Abstract
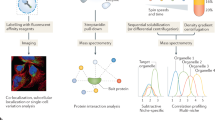
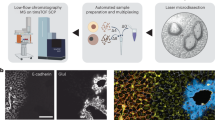
Protein subcellular localization is tightly controlled and intimately linked to protein function in health and disease. Capturing the spatial proteome — that is, the localizations of proteins and their dynamics at the subcellular level — is therefore essential for a complete understanding of cell biology. Owing to substantial advances in microscopy, mass spectrometry and machine learning applications for data analysis, the field is now mature for proteome-wide investigations of spatial cellular regulation. Studies of the human proteome have begun to reveal a complex architecture, including single-cell variations, dynamic protein translocations, changing interaction networks and proteins localizing to multiple compartments. Furthermore, several studies have successfully harnessed the power of comparative spatial proteomics as a discovery tool to unravel disease mechanisms. We are at the beginning of an era in which spatial proteomics finally integrates with cell biology and medical research, thereby paving the way for unbiased systems-level insights into cellular processes. Here, we discuss current methods for spatial proteomics using imaging or mass spectrometry and specifically highlight global comparative applications. The aim of this Review is to survey the state of the field and also to encourage more cell biologists to apply spatial proteomics approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mitrea, D. M. & Kriwacki, R. W. Phase separation in biology; functional organization of a higher order. Cell Commun. Signal 14, 1 (2016).
Wheeler, R. J. & Hyman, A. A. Controlling compartmentalization by non-membrane-bound organelles. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 373, 20170193 (2018).
Bauer, N. C., Doetsch, P. W. & Corbett, A. H. Mechanisms regulating protein localization. Traffic 16, 1039–1061 (2015).
Guardia, C. M., De Pace, R., Mattera, R. & Bonifacino, J. S. Neuronal functions of adaptor complexes involved in protein sorting. Curr. Opin. Neurobiol. 51, 103–110 (2018).
Banworth, M. J. & Li, G. Consequences of Rab GTPase dysfunction in genetic or acquired human diseases. Small GTPases 9, 158–181 (2018).
Bridges, R. J. & Bradbury, N. A. Cystic fibrosis, cystic fibrosis transmembrane conductance regulator and drugs: insights from cellular trafficking. Handb Exp. Pharmacol. 245, 385–425 (2018).
Meyer, K. et al. Mutations in disordered regions can cause disease by creating dileucine motifs. Cell 175, 239–253 (2018).
Mattiazzi Usaj, M. et al. High-content screening for quantitative cell biology. Trends Cell Biol. 26, 598–611 (2016).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 (2016).
Hosp, F. & Mann, M. A primer on concepts and applications of proteomics in neuroscience. Neuron 96, 558–571 (2017).
Larance, M. & Lamond, A. I. Multidimensional proteomics for cell biology. Nat. Rev. Mol. Cell Biol. 16, 269–280 (2015).
Breker, M. & Schuldiner, M. The emergence of proteome-wide technologies: systematic analysis of proteins comes of age. Nat. Rev. Mol. Cell Biol. 15, 453–464 (2014).
Kim, D. I. & Roux, K. J. Filling the void: proximity-based labeling of proteins in living cells. Trends Cell Biol. 26, 804–817 (2016).
Lonn, P. & Landegren, U. Close encounters - probing proximal proteins in live or fixed cells. Trends Biochem. Sci. 42, 504–515 (2017).
Sullivan, D. P. et al. Deep learning is combined with massive-scale citizen science to improve large-scale image classification. Nat. Biotechnol. 36, 820–828 (2018).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Chong, Y. T. et al. Yeast proteome dynamics from single cell imaging and automated analysis. Cell 162, 1413–1424 (2015).
Itzhak, D. N. et al. A mass spectrometry-based approach for mapping protein subcellular localization reveals the spatial proteome of mouse primary neurons. Cell Rep. 20, 2706–2718 (2017).
Itzhak, D. N., Tyanova, S., Cox, J. & Borner, G. H. Global, quantitative and dynamic mapping of protein subcellular localization. eLife 5, e16950 (2016). This study presents the first demonstration that MS-based organellar maps of the cell can be used as an unbiased discovery tool, exemplified by EGF signalling. The study also includes a detailed description of how to generate maps and a database of the subcellular localization and copy number information for 8,700 human proteins.
Walther, N. et al. A quantitative map of human Condensins provides new insights into mitotic chromosome architecture. J. Cell Biol. 217, 2309–2328 (2018).
Ma, J. et al. Using deep learning to model the hierarchical structure and function of a cell. Nat. Methods 15, 290–298 (2018).
Jean Beltran, P. M., Mathias, R. A. & Cristea, I. M. A portrait of the human organelle proteome in space and time during cytomegalovirus infection. Cell Syst. 3, 361–373 (2016). This is the first study to charter the dynamics of a spatial proteome by MS over time during viral infection (with HCMV), revealing new insights into the interplay between the virus and the host cell.
Krahmer, N. et al. Organellar proteomics and phospho-proteomics reveal subcellular reorganization in diet-induced hepatic steatosis. Dev. Cell 47, 205–221 (2018). This is the first application of comparative MS-based organellar mapping to a mammalian tissue, the mouse liver. Organellar rearrangements caused by a high-fat diet were captured, providing a holistic view of the pathological changes during hepatic steatosis.
Davies, A. K. et al. AP-4 vesicles contribute to spatial control of autophagy via RUSC-dependent peripheral delivery of ATG9A. Nat. Commun. 9, 3958 (2018). This is the first example of how MS-based organellar maps of the cell can be used to pinpoint the molecular basis of a genetic disorder, in this case the neurodegenerative AP-4 deficiency syndrome.
Hirst, J., Itzhak, D. N., Antrobus, R., Borner, G. H. H. & Robinson, M. S. Role of the AP-5 adaptor protein complex in late endosome-to-Golgi retrieval. PLOS Biol. 16, e2004411 (2018).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Sharma, K. et al. Cell type- and brain region-resolved mouse brain proteome. Nat. Neurosci. 18, 1819–1831 (2015).
Doll, S. et al. Region and cell-type resolved quantitative proteomic map of the human heart. Nat. Commun. 8, 1469 (2017).
Takamori, S. et al. Molecular anatomy of a trafficking organelle. Cell 127, 831–846 (2006).
Tharkeshwar, A. K., Gevaert, K. & Annaert, W. Organellar omics-a reviving strategy to untangle the biomolecular complexity of the cell. Proteomics 18, e1700113 (2018).
De Duve, C. Principles of tissue fractionation. J. Theor. Biol. 6, 33–59 (1964).
Andersen, J. S. et al. Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574 (2003).
Krahmer, N. et al. Protein correlation profiles identify lipid droplet proteins with high confidence. Mol. Cell Proteom. 12, 1115–1126 (2013).
Harner, M. et al. The mitochondrial contact site complex, a determinant of mitochondrial architecture. EMBO J. 30, 4356–4370 (2011).
Morgenstern, M. et al. Definition of a high-confidence mitochondrial proteome at quantitative scale. Cell Rep. 19, 2836–2852 (2017).
Wyant, G. A. et al. NUFIP1 is a ribosome receptor for starvation-induced ribophagy. Science 360, 751–758 (2018).
Borner, G. H. et al. Multivariate proteomic profiling identifies novel accessory proteins of coated vesicles. J. Cell Biol. 197, 141–160 (2012).
Borner, G. H. et al. Fractionation profiling: a fast and versatile approach for mapping vesicle proteomes and protein-protein interactions. Mol. Biol. Cell 25, 3178–3194 (2014).
Wuhr, M. et al. The nuclear proteome of a vertebrate. Curr. Biol. 25, 2663–2671 (2015).
Weekes, M. P. et al. Quantitative temporal viromics: an approach to investigate host-pathogen interaction. Cell 157, 1460–1472 (2014).
Peikert, C. D. et al. Charting organellar importomes by quantitative mass spectrometry. Nat. Commun. 8, 15272 (2017).
Gatto, L. et al. A foundation for reliable spatial proteomics data analysis. Mol. Cell Proteom. 13, 1937–1952 (2014).
Lund-Johansen, F. et al. MetaMass, a tool for meta-analysis of subcellular proteomics data. Nat. Methods 13, 837–840 (2016).
Breckels, L. M. et al. The effect of organelle discovery upon sub-cellular protein localisation. J. Proteom. 88, 129–140 (2013).
Christoforou, A. et al. A draft map of the mouse pluripotent stem cell spatial proteome. Nat. Commun. 7, 8992 (2016).
Dunkley, T. P., Watson, R., Griffin, J. L., Dupree, P. & Lilley, K. S. Localization of organelle proteins by isotope tagging (LOPIT). Mol. Cell Proteom. 3, 1128–1134 (2004).
Gilchrist, A. et al. Quantitative proteomics analysis of the secretory pathway. Cell 127, 1265–1281 (2006).
Jadot, M. et al. Accounting for protein subcellular localization: a compartmental map of the rat liver proteome. Mol. Cell Proteom. 16, 194–212 (2017).
Mardakheh, F. K. et al. Proteomics profiling of interactome dynamics by colocalisation analysis (COLA). Mol. Biosyst. 13, 92–105 (2016).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017). This study presents the first near proteome-wide spatial map of human cells, at unprecedented resolution in terms of the number of organelles and structures that were mapped, which represents the Cell Atlas component of the HPA. This study provides localization data for more than 12,000 human proteins and revealed a very high number of multilocalizing proteins and a considerable level of single-cell variability.
Hein, M. Y. et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163, 712–723 (2015).
Huttlin, E. L. et al. Architecture of the human interactome defines protein communities and disease networks. Nature 545, 505–509 (2017).
Mackinder, L. C. M. et al. A spatial interactome reveals the protein organization of the algal CO2-concentrating mechanism. Cell 171, 133–147 (2017). This is an example of how AP–MS interaction networks can be used in conjunction with imaging to obtain a high-resolution architectural map of a subcellular compartment, the algal pyrenoid.
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018). This study reports on the latest generation of enzymes for proximity labelling, which can be applied for spatial protein interaction mapping in cultured cells and also in vivo, as demonstrated for flies and worms.
Hung, V. et al. Proteomic mapping of the human mitochondrial intermembrane space in live cells via ratiometric APEX tagging. Mol. Cell 55, 332–341 (2014).
Rhee, H. W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Youn, J. Y. et al. High-density proximity mapping reveals the subcellular organization of mRNA-associated granules and bodies. Mol. Cell 69, 517–532 (2018).
Gupta, G. D. et al. A dynamic protein interaction landscape of the human centrosome-cilium interface. Cell 163, 1484–1499 (2015).
Han, S., Li, J. & Ting, A. Y. Proximity labeling: spatially resolved proteomic mapping for neurobiology. Curr. Opin. Neurobiol. 50, 17–23 (2018).
Liu, X. et al. An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations. Nat. Commun. 9, 1188 (2018).
Lobingier, B. T. et al. An approach to spatiotemporally resolve protein interaction networks in living cells. Cell 169, 350–360 (2017).
Paek, J. et al. Multidimensional tracking of GPCR signaling via peroxidase-catalyzed proximity labeling. Cell 169, 338–349 (2017).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003). This study presents the first proteome-wide spatial map of a eukaryotic cell, and revolutionized yeast biology. The yeast library created as part of this work has been used to study, for example, the proteome response to perturbations, single-cell variability, stochasticity and noise in gene expression.
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006).
Breker, M., Gymrek, M. & Schuldiner, M. A novel single-cell screening platform reveals proteome plasticity during yeast stress responses. J. Cell Biol. 200, 839–850 (2013).
Lu, A. X. et al. Integrating images from multiple microscopy screens reveals diverse patterns of change in the subcellular localization of proteins. eLife 7, e31872 (2018). This study reports a meta-analysis of 24 imaging-based yeast screens using unsupervised computational image analysis. Specific and general spatial relocalization responses to various environmental perturbations are identified. Importantly, this study shows that the proportion of the proteome that relocalizes is similar to the proportion showing changes in abundance and that these two responses constitute independent layers of cellular regulation.
Balazsi, G., van Oudenaarden, A. & Collins, J. J. Cellular decision making and biological noise: from microbes to mammals. Cell 144, 910–925 (2011).
Rubakhin, S. S., Lanni, E. J. & Sweedler, J. V. Progress toward single cell metabolomics. Curr. Opin. Biotechnol. 24, 95–104 (2013).
Narayanaswamy, R. et al. Widespread reorganization of metabolic enzymes into reversible assemblies upon nutrient starvation. Proc. Natl Acad. Sci. USA 106, 10147–10152 (2009).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Denervaud, N. et al. A chemostat array enables the spatio-temporal analysis of the yeast proteome. Proc. Natl Acad. Sci. USA 110, 15842–15847 (2013).
Tkach, J. M. et al. Dissecting DNA damage response pathways by analysing protein localization and abundance changes during DNA replication stress. Nat. Cell Biol. 14, 966–976 (2012).
Torres, N. P., Ho, B. & Brown, G. W. High-throughput fluorescence microscopic analysis of protein abundance and localization in budding yeast. Crit. Rev. Biochem. Mol. Biol. 51, 110–119 (2016).
Coons, A. H., Creech, H. J., Jones, R. N. & Berliner, E. The demonstration of pneumococcal antigen in tissues by the use of fluorescent antibody. J. Immunol. 45, 159–170 (1942).
Lazarides, E. & Weber, K. Actin antibody: the specific visualization of actin filaments in non-muscle cells. Proc. Natl Acad. Sci. USA 71, 2268–2272 (1974).
Schnell, U., Dijk, F., Sjollema, K. A. & Giepmans, B. N. Immunolabeling artifacts and the need for live-cell imaging. Nat. Methods 9, 152–158 (2012).
Stadler, C., Skogs, M., Brismar, H., Uhlen, M. & Lundberg, E. A single fixation protocol for proteome-wide immunofluorescence localization studies. J. Proteom. 73, 1067–1078 (2010).
Simeon, R. & Chen, Z. In vitro-engineered non-antibody protein therapeutics. Protein Cell 9, 3–14 (2018).
Stadler, C. et al. Immunofluorescence and fluorescent-protein tagging show high correlation for protein localization in mammalian cells. Nat. Methods 10, 315–323 (2013).
Baker, M. Reproducibility crisis: blame it on the antibodies. Nature 521, 274–276 (2015).
Uhlen, M. et al. A proposal for validation of antibodies. Nat. Methods 13, 823–827 (2016).
Bandrowski, A. et al. The Resource Identification Initiative: a cultural shift in publishing. F1000Res 4, 134 (2015).
Uhlen, M. et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 28, 1248–1250 (2010).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507 (2017).
Nilsson, P. et al. Towards a human proteome atlas: high-throughput generation of mono-specific antibodies for tissue profiling. Proteomics 5, 4327–4337 (2005).
Uhlen, M. et al. A human protein atlas for normal and cancer tissues based on antibody proteomics. Mol. Cell Proteom. 4, 1920–1932 (2005).
Algenas, C. et al. Antibody performance in western blot applications is context-dependent. Biotechnol. J. 9, 435–445 (2014).
Skogs, M. et al. Antibody validation in bioimaging applications based on endogenous expression of tagged proteins. J. Proteome Res. 16, 147–155 (2017).
Stadler, C. et al. Systematic validation of antibody binding and protein subcellular localization using siRNA and confocal microscopy. J. Proteom. 75, 2236–2251 (2012).
Angelo, M. et al. Multiplexed ion beam imaging of human breast tumors. Nat. Med. 20, 436–442 (2014).
Bodenmiller, B. Multiplexed epitope-based tissue imaging for discovery and healthcare applications. Cell Syst. 2, 225–238 (2016).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplex imaging. Cell 174, 968–981 (2018).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Meurer, M. et al. Genome-wide C-SWAT library for high-throughput yeast genome tagging. Nat. Methods 15, 598–600 (2018).
Weill, U. et al. Genome-wide SWAp-Tag yeast libraries for proteome exploration. Nat. Methods 15, 617–622 (2018).
Weill, U. et al. Toolbox: creating a systematic database of secretory pathway proteins uncovers new cargo for COPI. Traffic 19, 370–379 (2018).
Breker, M., Gymrek, M., Moldavski, O. & Schuldiner, M. LoQAtE — Localization and Quantitation ATlas of the yeast proteomE. A new tool for multiparametric dissection of single-protein behavior in response to biological perturbations in yeast. Nucleic Acids Res. 42, D726–D730 (2014).
Riffle, M. & Davis, T. N. The Yeast Resource Center Public Image Repository: a large database of fluorescence microscopy images. BMC Bioinformatics 11, 263 (2010).
Chuartzman, S. G. & Schuldiner, M. Database for High Throughput Screening Hits (dHITS): a simple tool to retrieve gene specific phenotypes from systematic screens done in yeast. Yeast 35, 477–483 (2018).
Cherry, J. M. et al. Saccharomyces Genome Database: the genomics resource of budding yeast. Nucleic Acids Res. 40, D700–D705 (2012).
Koh, J. L. et al. CYCLoPs: a comprehensive database constructed from automated analysis of protein abundance and subcellular localization patterns in Saccharomyces cerevisiae. G3 (Bethesda) 5, 1223–1232 (2015).
Dubreuil, B. et al. YeastRGB: comparing the abundance and localization of yeast proteins across cells and libraries. Nucleic Acids Res. https://doi.org/10.1093/nar/gky941 (2018).
Simpson, J. C., Wellenreuther, R., Poustka, A., Pepperkok, R. & Wiemann, S. Systematic subcellular localization of novel proteins identified by large-scale cDNA sequencing. EMBO Rep. 1, 287–292 (2000).
Frenkel-Morgenstern, M. et al. Dynamic proteomics: a database for dynamics and localizations of endogenous fluorescently-tagged proteins in living human cells. Nucleic Acids Res. 38, D508–D512 (2010).
Sigal, A. et al. Generation of a fluorescently labeled endogenous protein library in living human cells. Nat. Protoc. 2, 1515–1527 (2007).
Sigal, A. et al. Dynamic proteomics in individual human cells uncovers widespread cell-cycle dependence of nuclear proteins. Nat. Methods 3, 525–531 (2006).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Cho, W. K. et al. Super-resolution imaging of fluorescently labeled, endogenous RNA polymerase II in living cells with CRISPR/Cas9-mediated gene editing. Sci. Rep. 6, 35949 (2016).
Dambournet, D., Hong, S. H., Grassart, A. & Drubin, D. G. Tagging endogenous loci for live-cell fluorescence imaging and molecule counting using ZFNs, TALENs, and Cas9. Methods Enzymol. 546, 139–160 (2014).
Doyon, J. B. et al. Rapid and efficient clathrin-mediated endocytosis revealed in genome-edited mammalian cells. Nat. Cell Biol. 13, 331–337 (2011).
Ratz, M., Testa, I., Hell, S. W. & Jakobs, S. CRISPR/Cas9-mediated endogenous protein tagging for RESOLFT super-resolution microscopy of living human cells. Sci. Rep. 5, 9592 (2015).
Li-Kroeger, D. et al. An expanded toolkit for gene tagging based on MiMIC and scarless CRISPR tagging in Drosophila. eLife 7, e38709 (2018).
Merkle, F. T. et al. Efficient CRISPR-Cas9-mediated generation of knockin human pluripotent stem cells lacking undesired mutations at the targeted locus. Cell Rep. 11, 875–883 (2015).
Roberts, B. et al. Systematic gene tagging using CRISPR/Cas9 in human stem cells to illuminate cell organization. Mol. Biol. Cell 28, 2854–2874 (2017).
Feng, S. et al. Improved split fluorescent proteins for endogenous protein labeling. Nat. Commun. 8, 370 (2017).
Kamiyama, D. et al. Versatile protein tagging in cells with split fluorescent protein. Nat. Commun. 7, 11046 (2016).
Leonetti, M. D., Sekine, S., Kamiyama, D., Weissman, J. S. & Huang, B. A scalable strategy for high-throughput GFP tagging of endogenous human proteins. Proc. Natl Acad. Sci. USA 113, E3501–E3508 (2016).
Sahl, S. J., Hell, S. W. & Jakobs, S. Fluorescence nanoscopy in cell biology. Nat. Rev. Mol. Cell Biol. 18, 685–701 (2017).
Monnich, M. et al. CEP128 localizes to the subdistal appendages of the mother centriole and regulates TGFβ/BMP signaling at the primary cilium. Cell Rep. 22, 2584–2592 (2018).
Jakobsen, L. et al. Novel asymmetrically localizing components of human centrosomes identified by complementary proteomics methods. EMBO J. 30, 1520–1535 (2011).
Caicedo, J. C. et al. Data-analysis strategies for image-based cell profiling. Nat. Methods 14, 849–863 (2017).
Coelho, L. P. et al. Determining the subcellular location of new proteins from microscope images using local features. Bioinformatics 29, 2343–2349 (2013).
Li, J., Newberg, J. Y., Uhlen, M., Lundberg, E. & Murphy, R. F. Automated analysis and reannotation of subcellular locations in confocal images from the Human Protein Atlas. PLOS ONE 7, e50514 (2012).
Li, J., Xiong, L., Schneider, J. & Murphy, R. F. Protein subcellular location pattern classification in cellular images using latent discriminative models. Bioinformatics 28, i32–i39 (2012).
Schmidhuber, J. Deep learning in neural networks: an overview. Neural Netw. 61, 85–117 (2015).
Girshick, R., Donahue, J., Darrell, T. & Malik, J. in 2014 IEEE Conference on Computer Vision and Pattern Recognition 580–587 (IEEE, 2014).
He, K. M., Zhang, X. Y., Ren, S. Q. & Sun, J. in 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Parnamaa, T. & Parts, L. Accurate classification of protein subcellular localization from high-throughput microscopy images using deep learning. G3 (Bethesda) 7, 1385–1392 (2017).
Kraus, O. Z. et al. Automated analysis of high-content microscopy data with deep learning. Mol. Syst. Biol. 13, 924 (2017).
Kraus, O. Z., Ba, J. L. & Frey, B. J. Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 32, i52–i59 (2016).
Hughes, A. et al. Quanti.us: a tool for rapid, flexible, crowd-based annotation of images. Nat. Methods 15, 587–590 (2018).
Ronneberger, O., Fischer, P. & Brox, T. in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015 (Lecture Notes in Computer Science Book Series) Vol. 9351 234–241 (Springer, 2015).
Simon, A. A. et al. A probabilistic U-Net for segmentation of ambiguous images. Preprint at arXiv https://arxiv.org/abs/1806.05034 (2018).
Christiansen, E. M. et al. In silico labeling: predicting fluorescent labels in unlabeled images. Cell 173, 792–803 (2018).
Ounkomol, C., Seshamani, S., Maleckar, M. M., Collman, F. & Johnson, G. R. Label-free prediction of three-dimensional fluorescence images from transmitted light microscopy. Nat. Methods 15, 917–920 (2018).
Ellenberg, J. et al. Public archives for biological image data. Preprint at arXiv https://arxiv.org/abs/1801.10189 (2018).
Dunkley, T. P. et al. Mapping the Arabidopsis organelle proteome. Proc. Natl Acad. Sci. USA 103, 6518–6523 (2006).
Foster, L. J. et al. A mammalian organelle map by protein correlation profiling. Cell 125, 187–199 (2006).
Mazumder, A., Pesudo, L. Q., McRee, S., Bathe, M. & Samson, L. D. Genome-wide single-cell-level screen for protein abundance and localization changes in response to DNA damage in S. cerevisiae. Nucleic Acids Res. 41, 9310–9324 (2013).
Cai, Y. et al. Experimental and computational framework for a dynamic protein atlas of human cell division. Nature 561, 411–415 (2018). This is a seminal study that integrates imaging-based spatiotemporal protein mapping with absolute quantification to enable stoichiometric modelling of the spatial and temporal dynamics of cell division — this paper shows where the field is heading.
Politi, A. Z. et al. Quantitative mapping of fluorescently tagged cellular proteins using FCS-calibrated four-dimensional imaging. Nat. Protoc. 13, 1445–1464 (2018).
Larochelle, S. Tracking the proteome. Nat. Methods 13, 821–821 (2016).
Lu, A. X. & Moses, A. M. An unsupervised kNN method to systematically detect changes in protein localization in high-throughput microscopy images. PLOS ONE 11, e0158712 (2016).
Nesvizhskii, A. I. Proteogenomics: concepts, applications and computational strategies. Nat. Methods 11, 1114–1125 (2014).
Queiroz, R. M. L. et al. Unbiased dynamic characterization of RNA-protein interactions by OOPS. Preprint at bioRxiv https://doi.org/10.1101/333336 (2018).
Kaewsapsak, P., Shechner, D. M., Mallard, W., Rinn, J. L. & Ting, A. Y. Live-cell mapping of organelle-associated RNAs via proximity biotinylation combined with protein-RNA crosslinking. eLife 6, e29224 (2017).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-seq. Preprint at bioRxiv https://doi.org/10.1101/454470 (2018).
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
Jeffery, C. J. Moonlighting proteins. Trends Biochem. Sci. 24, 8–11 (1999).
Jeffery, C. J. Why study moonlighting proteins? Front. Genet. 6, 211 (2015).
Jeffery, C. J. Protein species and moonlighting proteins: very small changes in a protein’s covalent structure can change its biochemical function. J. Proteom. 134, 19–24 (2016).
Franco-Serrano, L. et al. MultitaskProtDB-II: an update of a database of multitasking/moonlighting proteins. Nucleic Acids Res. 46, D645–D648 (2018).
Chen, C., Zabad, S., Liu, H. P., Wang, F. & Jeffery, C. MoonProt 2.0: an expansion and update of the moonlighting proteins database. Nucleic Acids Res. 46, D640–D644 (2018).
Chapple, C. E. et al. Extreme multifunctional proteins identified from a human protein interaction network. Nat. Commun. 6, 7412 (2015).
Khan, I. K., Bhuiyan, M. & Kihara, D. DextMP: deep dive into text for predicting moonlighting proteins. Bioinformatics 33, i83–i91 (2017).
Smith, L. M. & Kelleher, N. L. & The Consortium for Top Down Proteomics. Proteoform: a single term describing protein complexity. Nat. Methods 10, 186–187 (2013).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699 (2018).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Ponomarenko, E. A. et al. The size of the human proteome: the width and depth. Int. J. Anal. Chem. 2016, 7436849 (2016).
Skinner, O. S. et al. Top-down characterization of endogenous protein complexes with native proteomics. Nat. Chem. Biol. 14, 36–41 (2018).
Aebersold, R. et al. How many human proteoforms are there? Nat. Chem. Biol. 14, 206–214 (2018).
Yang, X. et al. Widespread expansion of protein interaction capabilities by alternative splicing. Cell 164, 805–817 (2016).
Zhu, Y. et al. Nanodroplet processing platform for deep and quantitative proteome profiling of 10–100 mammalian cells. Nat. Commun. 9, 882 (2018).
Acknowledgements
E.L. acknowledges funding from the Knut and Alice Wallenberg foundation (KAW 2016.0204), the Swedish Research Council (2017-05327) and the Chan Zuckerberg Initiative (173965 (5022)). G.H.H.B. received funding from the German Research Foundation (DFG/Gottfried Wilhelm Leibniz Prize MA 1764/2-1) and the Max Planck Society for the Advancement of Science.
Reviewer information
Nature Reviews Molecular Cell Biology thanks G. W. Brown and other anonymous reviewer(s), for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Both authors contributed equally to all aspects of preparing the article (researching data for the article, substantial contributions to the discussion of the content and writing, reviewing and editing of the manuscript before submission).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
HeLa Spatial Proteome: www.mapofthecell.org
Human Protein Atlas (HPA): www.proteinatlas.org
Obesity-induced non-alcoholic fatty liver disease (NAFLD): http://nafld-organellemap.org/
SpatialMap: http://spatialmap.org/
Supplementary information
Glossary
- Dynamic protein translocation
-
Translocation describes the movement of a protein between cellular compartments. Dynamic translocation refers to a constant change in translocation activity.
- Multimodal organellar distribution
-
Refers to the distribution of proteins that simultaneously localize to multiple compartments within a cell.
- Affinity reagents
-
Molecules, such as an antibody, protein, peptide or nucleic acid, that bind specifically to a target protein to enable the identification, visualization, capture or modulation of the target protein or its activity.
- Tandem mass tag multiplexing
-
A strategy for quantitative proteomic analyses. Peptides from multiple samples are labelled with different mass tags, pooled and analysed as a single sample by mass spectrometry. The tags can be distinguished by their mass and thus enable the simultaneous, relative quantification of peptide and protein abundances across several samples.
- Segmentation
-
Describes the process of partitioning a digital image into segments that represent, for example, a cell or a nucleus.
- Citizen science
-
Public participation in scientific research.
- Tagged yeast libraries
-
Genome-wide libraries of yeast cells, each expressing a protein fused to a fluorescent reporter protein (such as GFP).
- Proteogenomics
-
Integration of proteomics, transcriptomics and genomics for the discovery and identification of peptides using mass spectrometry. Practically, DNA or RNA sequence information is used to provide an experiment-specific or cell-type-specific tailored database for proteomic protein identification rather than a generic organism-specific database.
Rights and permissions
About this article
Cite this article
Lundberg, E., Borner, G.H.H. Spatial proteomics: a powerful discovery tool for cell biology. Nat Rev Mol Cell Biol 20, 285–302 (2019). https://doi.org/10.1038/s41580-018-0094-y
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-018-0094-y
This article is cited by
-
Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases
Genome Medicine (2024)
-
The interaction between ageing and Alzheimer's disease: insights from the hallmarks of ageing
Translational Neurodegeneration (2024)
-
Integration of pan-omics technologies and three-dimensional in vitro tumor models: an approach toward drug discovery and precision medicine
Molecular Cancer (2024)
-
Mapping cancer biology in space: applications and perspectives on spatial omics for oncology
Molecular Cancer (2024)
-
Cross-link assisted spatial proteomics to map sub-organelle proteomes and membrane protein topologies
Nature Communications (2024)