Abstract
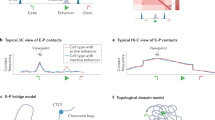
The spatial organization of the genome into compartments and topologically associated domains can have an important role in the regulation of gene expression. But could gene expression conversely regulate genome organization? Here, we review recent studies that assessed the requirement of transcription and/or the transcription machinery for the establishment or maintenance of genome topology. The results reveal different requirements at different genomic scales. Transcription is generally not required for higher-level genome compartmentalization, has only moderate effects on domain organization and is not sufficient to create new domain boundaries. However, on a finer scale, transcripts or transcription does seem to have a role in the formation of subcompartments and subdomains and in stabilizing enhancer–promoter interactions. Recent evidence suggests a dynamic, reciprocal interplay between fine-scale genome organization and transcription, in which each is able to modulate or reinforce the activity of the other.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Cavalli, G. & Misteli, T. Functional implications of genome topology. Nat. Struct. Mol. Biol. 20, 290–299 (2013).
Shachar, S. & Misteli, T. Causes and consequences of nuclear gene positioning. J. Cell Sci. 130, 1501–1508 (2017).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Hou, C., Li, L., Qin, Z. S. & Corces, V. G. Gene density, transcription, and insulators contribute to the partition of the Drosophila genome into physical domains. Mol. Cell 48, 471–484 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Brangwynne, C. P., Mitchison, T. J. & Hyman, A. A. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc. Natl Acad. Sci. USA 108, 4334–4339 (2011).
Sirri, V., Urcuqui-Inchima, S., Roussel, P. & Hernandez-Verdun, D. Nucleolus: the fascinating nuclear body. Histochem. Cell Biol. 129, 13–31 (2008).
Nemeth, A. & Grummt, I. Dynamic regulation of nucleolar architecture. Curr. Opin. Cell Biol. 52, 105–111 (2018).
Falahati, H., Pelham-Webb, B., Blythe, S. & Wieschaus, E. Nucleation by rRNA dictates the precision of nucleolus assembly. Curr. Biol. 26, 277–285 (2016).
Heyn, P., Salmonowicz, H., Rodenfels, J. & Neugebauer, K. M. Activation of transcription enforces the formation of distinct nuclear bodies in zebrafish embryos. RNA Biol. 14, 752–760 (2017).
Verheggen, C., Almouzni, G. & Hernandez-Verdun, D. The ribosomal RNA processing machinery is recruited to the nucleolar domain before RNA polymerase I during Xenopus laevis development. J. Cell Biol. 149, 293–306 (2000).
Mais, C., Wright, J. E., Prieto, J. L., Raggett, S. L. & McStay, B. UBF-binding site arrays form pseudo-NORs and sequester the RNA polymerase I transcription machinery. Genes Dev. 19, 50–64 (2005).
Hamdane, N. et al. Disruption of the UBF gene induces aberrant somatic nucleolar bodies and disrupts embryo nucleolar precursor bodies. Gene 612, 5–11 (2017).
Caudron-Herger, M. et al. Alu element-containing RNAs maintain nucleolar structure and function. EMBO J. 34, 2758–2774 (2015).
Heitz, E. Das heterochromatin der moose [German]. Jahrb. Wiss. Bot. 69, 762–818 (1928).
Filion, G. J. et al. Systematic protein location mapping reveals five principal chromatin types in Drosophila cells. Cell 143, 212–224 (2010).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
van Bemmel, J. G. et al. The insulator protein SU(HW) fine-tunes nuclear lamina interactions of the Drosophila genome. PLOS ONE 5, e15013 (2010).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Nemeth, A. et al. Initial genomics of the human nucleolus. PLOS Genet. 6, e1000889 (2010).
van Koningsbruggen, S. et al. High-resolution whole-genome sequencing reveals that specific chromatin domains from most human chromosomes associate with nucleoli. Mol. Biol. Cell 21, 3735–3748 (2010).
Dillinger, S., Straub, T. & Nemeth, A. Nucleolus association of chromosomal domains is largely maintained in cellular senescence despite massive nuclear reorganisation. PLOS ONE 12, e0178821 (2017).
Kind, J. et al. Single-cell dynamics of genome-nuclear lamina interactions. Cell 153, 178–192 (2013).
Ragoczy, T., Telling, A., Scalzo, D., Kooperberg, C. & Groudine, M. Functional redundancy in the nuclear compartmentalization of the late-replicating genome. Nucleus 5, 626–635 (2014).
Kalverda, B., Pickersgill, H., Shloma, V. V. & Fornerod, M. Nucleoporins directly stimulate expression of developmental and cell-cycle genes inside the nucleoplasm. Cell 140, 360–371 (2010).
Solovei, I. et al. Nuclear architecture of rod photoreceptor cells adapts to vision in mammalian evolution. Cell 137, 356–368 (2009).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Vieux-Rochas, M., Fabre, P. J., Leleu, M., Duboule, D. & Noordermeer, D. Clustering of mammalian Hox genes with other H3K27me3 targets within an active nuclear domain. Proc. Natl Acad. Sci. USA 112, 4672–4677 (2015).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Pinheiro, I. & Heard, E. X chromosome inactivation: new players in the initiation of gene silencing. F1000Res 6, 344 (2017).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Probst, A. V. et al. A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development. Dev. Cell 19, 625–638 (2010).
Velazquez Camacho, O. et al. Major satellite repeat RNA stabilize heterochromatin retention of Suv39h enzymes by RNA-nucleosome association and RNA: DNA hybrid formation. eLife 6, e25293 (2017).
Martienssen, R. & Moazed, D. RNAi and heterochromatin assembly. Cold Spring Harb. Perspect. Biol. 7, a019323 (2015).
Yuan, K. & O’Farrell, P. H. TALE-light imaging reveals maternally guided, H3K9me2/3-independent emergence of functional heterochromatin in Drosophila embryos. Genes Dev. 30, 579–593 (2016).
Brackley, C. A., Johnson, J., Kelly, S., Cook, P. R. & Marenduzzo, D. Simulated binding of transcription factors to active and inactive regions folds human chromosomes into loops, rosettes and topological domains. Nucleic Acids Res. 44, 3503–3512 (2016).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell 169, 216–228 (2017).
Battulin, N. et al. Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach. Genome Biol. 16, 77 (2015).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Jung, Y. H. et al. Chromatin states in mouse sperm correlate with embryonic and adult regulatory landscapes. Cell Rep. 18, 1366–1382 (2017).
Carone, B. R. et al. High-resolution mapping of chromatin packaging in mouse embryonic stem cells and sperm. Dev. Cell 30, 11–22 (2014).
Palstra, R. J. et al. Maintenance of long-range DNA interactions after inhibition of ongoing RNA polymerase II transcription. PLOS ONE 3, e1661 (2008).
Lund, E. et al. Lamin A/C-promoter interactions specify chromatin state-dependent transcription outcomes. Genome Res. 23, 1580–1589 (2013).
Kohwi, M., Lupton, J. R., Lai, S. L., Miller, M. R. & Doe, C. Q. Developmentally regulated subnuclear genome reorganization restricts neural progenitor competence in Drosophila. Cell 152, 97–108 (2013).
Tumbar, T. & Belmont, A. S. Interphase movements of a DNA chromosome region modulated by VP16 transcriptional activator. Nat. Cell Biol. 3, 134–139 (2001).
Chuang, C. H. et al. Long-range directional movement of an interphase chromosome site. Curr. Biol. 16, 825–831 (2006).
Bensaude, O. Inhibiting eukaryotic transcription: which compound to choose? How to evaluate its activity? Transcription 2, 103–108 (2011).
Therizols, P. et al. Chromatin decondensation is sufficient to alter nuclear organization in embryonic stem cells. Science 346, 1238–1242 (2014).
Isoda, T. et al. Non-coding transcription instructs chromatin folding and compartmentalization to dictate enhancer-promoter communication and T cell fate. Cell 171, 103–119 (2017).
Heinz, S. et al. Transcription elongation can affect genome 3D structure. Cell 174, 1522–1536 (2018).
Hu, Y., Plutz, M. & Belmont, A. S. Hsp70 gene association with nuclear speckles is Hsp70 promoter specific. J. Cell Biol. 191, 711–719 (2010).
Khanna, N., Hu, Y. & Belmont, A. S. HSP70 transgene directed motion to nuclear speckles facilitates heat shock activation. Curr. Biol. 24, 1138–1144 (2014).
Chen, Y. et al. TSA-Seq mapping of nuclear genome organization. J. Cell Biol. (in the press).
Brickner, J. Genetic and epigenetic control of the spatial organization of the genome. Mol. Biol. Cell 28, 364–369 (2017).
Osborne, C. S. et al. Myc dynamically and preferentially relocates to a transcription factory occupied by Igh. PLOS Biol. 5, e192 (2007).
Schoenfelder, S. et al. Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 42, 53–61 (2010).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174, 744–757 (2018).
Beagrie, R. A. et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature 543, 519–524 (2017).
Denholtz, M. et al. Long-range chromatin contacts in embryonic stem cells reveal a role for pluripotency factors and polycomb proteins in genome organization. Cell Stem Cell 13, 602–616 (2013).
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Ghavi-Helm, Y. et al. Enhancer loops appear stable during development and are associated with paused polymerase. Nature 512, 96–100 (2014).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Szabo, Q. et al. TADs are 3D structural units of higher-order chromosome organization in Drosophila. Sci. Adv. 4, eaar8082 (2018).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Symmons, O. et al. The Shh topological domain facilitates the action of remote enhancers by reducing the effects of genomic distances. Dev. Cell 39, 529–543 (2016).
Narendra, V. et al. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 347, 1017–1021 (2015).
Sexton, T. & Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell 160, 1049–1059 (2015).
Dekker, J., Guttman, M. & Lomvardas, S. A guide to packing your DNA. Cell 165, 259–261 (2016).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
van Ruiten, M. S. & Rowland, B. D. SMC complexes: universal DNA looping machines with distinct regulators. Trends Genet. 34, 477–487 (2018).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Sofueva, S. et al. Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J. 32, 3119–3129 (2013).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Van Bortle, K. & Corces, V. G. tDNA insulators and the emerging role of TFIIIC in genome organization. Transcription 3, 277–284 (2012).
Van Bortle, K. et al. Insulator function and topological domain border strength scale with architectural protein occupancy. Genome Biol. 15, R82 (2014).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
de Wit, E. et al. CTCF binding polarity determines chromatin looping. Mol. Cell 60, 676–684 (2015).
Hanssen, L. L. P. et al. Tissue-specific CTCF-cohesin-mediated chromatin architecture delimits enhancer interactions and function in vivo. Nat. Cell Biol. 19, 952–961 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Gambetta, M. C. & Furlong, E. E. M. The insulator protein CTCF is required for correct hox gene expression, but not for embryonic development in Drosophila. Genetics 210, 129–136 (2018).
Hsieh, T. H. et al. Mapping nucleosome resolution chromosome folding in yeast by Micro-C. Cell 162, 108–119 (2015).
Donze, D. & Kamakaka, R. T. RNA polymerase III and RNA polymerase II promoter complexes are heterochromatin barriers in Saccharomyces cerevisiae. EMBO J. 20, 520–531 (2001).
Yuen, K. C., Slaughter, B. D. & Gerton, J. L. Condensin II is anchored by TFIIIC and H3K4me3 in the mammalian genome and supports the expression of active dense gene clusters. Sci. Adv. 3, e1700191 (2017).
Le, T. B., Imakaev, M. V., Mirny, L. A. & Laub, M. T. High-resolution mapping of the spatial organization of a bacterial chromosome. Science 342, 731–734 (2013).
Marbouty, M. et al. Condensin- and replication-mediated bacterial chromosome folding and origin condensation revealed by Hi-C and super-resolution imaging. Mol. Cell 59, 588–602 (2015).
Le, T. B. & Laub, M. T. Transcription rate and transcript length drive formation of chromosomal interaction domain boundaries. EMBO J. 35, 1582–1595 (2016).
Liu, C. et al. Genome-wide analysis of chromatin packing in Arabidopsis thaliana at single-gene resolution. Genome Res. 26, 1057–1068 (2016).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Ramirez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Ulianov, S. V. et al. Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res. 26, 70–84 (2016).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Li, L. et al. Widespread rearrangement of 3D chromatin organization underlies polycomb-mediated stress-induced silencing. Mol. Cell 58, 216–231 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Darrow, E. M. et al. Deletion of DXZ4 on the human inactive X chromosome alters higher-order genome architecture. Proc. Natl Acad. Sci. USA 113, E4504–E4512 (2016).
Minajigi, A. et al. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015).
Ke, Y. et al. 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170, 367–381 (2017).
Kaaij, L. J. T., van der Weide, R. H., Ketting, R. F. & de Wit, E. Systemic loss and gain of chromatin architecture throughout zebrafish development. Cell Rep. 24, 1–10 (2018).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell 67, 837–852 (2017).
El-Sharnouby, S. et al. Regions of very low H3K27me3 partition the Drosophila genome into topological domains. PLOS ONE 12, e0172725 (2017).
Long, H. K., Prescott, S. L. & Wysocka, J. Ever-changing landscapes: transcriptional enhancers in development and evolution. Cell 167, 1170–1187 (2016).
Spurrell, C. H., Dickel, D. E. & Visel, A. The ties that bind: mapping the dynamic enhancer-promoter interactome. Cell 167, 1163–1166 (2016).
Andrey, G. & Mundlos, S. The three-dimensional genome: regulating gene expression during pluripotency and development. Development 144, 3646–3658 (2017).
Furlong, E. E. M. & Levine, M. Developmental enhancers and chromosome topology. Science 361, 1341–1345 (2018).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012).
Deng, W. et al. Reactivation of developmentally silenced globin genes by forced chromatin looping. Cell 158, 849–860 (2014).
Spilianakis, C. G. & Flavell, R. A. Long-range intrachromosomal interactions in the T helper type 2 cytokine locus. Nat. Immunol. 5, 1017–1027 (2004).
Andrey, G. et al. A switch between topological domains underlies HoxD genes collinearity in mouse limbs. Science 340, 1234167 (2013).
Rubin, A. J. et al. Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat. Genet. 49, 1522–1528 (2017).
Alexander, J. M., Guan, J., Huang, B., Lomvardas, S. & Weiner, O. D. Live-cell imaging reveals enhancer-dependent Sox2 transcription in the absence of enhancer proximity. Preprint at bioRxiv. https://doi.org/10.1101/409672 (2018).
Benabdallah, N. S. et al. PARP mediated chromatin unfolding is coupled to long- range enhancer activation. Preprint at bioRxiv. https://doi.org/10.1101/155325 (2017).
Chen, H. et al. Dynamic interplay between enhancer-promoter topology and gene activity. Nat. Genet. 50, 1296–1303 (2018).
Lefevre, P., Witham, J., Lacroix, C. E., Cockerill, P. N. & Bonifer, C. The LPS-induced transcriptional upregulation of the chicken lysozyme locus involves CTCF eviction and noncoding RNA transcription. Mol. Cell 32, 129–139 (2008).
Lengronne, A. et al. Cohesin relocation from sites of chromosomal loading to places of convergent transcription. Nature 430, 573–578 (2004).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
Chernukhin, I. et al. CTCF interacts with and recruits the largest subunit of RNA polymerase II to CTCF target sites genome-wide. Mol. Cell. Biol. 27, 1631–1648 (2007).
Ruiz-Velasco, M. et al. CTCF-mediated chromatin loops between promoter and gene body regulate alternative splicing across individuals. Cell Syst. 5, 628–637 (2017).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A. Phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Boehning, M. et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Biol. 25, 833–840 (2018).
Jackson, D. A., Hassan, A. B., Errington, R. J. & Cook, P. R. Visualization of focal sites of transcription within human nuclei. EMBO J. 12, 1059–1065 (1993).
van Steensel, B. et al. Localization of the glucocorticoid receptor in discrete clusters in the cell nucleus. J. Cell Sci. 108, 3003–3011 (1995).
Cho, W. K. et al. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361, 412–415 (2018).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Nuebler, J., Fudenberg, G., Imakaev, M., Abdennur, N. & Mirny, L. A. Chromatin organization by an interplay of loop extrusion and compartmental segregation. Proc. Natl Acad. Sci. USA 115, E6697–E6706 (2018).
Kueng, S., Oppikofer, M. & Gasser, S. M. SIR proteins and the assembly of silent chromatin in budding yeast. Annu. Rev. Genet. 47, 275–306 (2013).
Tolhuis, B. et al. Interactions among polycomb domains are guided by chromosome architecture. PLOS Genet. 7, e1001343 (2011).
Bantignies, F. et al. Polycomb-dependent regulatory contacts between distant Hox loci in Drosophila. Cell 144, 214–226 (2011).
Ogiyama, Y., Schuettengruber, B., Papadopoulos, G. L., Chang, J. M. & Cavalli, G. Polycomb-dependent chromatin looping contributes to gene silencing during Drosophila development. Mol. Cell 71, 73–88 (2018).
Zhu, Y. et al. Comprehensive characterization of neutrophil genome topology. Genes Dev. 31, 141–153 (2017).
Csink, A. K. & Henikoff, S. Genetic modification of heterochromatic association and nuclear organization in Drosophila. Nature 381, 529–531 (1996).
Seum, C., Delattre, M., Spierer, A. & Spierer, P. Ectopic HP1 promotes chromosome loops and variegated silencing in Drosophila. EMBO J. 20, 812–818 (2001).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Cabianca, D. S. & Gasser, S. M. Spatial segregation of heterochromatin: Uncovering functionality in a multicellular organism. Nucleus 7, 301–307 (2016).
Boumendil, C., Hari, P., Olsen, K. C. F., Acosta, J. C. & Bickmore, W. A. Nuclear pore density controls heterochromatin reorganization during senescence. Genes Dev. 33, 144–149 (2019).
Allshire, R. C. & Madhani, H. D. Ten principles of heterochromatin formation and function. Nat. Rev. Mol. Cell Biol. 19, 229–244 (2018).
Acknowledgements
The authors thank members of their laboratories, E. de Wit and anonymous reviewers for helpful comments. B.v.S. and E.E.M.F. are supported by European Research Council (ERC) Advanced Grants, GoCADiSC (694466) and DeCRyPT (787611), respectively. The Oncode Institute is supported by KWF Dutch Cancer Society.
Reviewer information
Nature Reviews Molecular Cell Biology thanks B. Bruneau, G. Almouzni and other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Alu elements
-
A type of short and highly abundant transposable element found throughout primate genomes.
- Histone modifications
-
A generic term for a wide range of post-translational modifications of histone residues. Histone modifications have a variety of functions, including in the packaging of chromatin and regulation of transcription.
- Nuclear lamina
-
A layer of proteins coating the inner nuclear membrane and thought to form a large contact surface for lamina-associated domains.
- CCCTC-binding factor
-
(CTCF). A DNA-binding protein that often marks borders of lamina-associated domains, topologically associated domains and chromatin loops and can act as a transcriptional insulator.
- Hi-C
-
A chromosome conformation capture method that systematically identifies genomic sequences that are in close proximity to one another inside cell nuclei.
- Super enhancers
-
A somewhat arbitrary definition of genomic regions that contain a high density of active enhancers.
Rights and permissions
About this article
Cite this article
van Steensel, B., Furlong, E.E.M. The role of transcription in shaping the spatial organization of the genome. Nat Rev Mol Cell Biol 20, 327–337 (2019). https://doi.org/10.1038/s41580-019-0114-6
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-019-0114-6
This article is cited by
-
Regulation of chromatin organization during animal regeneration
Cell Regeneration (2023)
-
Human artificial chromosome carrying 3p21.3-p22.2 region suppresses hTERT transcription in oral cancer cells
Chromosome Research (2023)
-
Research Progress of Long Non-coding RNAs in Spinal Cord Injury
Neurochemical Research (2023)
-
A comparison of topologically associating domain callers over mammals at high resolution
BMC Bioinformatics (2022)
-
The role of epigenetic mechanisms in the regulation of gene expression in the cyclical endometrium
Clinical Epigenetics (2021)