Abstract
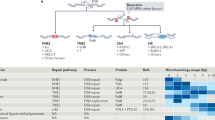
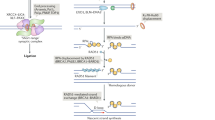
Non-homologous DNA end joining (NHEJ) is the predominant repair mechanism of any type of DNA double-strand break (DSB) during most of the cell cycle and is essential for the development of antigen receptors. Defects in NHEJ result in sensitivity to ionizing radiation and loss of lymphocytes. The most critical step of NHEJ is synapsis, or the juxtaposition of the two DNA ends of a DSB, because all subsequent steps rely on it. Recent findings show that, like the end processing step, synapsis can be achieved through several mechanisms. In this Review, we first discuss repair pathway choice between NHEJ and other DSB repair pathways. We then integrate recent insights into the mechanisms of NHEJ synapsis with updates on other steps of NHEJ, such as DNA end processing and ligation. Finally, we discuss NHEJ-related human diseases, including inherited disorders and neoplasia, which arise from rare failures at different NHEJ steps.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Daley, J. M., Niu, H., Miller, A. S. & Sung, P. Biochemical mechanism of DSB end resection and its regulation. DNA Repair 32, 66–74 (2015).
Kaniecki, K., De Tullio, L. & Greene, E. C. A change of view: homologous recombination at single-molecule resolution. Nat. Rev. Genet. 19, 191–207 (2018).
Wright, W. D., Shah, S. S. & Heyer, W.-D. Homologous recombination and the repair of DNA double-strand breaks. J. Biol. Chem. 293, 10524–10535 (2018).
Piazza, A. & Heyer, W.-D. Homologous recombination and the formation of complex genomic rearrangements. Trends Cell Biol. 29, 135–149 (2019).
Bell, J. C. & Kowalczykowski, S. C. Mechanics and single-molecule interrogation of DNA recombination. Annu. Rev. Biochem. 85, 193–226 (2016).
Scully, R., Panday, A., Elango, R. & Willis, N. A. DNA double-strand break repair-pathway choice in somatic mammalian cells. Nat. Rev. Mol. Cell Biol. 20, 698–714 (2019).
Chang, H. H. Y. et al. Different DNA end configurations dictate which NHEJ components are most important for joining efficiency. J. Biol. Chem. 291, 24377–24389 (2016). This article describes a comprehensive in vitro reconstitution of joining by NHEJ of diverse DNA end structures.
Waters, C. A. et al. The fidelity of the ligation step determines how ends are resolved during nonhomologous end joining. Nat. Commun. 5, 4286 (2014). This article provides evidence for a ligation-first hierarchy of attempted NHEJ factor engagement at DSBs, resulting in parsimony in the number of processing steps required for resolution.
Reid, D. A. et al. Organization and dynamics of the nonhomologous end-joining machinery during DNA double-strand break repair. Proc. Natl Acad. Sci. USA 112, E2575–E2584 (2015). This article reports the first reconstituted NHEJ synapsis using purified NHEJ proteins and using a single-molecule Förster resonance energy transfer method.
Zhao, B. et al. The essential elements for the noncovalent association of two DNA ends during NHEJ synapsis. Nat. Commun. 10, 3588 (2019). This article reports that two kinds of synaptic complexes (flexible synapsis and close synapsis) are formed for NHEJ synapsis, and that DNA-PKcs is not required for synapsis.
Zhao, B., Watanabe, G. & Lieber, M. R. Polymerase μ in non-homologous DNA end joining: importance of the order of arrival at a double-strand break in a purified system. Nucleic Acids Res. 48, 3605–3618 (2020). This article reports that Polμ can mediate NHEJ synapsis by itself, and describes for the first time that synapsis can be flexible.
Reid, D. A. et al. Bridging of double-stranded breaks by the nonhomologous end-joining ligation complex is modulated by DNA end chemistry. Nucleic Acids Res. 45, 1872–1878 (2017).
Conlin, M. P. et al. DNA ligase IV guides end-processing choice during nonhomologous end joining. Cell Rep. 20, 2810–2819 (2017). This article determines that tolerance of DNA ends with nucleotide damage requires XRCC4–LIG4-dependent remodelling of the ends within a synaptic complex, and that this is important for cell resistance to ionizing radiation.
Pannunzio, N. R., Watanabe, G. & Lieber, M. R. Nonhomologous DNA end joining for repair of DNA double-strand breaks. J. Biol. Chem. 293, 10512–10523 (2018).
Chang, H. H. Y., Pannunzio, N. R., Adachi, N. & Lieber, M. R. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 18, 495 (2017).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Beucher, A. et al. ATM and Artemis promote homologous recombination of radiation-induced DNA double-strand breaks in G2. EMBO J. 28, 3413–3427 (2009).
Rothkamm, K., Krüger, I., Thompson, L. H. & Löbrich, M. Pathways of DNA double-strand break repair during the mammalian cell cycle. Mol. Cell. Biol. 23, 5706–5715 (2003).
Karanam, K., Kafri, R., Loewer, A. & Lahav, G. Quantitative live cell imaging reveals a gradual shift between DNA repair mechanisms and a maximal use of HR in mid S phase. Mol. Cell 47, 320–329 (2012).
Shibata, A. et al. Factors determining DNA double-strand break repair pathway choice in G2 phase. EMBO J. 30, 1079–1092 (2011).
Schatz, D. G. & Swanson, P. C. V(D)J recombination: mechanisms of initiation. Annu. Rev. Genet. 45, 167–202 (2011).
Wang, X. S., Lee, B. J. & Zha, S. The recent advances in non-homologous end-joining through the lens of lymphocyte development. DNA Repair 94, 102874 (2020).
Taccioli, G. E. et al. Impairment of V(D)J recombination in double-strand break repair mutants. Science 260, 207–210 (1993).
Chaudhuri, J. & Alt, F. W. Class-switch recombination: interplay of transcription, DNA deamination and DNA repair. Nat. Rev. Immunol. 4, 541–552 (2004).
Mimitou, E. P. & Symington, L. S. Ku prevents Exo1 and Sgs1-dependent resection of DNA ends in the absence of a functional MRX complex or Sae2. EMBO J. 29, 3358–3369 (2010).
van Schendel, R., Roerink, S. F., Portegijs, V., van den Heuvel, S. & Tijsterman, M. Polymerase Θ is a key driver of genome evolution and of CRISPR/Cas9-mediated mutagenesis. Nat. Commun. 6, 7394 (2015).
van Kregten, M. et al. T-DNA integration in plants results from polymerase-θ-mediated DNA repair. Nat. Plants 2, 16164 (2016).
Chan, S. H., Yu, A. M. & McVey, M. Dual roles for DNA polymerase theta in alternative end-joining repair of double-strand breaks in Drosophila. PLoS Genet. 6, e1001005 (2010).
Saito, S., Maeda, R. & Adachi, N. Dual loss of human POLQ and LIG4 abolishes random integration. Nat. Commun. 8, 16112 (2017).
Schimmel, J., Kool, H., van Schendel, R. & Tijsterman, M. Mutational signatures of non-homologous and polymerase theta-mediated end-joining in embryonic stem cells. EMBO J. 36, 3634–3649 (2017).
Carvajal-Garcia, J. et al. Mechanistic basis for microhomology identification and genome scarring by polymerase theta. Proc. Natl Acad. Sci. USA 117, 8476–8485 (2020).
Wyatt, D. W. et al. Essential roles for polymerase θ-mediated end joining in the repair of chromosome breaks. Mol. Cell 63, 662–673 (2016).
He, P. & Yang, W. Template and primer requirements for DNA Pol θ-mediated end joining. Proc. Natl Acad. Sci. USA 115, 7747–7752 (2018).
Zhang, Y. & Jasin, M. An essential role for CtIP in chromosomal translocation formation through an alternative end-joining pathway. Nat. Struct. Mol. Biol. 18, 80–84 (2011).
Lee-Theilen, M., Matthews, A. J., Kelly, D., Zheng, S. & Chaudhuri, J. CtIP promotes microhomology-mediated alternative end joining during class-switch recombination. Nat. Struct. Mol. Biol. 18, 75–79 (2011).
Mateos-Gomez, P. A. et al. Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Bhargava, R., Onyango, D. O. & Stark, J. M. Regulation of single-strand annealing and its role in genome maintenance. Trends Genet. 32, 566–575 (2016).
Kelso, A. A., Lopezcolorado, F. W., Bhargava, R. & Stark, J. M. Distinct roles of RAD52 and POLQ in chromosomal break repair and replication stress response. PLoS Genet. 15, e1008319 (2019).
Motycka, T. A., Bessho, T., Post, S. M., Sung, P. & Tomkinson, A. E. Physical and functional interaction between the XPF/ERCC1 endonuclease and hRad52. J. Biol. Chem. 279, 13634–13639 (2004).
Lisby, M. & Rothstein, R. Cell biology of mitotic recombination. Cold Spring Harb. Perspect. Biol. 7, a016535 (2015).
Moore, J. K. & Haber, J. E. Cell cycle and genetic requirements of two pathways of nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae. Mol. Cell. Biol. 16, 2164–2173 (1996).
Gauss, G. H. & Lieber, M. R. Mechanistic constraints on diversity in human V(D)J recombination. Mol. Cell. Biol. 16, 258–269 (1996).
Chapman, J. R., Taylor, Martin, R. G. & Boulton, Simon J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Symington, L. S. & Gautier, J. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 45, 247–271 (2011).
Xu, G. T. et al. REV7 counteracts DNA double-strand break resection and affects PARP inhibition. Nature 521, 541–U308 (2015).
Boersma, V. et al. MAD2L2 controls DNA repair at telomeres and DNA breaks by inhibiting 5′ end resection. Nature 521, 537–540 (2015).
Zimmermann, M., Lottersberger, F., Buonomo, S. B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to Control 5′ end resection. Science 339, 700–704 (2013).
Escribano-Diaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol. Cell 49, 872–883 (2013).
Di Virgilio, M. et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339, 711–715 (2013).
Daley, J. M. & Sung, P. RIF1 in DNA break repair pathway choice. Mol. Cell 49, 840–841 (2013).
Chapman, J. R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol. Cell 49, 858–871 (2013).
Bunting, S. F. et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 141, 243–254 (2010).
Setiaputra, D. & Durocher, D. Shieldin – the protector of DNA ends. EMBO Rep. 20, e47560 (2019).
Tomida, J. et al. FAM35A associates with REV7 and modulates DNA damage responses of normal and BRCA1-defective cells. EMBO J. 37, e99543 (2018).
Noordermeer, S. M. et al. The shieldin complex mediates 53BP1-dependent DNA repair. Nature 560, 117–121 (2018).
Mirman, Z. et al. 53BP1-RIF1-shieldin counteracts DSB resection through CST- and Polalpha-dependent fill-in. Nature 560, 112–116 (2018).
Gupta, R. et al. DNA repair network analysis reveals shieldin as a key regulator of NHEJ and PARP inhibitor sensitivity. Cell 173, 972–988 e923 (2018).
Ghezraoui, H. et al. 53BP1 cooperation with the REV7–shieldin complex underpins DNA structure-specific NHEJ. Nature 560, 122–127 (2018).
Dev, H. et al. Shieldin complex promotes DNA end-joining and counters homologous recombination in BRCA1-null cells. Nat. Cell Biol. 20, 954–965 (2018).
Gao, S. et al. An OB-fold complex controls the repair pathways for DNA double-strand breaks. Nat. Commun. 9, 3925 (2018). Together with references 54–59, this article describes the discovery of the shieldin complex, which is the effector of 53BP1–RIF1.
Panier, S. & Boulton, S. J. Double-strand break repair: 53BP1 comes into focus. Nat. Rev. Mol. Cell Biol. 15, 7–18 (2014).
Botuyan, M. V. et al. Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006).
Fradet-Turcotte, A. et al. 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark. Nature 499, 50–54 (2013).
Ochs, F. et al. 53BP1 fosters fidelity of homology-directed DNA repair. Nat. Struct. Mol. Biol. 23, 714–721 (2016).
Tsai, L. J. et al. RNF8 has both KU-dependent and independent roles in chromosomal break repair. Nucleic Acids Res. 48, 6032–6052 (2020).
Guirouilh-Barbat, J. et al. 53BP1 protects against CtIP-dependent capture of ectopic chromosomal sequences at the junction of distant double-strand breaks. PLoS Genet. 12, e1006230 (2016).
Clairmont, C. S. et al. TRIP13 regulates DNA repair pathway choice through REV7 conformational change. Nat. Cell Biol. 22, 87–96 (2020).
Callen, E. et al. 53BP1 enforces distinct pre- and post-resection blocks on homologous recombination. Mol. Cell 77, 26–38.e27 (2020).
Ling, A. K. et al. SHLD2 promotes class switch recombination by preventing inactivating deletions within the Igh locus. EMBO Rep. 21, e49823 (2020).
Slavoff, S. A., Heo, J., Budnik, B. A., Hanakahi, L. A. & Saghatelian, A. A human short open reading frame (sORF)-encoded polypeptide that stimulates DNA end joining. J. Biol. Chem. 289, 10950–10957 (2014).
Arnoult, N. et al. Regulation of DNA repair pathway choice in S and G2 phases by the NHEJ inhibitor CYREN. Nature 549, 548–552 (2017).
Hung, P. J. et al. MRI is a DNA damage response adaptor during classical non-homologous end joining. Mol. Cell 71, 332–342 (2018).
Castañeda-Zegarra, S. et al. Generation of a mouse model lacking the non-homologous end-joining factor Mri/Cyren. Biomolecules 9, 798 (2019).
Tadi, Satish K. et al. PAXX is an accessory c-NHEJ factor that associates with Ku70 and has overlapping functions with XLF. Cell Rep. 17, 541–555 (2016).
Ahnesorg, P., Smith, P. & Jackson, S. P. XLF interacts with the XRCC4-DNA ligase IV complex to promote DNA nonhomologous end-joining. Cell 124, 301–313 (2006).
Lu, H., Pannicke, U., Schwarz, K. & Lieber, M. R. Length-dependent binding of human XLF to DNA and stimulation of XRCC4.DNA ligase IV activity. J. Biol. Chem. 282, 11155–11162 (2007).
Hammel, M., Yu, Y. P., Fang, S. J., Lees-Miller, S. P. & Tainer, J. A. XLF regulates filament architecture of the XRCC4·ligase IV complex. Structure 18, 1431–1442 (2010).
Yano, K.-i., Morotomi-Yano, K., Lee, K.-J. & Chen, D. J. Functional significance of the interaction with Ku in DNA double-strand break recognition of XLF. FEBS Lett. 585, 841–846 (2011).
Nemoz, C. et al. XLF and APLF bind Ku80 at two remote sites to ensure DNA repair by non-homologous end joining. Nat. Struct. Mol. Biol. 25, 971–980 (2018).
Ochi, T. et al. DNA repair. PAXX, a paralog of XRCC4 and XLF, interacts with Ku to promote DNA double-strand break repair. Science 347, 185–188 (2015).
Xing, M. et al. Interactome analysis identifies a new paralogue of XRCC4 in non-homologous end joining DNA repair pathway. Nat. Commun. 6, 6233 (2015). Together with reference 81, this srticle describes the discovery of PAXX.
Kumar, V., Alt, F. W. & Frock, R. L. PAXX and XLF DNA repair factors are functionally redundant in joining DNA breaks in a G1-arrested progenitor B-cell line. Proc. Natl Acad. Sci. USA 113, 10619–10624 (2016).
Li, W. et al. The nucleoskeleton protein IFFO1 immobilizes broken DNA and suppresses chromosome translocation during tumorigenesis. Nat. Cell Biol. 21, 1273–1285 (2019).
Graham, T. G., Walter, J. C. & Loparo, J. J. Two-stage synapsis of DNA ends during non-homologous end joining. Mol. Cell 61, 850–858 (2016).
Graham, T. G. W., Carney, S. M., Walter, J. C. & Loparo, J. J. A single XLF dimer bridges DNA ends during nonhomologous end joining. Nat. Struct. Mol. Biol. 25, 877–884 (2018).
Stinson, B. M., Moreno, A. T., Walter, J. C. & Loparo, J. J. A mechanism to minimize errors during non-homologous end joining. Mol. Cell 77, 1–12 (2020).
Wang, J. L. et al. Dissection of DNA double-strand-break repair using novel single-molecule forceps. Nat. Struct. Mol. Biol. 25, 482–487 (2018).
Kulesza, P. & Lieber, M. R. DNA-PK is essential only for coding joint formation in V(D)J recombination. Nucleic Acids Res. 26, 3944–3948 (1998).
Gao, Y. et al. A targeted DNA-PKcs-null mutation reveals DNA-PK-independent functions for KU in V(D)J recombination. Immunity 9, 367–376 (1998).
Jiang, W. et al. Differential phosphorylation of DNA-PKcs regulates the interplay between end-processing and end-ligation during nonhomologous end-joining. Mol. Cell 58, 172–185 (2015).
Davis, B. J., Havener, J. M. & Ramsden, D. A. End-bridging is required for pol μ to efficiently promote repair of noncomplementary ends by nonhomologous end joining. Nucleic Acids Res. 36, 3085–3094 (2008).
Andrade, P., Martín, M. J., Juárez, R., López de Saro, F. & Blanco, L. Limited terminal transferase in human DNA polymerase μ defines the required balance between accuracy and efficiency in NHEJ. Proc. Natl Acad. Sci. USA 106, 16203–16208 (2009).
Kaminski, A. M. et al. Structural snapshots of human DNA polymerase μ engaged on a DNA double-strand break. Nat. Commun. 11, 4784 (2020).
Loc’h, J. et al. Structural evidence for an in trans base selection mechanism involving Loop1 in polymerase μ at an NHEJ double-strand break junction. J. Biol. Chem. 294, 10579–10595 (2019).
Nick McElhinny, S. A. et al. A gradient of template dependence defines distinct biological roles for family X polymerases in nonhomologous end joining. Mol. Cell 19, 357–366 (2005).
Loc’h, J. & Delarue, M. Terminal deoxynucleotidyltransferase: the story of an untemplated DNA polymerase capable of DNA bridging and templated synthesis across strands. Curr. Opin. Struct. Biol. 53, 22–31 (2018).
Gouge, J. et al. Structural basis for a novel mechanism of DNA bridging and alignment in eukaryotic DSB DNA repair. EMBO J. 34, 1126–1142 (2015).
Loc’h, J., Rosario, S. & Delarue, M. Structural basis for a new templated activity by terminal deoxynucleotidyl transferase: implications for V(D)J recombination. Structure 24, 1452–1463 (2016).
Ma, Y. et al. A biochemically defined system for mammalian nonhomologous DNA end joining. Mol. Cell 16, 701–713 (2004).
Ma, Y., Lu, H., Schwarz, K. & Lieber, M. R. Repair of double-strand DNA breaks by the human nonhomologous DNA end joining pathway: the iterative processing model. Cell Cycle 4, 1193–2000 (2005).
Simsek, D. & Jasin, M. Alternative end-joining is suppressed by the canonical NHEJ component Xrcc4–ligase IV during chromosomal translocation formation. Nat. Struct. Mol. Biol. 17, 410–416 (2010).
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018).
Li, S. et al. Evidence that the DNA endonuclease ARTEMIS also has intrinsic 5′-exonuclease activity. J. Biol. Chem. 289, 7825–7834 (2014).
Ma, Y., Pannicke, U., Schwarz, K. & Lieber, M. R. Hairpin opening and overhang processing by an Artemis:DNA-PKcs complex in V(D)J recombination and in nonhomologous end joining. Cell 108, 781–794 (2002).
Gerodimos, C. A., Chang, H. H. Y., Watanabe, G. & Lieber, M. R. Effects of DNA end configuration on XRCC4:DNA ligase IV and its stimulation of Artemis activity. J. Biol. Chem. 292, 13914–13924 (2017).
Chang, H. H., Watanabe, G. & Lieber, M. R. Unifying the DNA end-processing roles of the Artemis nuclease: Ku-dependent Artemis resection at blunt DNA ends. J. Biol. Chem. 290, 24036–24050 (2015).
Lu, H. et al. A biochemically defined system for coding joint formation in human V(D)J recombination. Mol. Cell 31, 485–497 (2008).
Henner, W. D., Grunberg, S. M. & Haseltine, W. A. Enzyme action at 3′ termini of ionizing radiation-induced DNA strand breaks. J. Biol. Chem. 258, 15198–15205 (1983).
Henner, W. D., Rodriguez, L. O., Hecht, S. M. & Haseltine, W. A. Gamma ray induced deoxyribonucleic acid strand breaks. 3′ glycolate termini. J. Biol. Chem. 258, 711–713 (1983).
Valerie, K. & Povirk, L. F. Regulation and mechanisms of mammalian double-strand break repair. Oncogene 22, 5792–5812 (2003).
Zhou, T. et al. Deficiency in 3′-phosphoglycolate processing in human cells with a hereditary mutation in tyrosyl-DNA phosphodiesterase (TDP1). Nucleic Acids Res. 33, 289–297 (2005).
Povirk, L. F., Zhou, T., Zhou, R., Cowan, M. J. & Yannone, S. M. Processing of 3′-phosphoglycolate-terminated DNA double strand breaks by Artemis nuclease. J. Biol. Chem. 282, 3547–3558 (2007).
Yannone, S. M. et al. Coordinate 5′ and 3′ endonucleolytic trimming of terminally blocked blunt DNA double-strand break ends by Artemis nuclease and DNA-dependent protein kinase. Nucleic Acids Res. 36, 3354–3365 (2008).
Anne Esguerra, Z., Watanabe, G., Okitsu, C. Y., Hsieh, C.-L. & Lieber, M. R. DNA-PKcs chemical inhibition versus genetic mutation: impact on the junctional repair steps of V(D)J recombination. Mol. Immunol. 120, 93–100 (2020).
Moshous, D. et al. Artemis, a novel DNA double-strand break repair/V(D)J recombination protein, is mutated in human severe combined immune deficiency. Cell 105, 177–186 (2001).
Blunt, T. et al. Defective DNA-dependent protein kinase activity is linked to V(D)J recombination and DNA repair defects associated with the murine scid mutation. Cell 80, 813–823 (1995).
Woodbine, L. et al. PRKDC mutations in a SCID patient with profound neurological abnormalities. J. Clin. Invest. 123, 2969–2980 (2013).
Li, S. et al. Polynucleotide kinase and aprataxin-like forkhead-associated protein (PALF) acts as both a single-stranded DNA endonuclease and a single-stranded DNA 3′ exonuclease and can participate in DNA end joining in a biochemical system. J. Biol. Chem. 286, 36368–36377 (2011).
Pryor, J. M. et al. Essential role for polymerase specialization in cellular nonhomologous end joining. Proc. Natl Acad. Sci. USA 112, E4537–4545 (2015). This article demonstrates that in NHEJ, Polµ or Polλ is partially redundant; loss of both ablates almost all synthesis during NHEJ and severely impairs the pathway.
Mahajan, K. N., Nick McElhinny, S. A., Mitchell, B. S. & Ramsden, D. A. Association of DNA polymerase μ (pol μ) with Ku and ligase IV: role for pol μ in end-joining double-strand break repair. Mol. Cell. Biol. 22, 5194–5202 (2002).
Martin, M. J., Juarez, R. & Blanco, L. DNA-binding determinants promoting NHEJ by human Polµ. Nucleic Acids Res. 40, 11389–11403 (2012).
Gilfillan, S., Dierich, A., Lemeur, M., Benoist, C. & Mathis, D. Mice lacking TdT: mature animals with an immature lymphocyte repertoire. Science 261, 1175–1178 (1993).
Lange, S. S., Takata, K. & Wood, R. D. DNA polymerases and cancer. Nat. Rev. Cancer 11, 96–110 (2011).
Moon, A. F. et al. Structural insight into the substrate specificity of DNA polymerase μ. Nat. Struct. Mol. Biol. 14, 45–53 (2007).
Domínguez, O. et al. DNA polymerase mu (Pol mu), homologous to TdT, could act as a DNA mutator in eukaryotic cells. EMBO J. 19, 1731–1742 (2000).
Gu, J. et al. XRCC4:DNA ligase IV can ligate incompatible DNA ends and can ligate across gaps. EMBO J. 26, 1010–1023 (2007).
Moon, A. F. et al. The X family portrait: structural insights into biological functions of X family polymerases. DNA Repair 6, 1709–1725 (2007).
Kato, K. I., Gonçalves, J. M., Houts, G. E. & Bollum, F. J. Deoxynucleotide-polymerizing enzymes of calf thymus gland. II. Properties of the terminal deoxynucleotidyltransferase. J. Biol. Chem. 242, 2780–2789 (1967).
Boulé, J. B., Rougeon, F. & Papanicolaou, C. Terminal deoxynucleotidyl transferase indiscriminately incorporates ribonucleotides and deoxyribonucleotides. J. Biol. Chem. 276, 31388–31393 (2001).
Ruiz, J. F. et al. Lack of sugar discrimination by human Pol µ requires a single glycine residue. Nucleic Acids Res. 31, 4441–4449 (2003).
Martin, M. J., Garcia-Ortiz, M. V., Esteban, V. & Blanco, L. Ribonucleotides and manganese ions improve non-homologous end joining by human Polµ. Nucleic Acids Res. 41, 2428–2436 (2012).
Nick McElhinny, S. A. & Ramsden, D. A. Polymerase Mu is a DNA-directed DNA/RNA polymerase. Mol. Cell. Biol. 23, 2309–2315 (2003).
Pryor, J. M. et al. Ribonucleotide incorporation enables repair of chromosome breaks by nonhomologous end joining. Science 361, 1126–1129 (2018). This article shows that polymerases in cells very often incorporate ribonucleotides during NHEJ, and that this facilitates the ligation step.
Lieber, M. R., Hesse, J. E., Mizuuchi, K. & Gellert, M. Lymphoid V(D)J recombination: nucleotide insertion at signal joints as well as coding joints. Proc. Natl Acad. Sci. USA 85, 8588–8592 (1988).
Duvauchelle, J.-B., Blanco, L., Fuchs, R. P. P. & Cordonnier, A. M. Human DNA polymerase mu (Pol µ) exhibits an unusual replication slippage ability at AAF lesion. Nucleic Acids Res. 30, 2061–2067 (2002).
Tippin, B., Kobayashi, S., Bertram, J. G. & Goodman, M. F. To slip or skip, visualizing frameshift mutation dynamics for error-prone DNA polymerases. J. Biol. Chem. 279, 45360–45368 (2004).
Maga, G. et al. DNA elongation by the human DNA polymerase λ polymerase and terminal transferase activities are differentially coordinated by proliferating cell nuclear antigen and replication protein A. J. Biol. Chem. 280, 1971–1981 (2005).
Jäger, U. et al. Follicular lymphomas BCL-2/IgH junctions contain templated nucleotide insertions: novel insights into the mechanism of t(14;18) translocation. Blood 95, 3520–3529 (2000).
Welzel, N. et al. Templated nucleotide addition and immunoglobulin JH-gene utilization in t(11;14) junctions: implications for the mechanism of translocation and the origin of mantle cell lymphoma. Cancer Res. 61, 1629–1636 (2001).
Grawunder, U. et al. Activity of DNA ligase IV stimulated by complex formation with XRCC4 protein in mammalian cells. Nature 388, 492–495 (1997).
Wilson, T. E., Grawunder, U. & Lieber, M. R. Yeast DNA ligase IV mediates non-homologous DNA end joining. Nature 388, 495–498 (1997).
Schar, P., Herrmann, G., Daly, G. & Lindahl, T. A newly identified DNA ligase of Saccharomyces cerevisiae involved in RAD52-independent repair of DNA double-strand breaks. Genes Dev. 11, 1912–1924 (1997).
Chen, S.-H. & Yu, X. Human DNA ligase IV is able to use NAD+ as an alternative adenylation donor for DNA ends ligation. Nucleic Acids Res. 47, 1321–1334 (2019).
Liu, X., Shao, Z., Jiang, W., Lee, B. J. & Zha, S. PAXX promotes KU accumulation at DNA breaks and is essential for end-joining in XLF-deficient mice. Nat. Commun. 8, 13816 (2017).
Balmus, G. et al. Synthetic lethality between PAXX and XLF in mammalian development. Genes Dev. 30, 2152–2157 (2016).
Lescale, C. et al. Specific roles of XRCC4 paralogs PAXX and XLF during V(D)J recombination. Cell Rep. 16, 2967–2979 (2016).
Ropars, V. et al. Structural characterization of filaments formed by human Xrcc4-Cernunnos/XLF complex involved in nonhomologous DNA end-joining. Proc. Natl Acad. Sci. USA 108, 12663–12668 (2011).
Brouwer, I. et al. Sliding sleeves of XRCC4-XLF bridge DNA and connect fragments of broken DNA. Nature 535, 566–569 (2016).
Bermudez-Hernandez, K. et al. A method for quantifying molecular interactions using stochastic modelling and super-resolution microscopy. Sci. Rep. 7, 14882 (2017).
Wang, Y., Lamarche, B. J. & Tsai, M. D. Human DNA ligase IV and the ligase IV/XRCC4 complex: analysis of nick ligation fidelity. Biochemistry 46, 4962–4976 (2007).
Gu, J., Lu, H., Tsai, A. G., Schwarz, K. & Lieber, M. R. Single-stranded DNA ligation and XLF-stimulated incompatible DNA end ligation by the XRCC4-DNA ligase IV complex: influence of terminal DNA sequence. Nucleic Acids Res. 35, 5755–5762 (2007).
Tsai, C. J., Kim, S. A. & Chu, G. Cernunnos/XLF promotes the ligation of mismatched and noncohesive DNA ends. Proc. Natl Acad. Sci. USA 104, 7851–7856 (2007).
Roberts, S. A. & Ramsden, D. A. Loading of the nonhomologous end joining factor, Ku, on protein-occluded DNA ends. J. Biol. Chem. 282, 10605–10613 (2007).
Batenburg, N. L. et al. ATM and CDK2 control chromatin remodeler CSB to inhibit RIF1 in DSB repair pathway choice. Nat. Commun. 8, 1921 (2017).
Dos Santos, M. et al. Influence of chromatin condensation on the number of direct DSB damages induced by ions studied using a Monte Carlo code. Radiat. Prot. Dosimetry 161, 469–473 (2014).
Falk, M., Lukasova, E. & Kozubek, S. Higher-order chromatin structure in DSB induction, repair and misrepair. Mutat. Res. 704, 88–100 (2010).
Falk, M., Lukasova, E., Gabrielova, B., Ondrej, V. & Kozubek, S. Chromatin dynamics during DSB repair. Biochim. Biophys. Acta 1773, 1534–1545 (2007).
Altmeyer, M. et al. Liquid demixing of intrinsically disordered proteins is seeded by poly(ADP-ribose). Nat. Commun. 6, 8088 (2015).
Menolfi, D., Zha, S. & ATM, A. T. R. and DNA-PKcs kinases-the lessons from the mouse models: inhibition not equal deletion. Cell Biosci. 10, 8 (2020).
Stucki, M. & Jackson, S. P. gammaH2AX and MDC1: anchoring the DNA-damage-response machinery to broken chromosomes. DNA Repair 5, 534–543 (2006).
Doil, C. et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell 136, 435–446 (2009).
Stewart, G. S. et al. The RIDDLE syndrome protein mediates a ubiquitin-dependent signaling cascade at sites of DNA damage. Cell 136, 420–434 (2009).
Baranes-Bachar, K. et al. The ubiquitin E3/E4 ligase UBE4A adjusts protein ubiquitylation and accumulation at sites of DNA damage, facilitating double-strand break repair. Mol. Cell 69, 866–878 e867 (2018).
Kilic, S. et al. Phase separation of 53BP1 determines liquid-like behavior of DNA repair compartments. EMBO J. 38, e101379 (2019).
Ariumi, Y. et al. Suppression of the poly(ADP-ribose) polymerase activity by DNA-dependent protein kinase in vitro. Oncogene 18, 4616–4625 (1999).
Veuger, S. J., Curtin, N. J., Smith, G. C. & Durkacz, B. W. Effects of novel inhibitors of poly(ADP-ribose) polymerase-1 and the DNA-dependent protein kinase on enzyme activities and DNA repair. Oncogene 23, 7322–7329 (2004).
Hochegger, H. et al. Parp-1 protects homologous recombination from interference by Ku and ligase IV in vertebrate cells. EMBO J. 25, 1305–1314 (2006).
Wang, M. et al. PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res. 34, 6170–6182 (2006).
Cheng, Q. et al. Ku counteracts mobilization of PARP1 and MRN in chromatin damaged with DNA double-strand breaks. Nucleic Acids Res. 39, 9605–9619 (2011).
Frit, P., Barboule, N., Yuan, Y., Gomez, D. & Calsou, P. Alternative end-joining pathway(s): bricolage at DNA breaks. DNA Repair 17, 81–97 (2014).
Yang, G. et al. Super-resolution imaging identifies PARP1 and the Ku complex acting as DNA double-strand break sensors. Nucleic Acids Res. 46, 3446–3457 (2018).
Couto, C. A. et al. PARP regulates nonhomologous end joining through retention of Ku at double-strand breaks. J. Cell Biol. 194, 367–375 (2011).
Caron, M. C. et al. Poly(ADP-ribose) polymerase-1 antagonizes DNA resection at double-strand breaks. Nat. Commun. 10, 2954 (2019).
Lee, K. J. et al. Phosphorylation of Ku dictates DNA double-strand break (DSB) repair pathway choice in S phase. Nucleic Acids Res. 44, 1732–1745 (2016).
Kakarougkas, A. & Jeggo, P. A. DNA DSB repair pathway choice: an orchestrated handover mechanism. Br. J. Radiol. 87, 20130685 (2014).
Gigi, V. et al. RAG2 mutants alter DSB repair pathway choice in vivo and illuminate the nature of ‘alternative NHEJ’. Nucleic Acids Res. 42, 6352–6364 (2014).
Delacote, F. & Lopez, B. S. Importance of the cell cycle phase for the choice of the appropriate DSB repair pathway, for genome stability maintenance: the trans-S double-strand break repair model. Cell Cycle 7, 33–38 (2008).
Pellegrino, S., Michelena, J., Teloni, F., Imhof, R. & Altmeyer, M. Replication-coupled dilution of H4K20me2 guides 53BP1 to pre-replicative chromatin. Cell Rep. 19, 1819–1831 (2017).
Ochs, F. et al. Stabilization of chromatin topology safeguards genome integrity. Nature 574, 571–574 (2019).
Spies, J. et al. 53BP1 nuclear bodies enforce replication timing at under-replicated DNA to limit heritable DNA damage. Nat. Cell Biol. 21, 487–497 (2019).
Hu, J. et al. Chromosomal loop domains direct the recombination of antigen receptor genes. Cell 163, 947–959 (2015).
Lin, S. G., Ba, Z., Alt, F. W. & Zhang, Y. RAG chromatin scanning during V(D)J recombination and chromatin loop extrusion are related processes. Adv. Immunol. 139, 93–135 (2018).
Zhang, Y. et al. The fundamental role of chromatin loop extrusion in physiological V(D)J recombination. Nature 573, 600–604 (2019).
Zhang, X. et al. Fundamental roles of chromatin loop extrusion in antibody class switching. Nature 575, 385–389 (2019).
Lottersberger, F., Karssemeijer, R. A., Dimitrova, N. & de Lange, T. 53BP1 and the LINC complex promote microtubule-dependent DSB mobility and DNA repair. Cell 163, 880–893 (2015).
Schrank, B. R. et al. Nuclear ARP2/3 drives DNA break clustering for homology-directed repair. Nature 559, 61–66 (2018).
Dimitrova, N., Chen, Y. C., Spector, D. L. & de Lange, T. 53BP1 promotes non-homologous end joining of telomeres by increasing chromatin mobility. Nature 456, 524–528 (2008).
Raschellà, G., Melino, G. & Malewicz, M. New factors in mammalian DNA repair—the chromatin connection. Oncogene 36, 4673–4681 (2017).
Mirman, Z. & de Lange, T. 53BP1: a DSB escort. Genes Dev. 34, 7–23 (2020).
Dutertre, M. & Vagner, S. DNA-damage response RNA-binding proteins (DDRBPs): perspectives from a new class of proteins and their RNA targets. J. Mol. Biol. 429, 3139–3145 (2017).
D’Alessandro, G. & d’Adda di Fagagna, F. Transcription and DNA damage: holding hands or crossing swords? J. Mol. Biol. 429, 3215–3229 (2017).
D’Alessandro, G. et al. BRCA2 controls DNA:RNA hybrid level at DSBs by mediating RNase H2 recruitment. Nat. Commun. 9, 5376 (2018).
Chakraborty, A. et al. Classical non-homologous end-joining pathway utilizes nascent RNA for error-free double-strand break repair of transcribed genes. Nat. Commun. 7, 13049 (2016).
Zhang, Y. et al. Long noncoding RNA LINP1 regulates repair of DNA double-strand breaks in triple-negative breast cancer. Nat. Struct. Mol. Biol. 23, 522–530 (2016).
Michelini, F. et al. Damage-induced lncRNAs control the DNA damage response through interaction with DDRNAs at individual double-strand breaks. Nat. Cell Biol. 19, 1400–1411 (2017).
Pessina, F. et al. Functional transcription promoters at DNA double-strand breaks mediate RNA-driven phase separation of damage-response factors. Nat. Cell Biol. 21, 1286–1299 (2019).
Jaafar, L., Li, Z., Li, S. & Dynan, W. S. SFPQ*NONO and XLF function separately and together to promote DNA double-strand break repair via canonical nonhomologous end joining. Nucleic Acids Res. 45, 1848–1859 (2017).
Li, S. et al. Cell-type specific role of the RNA-binding protein, NONO, in the DNA double-strand break response in the mouse testes. DNA Repair 51, 70–78 (2017).
Nasmyth, K. & Haering, C. H. Cohesin: its roles and mechanisms. Annu. Rev. Genet. 43, 525–558 (2009).
Uhlmann, F. SMC complexes: from DNA to chromosomes. Nat. Rev. Mol. Cell Biol. 17, 399–412 (2016).
Bauerschmidt, C. et al. Cohesin promotes the repair of ionizing radiation-induced DNA double-strand breaks in replicated chromatin. Nucleic Acids Res. 38, 477–487 (2009).
Kim, J.-S., Krasieva, T. B., LaMorte, V., Taylor, A. M. R. & Yokomori, K. Specific recruitment of human cohesin to laser-induced DNA damage. J. Biol. Chem. 277, 45149–45153 (2002).
Kong, X. et al. Distinct functions of human cohesin-SA1 and cohesin-SA2 in double-strand break repair. Mol. Cell. Biol. 34, 685–698 (2014).
Caron, P. et al. Cohesin protects genes against γH2AX induced by DNA double-strand breaks. PLoS Genet. 8, e1002460 (2012).
Schär, P., Fäsi, M. & Jessberger, R. SMC1 coordinates DNA double-strand break repair pathways. Nucleic Acids Res. 32, 3921–3929 (2004).
Potts, P. R., Porteus, M. H. & Yu, H. Human SMC5/6 complex promotes sister chromatid homologous recombination by recruiting the SMC1/3 cohesin complex to double-strand breaks. EMBO J. 25, 3377–3388 (2006).
Enervald, E. et al. A regulatory role for the cohesin loader NIPBL in nonhomologous end joining during immunoglobulin class switch recombination. J. Exp. Med. 210, 2503–2513 (2013).
Thomas-Claudepierre, A.-S. et al. The cohesin complex regulates immunoglobulin class switch recombination. J. Exp. Med. 210, 2495–2502 (2013).
Gelot, C. et al. The cohesin complex prevents the end joining of distant DNA double-strand ends. Mol. Cell 61, 15–26 (2016).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Ba, Z. et al. CTCF orchestrates long-range cohesin-driven V(D)J recombinational scanning. Nature https://doi.org/10.1038/s41586-41020-42578-41580 (2020).
Zhu, C., Bogue, M. A., Lim, D.-S., Hasty, P. & Roth, D. B. Ku86-deficient mice exhibit severe combined immunodeficiency and defective processing of V(D)J recombination intermediates. Cell 86, 379–389 (1996).
Taccioli, G. E. et al. Targeted disruption of the catalytic subunit of the DNA-PK gene in mice confers severe combined immunodeficiency and radiosensitivity. Immunity 9, 355–366 (1998).
Rooney, S. et al. Leaky scid phenotype associated with defective V(D)J coding end processing in Artemis-deficient mice. Mol. Cell 10, 1379–1390 (2002).
Frank, K. M. et al. Late embryonic lethality and impaired V (D)J recombination in mice lacking DNA ligase IV. Nature 396, 173–177 (1998).
Altmann, T. & Gennery, A. R. DNA ligase IV syndrome; a review. Orphanet J. Rare Dis. 11, 137–137 (2016).
Saito, S., Kurosawa, A. & Adachi, N. Mutations in XRCC4 cause primordial dwarfism without causing immunodeficiency. J. Hum. Genet. 61, 679–685 (2016).
Woodbine, L., Gennery, A. R. & Jeggo, P. A. The clinical impact of deficiency in DNA non-homologous end-joining. DNA Repair 16, 84–96 (2014).
Li, G. et al. Lymphocyte-specific compensation for XLF/Cernunnos end-joining functions in V(D)J recombination. Mol. Cell 31, 631–640 (2008).
Grawunder, U., Zimmer, D. & Lieber, M. R. DNA ligase IV binds to XRCC4 via a motif located between rather than within its BRCT domains. Curr. Biol. 8, 873–876 (1998).
Gao, Y. et al. A critical role for DNA end-joining proteins in both lymphogenesis and neurogenesis. Cell 95, 891–902 (1998).
Bee, L. et al. A nonsense mutation of human XRCC4 is associated with adult-onset progressive encephalocardiomyopathy. EMBO Mol. Med. 7, 918–929 (2015).
de Villartay, J.-P. When natural mutants do not fit our expectations: the intriguing case of patients with XRCC4 mutations revealed by whole-exome sequencing. EMBO Mol. Med. 7, 862–864 (2015).
Cui, X. et al. Both CpG Methylation and AID are required for the fragility of the human Bcl-2 major breakpoint region: implications for the timing of the breaks in the t(14;18). Mol. Cell Biol. 33, 947–957 (2013).
Daley, J. M., Laan, R. L., Suresh, A. & Wilson, T. E. DNA joint dependence of pol X family polymerase action in nonhomologous end joining. J. Biol. Chem. 280, 29030–29037 (2005).
Meek, K. Activation of DNA-PK by hairpinned DNA ends reveals a stepwise mechanism of kinase activation. Nucleic Acids Res. 48, 9098–9108 (2020).
Bétermier, M., Bertrand, P. & Lopez, B. S. Is non-homologous end-joining really an inherently error-prone process? PLoS Genet. 10, e1004086 (2014).
Bhargava, R. et al. C-NHEJ without indels is robust and requires synergistic function of distinct XLF domains. Nat. Commun. 9, 2484 (2018).
Felgentreff, K. et al. Functional analysis of naturally occurring DCLRE1C mutations and correlation with the clinical phenotype of ARTEMIS deficiency. J. Allergy Clin. Immunol. 136, 140–150.e7 (2015).
Ru, H., Zhang, P. & Wu, H. Structural gymnastics of RAG-mediated DNA cleavage in V(D)J recombination. Curr. Opin. Struct. Biol. 53, 178–186 (2018).
Chen, X. et al. Cutting antiparallel DNA strands in a single active site. Nat. Struct. Mol. Biol. 27, 119–126 (2020).
Corneo, B. et al. Rag mutations reveal robust alternative end joining. Nature 449, 483–486 (2007).
Acknowledgements
This work was supported by US National Institutes of Health grants (GM118009, CA196671, CA100504 and P30 CA014089 to M.R.L.; CA097096 and CA84442 to D.A.R; and R01 GM108119 to E.R.), American Cancer Society grant 130304-RSG-16-241-01-DMC (E.R.), and V Foundation for Cancer Research grant D2018-020 (E.R.).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks Sathees Raghavan, Dongyi Xu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Homologous recombination
-
(HR). An important DNA double-strand break repair mechanism, which usually requires long homologous sequences.
- Class switch recombination
-
(CSR). Recombination of the immunoglobulin heavy chain locus, which results in a switch of the expressed heavy chain isotype from IgM to IgA, IgE or IgG.
- Microhomology
-
Short stretches of base pairs of complementarity between two broken DNA ends.
- Synaptic complex
-
The complex formed by the two juxtaposed DNA ends of a double-strand break and related non-homologous end joining proteins.
- X polymerases
-
A subtype of DNA polymerases that includes terminal deoxynucleotidyl transferase, polymerase-µ (Polµ), Polλ and Polβ.
- N nucleotides
-
In V(D)J recombination, nucleotides added by the polymerase terminal deoxynucleotidyl transferase to the ends of coding DNA segments independently of a template.
- Inverted repeats
-
Nucleotides that are added at a DNA double-strand break repair junction and are sometimes copied inversely from either of the two broken DNA ends.
- T nucleotides
-
Nucleotides added at a DNA double-strand break repair junction, which are sometimes copied (directly or inversely) from either strand of either of the two broken DNA ends in a template-dependent manner.
Rights and permissions
About this article
Cite this article
Zhao, B., Rothenberg, E., Ramsden, D.A. et al. The molecular basis and disease relevance of non-homologous DNA end joining. Nat Rev Mol Cell Biol 21, 765–781 (2020). https://doi.org/10.1038/s41580-020-00297-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-020-00297-8
This article is cited by
-
RUVBL1 ubiquitination by DTL promotes RUVBL1/2-β-catenin-mediated transcriptional regulation of NHEJ pathway and enhances radiation resistance in breast cancer
Cell Death & Disease (2024)
-
APE1 promotes non-homologous end joining by initiating DNA double-strand break formation and decreasing ubiquitination of artemis following oxidative genotoxic stress
Journal of Translational Medicine (2023)
-
Recent advances and applications of CRISPR-Cas9 in cancer immunotherapy
Molecular Cancer (2023)
-
CRISPR/Cas9 system: recent applications in immuno-oncology and cancer immunotherapy
Experimental Hematology & Oncology (2023)
-
Extracellular vesicle-packaged circBIRC6 from cancer-associated fibroblasts induce platinum resistance via SUMOylation modulation in pancreatic cancer
Journal of Experimental & Clinical Cancer Research (2023)