Abstract
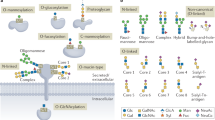
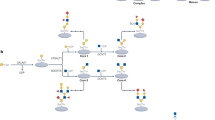
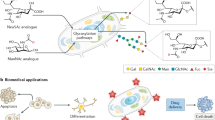
Glycosylation is the most abundant and diverse form of post-translational modification of proteins that is common to all eukaryotic cells. Enzymatic glycosylation of proteins involves a complex metabolic network and different types of glycosylation pathways that orchestrate enormous amplification of the proteome in producing diversity of proteoforms and its biological functions. The tremendous structural diversity of glycans attached to proteins poses analytical challenges that limit exploration of specific functions of glycosylation. Major advances in quantitative transcriptomics, proteomics and nuclease-based gene editing are now opening new global ways to explore protein glycosylation through analysing and targeting enzymes involved in glycosylation processes. In silico models predicting cellular glycosylation capacities and glycosylation outcomes are emerging, and refined maps of the glycosylation pathways facilitate genetic approaches to address functions of the vast glycoproteome. These approaches apply commonly available cell biology tools, and we predict that use of (single-cell) transcriptomics, genetic screens, genetic engineering of cellular glycosylation capacities and custom design of glycoprotein therapeutics are advancements that will ignite wider integration of glycosylation in general cell biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Fournet, M., Bonté, F. & Desmoulière, A. Glycation damage: a possible hub for major pathophysiological disorders and aging. Aging Dis. 9, 880–900 (2018).
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 32, 1478–1488 (2013). This paper presents a deep analysis of the human GalNAc-type O-glycoproteome.
Zielinska, D. F., Gnad, F., Wiśniewski, J. R. & Mann, M. Precision mapping of an in vivo N-glycoproteome reveals rigid topological and sequence constraints. Cell 141, 897–907 (2010). This paper presents a deep analysis of N-glycosites in the human proteome.
Hart, G. W. Nutrient regulation of signaling and transcription. J. Biol. Chem. 294, 2211–2231 (2019).
Hart, G. W. et al. Glycosylation of Nuclear and Cytoplasmic Proteins is as Abundant and as Dynamic as Phosphorylation. in Glyco- and Cellbiology (eds Wieland, F. & Reutter, W.) 91–103 (Springer, 1994).
Aebersold, R. et al. How many human proteoforms are there? Nat. Chem. Biol. 14, 206–214 (2018).
Varki, A. Biological roles of glycans. Glycobiology 27, 3–49 (2017).
Spiro, R. G. Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12, 43R–56R (2002).
Cummings, R. D. The repertoire of glycan determinants in the human glycome. Mol. Biosyst. 5, 1087–1104 (2009).
Joshi, H. J. et al. SnapShot: O-glycosylation pathways across kingdoms. Cell 172, 632–632.e2 (2018).
Springer, S. A. & Gagneux, P. Glycomics: revealing the dynamic ecology and evolution of sugar molecules. J. Proteom. 135, 90–100 (2016).
Chou, H. H. et al. A mutation in human CMP-sialic acid hydroxylase occurred after the Homo-Pan divergence. Proc. Natl Acad. Sci. USA 95, 11751–11756 (1998).
Larsen, R. D., Rivera-Marrero, C. A., Ernst, L. K., Cummings, R. D. & Lowe, J. B. Frameshift and nonsense mutations in a human genomic sequence homologous to a murine UDP-Gal:β-d-Gal(1,4)-d-GlcNAc α(1,3)-galactosyltransferase cDNA. J. Biol. Chem. 265, 7055–7061 (1990).
Christiansen, D. et al. Humans lack iGb3 due to the absence of functional iGb3-synthase: implications for NKT cell development and transplantation. PLoS Biol. 6, e172 (2008).
Stanley, P. What have we learned from glycosyltransferase knockouts in mice? J. Mol. Biol. 428, 3166–3182 (2016).
Lowe, J. B. & Marth, J. D. A genetic approach to mammalian glycan function. Annu. Rev. Biochem. 72, 643–691 (2003).
Freeze, H. H., Chong, J. X., Bamshad, M. J. & Ng, B. G. Solving glycosylation disorders: fundamental approaches reveal complicated pathways. Am. J. Hum. Genet. 94, 161–175 (2014).
Jaeken, J. & Péanne, R. What is new in CDG? J. Inherit. Metab. Dis. 40, 569–586 (2017).
Moremen, K. W., Tiemeyer, M. & Nairn, A. V. Vertebrate protein glycosylation: diversity, synthesis and function. Nat. Rev. Mol. Cell Biol. 13, 448–462 (2012).
Joshi, H. J. et al. Glycosyltransferase genes that cause monogenic congenital disorders of glycosylation are distinct from glycosyltransferase genes associated with complex diseases. Glycobiology 28, 284–294 (2018).
Hansen, L. et al. A mutation map for human glycoside hydrolase genes. Glycobiology 30, 500–515 (2020).
Cummings, R. D. & Pierce, J. M. The challenge and promise of glycomics. Chem. Biol. 21, 1–15 (2014).
Kornfeld, R. & Kornfeld, S. Assembly of asparagine-linked oligosaccharides. Annu. Rev. Biochem. 54, 631–664 (1985).
Narimatsu, H. Human glycogene cloning: focus on β3-glycosyltransferase and β4-glycosyltransferase families. Curr. Opin. Struct. Biol. 16, 567–575 (2006).
Bennett, E. P. et al. Control of mucin-type O-glycosylation: a classification of the polypeptide GalNAc-transferase gene family. Glycobiology 22, 736–756 (2012).
Tsuji, S., Datta, A. K. & Paulson, J. C. Systematic nomenclature for sialyltransferases. Glycobiology 6, v–vii (1996).
Larsen, I. S. B. et al. Discovery of an O-mannosylation pathway selectively serving cadherins and protocadherins. Proc. Natl Acad. Sci. USA 114, 11163–11168 (2017).
Yoshida-Moriguchi, T. & Campbell, K. P. Matriglycan: a novel polysaccharide that links dystroglycan to the basement membrane. Glycobiology 25, 702–713 (2015).
Praissman, J. L. et al. The functional O-mannose glycan on α-dystroglycan contains a phospho-ribitol primed for matriglycan addition. eLife 5, e14473 (2016).
Hirata, T. et al. Identification of a Golgi GPI-N-acetylgalactosamine transferase with tandem transmembrane regions in the catalytic domain. Nat. Commun. 9, 405 (2018).
Cejas, R. B., Lorenz, V., Garay, Y. C. & Irazoqui, F. J. Biosynthesis of O-N-acetylgalactosamine glycans in the human cell nucleus. J. Biol. Chem. 294, 2997–3011 (2019).
Tu, L., Chen, L. & Banfield, D. K. A conserved N-terminal arginine-motif in GOLPH3-family proteins mediates binding to coatomer. Traffic 13, 1496–1507 (2012).
Liu, L., Doray, B. & Kornfeld, S. Recycling of Golgi glycosyltransferases requires direct binding to coatomer. Proc. Natl Acad. Sci. USA 115, 8984–8989 (2018).
Kuhn, P.-H. et al. Secretome analysis identifies novel signal peptide peptidase-like 3 (Sppl3) substrates and reveals a role of Sppl3 in multiple Golgi glycosylation pathways. Mol. Cell. Proteom. 14, 1584–1598 (2015).
Schjoldager, K. T.-B. G. et al. A systematic study of site-specific GalNAc-type O-glycosylation modulating proprotein convertase processing. J. Biol. Chem. 286, 40122–40132 (2011).
Shifley, E. T. & Cole, S. E. Lunatic fringe protein processing by proprotein convertases may contribute to the short protein half-life in the segmentation clock. Biochim. Biophys. Acta 1783, 2384–2390 (2008).
Paulson, J. C. & Colley, K. J. Glycosyltransferases. Structure, localization, and control of cell type-specific glycosylation. J. Biol. Chem. 264, 17615–17618 (1989).
Liefhebber, J. M., Punt, S., Spaan, W. J. & van Leeuwen, H. C. The human collagen β(1-O)galactosyltransferase, GLT25D1, is a soluble endoplasmic reticulum localized protein. BMC Cell Biol. 11, 33 (2010).
Harvey, B. M. & Haltiwanger, R. S. Regulation of Notch function by O-glycosylation. Adv. Exp. Med. Biol. 1066, 59–78 (2018).
Ogawa, M. et al. GTDC2 modifies O-mannosylated α-dystroglycan in the endoplasmic reticulum to generate N-acetyl glucosamine epitopes reactive with CTD110.6 antibody. Biochem. Biophys. Res. Commun. 440, 88–93 (2013).
Snider, M. D. & Rogers, O. C. Intracellular movement of cell surface receptors after endocytosis: resialylation of asialo-transferrin receptor in human erythroleukemia cells. J. Cell Biol. 100, 826–834 (1985).
Duncan, J. R. & Kornfeld, S. Intracellular movement of two mannose 6-phosphate receptors: return to the Golgi apparatus. J. Cell Biol. 106, 617–628 (1988).
Litvinov, S. V. & Hilkens, J. The epithelial sialomucin, episialin, is sialylated during recycling. J. Biol. Chem. 268, 21364–21371 (1993).
Razawi, H. et al. Evidence for Core 2 to Core 1 O-glycan remodeling during the recycling of MUC1. Glycobiology 23, 935–45 (2013).
Gilmour, A. M. et al. A novel epidermal growth factor receptor-signaling platform and its targeted translation in pancreatic cancer. Cell Signal. 25, 2587–2603 (2013).
Haxho, F., Neufeld, R. J. & Szewczuk, M. R. Neuraminidase-1: a novel therapeutic target in multistage tumorigenesis. Oncotarget 7, 40860–40881 (2016).
Lillehoj, E. P. et al. NEU1 sialidase expressed in human airway epithelia regulates epidermal growth factor receptor (EGFR) and MUC1 protein signaling. J. Biol. Chem. 287, 8214–8231 (2012).
Welch, L. G. & Munro, S. A tale of short tails, through thick and thin: investigating the sorting mechanisms of Golgi enzymes. FEBS Lett. 593, 2452–2465 (2019).
Kellokumpu, S., Hassinen, A. & Glumoff, T. Glycosyltransferase complexes in eukaryotes: long-known, prevalent but still unrecognized. Cell. Mol. Life Sci. 73, 305–325 (2016).
Moremen, K. W. & Haltiwanger, R. S. Emerging structural insights into glycosyltransferase-mediated synthesis of glycans. Nat. Chem. Biol. 15, 853–864 (2019).
de Las Rivas, M., Lira-Navarrete, E., Gerken, T. A. & Hurtado-Guerrero, R. Polypeptide GalNAc-Ts: from redundancy to specificity. Curr. Opin. Struct. Biol. 56, 87–96 (2019).
Taujale, R. et al. Deep evolutionary analysis reveals the design principles of fold A glycosyltransferases. eLife 9, e54532 (2020).
Kinoshita, T. & Fujita, M. Biosynthesis of GPI-anchored proteins: special emphasis on GPI lipid remodeling. J. Lipid Res. 57, 6–24 (2016).
Wild, R. et al. Structure of the yeast oligosaccharyltransferase complex gives insight into eukaryotic N-glycosylation. Science 359, 545–550 (2018).
Ruiz-Canada, C., Kelleher, D. J. & Gilmore, R. Cotranslational and posttranslational N-glycosylation of polypeptides by distinct mammalian OST isoforms. Cell 136, 272–283 (2009).
Cherepanova, N. A. & Gilmore, R. Mammalian cells lacking either the cotranslational or posttranslocational oligosaccharyltransferase complex display substrate-dependent defects in asparagine linked glycosylation. Sci. Rep. 6, 20946 (2016).
Ramírez, A. S., Kowal, J. & Locher, K. P. Cryo–electron microscopy structures of human oligosaccharyltransferase complexes OST-A and OST-B. Science 366, 1372–1375 (2019).
Harada, Y., Masahara-Negishi, Y. & Suzuki, T. Cytosolic-free oligosaccharides are predominantly generated by the degradation of dolichol-linked oligosaccharides in mammalian cells. Glycobiology 25, 1196–1205 (2015).
Lu, H. et al. Mammalian STT3A/B oligosaccharyltransferases segregate N-glycosylation at the translocon from lipid-linked oligosaccharide hydrolysis. Proc. Natl Acad. Sci. USA 115, 9557–9562 (2018).
Shan, A. et al. Polypeptide N-acetylgalactosaminyltransferase 18 non-catalytically regulates the ER homeostasis and O-glycosylation. Biochim. Biophys. Acta Gen. Subj. 1863, 870–882 (2019).
Joshi, H. J. et al. GlycoDomainViewer: a bioinformatics tool for contextual exploration of glycoproteomes. Glycobiology 28, 131–136 (2018).
Schjoldager, K. T. et al. Deconstruction of O-glycosylation–GalNAc-T isoforms direct distinct subsets of the O-glycoproteome. EMBO Rep. 16, 1713–1722 (2015).
Narimatsu, Y. et al. Exploring regulation of protein O-glycosylation in isogenic human HEK293 cells by differential O-glycoproteomics. Mol. Cell. Proteom. 18, 1396–1409 (2019).
Bagdonaite, I. et al. O-glycan initiation directs distinct biological pathways and controls epithelial differentiation. EMBO Rep. 21, e48885 (2020).
Wang, S. et al. Site-specific O-glycosylation of members of the low-density lipoprotein receptor superfamily enhances ligand interactions. J. Biol. Chem. 293, 7408–7422 (2018). This paper describes a role of site-specific O-glycosylation in regulation of the affinity of LDLR-related receptors.
Tagliabracci, V. S. et al. A single kinase generates the majority of the secreted phosphoproteome. Cell 161, 1619–1632 (2015).
Tagliabracci, V. S. et al. Dynamic regulation of FGF23 by Fam20C phosphorylation, GalNAc-T3 glycosylation, and furin proteolysis. Proc. Natl Acad. Sci. USA 111, 5520–5 (2014). This paper is the first demonstration of cross-talk between extracellular protein phosphorylation and O-glycosylation.
Bordoli, M. R. et al. A secreted tyrosine kinase acts in the extracellular environment. Cell 158, 1033–1044 (2014).
Mehta, A. Y., Heimburg-Molinaro, J., Cummings, R. D. & Goth, C. K. Emerging patterns of tyrosine sulfation and O-glycosylation cross-talk and co-localization. Curr. Opin. Struct. Biol. 62, 102–111 (2020).
Yu, H. & Takeuchi, H. Protein O-glucosylation: another essential role of glucose in biology. Curr. Opin. Struct. Biol. 56, 64–71 (2019).
Holdener, B. C. & Haltiwanger, R. S. Protein O-fucosylation: structure and function. Curr. Opin. Struct. Biol. 56, 78–86 (2019).
Ogawa, M. & Okajima, T. Structure and function of extracellular O-GlcNAc. Curr. Opin. Struct. Biol. 56, 72–77 (2019).
Takeuchi, H. et al. Two novel protein O-glucosyltransferases that modify sites distinct from POGLUT1 and affect Notch trafficking and signaling. Proc. Natl Acad. Sci. USA 115, E8395–E8402 (2018). This paper describes the identification and differential functions of POGLUT isoenzymes in glycosylation of NOTCH EGF-like repeats and their regulation of NOTCH functions.
Sakaidani, Y. et al. O-linked-N-acetylglucosamine modification of mammalian Notch receptors by an atypical O-GlcNAc transferase Eogt1. Biochem. Biophys. Res. Commun. 419, 14–19 (2012).
Manya, H. et al. Demonstration of mammalian protein O-mannosyltransferase activity: coexpression of POMT1 and POMT2 required for enzymatic activity. Proc. Natl Acad. Sci. USA 101, 500–505 (2004).
Neubert, P. et al. Mapping the O-mannose glycoproteome in Saccharomyces cerevisiae. Mol. Cell. Proteom. 15, 1323–1337 (2016).
Shcherbakova, A., Tiemann, B., Buettner, F. F. R. & Bakker, H. Distinct C-mannosylation of netrin receptor thrombospondin type 1 repeats by mammalian DPY19L1 and DPY19L3. Proc. Natl Acad. Sci. USA 114, 2574–2579 (2017). This paper describes the DPY19L isoenzymes directing C-mannosylation and identifies distinct differences in their substrate preferences.
Roch, C., Kuhn, J., Kleesiek, K. & Götting, C. Differences in gene expression of human xylosyltransferases and determination of acceptor specificities for various proteoglycans. Biochem. Biophys. Res. Commun. 391, 685–691 (2010).
Noborn, F. et al. Identification of chondroitin sulfate linkage region glycopeptides reveals prohormones as a novel class of proteoglycans. Mol. Cell. Proteom. 14, 41–49 (2015).
Hennet, T. Collagen glycosylation. Curr. Opin. Struct. Biol. 56, 131–138 (2019).
Scietti, L. et al. Molecular architecture of the multifunctional collagen lysyl hydroxylase and glycosyltransferase LH3. Nat. Commun. 9, 3163 (2018).
Slawson, C. & Hart, G. W. O-GlcNAc signalling: implications for cancer cell biology. Nat. Rev. Cancer 11, 678–684 (2011).
Lazarus, M. B., Nam, Y., Jiang, J., Sliz, P. & Walker, S. Structure of human O-GlcNAc transferase and its complex with a peptide substrate. Nature 469, 564–567 (2011).
Tsuji, S., Datta, A. K. & Paulson, J. C. Systematic nomenclature for sialyltransferases. Glycobiology 6, v–vii (1996).
Oriol, R., Mollicone, R., Cailleau, A., Balanzino, L. & Breton, C. Divergent evolution of fucosyltransferase genes from vertebrates, invertebrates, and bacteria. Glycobiology 9, 323–334 (1999).
Nagae, M., Yamaguchi, Y., Taniguchi, N. & Kizuka, Y. 3D structure and function of glycosyltransferases involved in N-glycan maturation. Int J. Mol. Sci. 21, 437 (2020).
Honke, K. & Taniguchi, N. Sulfotransferases and sulfated oligosaccharides. Med. Res. Rev. 22, 637–654 (2002).
Esko, J. D. & Selleck, S. B. Order out of chaos: assembly of ligand binding sites in heparan sulfate. Annu. Rev. Biochem. 71, 435–471 (2002).
Mikami, T. & Kitagawa, H. Biosynthesis and function of chondroitin sulfate. Biochim. Biophys. Acta 1830, 4719–4733 (2013).
Kitayama, K., Hayashida, Y., Nishida, K. & Akama, T. O. Enzymes responsible for synthesis of corneal keratan sulfate glycosaminoglycans. J. Biol. Chem. 282, 30085–30096 (2007).
Yoshida-Moriguchi, T. et al. SGK196 is a glycosylation-specific O-mannose kinase required for dystroglycan function. Science 341, 896–899 (2013).
Sheikh, M. O., Halmo, S. M. & Wells, L. Recent advancements in understanding mammalian O-mannosylation. Glycobiology 27, 806–819 (2017).
Koike, T., Izumikawa, T., Tamura, J.-I. & Kitagawa, H. FAM20B is a kinase that phosphorylates xylose in the glycosaminoglycan–protein linkage region. Biochem. J. 421, 157–162 (2009).
Wen, J. et al. Xylose phosphorylation functions as a molecular switch to regulate proteoglycan biosynthesis. Proc. Natl Acad. Sci. USA 111, 15723–15728 (2014).
Duan, S. & Paulson, J. C. Siglecs as immune cell checkpoints in disease. Annu. Rev. Immunol. 38, 365–395 (2020).
Baumann, A.-M. T. et al. 9-O-Acetylation of sialic acids is catalysed by CASD1 via a covalent acetyl-enzyme intermediate. Nat. Commun. 6, 7673 (2015).
Orizio, F. et al. Human sialic acid acetyl esterase: towards a better understanding of a puzzling enzyme. Glycobiology 25, 992–1006 (2015).
Vlasak, R., Luytjes, W., Spaan, W. & Palese, P. Human and bovine coronaviruses recognize sialic acid-containing receptors similar to those of influenza C viruses. Proc. Natl Acad. Sci. USA 85, 4526–4529 (1988).
Barb, A. W. & Prestegard, J. H. NMR analysis demonstrates immunoglobulin G N-glycans are accessible and dynamic. Nat. Chem. Biol. 7, 147–153 (2011).
Krapp, S., Mimura, Y., Jefferis, R., Huber, R. & Sondermann, P. Structural analysis of human IgG-Fc glycoforms reveals a correlation between glycosylation and structural integrity. J. Mol. Biol. 325, 979–989 (2003).
Bowden, T. A. et al. Chemical and structural analysis of an antibody folding intermediate trapped during glycan biosynthesis. J. Am. Chem. Soc. 134, 17554–17563 (2012).
Patel, K. R., Roberts, J. T. & Barb, A. W. Multiple variables at the leukocyte cell surface impact Fcγ receptor-dependent mechanisms. Front. Immunol. 10, 223 (2019).
Ye, Z., Mao, Y., Clausen, H. & Vakhrushev, S. Y. Glyco-DIA: a method for quantitative O-glycoproteomics with in silico-boosted glycopeptide libraries. Nat. Methods 16, 902–910 (2019).
Kornfeld, S. Trafficking of lysosomal enzymes in normal and disease states. J. Clin. Invest. 77, 1–6 (1986).
Reitman, M. L. & Kornfeld, S. Lysosomal enzyme targeting. N-Acetylglucosaminylphosphotransferase selectively phosphorylates native lysosomal enzymes. J. Biol. Chem. 256, 11977–11980 (1981).
Schnaar, R. L., Gerardy-Schahn, R. & Hildebrandt, H. Sialic acids in the brain: gangliosides and polysialic acid in nervous system development, stability, disease, and regeneration. Physiol. Rev. 94, 461–518 (2014).
Bhide, G. P., Prehna, G., Ramirez, B. E. & Colley, K. J. The polybasic region of the polysialyltransferase ST8Sia-IV binds directly to the neural cell adhesion molecule, NCAM. Biochemistry 56, 1504–1517 (2017).
Bhide, G. P., Fernandes, N. R. J. & Colley, K. J. Sequence requirements for neuropilin-2 recognition by ST8SiaIV and polysialylation of its O-glycans. J. Biol. Chem. 291, 9444–9457 (2016).
Marcus, D. M. & Cass, L. E. Glycosphingolipids with Lewis blood group activity: uptake by human erythrocytes. Science 164, 553–555 (1969).
Parry, S. et al. The sperm agglutination antigen-1 (SAGA-1) glycoforms of CD52 are O-glycosylated. Glycobiology 17, 1120–1126 (2007).
Wandall, H. H. et al. The origin and function of platelet glycosyltransferases. Blood 120, 626–635 (2012).
Manhardt, C. T., Punch, P. R., Dougher, C. W. L. & Lau, J. T. Y. Extrinsic sialylation is dynamically regulated by systemic triggers in vivo. J. Biol. Chem. 292, 13514–13520 (2017).
Jones, M. B. et al. B-cell-independent sialylation of IgG. Proc. Natl Acad. Sci. USA 113, 7207–7212 (2016).
Irons, E. E. et al. B cells suppress medullary granulopoiesis by an extracellular glycosylation-dependent mechanism. eLife 8, e47328 (2019).
Zhang, Q. et al. Transfer of functional cargo in exomeres. Cell Rep. 27, 940–954.e6 (2019).
Lu, Q., Li, S. & Shao, F. Sweet talk: protein glycosylation in bacterial interaction with the host. Trends Microbiol. 23, 630–641 (2015).
El Qaidi, S. et al. NleB/SseK effectors from Citrobacter rodentium, Escherichia coli, and Salmonella enterica display distinct differences in host substrate specificity. J. Biol. Chem. 292, 11423–11430 (2017).
Li, S. et al. Pathogen blocks host death receptor signalling by arginine GlcNAcylation of death domains. Nature 501, 242–246 (2013).
Helenius, A. & Aebi, M. Roles of N-linked glycans in the endoplasmic reticulum. Annu. Rev. Biochem. 73, 1019–1049 (2004).
Vasudevan, D., Takeuchi, H., Johar, S. S., Majerus, E. & Haltiwanger, R. S. Peters plus syndrome mutations disrupt a noncanonical ER quality-control mechanism. Curr. Biol. 25, 286–295 (2015).
Takeuchi, H. et al. O-Glycosylation modulates the stability of epidermal growth factor-like repeats and thereby regulates Notch trafficking. J. Biol. Chem. 292, 15964–15973 (2017).
Ogawa, M., Tashima, Y., Sakaguchi, Y., Takeuchi, H. & Okajima, T. Contribution of extracellular O-GlcNAc to the stability of folded epidermal growth factor-like domains and Notch1 trafficking. Biochem. Biophys. Res. Commun. 526, 184–190 (2020).
Goth, C. K., Vakhrushev, S. Y., Joshi, H. J., Clausen, H. & Schjoldager, K. T. Fine-tuning limited proteolysis: a major role for regulated site-specific O-glycosylation. Trends Biochem. Sci. 43, 269–284 (2018).
Goth, C. K. et al. A systematic study of modulation of ADAM-mediated ectodomain shedding by site-specific O-glycosylation. Proc. Natl Acad. Sci. USA 112, 14623–14628 (2015).
Goth, C. K. et al. Site-specific O-glycosylation by polypeptide N-acetylgalactosaminyltransferase 2 (GalNAc-transferase T2) co-regulates β1-adrenergic receptor N-terminal cleavage. J. Biol. Chem. 292, 4714–4726 (2017).
Hansen, L. H. et al. Discovery of O-glycans on atrial natriuretic peptide (ANP) that affect both its proteolytic degradation and potency at its cognate receptor. J. Biol. Chem. 294, 12567–12578 (2019).
Madsen, T. D. et al. An atlas of O-linked glycosylation on peptide hormones reveals diverse biological roles. Nat. Commun. 11, 4033 (2020).
Hintze, J. et al. Probing the contribution of individual polypeptide GalNAc-transferase isoforms to the O-glycoproteome by inducible expression in isogenic cell lines. J. Biol. Chem. 293, 19064–19077 (2018).
Kato, K. et al. Polypeptide GalNAc-transferase T3 and familial tumoral calcinosis. Secretion of fibroblast growth factor 23 requires O-glycosylation. J. Biol. Chem. 281, 18370–18377 (2006).
Takashi, Y. et al. Activation of unliganded FGF receptor by extracellular phosphate potentiates proteolytic protection of FGF23 by its O-glycosylation. Proc. Natl Acad. Sci. USA 116, 11418–11427 (2019).
Larsen, I. S. B., Narimatsu, Y., Clausen, H., Joshi, H. J. & Halim, A. Multiple distinct O-mannosylation pathways in eukaryotes. Curr. Opin. Struct. Biol. 56, 171–178 (2019).
Rexach, J. E. et al. Dynamic O-GlcNAc modification regulates CREB-mediated gene expression and memory formation. Nat. Chem. Biol. 8, 253–261 (2012).
Tarbet, H. J. et al. Site-specific glycosylation regulates the form and function of the intermediate filament cytoskeleton. eLife 7, e31807 (2018).
Dennis, J. W. & Brewer, C. F. Density-dependent lectin-glycan interactions as a paradigm for conditional regulation by posttranslational modifications. Mol. Cell. Proteom. 12, 913–20 (2013).
Granovsky, M. et al. Suppression of tumor growth and metastasis in Mgat5-deficient mice. Nat. Med. 6, 306–312 (2000).
Martínez Allo, V. C. et al. Suppression of age-related salivary gland autoimmunity by glycosylation-dependent galectin-1-driven immune inhibitory circuits. Proc. Natl Acad. Sci. USA 117, 6630–6639 (2020).
Demetriou, M., Nabi, I. R., Coppolino, M., Dedhar, S. & Dennis, J. W. Reduced contact-inhibition and substratum adhesion in epithelial cells expressing GlcNAc-transferase V. J. Cell Biol. 130, 383–392 (1995).
Nakano, M. et al. Bisecting GlcNAc is a general suppressor of terminal modification of N-glycan. Mol. Cell. Proteom. 18, 2044–2057 (2019).
Isaji, T. et al. Introduction of bisecting GlcNAc into integrin α5β1 reduces ligand binding and down-regulates cell adhesion and cell migration. J. Biol. Chem. 279, 19747–19754 (2004).
Wang, X. et al. Core fucosylation regulates epidermal growth factor receptor-mediated intracellular signaling. J. Biol. Chem. 281, 2572–2577 (2006).
Liang, W. et al. Core fucosylation of the T cell receptor is required for T cell activation. Front. Immunol. 9, 78 (2018).
Shields, R. L. et al. Lack of fucose on human IgG1 N-linked oligosaccharide improves binding to human FcγRIII and antibody-dependent cellular toxicity. J. Biol. Chem. 277, 26733–26740 (2002). This paper identifies core fucose on the N-glycans of IgG1 in regulation of antibody effector functions.
Nguyen, J. T. et al. CD45 modulates galectin-1-induced T cell death: regulation by expression of Core 2 O-glycans. J. Immunol. 167, 5697–5707 (2001).
Lee, S. H. et al. Core2 O-glycan structure is essential for the cell surface expression of sucrase isomaltase and dipeptidyl peptidase-IV during intestinal cell differentiation. J. Biol. Chem. 285, 37683–37692 (2010).
Halmo, S. M. et al. Protein O-linked mannose β-1,4-N-acetylglucosaminyl-transferase 2 (POMGNT2) is a gatekeeper enzyme for functional glycosylation of α-dystroglycan. J. Biol. Chem. 292, 2101–2109 (2017). This paper describes the peptide substrate selectivity of the ER-located POMGNT2 that determines the O-Man glycosites that proceed towards biosynthesis of the elaborated matriglycan.
Varki, A. Glycan-based interactions involving vertebrate sialic-acid-recognizing proteins. Nature 446, 1023–1029 (2007).
Cohen, M. & Varki, A. Modulation of glycan recognition by clustered saccharide patches. Int. Rev. Cell Mol. Biol. 308, 75–125 (2014).
Giannini, S. et al. β4GALT1 controls β1 integrin function to govern thrombopoiesis and hematopoietic stem cell homeostasis. Nat. Commun. 11, 356 (2020).
Hennet, T., Chui, D., Paulson, J. C. & Marth, J. D. Immune regulation by the ST6Gal sialyltransferase. Proc. Natl Acad. Sci. USA 95, 4504–4509 (1998).
Comelli, E. M. et al. Activation of murine CD4+ and CD8+ T lymphocytes leads to dramatic remodeling of N-linked glycans. J. Immunol. 177, 2431–2440 (2006).
Priatel, J. J. et al. The ST3Gal-I sialyltransferase controls CD8+ T lymphocyte homeostasis by modulating O-glycan biosynthesis. Immunity 12, 273–283 (2000).
Ohtsubo, K. & Marth, J. D. Glycosylation in cellular mechanisms of health and disease. Cell 126, 855–867 (2006).
Mereiter, S., Balmaña, M., Campos, D., Gomes, J. & Reis, C. A. Glycosylation in the era of cancer-targeted therapy: where are we heading? Cancer Cell 36, 6–16 (2019).
Ng, B. G. & Freeze, H. H. Perspectives on glycosylation and its congenital disorders. Trends Genet. 34, 466–476 (2018).
Zilmer, M. et al. Novel congenital disorder of O-linked glycosylation caused by loss of function of GALNT2. Brain 143, 1114–1126 (2020).
Topaz, O. et al. Mutations in GALNT3, encoding a protein involved in O-linked glycosylation, cause familial tumoral calcinosis. Nat. Genet. 36, 579–581 (2004).
Tian, E. et al. Galnt11 regulates kidney function by glycosylating the endocytosis receptor megalin to modulate ligand binding. Proc. Natl Acad. Sci. USA 116, 25196–25202 (2019).
Hansen, L. et al. A glycogene mutation map for discovery of diseases of glycosylation. Glycobiology 25, 211–224 (2015).
Willer, C. J. et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat. Genet. 40, 161–169 (2008).
Khetarpal, S. A. et al. Loss of function of GALNT2 lowers high-density lipoproteins in humans, nonhuman primates, and rodents. Cell Metab. 24, 234–245 (2016). This paper describes validation of the first glycosyltransferase gene implicated as a GWAS candidate.
Roman, T. S. et al. Multiple hepatic regulatory variants at the GALNT2 GWAS locus associated with high-density lipoprotein cholesterol. Am. J. Hum. Genet. 97, 801–815 (2015).
Cavalli, M., Pan, G., Nord, H. & Wadelius, C. Looking beyond GWAS: allele-specific transcription factor binding drives the association of GALNT2 to HDL-C plasma levels. Lipids Health Dis. 15, 18 (2016).
Duncan, E. L. et al. Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLoS Genet. 7, e1001372 (2011).
de Las Rivas, M. et al. Molecular basis for fibroblast growth factor 23 O-glycosylation by GalNAc-T3. Nat. Chem. Biol. 16, 351–360 (2020).
Taniguchi, N. & Kizuka, Y. Glycans and cancer: role of N-glycans in cancer biomarker, progression and metastasis, and therapeutics. Adv. Cancer Res. 126, 11–51 (2015).
Schultz, M. J., Swindall, A. F. & Bellis, S. L. Regulation of the metastatic cell phenotype by sialylated glycans. Cancer Metastasis Rev. 31, 501–518 (2012).
Christiansen, M. N. et al. Cell surface protein glycosylation in cancer. Proteomics 14, 525–546 (2014).
Ju, T. et al. Human tumor antigens Tn and sialyl Tn arise from mutations in Cosmc. Cancer Res. 68, 1636–46 (2008).
Sun, X., Ju, T. & Cummings, R. D. Differential expression of Cosmc, T-synthase and mucins in Tn-positive colorectal cancers. BMC Cancer 18, 827 (2018).
Radhakrishnan, P. et al. Immature truncated O-glycophenotype of cancer directly induces oncogenic features. Proc. Natl Acad. Sci. USA 111, E4066–E4075 (2014).
Fernandez, A. J. et al. The structure of the colorectal cancer-associated enzyme GalNAc-T12 reveals how nonconserved residues dictate its function. Proc. Natl Acad. Sci. USA 116, 20404–20410 (2019).
Büll, C., Stoel, M. A., den Brok, M. H. & Adema, G. J. Sialic acids sweeten a tumor’s life. Cancer Res. 74, 3199–3204 (2014).
Dall’Olio, F. & Trinchera, M. Epigenetic bases of aberrant glycosylation in cancer. Int. J. Mol. Sci. 18, 998 (2017).
Ashkani, J. & Naidoo, K. J. Glycosyltransferase gene expression profiles classify cancer types and propose prognostic subtypes. Sci. Rep. 6, 26451 (2016).
Dusoswa, S. A. et al. Glioblastomas exploit truncated O-linked glycans for local and distant immune modulation via the macrophage galactose-type lectin. Proc. Natl Acad. Sci. USA 117, 3693–3703 (2020).
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Zeng, J. et al. Promoters of human Cosmc and T-synthase genes are similar in structure, yet different in epigenetic regulation. J. Biol. Chem. 290, 19018–19033 (2015).
Agrawal, P. et al. Mapping posttranscriptional regulation of the human glycome uncovers microRNA defining the glycocode. Proc. Natl Acad. Sci. USA 111, 4338–4343 (2014).
Kurcon, T. et al. miRNA proxy approach reveals hidden functions of glycosylation. Proc. Natl Acad. Sci. USA 112, 7327–7332 (2015).
Bhattacharyya, R., Bhaumik, M., Raju, T. S. & Stanley, P. Truncated, inactive N-acetylglucosaminyltransferase III (GlcNAc-TIII) induces neurological and other traits absent in mice that lack GlcNAc-TIII. J. Biol. Chem. 277, 26300–26309 (2002).
Matsumoto, K. et al. N-Glycan fucosylation of epidermal growth factor receptor modulates receptor activity and sensitivity to epidermal growth factor receptor tyrosine kinase inhibitor. Cancer Sci. 99, 1611–1617 (2008).
Agrawal, P. et al. A systems biology approach identifies FUT8 as a driver of melanoma metastasis. Cancer Cell 31, 804–819.e7 (2017).
Okada, M. et al. Blockage of core fucosylation reduces cell-surface expression of PD-1 and promotes anti-tumor immune responses of T cells. Cell Rep. 20, 1017–1028 (2017).
Marcos, N. T. et al. Role of the human ST6GalNAc-I and ST6GalNAc-II in the synthesis of the cancer-associated Sialyl-Tn antigen. Cancer Res. 64, 7050–7057 (2004).
Sewell, R. et al. The ST6GalNAc-I sialyltransferase localizes throughout the Golgi and is responsible for the synthesis of the tumor-associated sialyl-Tn O-glycan in human breast cancer. J. Biol. Chem. 281, 3586–3594 (2006).
Tarp, M. A. & Clausen, H. Mucin-type O-glycosylation and its potential use in drug and vaccine development. Biochim. Biophys. Acta 1780, 546–563 (2008).
Kudelka, M. R., Ju, T., Heimburg-Molinaro, J. & Cummings, R. D. Simple sugars to complex disease — mucin-type O-glycans in cancer. Adv. Cancer Res. 126, 53–135 (2015).
Sutherlin, M. E. et al. Expression of three UDP-N-acetyl-α-d-galactosamine:polypeptide GalNAc N-acetylgalactosaminyltransferases in adenocarcinoma cell lines. Cancer Res. 57, 4744–4748 (1997).
Freire, T. et al. UDP-N-acetyl-d-galactosamine:polypeptide N-acetylgalactosaminyltransferase 6 (ppGalNAc-T6) mRNA as a potential new marker for detection of bone marrow-disseminated breast cancer cells. Int. J. Cancer 119, 1383–1388 (2006).
Song, K.-H. et al. GALNT14 promotes lung-specific breast cancer metastasis by modulating self-renewal and interaction with the lung microenvironment. Nat. Commun. 7, 13796 (2016).
Kitada, S. et al. Polypeptide N-acetylgalactosaminyl transferase 3 independently predicts high-grade tumours and poor prognosis in patients with renal cell carcinomas. Br. J. Cancer 109, 472–481 (2013).
Lavrsen, K. et al. De novo expression of human polypeptide N-acetylgalactosaminyltransferase 6 (GalNAc-T6) in colon adenocarcinoma inhibits the differentiation of colonic epithelium. J. Biol. Chem. 293, 1298–1314 (2018).
Dalziel, M. et al. The relative activities of the C2GnT1 and ST3Gal-I glycosyltransferases determine O-glycan structure and expression of a tumor-associated epitope on MUC1. J. Biol. Chem. 276, 11007–11015 (2001).
Swindall, A. F. & Bellis, S. L. Sialylation of the Fas death receptor by ST6Gal-I provides protection against Fas-mediated apoptosis in colon carcinoma cells. J. Biol. Chem. 286, 22982–22990 (2011).
Holdbrooks, A. T., Britain, C. M. & Bellis, S. L. ST6Gal-I sialyltransferase promotes tumor necrosis factor (TNF)-mediated cancer cell survival via sialylation of the TNF receptor 1 (TNFR1) death receptor. J. Biol. Chem. 293, 1610–1622 (2018).
Barthel, S. R. et al. α1,3 Fucosyltransferases are master regulators of prostate cancer cell trafficking. Proc. Natl Acad. Sci. USA 106, 19491–19496 (2009).
Esposito, M. et al. Bone vascular niche E-selectin induces mesenchymal-epithelial transition and Wnt activation in cancer cells to promote bone metastasis. Nat. Cell Biol. 21, 627–639 (2019).
Engle, D. D. et al. The glycan CA19-9 promotes pancreatitis and pancreatic cancer in mice. Science 364, 1156–1162 (2019).
Guo, H. et al. O-Linked N-acetylglucosamine (O-GlcNAc) expression levels epigenetically regulate colon cancer tumorigenesis by affecting the cancer stem cell compartment via modulating expression of transcriptional factor MYBL1. J. Biol. Chem. 292, 4123–4137 (2017).
Hanover, J. A., Chen, W. & Bond, M. R. O-GlcNAc in cancer: an oncometabolism-fueled vicious cycle. J. Bioenerg. Biomembr. 50, 155–173 (2018).
Theocharis, A. D. & Karamanos, N. K. Proteoglycans remodeling in cancer: underlying molecular mechanisms. Matrix Biol. 75–76, 220–259 (2019).
Salanti, A. et al. Targeting human cancer by a glycosaminoglycan binding malaria protein. Cancer Cell 28, 500–514 (2015). This paper identifies a broadly expressed cancer-associated 4-O-sulfated chondroitin sulfate epitope using the malarial receptor VAR2CSA.
Chen, Y.-H. et al. The GAGOme: a cell-based library of displayed glycosaminoglycans. Nat. Methods 15, 881–888 (2018). This paper describes the development of a cell-based display of glycosaminoglycans.
Petrosyan, A. Unlocking Golgi: why does morphology matter? Biochemistry 84, 1490–1501 (2019).
Kulkarni-Gosavi, P., Makhoul, C. & Gleeson, P. A. Form and function of the Golgi apparatus: scaffolds, cytoskeleton and signalling. FEBS Lett. 593, 2289–2305 (2019).
Petrosyan, A. Onco-Golgi: is fragmentation a gate to cancer progression? Biochem. Mol. Biol. J. https://doi.org/10.21767/2471-8084.100006 (2015).
Chia, J., Goh, G. & Bard, F. Short O-GalNAc glycans: regulation and role in tumor development and clinical perspectives. Biochim. Biophys. Acta 1860, 1623–1639 (2016).
Gill, D. J., Clausen, H. & Bard, F. Location, location, location: new insights into O-GalNAc protein glycosylation. Trends Cell Biol. 21, 149–158 (2011).
Axelsson, M. A. B. et al. Neutralization of pH in the Golgi apparatus causes redistribution of glycosyltransferases and changes in the O-glycosylation of mucins. Glycobiology 11, 633–644 (2001).
Chatterjee, S. et al. Protein paucimannosylation is an enriched N-glycosylation signature of human cancers. Proteomics 19, e1900010 (2019).
Chia, J., Tay, F. & Bard, F. The GalNAc-T Activation (GALA) pathway: drivers and markers. PLoS ONE 14, e0214118 (2019).
Nguyen, A. T. et al. Organelle specific O-glycosylation drives MMP14 activation, tumor growth, and metastasis. Cancer Cell 32, 639–653.e6 (2017).
Yamaji, T. et al. A CRISPR screen using subtilase cytotoxin identifies SLC39A9 as a glycan-regulating factor. iScience 15, 407–420 (2019).
Narimatsu, Y. et al. A validated gRNA library for CRISPR/Cas9 targeting of the human glycosyltransferase genome. Glycobiology 28, 295–305 (2018).
Narimatsu, Y. et al. An atlas of human glycosylation pathways enables display of the human glycome by gene engineered cells. Mol. Cell 75, 394–407.e5 (2019). This paper describes the development and utility of a cell-based display of the human glycome.
Yang, Z. et al. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Nat. Biotechnol. 33, 842–844 (2015).
Patnaik, S. K. & Stanley, P. Lectin-resistant CHO glycosylation mutants. Methods Enzymol. 416, 159–182 (2006).
Steentoft, C. et al. Precision genome editing: a small revolution for glycobiology. Glycobiology 24, 663–680 (2014).
Jae, L. T. et al. Deciphering the glycosylome of dystroglycanopathies using haploid screens for lassa virus entry. Science 340, 479–483 (2013).
Han, J. et al. Genome-wide CRISPR/Cas9 screen identifies host factors essential for influenza virus replication. Cell Rep. 23, 596–607 (2018).
Ye, L. et al. In vivo CRISPR screening in CD8 T cells with AAV–Sleeping Beauty hybrid vectors identifies membrane targets for improving immunotherapy for glioblastoma. Nat. Biotechnol. 37, 1302–1313 (2019).
Miller, R. L. et al. Shotgun ion mobility mass spectrometry sequencing of heparan sulfate saccharides. Nat. Commun. 11, 1481 (2020).
Stopschinski, B. E. et al. Specific glycosaminoglycan chain length and sulfation patterns are required for cell uptake of tau versus α-synuclein and β-amyloid aggregates. J. Biol. Chem. 293, 10826–10840 (2018).
Tian, S. et al. Genome-wide CRISPR screens for Shiga toxins and ricin reveal Golgi proteins critical for glycosylation. PLoS Biol. 16, e2006951 (2018).
Dabelsteen, S. et al. Essential functions of glycans in human epithelia dissected by a CRISPR–Cas9-engineered human organotypic skin model. Dev. Cell 54, 669–684 (2020).
Büll, C., Joshi, H. J., Clausen, H. & Narimatsu, Y. Cell-based glycan arrays — a practical guide to dissect the human glycome. STAR Protoc. 1, 100017 (2020).
Naegeli, A. & Aebi, M. Current approaches to engineering N-linked protein glycosylation in bacteria. Methods Mol. Biol. 1321, 3–16 (2015).
Hudak, J. E. & Bertozzi, C. R. Glycotherapy: new advances inspire a reemergence of glycans in medicine. Chem. Biol. 21, 16–37 (2014).
Van Landuyt, L., Lonigro, C., Meuris, L. & Callewaert, N. Customized protein glycosylation to improve biopharmaceutical function and targeting. Curr. Opin. Biotechnol. 60, 17–28 (2019).
Montero-Morales, L. & Steinkellner, H. Advanced plant-based glycan engineering. Front. Bioeng. Biotechnol. 6, 81 (2018).
Walsh, G. Post-translational modifications of protein biopharmaceuticals. Drug Discov. Today 15, 773–780 (2010).
Čaval, T., Tian, W., Yang, Z., Clausen, H. & Heck, A. J. R. Direct quality control of glycoengineered erythropoietin variants. Nat. Commun. 9, 3342 (2018).
Grabowski, G. A. et al. Enzyme therapy in type 1 Gaucher disease: comparative efficacy of mannose-terminated glucocerebrosidase from natural and recombinant sources. Ann. Intern. Med. 122, 33–39 (1995).
Umaña, P., Jean-Mairet, J., Moudry, R., Amstutz, H. & Bailey, J. E. Engineered glycoforms of an antineuroblastoma IgG1 with optimized antibody-dependent cellular cytotoxic activity. Nat. Biotechnol. 17, 176–180 (1999).
Yamane-Ohnuki, N. et al. Establishment of FUT8 knockout Chinese hamster ovary cells: an ideal host cell line for producing completely defucosylated antibodies with enhanced antibody-dependent cellular cytotoxicity. Biotechnol. Bioeng. 87, 614–622 (2004).
Meuris, L. et al. GlycoDelete engineering of mammalian cells simplifies N-glycosylation of recombinant proteins. Nat. Biotechnol. 32, 485–489 (2014).
Chang, M. M. et al. Small-molecule control of antibody N-glycosylation in engineered mammalian cells. Nat. Chem. Biol. 15, 730–736 (2019).
Schulz, M. A. et al. Glycoengineering design options for IgG1 in CHO cells using precise gene editing. Glycobiology 28, 542–549 (2018).
Tian, W. et al. The glycosylation design space for recombinant lysosomal replacement enzymes produced in CHO cells. Nat. Commun. 10, 1785 (2019).
RodrÍguez, E., Schetters, S. T. T. & van Kooyk, Y. The tumour glyco-code as a novel immune checkpoint for immunotherapy. Nat. Rev. Immunol. 18, 204–211 (2018).
Mathiesen, C. B. K. et al. Genetically engineered cell factories produce glycoengineered vaccines that target antigen-presenting cells and reduce antigen-specific T-cell reactivity. J. Allergy Clin. Immunol. 142, 1983–1987 (2018).
Byrne, G. et al. CRISPR/Cas9 gene editing for the creation of an MGAT1-deficient CHO cell line to control HIV-1 vaccine glycosylation. PLoS Biol. 16, e2005817 (2018).
Zhang, R. et al. Reducing immunoreactivity of porcine bioprosthetic heart valves by genetically-deleting three major glycan antigens, GGTA1/β4GalNT2/CMAH. Acta Biomater. 72, 196–205 (2018).
Liu, Q. P. et al. Identification of a GH110 subfamily of α1,3-galactosidases: novel enzymes for removal of the α3Gal xenotransplantation antigen. J. Biol. Chem. 283, 8545–8554 (2008).
Ibrahim, A. M. S. et al. Acellular dermal matrices in breast surgery: a comprehensive review. Ann. Plast. Surg. 70, 732–738 (2013).
Neelamegham, S. et al. Updates to the Symbol Nomenclature for Glycans guidelines. Glycobiology 29, 620–624 (2019).
Parker, J. L. & Newstead, S. Gateway to the Golgi: molecular mechanisms of nucleotide sugar transporters. Curr. Opin. Struct. Biol. 57, 127–134 (2019).
Nabi, I. R., Shankar, J. & Dennis, J. W. The galectin lattice at a glance. J. Cell Sci. 128, 2213–2219 (2015).
Ilver, D. Helicobacter pylori adhesin binding fucosylated histo-blood group antigens revealed by retagging. Science 279, 373–377 (1998).
Mahdavi, J. Helicobacter pylori SabA adhesin in persistent infection and chronic inflammation. Science 297, 573–578 (2002).
Rossez, Y. et al. The LacdiNAc-specific adhesin LabA mediates adhesion of Helicobacter pylori to human gastric mucosa. J. Infect. Dis. 210, 1286–1295 (2014).
Nagae, M. et al. Structure and mechanism of cancer-associated N-acetylglucosaminyltransferase-V. Nat. Commun. 9, 3380 (2018).
Pinho, S. S. & Reis, C. A. Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer 15, 540–555 (2015).
Läubli, H. & Varki, A. Sialic acid-binding immunoglobulin-like lectins (Siglecs) detect self-associated molecular patterns to regulate immune responses. Cell. Mol. Life Sci. 77, 593–605 (2020).
Maxson, J. E. et al. Ligand-independence of the colony stimulating factor 3 receptor (CSF3R) T618I mutation results from loss of O-linked glycosylation and increased receptor dimerization. J. Biol. Chem. 289, 5820–5827.(2014).
Jiang, Y. et al. O-glycans on death receptors in cells modulate their sensitivity to TRAIL-induced apoptosis through affecting on their stability and oligomerization. FASEB J. 34, 11786–11801 (2020).
Rillahan, C. D. & Paulson, J. C. Glycan microarrays for decoding the glycome. Annu. Rev. Biochem. 80, 797–823 (2011).
Blixt, O. et al. Printed covalent glycan array for ligand profiling of diverse glycan binding proteins. Proc. Natl Acad. Sci. USA 101, 17033–17038 (2004).
Acknowledgements
The authors are grateful to R. Schnaar and B. Henrissat for discussions and critical comments on the manuscript. They thank H. Wandall, A. Halim, C. Büll, Y. Zhang and L. Hansen for help with the manuscript, and all members of the Copenhagen Center for Glycomics for discussions. Supported by the Lundbeck Foundation, the Novo Nordisk Foundation and the Danish National Research Foundation (DNRF107).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The University of Copenhagen has filed patent applications related to engineering cellular glycosylation capacities and cell-based glycan arrays. H.C. and Y.N. are named inventors on these applications. GlycoDisplay Aps has license rights to these patent applications, and H.C. and Y.N. hold shares in GlycoDisplay Aps. The remaining authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks S. Flitsch, C. Reis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Carbohydrate-Active enZYmes database: http://www.cazy.org
Copenhagen Center for Glycomics, University of Copenhagen: https://glycomics.ku.dk
Essentials of Glycobiology, 3rd edition: https://www.ncbi.nlm.nih.gov/books/NBK310274/
ExPADSy glycomics: https://www.expasy.org/glycomics
GlycoDomainViewer: https://glycodomain.glycomics.ku.dk
GlyGen: https://www.glygen.org
GlyTouCan: https://glytoucan.org
Handbook of Glycosyltransferases and Related Genes. Editors: Taniguchi, N., Honke, K., Fukuda, M., Narimatsu, H., Yamaguchi, Y., Angata, T. (Eds.) Springer Nature, 2014. Society for Glycobiology: http://glycobiology.org
International Glycoconjugate Organization: https://intl-glyco.org
NetOGlyc 4.0: http://www.cbs.dtu.dk/services/NetOGlyc/
Supplementary information
Glossary
- O-GlcNAcylation
-
The enzymatic process directed by the N-acetyl-d-glucosamine (GlcNAc) glycosyltransferase (OGT) that transfers GlcNAc to proteins (Ser and Thr residues) occurring in the cytosol and nucleus of cells.
- Isoenzymes
-
Enzymes that catalyse the same reactions but differ in amino acid sequence and often have partially distinct (non-redundant) functions.
- Glycosylphosphatidylinositol (GPI)-anchored glycoproteins
-
A class of proteins that are attached to the membrane lipid bilayer via a carboxy-terminal glycolipid anchor consisting of phosphoethanolamine, an oligosaccharide core and phosphatidylinositol.
- Proteoglycans
-
Proteins carrying one or more glycosaminoglycan chains attached covalently.
- Glycocalyx
-
The cell coat comprising glycans and glycoconjugates surrounding animal cells found as an electron-dense layer by electron microscopy. It protects the cell from physical stress and mediates a plethora of macromolecular and cell–cell interactions.
- Xenoantigens
-
Antigens found in multiple species that elicit antibodies in a species without the antigen after transplantation of tissues and organs. A major xenoantigen in porcine to human transplantation is the Galα1–3Galβ1–R glycan epitope (αGal).
- Lectins
-
Proteins that bind to glycans. Major animal lectin families include Galectin, C-type, P-type and I-type lectins. Lectins, mostly from plants, with well-characterized binding specificities are frequently used as tools in glycobiology. Lectins are often multivalent with binding affinities in the low micromolar range and binding avidities approaching the nanomolar range for larger glycans with multiple epitopes.
- Dolichol
-
(Dol). A polyisoprenol lipid that serves as an acceptor for the lipid-linked oligosaccharides in N-glycan biosynthesis.
- Type II transmembrane glycoproteins
-
Single-pass transmembrane glycoproteins with the amino terminus oriented towards the cytosol and the carboxyl terminus facing the lumen of the secretory pathway or cell exterior.
- COPI-coated vesicles
-
Coat protein complex I-coated vesicles that mediate intra-Golgi and Golgi-to-endoplasmic reticulum retrograde transport.
- Multipass transmembrane proteins
-
Proteins spanning the membrane more than once.
- KDEL signals
-
A carboxy-terminal Lys-Asp-Glu-Leu (KDEL) retention sequence found on endoplasmic reticulum (ER)-resident proteins. The KDEL receptors recognizing this signal facilitate the retrograde movement of ER-based proteins from the Golgi and back to the ER by coat protein complex I (COPI) vesicles.
- Sialylation
-
Modification by the addition of sialic acids, which are a large family of glycans derived from the neuraminic acid (Neu) monosaccharide with a nine-carbon backbone. In humans, N-acetylneuraminic acid (Neu5Ac) is the most common sialic acid, often found in the non-reducing terminal of glycoconjugates.
- O-Glycan core structures
-
The initiating O-GalNAc glycan can be extended to form four different common core structures. Core1, Galβ1–3GalNAcα1–O-Ser/Thr; Core2, GlcNAcβ1–6(Galβ1–3)GalNAcα1–O-Ser/Thr; Core3, GlcNAcβ1–3GalNAcα1–O-Ser/Thr; and Core4, GlcNAcβ1–6(GlcNAcβ1–3)GalNAcα1–O-Ser/Thr. The core structures can be further elongated or branched.
- Oligosaccharyltransferase (OST) complex
-
A membrane protein complex in the endoplasmic reticulum that transfers an oligosaccharide from a dolichol pyrophosphate-activated donor to N-linked acceptor sequences on secreted proteins.
- Translocon
-
A protein complex that mediates translocation of newly synthesized polypeptides from the cytosol across the endoplasmic reticulum membrane.
- Oxidoreductase
-
An enzyme that catalyses thiol–disulphide exchange reactions. In vivo oxidoreductases are important in the oxidative protein folding that takes place in the endoplasmic reticulum. Well-known examples are PDI and ERp57.
- Lectin domains
-
Carbohydrate-binding protein domains.
- Sulfation
-
An enzymatic process that transfers a sulfo group to another molecule, for example a glycan, by modifying a hydroxyl group on a monosaccharide by addition of a sulfo group. Sulfotransferases catalyse the reaction using 3′-phospho-5′-adenylyl sulfate (PAPS) as a donor.
- EGF-like repeats
-
Common motifs of 30–40 amino acids found in the extracellular domain of transmembrane proteins or in proteins known to be secreted. The epidermal growth factor (EGF)-like repeats include six conserved cysteines forming three disulfide bonds.
- Thrombospondin type 1 repeats
-
(TSRs) Common protein motifs of 50–60 amino acids (6 conserved cysteines forming 3 disulfide bonds) found on transmembrane proteins and proteins in the extracellular matrix.
- Mucin
-
A large viscous heavily O-glycosylated protein. Mucins are the most abundant macromolecule in biofluids and mucus, covering most epithelial surfaces in the body.
- α-Dystroglycan
-
Dystroglycan is encoded by the DAG1 gene and comprises two non-covalently-bound subunits (α and β). The extracellular α-subunit with the O-Man matriglycan provides binding to laminin, and the transmembrane β-subunit provides binding to dystrophin and the cytoskeleton.
- IPT domains
-
(also known as TIG domains). Immunoglobulin–plexin–transcription (IPT) protein domains found on cell-surface transmembrane receptors and intracellular transcription factors with an immunoglobulin-like fold.
- Glycosaminoglycan chains
-
(GAGs). Extended linear polysaccharides comprising repeating disaccharides.
- Stoichiometry
-
The fraction of a glycosylation site in a glycoprotein that is occupied by a glycan.
- ABH
-
Carbohydrate antigens of the ABO blood group system.
- Lipoproteins
-
Large complexes of lipids and proteins that transport lipids in the blood.
- Lipopolysaccharide
-
A bacterial glycolipid endotoxin and a major constituent of the outer membrane of Gram-negative bacteria.
- Proprotein convertases
-
A family of seven secretory mammalian serine proteinases that post-translationally activate proproteins in the secretory pathway by limited proteolysis after multiple basic residues with a general recognition motif, (R/K)Xn(R/K). A prototypical proprotein convertase is furin.
- Epithelial–mesenchymal transition
-
The physiological process in which epithelial cells transition to a mesenchymal state by loss of polarity and other characteristics, and acquire migratory and invasive properties. This occurs during embryonic development and during cancer progression.
- Immune checkpoint
-
An inhibitory pathway in the immune system that is crucial for maintaining self-tolerance and modulating the duration and amplitude of physiological immune responses.
- Sialyl-Lewisx antigen
-
A glycan determinant/epitope and selectin ligand (NeuAcα2–3Galβ1–4(Fucα1–3)GlcNAcβ1-R) found on glycoproteins and glycolipids.
- Sialyl-Lewisa antigen
-
A glycan determinant/epitope and selectin ligand (NeuAcα2–3Galβ1–3(Fucα1–4)GlcNAcβ1-R) found on glycoproteins and glycolipids.
- Warburg effect
-
The capacity for tumours to metabolize glucose anaerobically when oxygen is available, increase glucose uptake and produce lactate. The enhanced flux of metabolites through the hexosamine biosynthetic pathway may lead to increased O-GlcNAcylation.
- Paucimannose
-
An N-glycan structure consisting of two N-acetyl-d-glucosamines (GlcNAcs) and only one to three mannose residues (core Fuc optional).
- Haploid genetic screen
-
A genetic screen performed in haploid cells to ease the search for recessive functions in mammalian cells.
- Tau protein
-
Tubulin binding protein involved in neurodegenerative diseases called tauopathies where Tau aggregates into intracellular neurofibrillary tangles.
- Native mass spectrometry
-
An analytical technique used to study non-denatured proteoforms and protein complexes in the gas phase.
- Glucocerebrosidase
-
(GBA). A lysosomal enzyme that hydrolyses glucosylceramide (GlcCer) cerobroside into glucose and free ceramide.
- Gaucher disease
-
A rare genetic lysosomal storage disease caused by mutations in the GBA gene encoding glucocerebrosidase (GBA). Damaging mutations in GBA cause progressive accumulation of the glycolipid glucosylceramide (GlcCer) substrate for the enzyme with damaging effects on tissues and organs. Enzyme replacement therapy is currently the treatment of choice for this disease.
- α-Galactosidase
-
(GLA). A lysosomal enzyme that hydrolyses α-galactose from glycolipids and glycoproteins.
- Fabry disease
-
A rare, genetic X-linked lysosomal storage disease caused by damaging mutations in the GLA gene encoding α-galactosidase A, which catalyses the hydrolysis of glycolipid globotriacylceramide (Gb3).
- αGal epitope
-
A Galα1–3Galβ1–4GlcNAc-R glycan epitope (also known as the Galili antigen) found on glycoproteins and glycolipids of non-primate mammals and new world monkeys, but absent in humans, apes and old world monkeys because of an inactivated GGTA1 gene.
Rights and permissions
About this article
Cite this article
Schjoldager, K.T., Narimatsu, Y., Joshi, H.J. et al. Global view of human protein glycosylation pathways and functions. Nat Rev Mol Cell Biol 21, 729–749 (2020). https://doi.org/10.1038/s41580-020-00294-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-020-00294-x
This article is cited by
-
O-GlcNAcylation: a pro-survival response to acute stress in the cardiovascular and central nervous systems
European Journal of Medical Research (2024)
-
Impaired bisecting GlcNAc reprogrammed M1 polarization of macrophage
Cell Communication and Signaling (2024)
-
Decoding the glycoproteome: a new frontier for biomarker discovery in cancer
Journal of Hematology & Oncology (2024)
-
Identification and characterization of intact glycopeptides in human urine
Scientific Reports (2024)
-
Protein glycosylation in cardiovascular health and disease
Nature Reviews Cardiology (2024)