Abstract
Lipid droplets are storage organelles at the centre of lipid and energy homeostasis. They have a unique architecture consisting of a hydrophobic core of neutral lipids, which is enclosed by a phospholipid monolayer that is decorated by a specific set of proteins. Originating from the endoplasmic reticulum, lipid droplets can associate with most other cellular organelles through membrane contact sites. It is becoming apparent that these contacts between lipid droplets and other organelles are highly dynamic and coupled to the cycles of lipid droplet expansion and shrinkage. Importantly, lipid droplet biogenesis and degradation, as well as their interactions with other organelles, are tightly coupled to cellular metabolism and are critical to buffer the levels of toxic lipid species. Thus, lipid droplets facilitate the coordination and communication between different organelles and act as vital hubs of cellular metabolism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Schaffer, J. E. Lipotoxicity: when tissues overeat. Curr. Opin. Lipidol. 14, 281–287 (2003).
Fawcett, D. W. An Atlas of Fine Structure: The Cell, Its Organelles, and Inclusions (Saunders, 1966).
Farese, R. V. & Walther, T. C. Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 139, 855–860 (2009).
Sorger, D., Athenstaedt, K., Hrastnik, C. & Daum, G. A yeast strain lacking lipid particles bears a defect in ergosterol formation. J. Biol. Chem. 279, 31190–31196 (2004).
Sandager, L. et al. Storage lipid synthesis is non-essential in yeast. J. Biol. Chem. 277, 6478–6482 (2002).
Petschnigg, J. et al. Good fat, essential cellular requirements for triacylglycerol synthesis to maintain membrane homeostasis in yeast. J. Biol. Chem. 284, 30981–30993 (2009).
Velázquez, A. P., Tatsuta, T., Ghillebert, R., Drescher, I. & Graef, M. Lipid droplet-mediated ER homeostasis regulates autophagy and cell survival during starvation. J. Cell Biol. 212, 621–631 (2016).
Gaspar, M. L., Hofbauer, H. F., Kohlwein, S. D. & Henry, S. A. Coordination of storage lipid synthesis and membrane biogenesis: evidence for cross-talk between triacylglycerol metabolism and phosphatidylinositol synthesis. J. Biol. Chem. 286, 1696–1708 (2011).
Khandelia, H., Duelund, L., Pakkanen, K. I. & Ipsen, J. H. Triglyceride blisters in lipid bilayers: implications for lipid droplet biogenesis and the mobile lipid signal in cancer cell membranes. PLOS ONE 5, e12811 (2010).
Duelund, L. et al. Composition, structure and properties of POPC-triolein mixtures. Evidence of triglyceride domains in phospholipid bilayers. Biochim. Biophys. Acta 1828, 1909–1917 (2013).
Thiam, A. R. & Forêt, L. The physics of lipid droplet nucleation, growth and budding. Biochim. Biophys. Acta 1861, 715–722 (2016).
Choudhary, V., Ojha, N., Golden, A. & Prinz, W. A. A conserved family of proteins facilitates nascent lipid droplet budding from the ER. J. Cell Biol. 211, 261–271 (2015). This study uses electron tomography to visualize the lens-like structure in early stages of lipid droplet biogenesis.
Wang, H. et al. Seipin is required for converting nascent to mature lipid droplets. eLife 5, e16582 (2016).
Kassan, A. et al. Acyl-CoA synthetase 3 promotes lipid droplet biogenesis in ER microdomains. J. Cell Biol. 203, 985–1001 (2013).
Gomez-Navarro, N. & Miller, E. A. COP-coated vesicles. Curr. Biol. 26, R54–R57 (2016).
Adeyo, O. et al. The yeast lipin orthologue Pah1p is important for biogenesis of lipid droplets. J. Cell Biol. 192, 1043–1055 (2011).
Fei, W. et al. A role for phosphatidic acid in the formation of “supersized” lipid droplets. PLOS Genet. 7, e1002201 (2011).
Skinner, J. R. et al. Diacylglycerol enrichment of endoplasmic reticulum or lipid droplets recruits perilipin 3/TIP47 during lipid storage and mobilization. J. Biol. Chem. 284, 30941–30948 (2009).
Ben M’barek, K. et al. ER membrane phospholipids and surface tension control cellular lipid droplet formation. Dev. Cell 41, 591–604 (2017). This study uses artificial lipid droplets to demonstrate an important role for phospholipid composition and membrane surface tension in lipid droplet budding.
Choudhary, V. et al. Architecture of lipid droplets in endoplasmic reticulum is determined by phospholipid intrinsic curvature. Curr. Biol. 28, 915–926 (2018).
Chorlay, A. & Thiam, A. R. An asymmetry in monolayer tension regulates lipid droplet budding direction. Biophys. J. 114, 631–640 (2018).
Mishra, S. et al. Mature lipid droplets are accessible to ER luminal proteins. J. Cell Sci. 129, 3803–3815 (2016).
Kadereit, B. et al. Evolutionarily conserved gene family important for fat storage. Proc. Natl Acad. Sci. USA 105, 94–99 (2008).
Gross, D. A., Zhan, C. & Silver, D. L. Direct binding of triglyceride to fat storage-inducing transmembrane proteins 1 and 2 is important for lipid droplet formation. Proc. Natl Acad. Sci. USA 108, 19581–19586 (2011).
Hayes, M. et al. Fat storage-inducing transmembrane (FIT or FITM) proteins are related to lipid phosphatase/phosphotransferase enzymes. Microb. Cell 5, 88–103 (2017).
Becuwe, M. et al. FIT2 is a lipid phosphate phosphatase crucial for endoplasmic reticulum homeostasis. Preprint at https://www.biorxiv.org/content/early/2018/03/29/291765 (2018). References 25 and 26 identify the FIT family of proteins as lipid phosphatases that may affect lipid droplets through regulation of ER homeostasis.
Goh, V. J. et al. Postnatal deletion of fat storage-inducing transmembrane protein 2 (FIT2/FITM2) causes lethal enteropathy. J. Biol. Chem. 290, 25686–25699 (2015).
Miranda, D. A. et al. Fat storage-inducing transmembrane protein 2 is required for normal fat storage in adipose tissue. J. Biol. Chem. 289, 9560–9572 (2014).
Pomorski, T. G. & Menon, A. K. Lipid somersaults: uncovering the mechanisms of protein-mediated lipid flipping. Prog. Lipid Res. 64, 69–84 (2016).
Gao, Q. et al. Pet10p is a yeast perilipin that stabilizes lipid droplets and promotes their assembly. J. Cell Biol. 216, 3199–3217 (2017).
Sztalryd, C. & Brasaemle, D. L. The perilipin family of lipid droplet proteins: gatekeepers of intracellular lipolysis. Biochim. Biophys. Acta 1862, 1221–1232 (2017).
Bulankina, A. V. et al. TIP47 functions in the biogenesis of lipid droplets. J. Cell Biol. 185, 641–655 (2009).
Magré, J. et al. Identification of the gene altered in Berardinelli-Seip congenital lipodystrophy on chromosome 11q13. Nat. Genet. 28, 365–370 (2001).
Fei, W. et al. Fld1p, a functional homologue of human seipin, regulates the size of lipid droplets in yeast. J. Cell Biol. 180, 473–482 (2008).
Szymanski, K. M. et al. The lipodystrophy protein seipin is found at endoplasmic reticulum lipid droplet junctions and is important for droplet morphology. Proc. Natl Acad. Sci. USA 104, 20890–20895 (2007). References 34 and 35 identify seipin as a critical regulator of lipid droplet size that localizes to the ER–lipid droplet junction.
Cartwright, B. R. et al. Seipin performs dissectible functions in promoting lipid droplet biogenesis and regulating droplet morphology. Mol. Biol. Cell 26, 726–739 (2015).
Salo, V. T. et al. Seipin regulates ER-lipid droplet contacts and cargo delivery. EMBO J. 35, 2699–2716 (2016).
Grippa, A. et al. The seipin complex Fld1/Ldb16 stabilizes ER-lipid droplet contact sites. J. Cell Biol. 211, 829–844 (2015).
Wang, C.-W., Miao, Y.-H. & Chang, Y.-S. Control of lipid droplet size in budding yeast requires the collaboration between Fld1 and Ldb16. J. Cell Sci. 127, 1214–1228 (2014).
Wolinski, H. et al. Seipin is involved in the regulation of phosphatidic acid metabolism at a subdomain of the nuclear envelope in yeast. Biochim. Biophys. Acta 1851, 1450–1464 (2015).
Han, S., Binns, D. D., Chang, Y.-F. & Goodman, J. M. Dissecting seipin function: the localized accumulation of phosphatidic acid at ER/LD junctions in the absence of seipin is suppressed by Sei1p(ΔNterm) only in combination with Ldb16p. BMC Cell Biol. 16, 29 (2015).
Joshi, A. S. et al. Lipid droplet and peroxisome biogenesis occur at the same ER subdomains. Nat. Commun. 9, 2940 (2018).
Wang, S. et al. Seipin and the membrane-shaping protein Pex30 cooperate in organelle budding from the endoplasmic reticulum. Nat. Commun. 9, 2939 (2018). References 42 and 43 discover a role for Pex30 in regulating membrane domains that are required for the biogenesis of both lipid droplets and peroxisomes.
Joshi, A. S. et al. A family of membrane-shaping proteins at ER subdomains regulates pre-peroxisomal vesicle biogenesis. J. Cell Biol. 215, 515–529 (2016).
Schrul, B. & Kopito, R. R. Peroxin-dependent targeting of a lipid-droplet-destined membrane protein to ER subdomains. Nat. Cell Biol. 18, 740–751 (2016). This study demonstrates a role for PEX19 and PEX3 in the ER insertion of UBXD8, a hairpin protein that traffics from the ER to lipid droplets.
Pagac, M. et al. SEIPIN regulates lipid droplet expansion and adipocyte development by modulating the activity of glycerol-3-phosphate acyltransferase. Cell Rep. 17, 1546–1559 (2016).
Binns, D., Lee, S., Hilton, C. L., Jiang, Q.-X. & Goodman, J. M. Seipin is a discrete homooligomer. Biochemistry 49, 10747–10755 (2010).
Yan, R. et al. Human SEIPIN binds anionic phospholipids. Dev. Cell 47, 248–256 (2018).
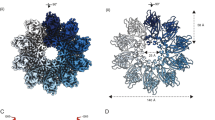
Sui, X. et al. Cryo-electron microscopy structure of the lipid droplet-formation protein seipin. J. Cell. Biol. https://doi.org/10.1083/jcb.201809067 (2018). References 48 and 49 report the cryo-electron microscopy structure of human and Drosophila melanogaster seipin, revealing an oligomeric ring conformation and a luminal lipid-binding domain.
Wilfling, F. et al. Triacylglycerol synthesis enzymes mediate lipid droplet growth by relocalizing from the ER to lipid droplets. Dev. Cell 24, 384–399 (2013). This study describes a role for lipid droplet-localized triacylglycerol synthesis enzymes in local triacylglycerol synthesis and lipid droplet expansion.
Krahmer, N. et al. Phosphatidylcholine synthesis for lipid droplet expansion is mediated by localized activation of CTP:phosphocholine cytidylyltransferase. Cell Metab. 14, 504–515 (2011).
Haider, A. et al. PCYT1A regulates phosphatidylcholine homeostasis from the inner nuclear membrane in response to membrane stored curvature elastic stress. Dev. Cell 45, 481–495 (2018).
Aitchison, A. J., Arsenault, D. J. & Ridgway, N. D. Nuclear-localized CTP:phosphocholine cytidylyltransferase α regulates phosphatidylcholine synthesis required for lipid droplet biogenesis. Mol. Biol. Cell 26, 2927–2938 (2015).
Wilfling, F. et al. Arf1/COPI machinery acts directly on lipid droplets and enables their connection to the ER for protein targeting. eLife 3, e01607 (2014).
Thiam, A. R. et al. COPI buds 60-nm lipid droplets from reconstituted water-phospholipid-triacylglyceride interfaces, suggesting a tension clamp function. Proc. Natl Acad. Sci. USA 110, 13244–13249 (2013). References 54 and 55 demonstrate the formation of ER–lipid droplet bridges through a mechanism involving ARF1–COPI-mediated regulation of lipid droplet surface tension via budding of phospholipid-rich nano-droplets.
Brasaemle, D. L., Dolios, G., Shapiro, L. & Wang, R. Proteomic analysis of proteins associated with lipid droplets of basal and lipolytically stimulated 3T3-L1 adipocytes. J. Biol. Chem. 279, 46835–46842 (2004).
Liu, P. et al. Chinese hamster ovary K2 cell lipid droplets appear to be metabolic organelles involved in membrane traffic. J. Biol. Chem. 279, 3787–3792 (2004).
Krahmer, N. et al. Protein correlation profiles identify lipid droplet proteins with high confidence. Mol. Cell. Proteomics 12, 1115–1126 (2013).
Bersuker, K. & Olzmann, J. A. Establishing the lipid droplet proteome: mechanisms of lipid droplet protein targeting and degradation. Biochim. Biophys. Acta 1862, 1166–1177 (2017).
Bersuker, K. et al. A proximity labeling strategy provides insights into the composition and dynamics of lipid droplet proteomes. Dev. Cell 44, 97–112 (2018). This study employs proximity labelling approaches in human cells to define high-confidence lipid droplet proteomes and implicates ERAD as a pathway for lipid droplet protein degradation.
Currie, E. et al. High confidence proteomic analysis of yeast LDs identifies additional droplet proteins and reveals connections to dolichol synthesis and sterol acetylation. J. Lipid Res. 55, 1465–1477 (2014).
Ingelmo-Torres, M. et al. Hydrophobic and basic domains target proteins to lipid droplets. Traffic 10, 1785–1801 (2009).
Jacquier, N. et al. Lipid droplets are functionally connected to the endoplasmic reticulum in Saccharomyces cerevisiae. J. Cell Sci. 124, 2424–2437 (2011).
Olzmann, J. A., Richter, C. M. & Kopito, R. R. Spatial regulation of UBXD8 and p97/VCP controls ATGL-mediated lipid droplet turnover. Proc. Natl Acad. Sci. USA 110, 1345–1350 (2013). This study demonstrates that UBAC2 can function as an ER-restricted tether that controls the ER–lipid droplet distribution of UBXD8.
Ruggiano, A., Mora, G., Buxó, L. & Carvalho, P. Spatial control of lipid droplet proteins by the ERAD ubiquitin ligase Doa10. EMBO J. 35, 1644–1655 (2016).
Ruggiano, A., Foresti, O. & Carvalho, P. ER-associated degradation: protein quality control and beyond. J. Cell Biol. 204, 869–879 (2014).
Stevenson, J., Huang, E. Y. & Olzmann, J. A. Endoplasmic reticulum-associated degradation and lipid homeostasis. Annu. Rev. Nutr. 36, 511–542 (2016).
Hinson, E. R. & Cresswell, P. The antiviral protein, viperin, localizes to lipid droplets via its N-terminal amphipathic alpha-helix. Proc. Natl Acad. Sci. USA 106, 20452–20457 (2009).
Barneda, D. et al. The brown adipocyte protein CIDEA promotes lipid droplet fusion via a phosphatidic acid-binding amphipathic helix. eLife 4, e07485 (2015).
Prévost, C. et al. Mechanism and determinants of amphipathic helix-containing protein targeting to lipid droplets. Dev. Cell 44, 73–86 (2018). This study demonstrates that proteins containing amphipathic helices target phospholipid packing defects in the lipid droplet surface.
Copic, A. et al. A giant amphipathic helix from a perilipin that is adapted for coating lipid droplets. Nat. Commun. 9, 1332 (2018).
Rowe, E. R. et al. Conserved amphipathic helices mediate lipid droplet targeting of perilipins 1–3. J. Biol. Chem. 291, 6664–6678 (2016).
Pataki, C. I. et al. Proteomic analysis of monolayer-integrated proteins on lipid droplets identifies amphipathic interfacial α-helical membrane anchors. Proc. Natl Acad. Sci. USA 115, E8172–E8180 (2018).
Giménez-Andrés, M., Copic, A. & Antonny, B. The many faces of amphipathic helices. Biomolecules 8, E45 (2018).
Bacle, A., Gautier, R., Jackson, C. L., Fuchs, P. F. J. & Vanni, S. Interdigitation between triglycerides and lipids modulates surface properties of lipid droplets. Biophys. J. 112, 1417–1430 (2017).
Valm, A. M. et al. Applying systems-level spectral imaging and analysis to reveal the organelle interactome. Nature 546, 162–167 (2017). This study employs multispectral imaging to simultaneously visualize six organelles and elucidate their interactions over time.
Prinz, W. A. Bridging the gap: membrane contact sites in signaling, metabolism, and organelle dynamics. J. Cell Biol. 205, 759–769 (2014).
Shai, N. et al. Systematic mapping of contact sites reveals tethers and a function for the peroxisome-mitochondria contact. Nat. Commun. 9, 1761 (2018).
Thazar-Poulot, N., Miquel, M., Fobis-Loisy, I. & Gaude, T. Peroxisome extensions deliver the Arabidopsis SDP1 lipase to oil bodies. Proc. Natl Acad. Sci. USA 112, 4158–4163 (2015).
Xu, N. et al. The FATP1-DGAT2 complex facilitates lipid droplet expansion at the ER-lipid droplet interface. J. Cell Biol. 198, 895–911 (2012).
Xu, D. et al. Rab18 promotes lipid droplet (LD) growth by tethering the ER to LDs through SNARE and NRZ interactions. J. Cell Biol. 217, 975–995 (2018).
Markgraf, D. F. et al. An ER protein functionally couples neutral lipid metabolism on lipid droplets to membrane lipid synthesis in the ER. Cell Rep. 6, 44–55 (2014).
Jayson, C. B. K. et al. Rab18 is not necessary for lipid droplet biogenesis or turnover in human mammary carcinoma cells. Mol. Biol. Cell 29, 2045–2054 (2018).
Ohsaki, Y. et al. PML isoform II plays a critical role in nuclear lipid droplet formation. J. Cell Biol. 212, 29–38 (2016).
Romanauska, A. & Köhler, A. The inner nuclear membrane is a metabolically active territory that generates nuclear lipid droplets. Cell 174, 700–715 (2018).
Gallardo-Montejano, V. I. et al. Nuclear perilipin 5 integrates lipid droplet lipolysis with PGC-1α/SIRT1-dependent transcriptional regulation of mitochondrial function. Nat. Commun. 7, 12723 (2016).
Gao, G. et al. Control of lipid droplet fusion and growth by CIDE family proteins. Biochim. Biophys. Acta 1862, 1197–1204 (2017).
Jambunathan, S., Yin, J., Khan, W., Tamori, Y. & Puri, V. FSP27 promotes lipid droplet clustering and then fusion to regulate triglyceride accumulation. PLOS ONE 6, e28614 (2011).
Gong, J. et al. Fsp27 promotes lipid droplet growth by lipid exchange and transfer at lipid droplet contact sites. J. Cell Biol. 195, 953–963 (2011). References 88 and 89 demonstrate that CIDEC has a key role in lipid droplet fusion by promoting lipid exchange between associated lipid droplets.
Klemm, E. J., Spooner, E. & Ploegh, H. L. Dual role of ancient ubiquitous protein 1 (AUP1) in lipid droplet accumulation and endoplasmic reticulum (ER) protein quality control. J. Biol. Chem. 286, 37602–37614 (2011).
Spandl, J., Lohmann, D., Kuerschner, L., Moessinger, C. & Thiele, C. Ancient ubiquitous protein 1 (AUP1) localizes to lipid droplets and binds the E2 ubiquitin conjugase G2 (Ube2g2) via its G2 binding region. J. Biol. Chem. 286, 5599–5606 (2011).
Lohmann, D. et al. Monoubiquitination of ancient ubiquitous protein 1 promotes lipid droplet clustering. PLOS ONE 8, e72453 (2013).
Thiam, A. R., Farese, R. V. & Walther, T. C. The biophysics and cell biology of lipid droplets. Nat. Rev. Mol. Cell. Biol. 14, 775–786 (2013).
Grahn, T. H. M. et al. FSP27 and PLIN1 interaction promotes the formation of large lipid droplets in human adipocytes. Biochem. Biophys. Res. Commun. 432, 296–301 (2013).
Sun, Z. et al. Perilipin1 promotes unilocular lipid droplet formation through the activation of Fsp27 in adipocytes. Nat. Commun. 4, 1594 (2013).
Wu, L. et al. Rab8a-AS160-MSS4 regulatory circuit controls lipid droplet fusion and growth. Dev. Cell 30, 378–393 (2014).
Qian, H. et al. HDAC6-mediated acetylation of lipid droplet-binding protein CIDEC regulates fat-induced lipid storage. J. Clin. Invest. 127, 1353–1369 (2017).
Rubio-Cabezas, O. et al. Partial lipodystrophy and insulin resistant diabetes in a patient with a homozygous nonsense mutation in CIDEC. EMBO Mol. Med. 1, 280–287 (2009).
Nguyen, T. B. et al. DGAT1-dependent lipid droplet biogenesis protects mitochondrial function during starvation-induced autophagy. Dev. Cell 42, 9–21 (2017). This study demonstrates a role for lipid droplets in protecting mitochondria from lipotoxic damage during autophagy.
Rambold, A. S., Cohen, S. & Lippincott-Schwartz, J. Fatty acid trafficking in starved cells: regulation by lipid droplet lipolysis, autophagy, and mitochondrial fusion dynamics. Dev. Cell 32, 678–692 (2015). This study employs a fluorescent fatty acid probe to visualize fatty acid trafficking from lipid droplets to mitochondria and identifies an important role for mitochondrial fusion dynamics in fatty acid trafficking and metabolism.
Herms, A. et al. AMPK activation promotes lipid droplet dispersion on detyrosinated microtubules to increase mitochondrial fatty acid oxidation. Nat. Commun. 6, 7176 (2015).
Tarnopolsky, M. A. et al. Influence of endurance exercise training and sex on intramyocellular lipid and mitochondrial ultrastructure, substrate use, and mitochondrial enzyme activity. Am. J. Physiol. Regul. Integr. Comp. Physiol. 292, R1271–R1278 (2007).
Benador, I. Y. et al. Mitochondria bound to lipid droplets have unique bioenergetics, composition, and dynamics that support lipid droplet expansion. Cell Metab. 27, 869–885 (2018).
Wang, H. et al. Perilipin 5, a lipid droplet-associated protein, provides physical and metabolic linkage to mitochondria. J. Lipid Res. 52, 2159–2168 (2011). This study implicates PLIN5 as a lipid droplet–mitochondrial tether.
Granneman, J. G., Moore, H.-P. H., Mottillo, E. P., Zhu, Z. & Zhou, L. Interactions of perilipin-5 (Plin5) with adipose triglyceride lipase. J. Biol. Chem. 286, 5126–5135 (2011).
Boutant, M. et al. Mfn2 is critical for brown adipose tissue thermogenic function. EMBO J. 36, 1543–1558 (2017).
Dirkx, R. et al. Absence of peroxisomes in mouse hepatocytes causes mitochondrial and ER abnormalities. Hepatology 41, 868–878 (2005).
Binns, D. et al. An intimate collaboration between peroxisomes and lipid bodies. J. Cell Biol. 173, 719–731 (2006).
Schrader, M. Tubulo-reticular clusters of peroxisomes in living COS-7 cells: dynamic behavior and association with lipid droplets. J. Histochem. Cytochem. 49, 1421–1429 (2001).
Kaushik, S. & Cuervo, A. M. Degradation of lipid droplet-associated proteins by chaperone-mediated autophagy facilitates lipolysis. Nat. Cell Biol. 17, 759–770 (2015). This study finds that CMA degrades select lipid droplet proteins and thereby affects lipid droplet degradation by lipolysis.
Schroeder, B. et al. The small GTPase Rab7 as a central regulator of hepatocellular lipophagy. Hepatology 61, 1896–1907 (2015).
Kaushik, S. & Cuervo, A. M. The coming of age of chaperone-mediated autophagy. Nat. Rev. Mol. Cell. Biol. 19, 365–381 (2018).
Schneider, J. L., Suh, Y. & Cuervo, A. M. Deficient chaperone-mediated autophagy in liver leads to metabolic dysregulation. Cell Metab. 20, 417–432 (2014).
Kaushik, S. & Cuervo, A. M. AMPK-dependent phosphorylation of lipid droplet protein PLIN2 triggers its degradation by CMA. Autophagy 12, 432–438 (2016).
Takahashi, Y. et al. Perilipin2 plays a positive role in adipocytes during lipolysis by escaping proteasomal degradation. Sci. Rep. 6, 20975 (2016).
Xu, G., Sztalryd, C. & Londos, C. Degradation of perilipin is mediated through ubiquitination-proteasome pathway. Biochim. Biophys. Acta 1761, 83–90 (2006).
Masuda, Y. et al. ADRP/adipophilin is degraded through the proteasome-dependent pathway during regression of lipid-storing cells. J. Lipid Res. 47, 87–98 (2006).
Eastman, S. W., Yassaee, M. & Bieniasz, P. D. A role for ubiquitin ligases and Spartin/SPG20 in lipid droplet turnover. J. Cell Biol. 184, 881–894 (2009).
Hariri, H. et al. Lipid droplet biogenesis is spatially coordinated at ER-vacuole contacts under nutritional stress. EMBO Rep. 19, 57–72 (2018).
Henne, W. M. et al. Mdm1/Snx13 is a novel ER-endolysosomal interorganelle tethering protein. J. Cell Biol. 210, 541–551 (2015).
Bryant, D. et al. SNX14 mutations affect endoplasmic reticulum-associated neutral lipid metabolism in autosomal recessive spinocerebellar ataxia 20. Hum. Mol. Genet. 27, 1927–1940 (2018).
Eisenberg-Bord, M. et al. Identification of seipin-linked factors that act as determinants of a lipid droplet subpopulation. J. Cell Biol. 217, 269–282 (2018).
Teixeira, V. et al. Regulation of lipid droplets by metabolically controlled Ldo isoforms. J. Cell Biol. 217, 127–138 (2018).
Toulmay, A. & Prinz, W. A. Direct imaging reveals stable, micrometer-scale lipid domains that segregate proteins in live cells. J. Cell Biol. 202, 35–44 (2013).
Wang, C.-W., Miao, Y.-H. & Chang, Y.-S. A sterol-enriched vacuolar microdomain mediates stationary phase lipophagy in budding yeast. J. Cell Biol. 206, 357–366 (2014).
Seo, A. Y. et al. AMPK and vacuole-associated Atg14p orchestrate μ-lipophagy for energy production and long-term survival under glucose starvation. eLife 6, e21690 (2017).
Zechner, R., Madeo, F. & Kratky, D. Cytosolic lipolysis and lipophagy: two sides of the same coin. Nat. Rev. Mol. Cell. Biol. 18, 671–684 (2017).
Schulze, R. J., Sathyanarayan, A. & Mashek, D. G. Breaking fat: the regulation and mechanisms of lipophagy. Biochim. Biophys. Acta 1862B, 1178–1187 (2017).
Martinez-Lopez, N. & Singh, R. Autophagy and lipid droplets in the liver. Annu. Rev. Nutr. 35, 215–237 (2015).
Krahmer, N., Farese, R. V. & Walther, T. C. Balancing the fat: lipid droplets and human disease. EMBO Mol. Med. 5, 973–983 (2013).
Greenberg, A. S. et al. The role of lipid droplets in metabolic disease in rodents and humans. J. Clin. Invest. 121, 2102–2110 (2011).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Olzmann, J. A. & Kopito, R. R. Lipid droplet formation is dispensable for endoplasmic reticulum-associated degradation. J. Biol. Chem. 286, 27872–27874 (2011).
Marcinkiewicz, A., Gauthier, D., Garcia, A. & Brasaemle, D. L. The phosphorylation of serine 492 of perilipin a directs lipid droplet fragmentation and dispersion. J. Biol. Chem. 281, 11901–11909 (2006).
Chitraju, C. et al. Triglyceride synthesis by DGAT1 protects adipocytes from lipid-induced ER stress during lipolysis. Cell Metab. 26, 407–418 (2017). This study demonstrates that DGAT1-dependent triacylglycerol synthesis protects adipocytes from ER stress during lipolysis.
Paar, M. et al. Remodeling of lipid droplets during lipolysis and growth in adipocytes. J. Biol. Chem. 287, 11164–11173 (2012).
Hashimoto, T. et al. Active involvement of micro-lipid droplets and lipid-droplet-associated proteins in hormone-stimulated lipolysis in adipocytes. J. Cell Sci. 125, 6127–6136 (2012).
Ariotti, N. et al. Postlipolytic insulin-dependent remodeling of micro lipid droplets in adipocytes. Mol. Biol. Cell 23, 1826–1837 (2012).
Fu, S. et al. Aberrant lipid metabolism disrupts calcium homeostasis causing liver endoplasmic reticulum stress in obesity. Nature 473, 528–531 (2011).
Ozcan, U. et al. Chemical chaperones reduce ER stress and restore glucose homeostasis in a mouse model of type 2 diabetes. Science 313, 1137–1140 (2006).
Volmer, R., van der Ploeg, K. & Ron, D. Membrane lipid saturation activates endoplasmic reticulum unfolded protein response transducers through their transmembrane domains. Proc. Natl Acad. Sci. USA 110, 4628–4633 (2013).
Promlek, T. et al. Membrane aberrancy and unfolded proteins activate the endoplasmic reticulum stress sensor Ire1 in different ways. Mol. Biol. Cell 22, 3520–3532 (2011). References 141 and 142 demonstrate that disruptions in lipid homeostasis are sufficient to activate UPR sensors, independently of protein misfolding.
Halbleib, K. et al. Activation of the unfolded protein response by lipid bilayer stress. Mol. Cell 67, 673–684 (2017).
Fei, W., Wang, H., Fu, X., Bielby, C. & Yang, H. Conditions of endoplasmic reticulum stress stimulate lipid droplet formation in Saccharomyces cerevisiae. Biochem. J. 424, 61–67 (2009).
Lee, J.-S., Mendez, R., Heng, H. H., Yang, Z.-Q. & Zhang, K. Pharmacological ER stress promotes hepatic lipogenesis and lipid droplet formation. Am. J. Transl Res. 4, 102–113 (2012).
Rutkowski, D. T. et al. UPR pathways combine to prevent hepatic steatosis caused by ER stress-mediated suppression of transcriptional master regulators. Dev. Cell 15, 829–840 (2008).
Moldavski, O. et al. Lipid droplets are essential for efficient clearance of cytosolic inclusion bodies. Dev. Cell 33, 603–610 (2015).
Vevea, J. D. et al. Role for lipid droplet biogenesis and microlipophagy in adaptation to lipid imbalance in yeast. Dev. Cell 35, 584–599 (2015).
Jo, Y., Hartman, I. Z. & DeBose-Boyd, R. A. Ancient ubiquitous protein-1 mediates sterol-induced ubiquitination of 3-hydroxy-3-methylglutaryl CoA reductase in lipid droplet-associated endoplasmic reticulum membranes. Mol. Biol. Cell 24, 169–183 (2013).
Hartman, I. Z. et al. Sterol-induced dislocation of 3-hydroxy-3-methylglutaryl coenzyme A reductase from endoplasmic reticulum membranes into the cytosol through a subcellular compartment resembling lipid droplets. J. Biol. Chem. 285, 19288–19298 (2010).
Ohsaki, Y., Cheng, J., Suzuki, M., Fujita, A. & Fujimoto, T. Lipid droplets are arrested in the ER membrane by tight binding of lipidated apolipoprotein B-100. J. Cell Sci. 121, 2415–2422 (2008).
Suzuki, M. et al. Derlin-1 and UBXD8 are engaged in dislocation and degradation of lipidated ApoB-100 at lipid droplets. Mol. Biol. Cell 23, 800–810 (2012).
Ploegh, H. L. A lipid-based model for the creation of an escape hatch from the endoplasmic reticulum. Nature 448, 435–438 (2007).
To, M. et al. Lipid disequilibrium disrupts ER proteostasis by impairing ERAD substrate glycan trimming and dislocation. Mol. Biol. Cell 28, 270–284 (2017).
Zoncu, R. et al. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science 334, 678–683 (2011).
Efeyan, A., Comb, W. C. & Sabatini, D. M. Nutrient-sensing mechanisms and pathways. Nature 517, 302–310 (2015).
Yen, C.-L. E., Stone, S. J., Koliwad, S., Harris, C. & Farese, R. V. Thematic review series: glycerolipids. DGAT enzymes and triacylglycerol biosynthesis. J. Lipid Res. 49, 2283–2301 (2008).
Roussel, J. et al. Palmitoyl-carnitine increases RyR2 oxidation and sarcoplasmic reticulum Ca2+ leak in cardiomyocytes: role of adenine nucleotide translocase. Biochim. Biophys. Acta 1852, 749–758 (2015).
Requero, M. A., Goñi, F. M. & Alonso, A. The membrane-perturbing properties of palmitoyl-coenzyme A and palmitoylcarnitine. A comparative study. Biochemistry 34, 10400–10405 (1995).
Requero, M. A., González, M., Goñi, F. M., Alonso, A. & Fidelio, G. Differential penetration of fatty acyl-coenzyme A and fatty acylcarnitines into phospholipid monolayers. FEBS Lett. 357, 75–78 (1995).
Long, J. Z. et al. The secreted enzyme PM20D1 regulates lipidated amino acid uncouplers of mitochondria. Cell 166, 424–435 (2016).
Perera, R. M. et al. Transcriptional control of autophagy-lysosome function drives pancreatic cancer metabolism. Nature 524, 361–365 (2015).
Yang, S. et al. Pancreatic cancers require autophagy for tumor growth. Genes Dev. 25, 717–729 (2011).
Stolz, A., Ernst, A. & Dikic, I. Cargo recognition and trafficking in selective autophagy. Nat. Cell Biol. 16, 495–501 (2014).
Son, N.-H. et al. PPARγ-induced cardiolipotoxicity in mice is ameliorated by PPARα deficiency despite increases in fatty acid oxidation. J. Clin. Invest. 120, 3443–3454 (2010).
McCoin, C. S., Knotts, T. A. & Adams, S. H. Acylcarnitines—old actors auditioning for new roles in metabolic physiology. Nat. Rev. Endocrinol. 11, 617–625 (2015).
Wajner, M. & Amaral, A. U. Mitochondrial dysfunction in fatty acid oxidation disorders: insights from human and animal studies. Biosci. Rep. 36, e00281 (2015).
Liu, L. et al. Cardiomyocyte-specific loss of diacylglycerol acyltransferase 1 (DGAT1) reproduces the abnormalities in lipids found in severe heart failure. J. Biol. Chem. 289, 29881–29891 (2014).
Liu, L. et al. DGAT1 expression increases heart triglyceride content but ameliorates lipotoxicity. J. Biol. Chem. 284, 36312–36323 (2009).
Liu, L. et al. Diacylglycerol acyl transferase 1 overexpression detoxifies cardiac lipids in PPARγ transgenic mice. J. Lipid Res. 53, 1482–1492 (2012). References 168–170 demonstrate a protective role for DGAT1 in mouse models of lipotoxicity and cardiac dysfunction.
Tauchi-Sato, K., Ozeki, S., Houjou, T., Taguchi, R. & Fujimoto, T. The surface of lipid droplets is a phospholipid monolayer with a unique fatty acid composition. J. Biol. Chem. 277, 44507–44512 (2002).
Wolins, N. E. et al. S3-12, adipophilin, and TIP47 package lipid in adipocytes. J. Biol. Chem. 280, 19146–19155 (2005).
Wolins, N. E., Brasaemle, D. L. & Bickel, P. E. A proposed model of fat packaging by exchangeable lipid droplet proteins. FEBS Lett. 580, 5484–5491 (2006).
Herms, A. et al. Cell-to-cell heterogeneity in lipid droplets suggests a mechanism to reduce lipotoxicity. Curr. Biol. 23, 1489–1496 (2013).
Bailey, A. P. et al. Antioxidant role for lipid droplets in a stem cell niche of drosophila. Cell 163, 340–353 (2015).
Liu, L., MacKenzie, K. R., Putluri, N., Maletic-Savatic, M. & Bellen, H. J. The glia-neuron lactate shuttle and elevated ROS promote lipid synthesis in neurons and lipid droplet accumulation in glia via APOE/D. Cell Metab. 26, 719–737 (2017).
Liu, L. et al. Glial lipid droplets and ROS induced by mitochondrial defects promote neurodegeneration. Cell 160, 177–190 (2015). References 175–177 implicate glial sequestration of toxic lipids as a mechanism to protect neurons from lipotoxic damage.
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Kozlitina, J. et al. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 46, 352–356 (2014).
Mahdessian, H. et al. TM6SF2 is a regulator of liver fat metabolism influencing triglyceride secretion and hepatic lipid droplet content. Proc. Natl Acad. Sci. USA 111, 8913–8918 (2014).
Schweiger, M., Lass, A., Zimmermann, R., Eichmann, T. O. & Zechner, R. Neutral lipid storage disease: genetic disorders caused by mutations in adipose triglyceride lipase/PNPLA2 or CGI-58/ABHD5. Am. J. Physiol. Endocrinol. Metab. 297, E289–E296 (2009).
Lord, C. C. et al. Regulation of hepatic triacylglycerol metabolism by CGI-58 does not require ATGL co-activation. Cell Rep. 16, 939–949 (2016).
Herranz, P., de Lucas, R., Pérez-España, L. & Mayor, M. Lipodystrophy syndromes. Dermatol. Clin. 26, 569–578 (2008).
Blackstone, C. Converging cellular themes for the hereditary spastic paraplegias. Curr. Opin. Neurobiol. 51, 139–146 (2018).
Renvoisé, B. et al. Reep1 null mice reveal a converging role for hereditary spastic paraplegia proteins in lipid droplet regulation. Hum. Mol. Genet. 25, 5111–5125 (2016).
Klemm, R. W. et al. A conserved role for atlastin GTPases in regulating lipid droplet size. Cell Rep. 3, 1465–1475 (2013).
Hooper, C., Puttamadappa, S. S., Loring, Z., Shekhtman, A. & Bakowska, J. C. Spartin activates atrophin-1-interacting protein 4 (AIP4) E3 ubiquitin ligase and promotes ubiquitination of adipophilin on lipid droplets. BMC Biol. 8, 72 (2010).
Edwards, T. L. et al. Endogenous spartin (SPG20) is recruited to endosomes and lipid droplets and interacts with the ubiquitin E3 ligases AIP4 and AIP5. Biochem. J. 423, 31–39 (2009).
Acknowledgements
The authors apologize to all scientists whose important contributions were not referenced in this review owing to space limitations. The authors thank Z. Li and L. Krshnan for careful reading and discussion of the manuscript. J.A.O. acknowledges funding from the US National Institutes of Health (R01GM112948 and R21AG056502) and the American Heart Association (16GRNT30870005). P.C. acknowledges funding from the European Research Council (starting grant DropFat 309477) and the Wellcome Trust.
Reviewer information
Nature Reviews Molecular Cell Biology thanks D. Mashek, S. Cohen and J. Goodman for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Both authors researched data for the article, contributed to discussion of the content, wrote the article and reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Lipotoxicity
-
Deleterious effects of improperly stored lipids on cellular health.
- COPII
-
Complex of proteins that coat vesicles budding from endoplasmic reticulum (ER) exit sites and facilitate anterograde protein transport towards the Golgi.
- Phospholipid scramblases
-
Proteins that mediate the bidirectional exchange of phospholipids between two leaflets of a lipid bilayer.
- Perilipins
-
Family of proteins that coat lipid droplets and regulate lipid droplet stability and turnover.
- COPI coatomer complex
-
Complex of proteins that coat vesicles budding from the Golgi and facilitate retrograde protein transport.
- Polytopic membrane proteins
-
Membrane-embedded proteins in which the polypeptide chain crosses the membrane multiple times.
- Farnesylation
-
Post-translational modification of a protein by addition of an isoprenoid farnesyl group to a cysteine residue.
- ER-associated protein degradation
-
(ERAD). A process that mediates the recognition and delivery of aberrant (for example, misfolded) proteins from the endoplasmic reticulum to the proteasome for degradation.
- Packing defects
-
Regions in which the neutral lipid or the hydrocarbon chains of phospholipids are exposed to the aqueous cytoplasm.
- SNAREs
-
Family of proteins that mediate membrane fusion.
- Brown adipocytes
-
Type of adipocyte that contains large numbers of mitochondria and expresses high amounts of uncoupling protein 1, resulting in the dissipation of the proton motive force and generation of heat.
- Insulin resistance
-
Condition in which cells fail to respond appropriately to insulin.
- Peroxidative damage
-
A form of oxidative damage, such as the formation of lipid peroxides.
- Ferroptosis
-
Regulated, iron-dependent form of cell death that is characterized by the accumulation of lipid peroxides.
- Apolipoprotein
-
Lipid-binding protein that enables lipid transport throughout the body.
- Retrotranslocation
-
Movement of endoplasmic reticulum-associated protein degradation substrates across the endoplasmic reticulum membrane back into the cytoplasm for degradation by the proteasome.
- Fatty acid oxidation diseases
-
Heterogeneous group of rare, autosomal recessive diseases characterized by defects in fatty acid catabolism.
- Ichthyosis
-
Condition characterized by skin that is thickened, dry and scaly.
Rights and permissions
About this article
Cite this article
Olzmann, J.A., Carvalho, P. Dynamics and functions of lipid droplets. Nat Rev Mol Cell Biol 20, 137–155 (2019). https://doi.org/10.1038/s41580-018-0085-z
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-018-0085-z
This article is cited by
-
Emerging role of senescent microglia in brain aging-related neurodegenerative diseases
Translational Neurodegeneration (2024)
-
Changes in lipid metabolism track with the progression of neurofibrillary pathology in tauopathies
Journal of Neuroinflammation (2024)
-
A transcriptomic examination of encased rotifer embryos reveals the developmental trajectory leading to long-term dormancy; are they “animal seeds”?
BMC Genomics (2024)
-
The role of DGAT1 and DGAT2 in regulating tumor cell growth and their potential clinical implications
Journal of Translational Medicine (2024)
-
A nanoemulsion targeting adipose hypertrophy and hyperplasia shows anti-obesity efficiency in female mice
Nature Communications (2024)