Abstract
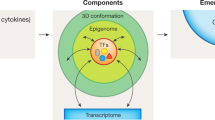
In eukaryotes, the genome does not exist as a linear molecule but instead is hierarchically packaged inside the nucleus. This complex genome organization includes multiscale structural units of chromosome territories, compartments, topologically associating domains, which are often demarcated by architectural proteins such as CTCF and cohesin, and chromatin loops. The 3D organization of chromatin modulates biological processes such as transcription, DNA replication, cell division and meiosis, which are crucial for cell differentiation and animal development. In this Review, we discuss recent progress in our understanding of the general principles of chromatin folding, its regulation and its functions in mammalian development. Specifically, we discuss the dynamics of 3D chromatin and genome organization during gametogenesis, embryonic development, lineage commitment and stem cell differentiation, and focus on the functions of chromatin architecture in transcription regulation. Finally, we discuss the role of 3D genome alterations in the aetiology of developmental disorders and human diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hug, C. B. & Vaquerizas, J. M. The birth of the 3D genome during early embryonic development. Trends Genet. 34, 903–914 (2018).
Gorkin, D. U., Leung, D. & Ren, B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell 14, 762–775 (2014).
Beagrie, R. A. et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature 543, 519–524 (2017).
Bonev, B. & Cavalli, G. Organization and function of the 3D genome. Nat. Rev. Genet. 17, 661–678 (2016).
Dekker, J. & Misteli, T. Long-range chromatin interactions. Cold Spring Harb. Perspect. Biol. 7, a019356 (2015).
Van Bortle, K. & Corces, V. G. Nuclear organization and genome function. Annu. Rev. Cell Dev. Biol. 28, 163–187 (2012).
Hagstrom, K. A. & Meyer, B. J. Condensin and cohesin: more than chromosome compactor and glue. Nat. Rev. Genet. 4, 520–534 (2003).
Lupianez, D. G., Spielmann, M. & Mundlos, S. Breaking TADs: how alterations of chromatin domains result in disease. Trends Genet. 32, 225–237 (2016).
Handel, M. A. & Schimenti, J. C. Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nat. Rev. Genet. 11, 124–136 (2010).
Schagdarsurengin, U. & Steger, K. Epigenetics in male reproduction: effect of paternal diet on sperm quality and offspring health. Nat. Rev. Urol. 13, 584–595 (2016).
Meistrich, M. L., Mohapatra, B., Shirley, C. R. & Zhao, M. Roles of transition nuclear proteins in spermiogenesis. Chromosoma 111, 483–488 (2003).
Balhorn, R., Gledhill, B. L. & Wyrobek, A. J. Mouse sperm chromatin proteins: quantitative isolation and partial characterization. Biochemistry 16, 4074–4080 (1977).
Gatewood, J. M., Cook, G. R., Balhorn, R., Schmid, C. W. & Bradbury, E. M. Isolation of four core histones from human sperm chromatin representing a minor subset of somatic histones. J. Biol. Chem. 265, 20662–20666 (1990).
Hilscher, B. et al. Kinetics of gametogenesis. I. Comparative histological and autoradiographic studies of oocytes and transitional prospermatogonia during oogenesis and prespermatogenesis. Cell Tissue Res. 154, 443–470 (1974).
MacLennan, M., Crichton, J. H., Playfoot, C. J. & Adams, I. R. Oocyte development, meiosis and aneuploidy. Semin. Cell Dev. Biol. 45, 68–76 (2015).
Eckersley-Maslin, M. A., Alda-Catalinas, C. & Reik, W. Dynamics of the epigenetic landscape during the maternal-to-zygotic transition. Nat. Rev. Mol. Cell. Biol. 19, 436–450 (2018).
Xu, Q. & Xie, W. Epigenome in early mammalian development: inheritance, reprogramming and establishment. Trends Cell Biol. 28, 237–253 (2018).
Schultz, R. M. The molecular foundations of the maternal to zygotic transition in the preimplantation embryo. Hum. Reprod. Update 8, 323–331 (2002).
Lee, M. T., Bonneau, A. R. & Giraldez, A. J. Zygotic genome activation during the maternal-to-zygotic transition. Annu. Rev. Cell Dev. Biol. 30, 581–613 (2014).
Lawson, K. A., Meneses, J. J. & Pedersen, R. A. Clonal analysis of epiblast fate during germ layer formation in the mouse embryo. Development 113, 891–911 (1991).
Gall, J. G. & Pardue, M. L. Formation and detection of RNA-DNA hybrid molecules in cytological preparations. Proc. Natl Acad. Sci. USA 63, 378–383 (1969).
Levsky, J. M. & Singer, R. H. Fluorescence in situ hybridization: past, present and future. J. Cell Sci. 116, 2833–2838 (2003).
Sigal, Y. M., Zhou, R. & Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 361, 880–887 (2018).
Wu, X., Mao, S., Ying, Y., Krueger, C. J. & Chen, A. K. Progress and challenges for live-cell imaging of genomic loci using CRISPR-based platforms. Genomics Proteomics Bioinformatics https://doi.org/10.1016/j.gpb.2018.10.001 (2019).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat. Genet. 38, 1341–1347 (2006).
Dostie, J. et al. Chromosome conformation capture carbon copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Fullwood, M. J. et al. An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462, 58–64 (2009).
Hughes, J. R. et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat. Genet. 46, 205–212 (2014).
Szalaj, P. & Plewczynski, D. Three-dimensional organization and dynamics of the genome. Cell Biol. Toxicol. 34, 381–404 (2018).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Nuebler, J., Fudenberg, G., Imakaev, M., Abdennur, N. & Mirny, L. A. Chromatin organization by an interplay of loop extrusion and compartmental segregation. Proc. Natl Acad. Sci. USA 115, E6697–E6706 (2018).
Cremer, T. & Cremer, M. Chromosome territories. Cold Spring Harb. Perspect. Biol. 2, a003889 (2010).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLOS Biol. 3, e157 (2005).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Misteli, T. Beyond the sequence: cellular organization of genome function. Cell 128, 787–800 (2007).
Bickmore, W. A. & van Steensel, B. Genome architecture: domain organization of interphase chromosomes. Cell 152, 1270–1284 (2013).
Shah, S. et al. Dynamics and spatial genomics of the nascent transcriptome by intron seqFISH. Cell 174, 363–376 (2018).
Boyle, S., Rodesch, M. J., Halvensleben, H. A., Jeddeloh, J. A. & Bickmore, W. A. Fluorescence in situ hybridization with high-complexity repeat-free oligonucleotide probes generated by massively parallel synthesis. Chromosome Res. 19, 901–909 (2011).
Osborne, C. S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 36, 1065–1071 (2004).
Schoenfelder, S. et al. Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 42, 53–61 (2010).
Morey, C., Da Silva, N. R., Perry, P. & Bickmore, W. A. Nuclear reorganisation and chromatin decondensation are conserved, but distinct, mechanisms linked to Hox gene activation. Development 134, 909–919 (2007).
Zink, D. et al. Transcription-dependent spatial arrangements of CFTR and adjacent genes in human cell nuclei. J. Cell Biol. 166, 815–825 (2004).
Branco, M. R. & Pombo, A. Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLOS Biol. 4, e138 (2006).
Padeken, J. & Heun, P. Nucleolus and nuclear periphery: Velcro for heterochromatin. Curr. Opin. Cell Biol. 28, 54–60 (2014).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Chen, Y. et al. TSA-seq mapping of nuclear genome organization. J. Cell Biol. 217, 4025–4048 (2018).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174, 744–757 (2018).
Pontvianne, F. et al. Identification of nucleolus-associated chromatin domains reveals a role for the nucleolus in 3D organization of the A. thaliana genome. Cell Rep. 16, 1574–1587 (2016).
Nemeth, A. et al. Initial genomics of the human nucleolus. PLOS Genet. 6, e1000889 (2010).
van Koningsbruggen, S. et al. High-resolution whole-genome sequencing reveals that specific chromatin domains from most human chromosomes associate with nucleoli. Mol. Biol. Cell 21, 3735–3748 (2010).
Solovei, I. et al. LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152, 584–598 (2013).
Larson, A. G. et al. Liquid droplet formation by HP1alpha suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Symmons, O. et al. Functional and topological characteristics of mammalian regulatory domains. Genome Res. 24, 390–400 (2014).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl Acad. Sci. USA 111, 996–1001 (2014).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Vietri Rudan, M. et al. Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Rep. 10, 1297–1309 (2015).
Ciosk, R. et al. Cohesin’s binding to chromosomes depends on a separate complex consisting of Scc2 and Scc4 proteins. Mol. Cell 5, 243–254 (2000).
Gandhi, R., Gillespie, P. J. & Hirano, T. Human Wapl is a cohesin-binding protein that promotes sister-chromatid resolution in mitotic prophase. Curr. Biol. 16, 2406–2417 (2006).
Kueng, S. et al. Wapl controls the dynamic association of cohesin with chromatin. Cell 127, 955–967 (2006).
Tedeschi, A. et al. Wapl is an essential regulator of chromatin structure and chromosome segregation. Nature 501, 564–568 (2013).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178 (2018).
Heinz, S. et al. Transcription elongation can affect genome 3D structure. Cell 174, 1522–1536 (2018).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456 (2015).
Hansen, A. S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq, X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. eLife 6, e25776 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Fudenberg, G., Abdennur, N., Imakaev, M., Goloborodko, A. & Mirny, L. A. Emerging evidence of chromosome folding by loop extrusion. Cold Spring Harb. Symp. Quant. Biol. 82, 45–55 (2018).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Sima, J. et al. Identifying cis elements for spatiotemporal control of mammalian DNA replication. Cell 176, 816–830 (2019).
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Sofueva, S. et al. Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J. 32, 3119–3129 (2013).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Tan, L. Z., Xing, D., Chang, C. H., Li, H. & Xie, S. Three-dimensional genome structures of single diploid human cells. Science 361, 924–928 (2018).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Szabo, Q. et al. TADs are 3D structural units of higher-order chromosome organization in Drosophila. Sci. Adv. 4, eaar8082 (2018).
Bintu, B. et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science 362, eaau1783 (2018).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell 67, 837–852 (2017).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A. Phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 (2018).
Hunt, P. A. Meiosis in mammals: recombination, non-disjunction and the environment. Biochem. Soc. Trans. 34, 574–577 (2006).
Li, L., Zheng, P. & Dean, J. Maternal control of early mouse development. Development 137, 859–870 (2010).
Battulin, N. et al. Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach. Genome Biol. 16, 77 (2015).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Jung, Y. H. et al. Chromatin states in mouse sperm correlate with embryonic and adult regulatory landscapes. Cell Rep. 18, 1366–1382 (2017).
Ke, Y. et al. 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170, 367–381 (2017).
Wang, Y. et al. Reprogramming of meiotic chromatin architecture during spermatogenesis. Mol. Cell 73, 547–561 (2019).
Alavattam, K. G. et al. Attenuated chromatin compartmentalization in meiosis and its maturation in sperm development. Nat. Struct. Mol. Biol. 26, 175–184 (2019).
Patel, L. et al. Dynamic reorganization of the genome shapes the recombination landscape in meiotic prophase. Nat. Struct. Mol. Biol. 26, 164–174 (2019). Wang et al. ( Mol. Cell , 2019), Alavattam et al. (2019) and Patel et al. (2019) describe 3D genome dynamics during spermatogenesis in the mouse and the rhesus monkey.
Hud, N. V., Allen, M. J., Downing, K. H., Lee, J. & Balhorn, R. Identification of the elemental packing unit of DNA in mammalian sperm cells by atomic force microscopy. Biochem. Biophys. Res. Commun. 193, 1347–1354 (1993).
Carone, B. R. et al. High-resolution mapping of chromatin packaging in mouse embryonic stem cells and sperm. Dev. Cell 30, 11–22 (2014).
Tang, W. W. C., Kobayashi, T., Irie, N., Dietmann, S. & Surani, M. A. Specification and epigenetic programming of the human germ line. Nat. Rev. Genet. 17, 585–600 (2016).
Zamudio, N. M., Chong, S. Y. & O’Bryan, M. K. Epigenetic regulation in male germ cells. Reproduction 136, 131–146 (2008).
Cloutier, J. M. & Turner, J. M. A. Meiotic sex chromosome inactivation. Curr. Biol. 20, 1823–1831 (2010).
Cobb, J. & Handel, M. A. Dynamics of meiotic prophase I during spermatogenesis: from pairing to division. Semin. Cell Dev. Biol. 9, 445–450 (1998).
Turner, J. M. A. Meiotic sex chromosome inactivation. Development 134, 1823–1831 (2007).
Hernandez-Hernandez, A., Lilienthal, I., Fukuda, N., Galjart, N. & Hoog, C. CTCF contributes in a critical way to spermatogenesis and male fertility. Sci. Rep. 6, 28355 (2016).
Jegu, T., Aeby, E. & Lee, J. T. The X chromosome in space. Nat. Rev. Genet. 18, 377–389 (2017).
Schalbetter, S. A., Fudenberg, G., Baxter, J., Pollard, K. S. & Neale, M. J. Principles of meiotic chromosome assembly. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/442038v2 (2019).
Muller, H. et al. Characterizing meiotic chromosomes’ structure and pairing using a designer sequence optimized for Hi-C. Mol. Syst. Biol. 14, e8293 (2018).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359, eaao6135 (2018).
Zuccotti, M., Piccinelli, A., Giorgi Rossi, P., Garagna, S. & Redi, C. A. Chromatin organization during mouse oocyte growth. Mol. Reprod. Dev. 41, 479–485 (1995).
De La Fuente, R. Chromatin modifications in the germinal vesicle (GV) of mammalian oocytes. Dev. Biol. 292, 1–12 (2006).
Miyara, F. et al. Chromatin configuration and transcriptional control in human and mouse oocytes. Mol. Reprod. Dev. 64, 458–470 (2003).
Bouniol-Baly, C. et al. Differential transcriptional activity associated with chromatin configuration in fully grown mouse germinal vesicle oocytes. Biol. Reprod. 60, 580–587 (1999).
Ahmed, K. et al. Global chromatin architecture reflects pluripotency and lineage commitment in the early mouse embryo. PLOS ONE 5, e10531 (2010).
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell 169, 216–228 (2017).
Gassler, J. et al. A mechanism of cohesin-dependent loop extrusion organizes zygotic genome architecture. EMBO J. 36, 3600–3618 (2017). Flyamer et al. (2017), Du et al. (2017), Ke et al. (2017) and Gassler et al. (2017) describe the conformation of chromatin in mouse oocytes and early embryos using low-input Hi-C methods.
Reichmann, J. et al. Dual-spindle formation in zygotes keeps parental genomes apart in early mammalian embryos. Science 361, 189–193 (2018).
Zhang, Y. et al. Dynamic epigenomic landscapes during early lineage specification in mouse embryos. Nat. Genet. 50, 96–105 (2018).
Matoba, S. et al. Embryonic development following somatic cell nuclear transfer impeded by persisting histone methylation. Cell 159, 884–895 (2014).
Liu, W. Q. et al. Identification of key factors conquering developmental arrest of somatic cell cloned embryos by combining embryo biopsy and single-cell sequencing. Cell Discov. 2, 16010 (2016).
Liu, Z. et al. Cloning of macaque monkeys by somatic cell nuclear transfer. Cell 172, 881–887 (2018).
Kishigami, S. et al. Significant improvement of mouse cloning technique by treatment with trichostatin A after somatic nuclear transfer. Biochem. Biophys. Res. Commun. 340, 183–189 (2006).
Becker, J. S., Nicetto, D. & Zaret, K. S. H3K9me3-dependent heterochromatin: barrier to cell fate changes. Trends Genet. 32, 29–41 (2016).
Gorisch, S. M., Wachsmuth, M., Toth, K. F., Lichter, P. & Rippe, K. Histone acetylation increases chromatin accessibility. J. Cell Sci. 118, 5825–5834 (2005).
Cuartero, S. et al. Control of inducible gene expression links cohesin to hematopoietic progenitor self-renewal and differentiation. Nat. Immunol. 19, 932–941 (2018).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010).
Schmitt, A. D. et al. A compendium of chromatin contact maps reveals spatially active regions in the human genome. Cell Rep. 17, 2042–2059 (2016).
Fraser, J. et al. Hierarchical folding and reorganization of chromosomes are linked to transcriptional changes in cellular differentiation. Mol. Syst. Biol. 11, 852 (2015).
Pekowska, A. et al. Gain of CTCF-anchored chromatin loops marks the exit from naive pluripotency. Cell Syst. 7, 482–495 (2018).
Smallwood, A. & Ren, B. Genome organization and long-range regulation of gene expression by enhancers. Curr. Opin. Cell Biol. 25, 387–394 (2013).
Fullwood, M. J., Wei, C. L., Liu, E. T. & Ruan, Y. Next-generation DNA sequencing of paired-end tags (PET) for transcriptome and genome analyses. Genome Res. 19, 521–532 (2009).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598–606 (2015).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919 (2016).
Fang, R. X. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP-seq. Cell Res. 26, 1345–1348 (2016).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384 (2016).
Visel, A., Rubin, E. M. & Pennacchio, L. A. Genomic views of distant-acting enhancers. Nature 461, 199–205 (2009).
Phillips, J. E. & Corces, V. G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009).
Bell, A. C., West, A. G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Hou, C., Zhao, H., Tanimoto, K. & Dean, A. CTCF-dependent enhancer-blocking by alternative chromatin loop formation. Proc. Natl Acad. Sci. USA 105, 20398–20403 (2008).
Dowen, J. M. et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell 159, 374–387 (2014).
Ji, X. et al. 3D chromosome regulatory landscape of human pluripotent cells. Cell Stem Cell 18, 262–275 (2016).
Weintraub, A. S. et al. YY1 Is a structural regulator of enhancer-promoter loops. Cell 171, 1573 (2017).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Rubin, A. J. et al. Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat. Genet. 49, 1522–1528 (2017). This study identified two classes of promoter–enhancer interaction in epidermal differentiation, and associated stable promoter–enhancer interactions with cohesin.
Freire-Pritchett, P. et al. Global reorganisation of cis-regulatory units upon lineage commitment of human embryonic stem cells. eLife 6, e21926 (2017).
Kieffer-Kwon, K. R. et al. Interactome maps of mouse gene regulatory domains reveal basic principles of transcriptional regulation. Cell 155, 1507–1520 (2013).
Zhang, Y. et al. Chromatin connectivity maps reveal dynamic promoter-enhancer long-range associations. Nature 504, 306–310 (2013).
Noordermeer, D. & de Laat, W. Joining the loops: beta-globin gene regulation. IUBMB Life 60, 824–833 (2008).
Deng, W. et al. Reactivation of developmentally silenced globin genes by forced chromatin looping. Cell 158, 849–860 (2014).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012). Deng et al. (2014) and Deng et al. (2012) show that targeting an active cis -regulatory element to a silenced gene can result in gene activation.
Ghavi-Helm, Y. et al. Enhancer loops appear stable during development and are associated with paused polymerase. Nature 512, 96–100 (2014).
Montavon, T. et al. A regulatory archipelago controls Hox genes transcription in digits. Cell 147, 1132–1145 (2011).
Lonfat, N., Montavon, T., Darbellay, F., Gitto, S. & Duboule, D. Convergent evolution of complex regulatory landscapes and pleiotropy at Hox loci. Science 346, 1004–1006 (2014).
Benabdallah, N. S. et al. PARP mediated chromatin unfolding is coupled to long-range enhancer activation. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/155325v1 (2017).
Fukaya, T., Lim, B. & Levine, M. Enhancer control of transcriptional bursting. Cell 166, 358–368 (2016).
Gu, B. et al. Transcription-coupled changes in nuclear mobility of mammalian cis-regulatory elements. Science 359, 1050–1055 (2018).
Chepelev, I., Wei, G., Wangsa, D., Tang, Q. & Zhao, K. Characterization of genome-wide enhancer-promoter interactions reveals co-expression of interacting genes and modes of higher order chromatin organization. Cell Res. 22, 490–503 (2012).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Diao, Y. R. et al. A tiling-deletion-based genetic screen for cis-regulatory element identification in mammalian cells. Nat. Methods 14, 629 (2017).
Apostolou, E. et al. Genome-wide chromatin interactions of the Nanog locus in pluripotency, differentiation, and reprogramming. Cell Stem Cell 12, 699–712 (2013).
Schoenfelder, S. et al. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. 25, 582–597 (2015).
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Denholtz, M. et al. Long-range chromatin contacts in embryonic stem cells reveal a role for pluripotency factors and polycomb proteins in genome organization. Cell Stem Cell 13, 602–616 (2013). Diao et al. (2017), Apostolou et al. (2013), Schoenfelder et al. ( Genome Res. , 2015), de Wit et al. (2013) and Denholtz et al. (2013) reported the existence of a pluripotency-specific interactome in embryonic stem cells.
Lanzuolo, C. & Orlando, V. Memories from the polycomb group proteins. Annu. Rev. Genet. 46, 561–589 (2012).
Eskeland, R. et al. Ring1B compacts chromatin structure and represses gene expression independent of histone ubiquitination. Mol. Cell 38, 452–464 (2010).
Margueron, R. et al. Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol. Cell 32, 503–518 (2008).
Kundu, S. et al. Polycomb repressive complex 1 generates discrete compacted domains that change during differentiation. Mol. Cell 65, 432–446 (2017).
Li, Y. et al. Genome-wide analyses reveal a role of Polycomb in promoting hypomethylation of DNA methylation valleys. Genome Biol. 19, 18 (2018).
Eagen, K. P., Aiden, E. L. & Kornberg, R. D. Polycomb-mediated chromatin loops revealed by a subkilobase-resolution chromatin interaction map. Proc. Natl Acad. Sci. USA 114, 8764–8769 (2017).
Bantignies, F. et al. Polycomb-dependent regulatory contacts between distant Hox loci in Drosophila. Cell 144, 214–226 (2011).
Joshi, O. et al. Dynamic reorganization of extremely long-range promoter-promoter interactions between two states of pluripotency. Cell Stem Cell 17, 748–757 (2015).
Wani, A. H. et al. Chromatin topology is coupled to Polycomb group protein subnuclear organization. Nat. Commun. 7, 10291 (2016).
Isono, K. et al. SAM domain polymerization links subnuclear clustering of PRC1 to gene silencing. Dev. Cell 26, 565–577 (2013).
Schoenfelder, S. et al. Polycomb repressive complex PRC1 spatially constrains the mouse embryonic stem cell genome. Nat. Genet. 47, 1179–1186 (2015). Bantignies et al. (2011), Joshi et al. (2015), Wani et al. (2016), Isono et al. (2013) and Schoenfelder et al. ( Nat. Genet. , 2015) identified Polycomb-mediated interaction clusters and showed that such interactions may contribute to the corepression of developmental genes.
Medrano-Fernandez, A. & Barco, A. Nuclear organization and 3D chromatin architecture in cognition and neuropsychiatric disorders. Mol. Brain 9, 83 (2016).
Davis, L., Onn, I. & Elliott, E. The emerging roles for the chromatin structure regulators CTCF and cohesin in neurodevelopment and behavior. Cell. Mol. Life Sci. 75, 1205–1214 (2018).
Rosa-Garrido, M. et al. High-resolution mapping of chromatin conformation in cardiac myocytes reveals structural remodeling of the epigenome in heart failure. Circulation 136, 1613 (2017).
Kaiser, V. B. & Semple, C. A. When TADs go bad: chromatin structure and nuclear organisation in human disease. F1000Res 6, 314 (2017).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015). This article reports on the different limb syndromes that are caused by altered chromatin structure at the EPHA4 locus.
Giorgio, E. et al. A large genomic deletion leads to enhancer adoption by the lamin B1 gene: a second path to autosomal dominant adult-onset demyelinating leukodystrophy (ADLD). Hum. Mol. Genet. 24, 3143–3154 (2015).
Benko, S. et al. Disruption of a long distance regulatory region upstream of SOX9 in isolated disorders of sex development. J. Med. Genet. 48, 825–830 (2011).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Redin, C. et al. The genomic landscape of balanced cytogenetic abnormalities associated with human congenital anomalies. Nat. Genet. 49, 36–45 (2017).
Ibn-Salem, J. et al. Deletions of chromosomal regulatory boundaries are associated with congenital disease. Genome Biol. 15, 423 (2014).
Valton, A. L. & Dekker, J. TAD disruption as oncogenic driver. Curr. Opin. Genet. Dev. 36, 34–40 (2016).
Groschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in leukemia. Cell 157, 369–381 (2014).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Beroukhim, R., Zhang, X. & Meyerson, M. Copy number alterations unmasked as enhancer hijackers. Nat. Genet. 49, 5–6 (2016).
Katainen, R. et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat. Genet. 47, 818–821 (2015).
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Weischenfeldt, J. et al. Pan-cancer analysis of somatic copy-number alterations implicates IRS4 and IGF2 in enhancer hijacking. Nat. Genet. 49, 65–74 (2017).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Beliveau, B. J. et al. Versatile design and synthesis platform for visualizing genomes with Oligopaint FISH probes. Proc. Natl Acad. Sci. USA 109, 21301–21306 (2012).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Ma, H. et al. Multiplexed labeling of genomic loci with dCas9 and engineered sgRNAs using CRISPRainbow. Nat. Biotechnol. 34, 528–530 (2016).
Payer, B. & Lee, J. T. X. Chromosome dosage compensation: how mammals keep the balance. Annu. Rev. Genet. 42, 733–772 (2008).
Galupa, R. & Heard, E. X-Chromosome inactivation: new insights into cis and trans regulation. Curr. Opin. Genet. Dev. 31, 57–66 (2015).
Finestra, T. R. & Gribnau, J. X chromosome inactivation: silencing, topology and reactivation. Curr. Opin. Cell Biol. 46, 54–61 (2017).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Minajigi, A. et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, eaab2276 (2015).
Chen, C. K. et al. Xist recruits the X chromosome to the nuclear lamina to enable chromosome-wide silencing. Science 354, 468–472 (2016).
Pollex, T. & Heard, E. Nuclear positioning and pairing of X-chromosome inactivation centers are not primary determinants during initiation of random X-inactivation. Nat. Genet. 51, 285–295 (2019).
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011).
Deng, X. et al. Bipartite structure of the inactive mouse X chromosome. Genome Biol. 16, 152 (2015).
Wang, C.-Y., Jégu, T., Chu, H.-P., Oh, H. J. & Lee, J. T. SMCHD1 merges chromosome compartments and assists formation of super-structures on the inactive X. Cell 174, 406–421 (2018).
Chadwick, B. P. DXZ4 chromatin adopts an opposing conformation to that of the surrounding chromosome and acquires a novel inactive X-specific role involving CTCF and antisense transcripts. Genome Res. 18, 1259–1269 (2008).
Darrow, E. M. et al. Deletion of DXZ4 on the human inactive X chromosome alters higher-order genome architecture. Proc. Natl Acad. Sci. USA 113, E4504–E4512 (2016).
Horakova, A. H., Moseley, S. C., McLaughlin, C. R., Tremblay, D. C. & Chadwick, B. P. The macrosatellite DXZ4 mediates CTCF-dependent long-range intrachromosomal interactions on the human inactive X chromosome. Hum. Mol. Genet. 21, 4367–4377 (2012).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Froberg, J. E., Pinter, S. F., Kriz, A. J., Jegu, T. & Lee, J. T. Megadomains and superloops form dynamically but are dispensable for X-chromosome inactivation and gene escape. Nat. Commun. 9, 5004 (2018).
Gdula, M. R. et al. The non-canonical SMC protein SmcHD1 antagonises TAD formation and compartmentalisation on the inactive X chromosome. Nat. Commun. 10, 30 (2019).
Jansz, N. et al. Long-range chromatin interactions on the inactive X and at Hox clusters are regulated by the non-canonical SMC protein Smchd1. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/342212v1 (2018).
Acknowledgements
The authors thank B. Ren and J. Xu for careful reading of the manuscript. The authors thank Z. Du, Y. Wang, Y. Zhang and other members of the Xie laboratory for valuable comments. This Review included only selected studies as an illustration of the recent progress of our understanding of the 3D genome in development; the authors apologize to researchers whose studies could not be cited owing to space limitations. The work was supported by the National Key R&D Program of China (2016YFC0900300 to W.X.), the National Basic Research Program of China (2015CB856201 to W.X.), the National Natural Science Foundation of China (31725018 and 31830047 to W.X.), the Beijing Municipal Science & Technology Commission (Z181100001318006 to W.X.), the THU–PKU Center for Life Sciences, and Beijing Advanced Innovation Center for Structural Biology (W.X.). H.Z. is supported by a postdoctoral fellowship from the THU–PKU Center for Life Sciences. W.X. is a recipient of HHMI International Research Scholar.
Reviewer information
Nature Reviews Molecular Cell Biology thanks P. Fraser and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Both authors equally contributed to researching data for the article, the discussion of content, writing of the manuscript and its editing before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
DECIPHER: https://decipher.sanger.ac.uk/
Glossary
- Gametogenesis
-
The process in which diploid gamete-precursor cells undergo meiotic division and differentiation to form mature sperm and oocytes.
- Primordial germ cells
-
(PGCs). The germline ancestor cells of both sperm and oocytes. PGCs are diploid and are first found in the primary ectoderm of the epiblast.
- Protamines
-
Small, arginine-rich nuclear proteins that are specifically found in the haploid phase of mature sperm and that largely replace histones as the DNA-packaging proteins.
- Diplotene
-
A stage of prophase in meiosis I. During diplotene, the synaptonemal complex degrades and two homologous chromosomes separate from each other and uncoil.
- Totipotent
-
A totipotent cell has the capacity to divide and produce all the differentiated cells of both embryonic and extraembryonic tissues.
- Zygotic genome activation
-
(ZGA). The activation of gene transcription from the zygote genome after fertilization.
- Gastrulation
-
A process of early embryonic development during which the single-layer blastula is reorganized to form a multilayer structure known as the gastrula.
- Heterochromatin
-
A condensed form of chromatin in which gene activity is usually repressed.
- Nuclear lamina
-
A mesh structure just inside the nuclear membrane that is composed of lamins and lamin-associated proteins.
- Nuclear speckles
-
Also known as splicing speckles, these are nuclear domains enriched in pre-mRNA splicing factors and located in interchromatin regions of the nucleoplasm.
- Superenhancers
-
Genomic regions comprising multiple enhancers that are collectively bound by multiple transcription factors to drive gene transcription.
- Synaptonemal complex
-
A structure that forms between homologous chromosomes during meiosis and functions in mediating chromosome pairing, synapsis and recombination.
- Pachytene
-
A stage of prophase in meiosis I during which the paired chromosomes shorten and thicken. Homologous recombination occurs during this stage.
- Meiotic sex chromosome inactivation
-
In spermatogenesis, the transcriptional silencing of the X and Y chromosomes during meiotic pachytene.
- Maternal-to-zygotic transition
-
The stage in early embryonic development when the zygotic genome takes control of development from the maternal genome. This transition requires zygotic genome activation and the degradation of maternal RNA and proteins.
- Germinal vesicle oocytes
-
Growing or grown oocytes arrested in prophase of meiosis I before ovulation. The germinal vesicle refers to their nucleus, which is clearly visible under the microscope.
- Pronuclei
-
The paternal and maternal nuclei just after fertilization, when they are still physically separated in the zygote.
- Somatic cell nuclear transfer
-
A technique for creating a viable embryo by transferring a donor nucleus of a somatic cell to an enucleated oocyte.
- Naive pluripotency
-
In preimplantation embryos, pluripotent stem cells in the epiblast are in a ‘naive’ state. They become ‘primed’ during postimplantation development.
- Insulators
-
cis-regulatory elements that can block the function of enhancers or the spreading of gene silencing. The word can also refer to the protein complexes that bind to these elements, such as CTCF.
- 2i ground-state pluripotent mouse ESCs
-
Mouse embryonic stem cells cultured in the presence of MEK and glycogen synthase kinase 3 inhibitors and thought to be in a naive ground state.
- Primed-like pluripotent state
-
Mouse embryonic stem cells cultured in serum that are thought to be epigenetically more restricted and developmentally primed than 2i ground-state pluripotent mouse embryonic stem cells and are similar to cells in the postimplantation epiblast.
Rights and permissions
About this article
Cite this article
Zheng, H., Xie, W. The role of 3D genome organization in development and cell differentiation. Nat Rev Mol Cell Biol 20, 535–550 (2019). https://doi.org/10.1038/s41580-019-0132-4
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-019-0132-4
This article is cited by
-
The N-terminal dimerization domains of human and Drosophila CTCF have similar functionality
Epigenetics & Chromatin (2024)
-
Integrative analysis of transcriptomic and epigenomic data reveals distinct patterns for developmental and housekeeping gene regulation
BMC Biology (2024)
-
The impact of DNA methylation on CTCF-mediated 3D genome organization
Nature Structural & Molecular Biology (2024)
-
scGHOST: identifying single-cell 3D genome subcompartments
Nature Methods (2024)
-
The PTM profiling of CTCF reveals the regulation of 3D chromatin structure by O-GlcNAcylation
Nature Communications (2024)