIC-seq for global in situ profiling of RNA–RNA spatial interactions. Nature https://doi.org/10.1038/s41586-020-2249-1 (2020). This paper presents 3D RNA interaction maps in human cells generated with RIC-seq showing lncRNA involvement in several structural interactions, including the connections between promoters and enhancers required for enhancer function.
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Flynn, R. L. et al. TERRA and hnRNPA1 orchestrate an RPA-to-POT1 switch on telomeric single-stranded DNA. Nature 471, 532–536 (2011).
Montero, J. J. et al. TERRA recruitment of Polycomb to telomeres is essential for histone trymethylation marks at telomeric heterochromatin. Nat. Commun. 9, 1548 (2018).
Deng, Z., Norseen, J., Wiedmer, A., Riethman, H. & Lieberman, P. M. TERRA RNA binding to TRF2 facilitates heterochromatin formation and ORC recruitment at telomeres. Mol. Cell 35, 403–413 (2009).
Balk, B. et al. Telomeric RNA–DNA hybrids affect telomere-length dynamics and senescence. Nat. Struct. Mol. Biol. 20, 1199–1205 (2013).
Arora, R. et al. RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat. Commun. 5, 5220 (2014).
Porro, A. et al. Functional characterization of the TERRA transcriptome at damaged telomeres. Nat. Commun. 5, 5379 (2014).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Blower, M. D. Centromeric transcription regulates Aurora-B localization and activation. Cell Rep. 15, 1624–1633 (2016).
Liu, H. et al. Mitotic transcription installs Sgo1 at centromeres to coordinate chromosome segregation. Mol. Cell 59, 426–436 (2015).
Donley, N., Smith, L. & Thayer, M. J. ASAR15, a cis-acting locus that controls chromosome-wide replication timing and stability of human chromosome 15. PLoS Genet. 11, e1004923 (2015).
Donley, N., Stoffregen, E. P., Smith, L., Montagna, C. & Thayer, M. J. Asynchronous replication, mono-allelic expression, and long range cis-effects of ASAR6. PLoS Genet. 9, e1003423 (2013).
Platt, E. J., Smith, L. & Thayer, M. J. L1 retrotransposon antisense RNA within ASAR lncRNAs controls chromosome-wide replication timing. J. Cell Biol. 217, 541–553 (2018).
Heskett, M., Smith, L. G., Spellman, P. & Thayer, M. Reciprocal monoallelic expression of ASAR lncRNA genes controls replication timing of human chromosome 6. RNA 26, 724–738 (2020).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Keskin, H., Meers, C. & Storici, F. Transcript RNA supports precise repair of its own DNA gene. RNA Biol. 13, 157–165 (2016).
Keskin, H. et al. Transcript-RNA-templated DNA recombination and repair. Nature 515, 436–439 (2014).
Chakraborty, A. et al. Classical non-homologous end-joining pathway utilizes nascent RNA for error-free double-strand break repair of transcribed genes. Nat. Commun. 7, 13049 (2016).
Wei, L. et al. DNA damage during the G0/G1 phase triggers RNA-templated, Cockayne syndrome B-dependent homologous recombination. Proc. Natl Acad. Sci. USA 112, E3495–E3504 (2015).
Michelini, F. et al. Damage-induced lncRNAs control the DNA damage response through interaction with DDRNAs at individual double-strand breaks. Nat. Cell Biol. 19, 1400–1411 (2017). This paper describes the production of DNA damage-induced lncRNAs by Pol II at DNA breaks, which serve as precursors for small RNAs and recruiters of 53BP1 for DNA repair.
Betts, J. A. et al. Long noncoding RNAs CUPID1 and CUPID2 mediate breast cancer risk at 11q13 by modulating the response to DNA damage. Am. J. Hum. Genet. 101, 255–266 (2017).
Sakthianandeswaren, A., Liu, S. & Sieber, O. M. Long noncoding RNA LINP1: scaffolding non-homologous end joining. Cell Death Discov. 2, 16059 (2016).
Chaudhary, R. et al. Prosurvival long noncoding RNA PINCR regulates a subset of p53 targets in human colorectal cancer cells by binding to Matrin 3. eLife 6, e23244 (2017).
Sharma, V. et al. A BRCA1-interacting lncRNA regulates homologous recombination. EMBO Rep. 16, 1520–1534 (2015).
Bharti, S. K. et al. Molecular functions and cellular roles of the ChlR1 (DDX11) helicase defective in the rare cohesinopathy Warsaw breakage syndrome. Cell. Mol. Life Sci. 71, 2625–2639 (2014).
Acknowledgements
M.H. is supported by Fondo Europeo de Desarrollo Regional/Ministerio de Ciencia, Innovación y Universidades — Agencia Estatal de Investigación grant BFU2017–82773-P, European Research Council Consolidator grant 771425 and Worldwide Cancer Research grant 20-0204. L.-L. C. is supported by National Natural Science Foundation of China grants 91940303, 31861143025 and 31725009 and HHMI International Research Scholar Program grant 55008728.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks Tetsuro Hirose, Rory Johnson and Igor Ulitsky for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Human GENCODE: https://www.gencodegenes.org/human/
Supplementary information
Glossary
- Enhancer RNAs
-
Relatively short (≤2 kb) long non-coding RNAs that are bidirectionally transcribed from enhancer regions, are not consistently spliced or polyadenylated and are rapidly degraded.
- Promoter upstream transcripts
-
Transcripts that are divergently transcribed upstream of active gene promoters and rapidly degraded by the RNA exosome complex.
- RNA exosome
-
A complex of ribonucleases that regulates the degradation of multiple types of RNA in eukaryotes.
- Integrator
-
A multi-protein complex associated with RNA polymerase II that modulates the fate of many nascent RNAs, including mRNAs, enhancer RNAs and promoter upstream transcripts.
- R-loops
-
RNA–DNA hybrids formed by Watson–Crick interactions predominantly in cis, but also in trans through homologous recombination.
- Exosomes
-
Small vesicles secreted from cells that contain various RNAs and proteins.
- Triple helices
-
RNA–DNA structures forming when a polypurine motif of an RNA interacts with the major groove of a DNA double helix through Hoogsteen or reverse Hoogsteen base pairing; stabilized by nucleosomal organization.
- Paraspeckles
-
Nuclear condensates enriched in several proteins, a subset of mRNAs and even some primary microRNAs, which are involved in various biological processes.
- Nuclear speckles
-
Nuclear domains in mammalian cells enriched in pre-mRNA splicing factors, located in inter-chromosomal regions.
- RNA in situ conformation sequencing
-
(RIC-seq). A method of determining intramolecular and intermolecular RNA–RNA interactions.
- Small nucleolar RNA-related lncRNAs
-
Unusually processed long non-coding RNAs (lncRNAs) that are derived from introns and contain small nucleolar RNAs at both ends.
- 5′ small nucleolar RNA-capped and 3′ polyadenylated lncRNAs
-
(SPAs). Unusually processed long non-coding RNAs (lncRNAs) that are derived from polycistronic transcripts, capped by small nucleolar RNAs at their 5′ end and polyadenylated at their 3′ end.
- Perinucleolar compartment
-
A nuclear body at the periphery of the nucleolus. The perinucleolar compartment is a dynamic structure and is highly enriched in RNA-binding proteins and RNA polymerase III.
- Nuclear stress bodies
-
Nuclear condensates that are formed under different stresses, such as heat shock and DNA damage.
- Serine and arginine-rich (SR) proteins
-
RNA binding proteins known as regulators of constitutive and alternative splicing.
- Antisense oligonucleotides
-
(ASOs). Single-stranded deoxyribonucleotides complementary to a target RNA. Antisense oligonucleotides can affect RNA expression through various mechanisms, including modulation of splicing, steric blockade and cleavage by RNase H.
- Fused aptamers
-
Chimeric molecules comprising at least one aptamer — a single-stranded synthetic oligonucleotide that folds into a defined architecture to specifically bind with high affinity to an RNA, a protein or another ligand — and another molecule with different biological properties.
- k-mer
-
A short (3–8 nucleotides) sequence motif, often mediating the interaction of nucleic acids with proteins or other molecules. k specifies the length of the motif.
Rights and permissions
About this article
Cite this article
Statello, L., Guo, CJ., Chen, LL. et al. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol 22, 96–118 (2021). https://doi.org/10.1038/s41580-020-00315-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-020-00315-9
This article is cited by
-
Cuproptosis-related ceRNA axis triggers cell proliferation and cell cycle through CBX2 in lung adenocarcinoma
BMC Pulmonary Medicine (2024)
-
Multiple transcriptome analyses reveal mouse testis developmental dynamics
BMC Genomics (2024)
-
Role of ferroptosis and ferroptosis-related long non'coding RNA in breast cancer
Cellular & Molecular Biology Letters (2024)
-
MicroRNAs and long non-coding RNAs during transcriptional regulation and latency of HIV and HTLV
Retrovirology (2024)
-
Lnc-PSMA8-1 activated by GEFT promotes rhabdomyosarcoma progression via upregulation of mTOR expression by sponging miR-144-3p
BMC Cancer (2024)