Abstract
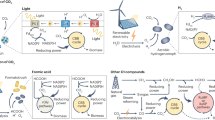
Ruminants produce edible products and contribute to food security. They house a complex rumen microbial community that enables the host to digest their plant feed through microbial-mediated fermentation. However, the rumen microbiome is also responsible for the production of one of the most potent greenhouse gases, methane, and contributes about 18% of its total anthropogenic emissions. Conventional methods to lower methane production by ruminants have proved successful, but to a limited and often temporary extent. An increased understanding of the host–microbiome interactions has led to the development of new mitigation strategies. In this Review we describe the composition, ecology and metabolism of the rumen microbiome, and the impact on host physiology and the environment. We also discuss the most pertinent methane mitigation strategies that emerged to balance food security and environmental impacts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ley, R. E., Lozupone, C. A., Hamady, M., Knight, R. & Gordon, J. I. Worlds within worlds: evolution of the vertebrate gut microbiota. Nat. Rev. Microbiol. 6, 776–788 (2008).
Mizrahi, I. in The Prokaryotes (eds Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E. & Thompson, F.) 533–544 (Springer, 2013).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nat. Genet. 39, 31–40 (2007).
Luckey, T. Germfree Life and Gnotobiology (Elsevier, 2012).
Hobson, P. N. & Stewart, C. S. The Rumen Microbial Ecosystem (Springer Science & Business Media, 2012).
Lin, L. et al. Ruminal microbiome–host crosstalk stimulates the development of the ruminal epithelium in a lamb model. Microbiome 7, 83 (2019).
Mizrahi, I. & Jami, E. Review: The compositional variation of the rumen microbiome and its effect on host performance and methane emission. Animal. 12, s220–s232 (2018).
Malmuthuge, N., Liang, G. & Guan, L. L. Regulation of rumen development in neonatal ruminants through microbial metagenomes and host transcriptomes. Genome Biol. 20, 172 (2019).
Eshel, G., Shepon, A., Makov, T. & Milo, R. Land, irrigation water, greenhouse gas, and reactive nitrogen burdens of meat, eggs, and dairy production in the United States. Proc. Natl Acad. Sci. USA 111, 11996–12001 (2014).
Maasakkers, J. D. et al. Global distribution of methane emissions, emission trends, and OH concentrations and trends inferred from an inversion of GOSAT satellite data for 2010–2015. Atmos. Chem. Phys. 19, 7859–7881 (2019).
Dong, H. et al. Emissions from livestock and manure management. Embrapa Meio Ambiente-Capítulo em Livro Científico (ALICE) Vol. 4, 1–87 (iGES, Kanagawa, 2006).
Le Quéré, C. et al. Global carbon budget 2018. Earth Syst. Sci. Data 10, 2141–2194 (2018).
Huws, S. A. et al. Addressing global ruminant agricultural challenges through understanding the rumen microbiome: past, present, and future. Front. Microbiol. 9, 2161 (2018).
Shabat, S. K. B. et al. Specific microbiome-dependent mechanisms underlie the energy harvest efficiency of ruminants. ISME J. 10, 2958–2972 (2016). This study links the compositional states of the cow rumen microbiome to feed efficiency and methane emission, with an emphasis on microbial lactic acid and hydrogen metabolism.
Shaani, Y., Zehavi, T., Eyal, S., Miron, J. & Mizrahi, I. Microbiome niche modification drives diurnal rumen community assembly, overpowering individual variability and diet effects. ISME J. 12, 2446–2457 (2018).
Friedman, N., Shriker, E. & Gold, B. Diet-induced changes of redox potential underlie compositional shifts in the rumen archaeal community. Environ. Microbiol. 19, 174–184 (2017).
Sasson, G. et al. Heritable bovine rumen bacteria are phylogenetically related and correlated with the cow’s capacity to harvest energy from its feed. mBio 8, e00703-17 (2017).
Friedman, N., Jami, E. & Mizrahi, I. Compositional and functional dynamics of the bovine rumen methanogenic community across different developmental stages. Environ. Microbiol. 19, 3365–3373 (2017).
Li, F. et al. Host genetics influence the rumen microbiota and heritable rumen microbial features associate with feed efficiency in cattle. Microbiome 7, 92 (2019).
Li, F., Hitch, T. C. A., Chen, Y., Creevey, C. J. & Guan, L. L. Comparative metagenomic and metatranscriptomic analyses reveal the breed effect on the rumen microbiome and its associations with feed efficiency in beef cattle. Microbiome 7, 6 (2019).
Nkrumah, J. D. et al. Relationships of feedlot feed efficiency, performance, and feeding behavior with metabolic rate, methane production, and energy partitioning in beef cattle. J. Anim. Sci. 84, 145–153 (2006).
Mizrahi, I. in Beneficial Microorganisms in Multicellular Life Forms (eds Rosenberg, E. & Gophna, U.) 203–210 (Springer, 2012).
Hart, E. H., Creevey, C. J., Hitch, T. & Kingston-Smith, A. H. Meta-proteomics of rumen microbiota indicates niche compartmentalisation and functional dominance in a limited number of metabolic pathways between abundant bacteria. Sci. Rep. 8, 10501 (2018).
Snelling, T. J. & Wallace, R. J. The rumen microbial metaproteome as revealed by SDS-PAGE. BMC Microbiol. 17, 9 (2017).
Jami, E. & Mizrahi, I. Composition and similarity of bovine rumen microbiota across individual animals. PLoS ONE 7, e33306 (2012).
Morais, S. & Mizrahi, I. Islands in the stream: from individual to communal fiber degradation in the rumen ecosystem. FEMS Rev. Microbiol. 43, 362–379 (2019). This review summarizes the enzymological fundamentals of fibre degradation with an emphasis on the community perspective, from individual genetic information to microbial interactions.
Henderson, G. et al. Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci. Rep. 5, 14567 (2015). This important study defines the rumen core microbiome across ruminant species with relation to geography and diet.
Wallace, R. J. et al. A heritable subset of the core rumen microbiome dictates dairy cow productivity and emissions. Sci. Adv. 5, eaav8391 (2019). This 1,000-animal study provides deep understanding of the extent to which ruminant microbiomes can be controlled by the host animal. The study identifies the characteristics of the host rumen microbiome axis that determine animal productivity and methane emissions.
Huws, S. A. et al. Temporal dynamics of the metabolically active rumen bacteria colonizing fresh perennial ryegrass. FEMS Microbiol. Ecol. 92, fiv137 (2016).
Piao, H. et al. Temporal dynamics of fibrolytic and methanogenic rumen microorganisms during in situ incubation of switchgrass determined by 16S rRNA gene profiling. Front. Microbiol. 5, 307 (2014).
Liu, J., Zhang, M., Xue, C., Zhu, W. & Mao, S. Characterization and comparison of the temporal dynamics of ruminal bacterial microbiota colonizing rice straw and alfalfa hay within ruminants. J. Dairy. Sci. 99, 9668–9681 (2016).
Jin, W., Wang, Y., Li, Y., Cheng, Y. & Zhu, W. Temporal changes of the bacterial community colonizing wheat straw in the cow rumen. Anaerobe 50, 1–8 (2018).
Stanton, T. B. & Canale-Parola, E. Treponema bryantii sp. nov., a rumen spirochete that interacts with cellulolytic bacteria. Arch. Microbiol. 127, 145–156 (1980).
Blackburn, T. H. & Hungate, R. E. Succinic acid turnover and propionate production in the bovine rumen. Appl. Microbiol. 11, 132–135 (1963).
Scheifinger, C. C. & Wolin, M. J. Propionate formation from cellulose and soluble sugars by combined cultures of Bacteroides succinogenes and Selenomonas ruminantium. Appl. Microbiol. 26, 789–795 (1973).
Kim, M., Morrison, M. & Yu, Z. Status of the phylogenetic diversity census of ruminal microbiomes. FEMS Microbiol. Ecol. 76, 49–63 (2011).
Arntzen, M. Ø., Várnai, A., Mackie, R. I., Eijsink, V. G. H. & Pope, P. B. Outer membrane vesicles from Fibrobacter succinogenes S85 contain an array of carbohydrate-active enzymes with versatile polysaccharide-degrading capacity. Environ. Microbiol. 19, 2701–2714 (2017).
Suen, G. et al. The complete genome sequence of Fibrobacter succinogenes S85 reveals a cellulolytic and metabolic specialist. PLoS ONE 6, e18814 (2011).
O’Hara, E. et al. Investigating temporal microbial dynamics in the rumen of beef calves raised on two farms during early life. FEMS Microbiol. Ecol. 96, fiz203 (2020).
Furman, O. et al. Stochasticity constrained by deterministic effects of diet and age drive rumen microbiome assembly dynamics. Nat. Commun. 11, 1904 (2020). This study demonstrates that stochastic colonization in early life, together with strong deterministic constraints imposed by diet and age, exhibits long-lasting impact on the development of animal microbiomes.
Moraïs, S. & Mizrahi, I. The road not taken: the rumen microbiome, functional groups, and community states. Trends Microbiol. 27, 538–549 (2019). This review takes an ecological approach to introduce the concept of the rumen functional group and community states to guide the interpretation of rumen microbiome data.
Weimer, P. J. Redundancy, resilience, and host specificity of the ruminal microbiota: implications for engineering improved ruminal fermentations. Front. Microbiol. 6, 296 (2015). This excellent review provides an ecological perspective of rumen metabolism.
Seshadri, R. et al. Cultivation and sequencing of rumen microbiome members from the Hungate1000 Collection. Nat. Biotechnol. 36, 359–367 (2018). This fundamental publication describes the genome resource of the Hungate1000 rumen isolates.
Taxis, T. M. et al. The players may change but the game remains: network analyses of ruminal microbiomes suggest taxonomic differences mask functional similarity. Nucleic Acids Res. 43, 9600–9612 (2015).
Hackmann, T. J., Ngugi, D. K. & Firkins, J. L. Genomes of rumen bacteria encode atypical pathways for fermenting hexoses to short-chain fatty acids. Environmentalist 19, 4670–4683 (2017).
Matte, A., Forsberg, C. W. & Verrinder Gibbins, A. M. Enzymes associated with metabolism of xylose and other pentoses by Prevotella (Bacteroides) ruminicola strains, Selenomonas ruminantium D, and Fibrobacter succinogenes S85. Can. J. Microbiol. 38, 370–S376 (1992).
Louis, P. & Flint, H. J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 19, 29–41 (2017).
Reichardt, N. et al. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 8, 1323–1335 (2014).
Laverde Gomez, J. A. et al. Formate cross-feeding and cooperative metabolic interactions revealed by transcriptomics in co-cultures of acetogenic and amylolytic human colonic bacteria. Environ. Microbiol. 21, 259–271 (2019).
Le Van, T. D. et al. Assessment of reductive acetogenesis with indigenous ruminal bacterium populations and Acetitomaculum ruminis. Appl. Environ. Microbiol. 64, 3429–3436 (1998).
Fonty, G. et al. Establishment and development of ruminal hydrogenotrophs in methanogen-free lambs. Appl. Environ. Microbiol. 73, 6391–6403 (2007).
Valdés, C., Newbold, C. J., Hillman, K. & Wallace, R. J. Evidence for methane oxidation in rumen fluid in vitro. Ann. Zootech. 45, 351–351 (1996).
Janssen, P. H. & Kirs, M. Structure of the archaeal community of the rumen. Appl. Environ. Microbiol. 74, 3619–3625 (2008).
Demirel, B. & Scherer, P. The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Rev. Environ. Sci. Technol. 7, 173–190 (2008).
Ferry, J. G. Enzymology of one-carbon metabolism in methanogenic pathways. FEMS Microbiol. Rev. 23, 13–38 (1999).
Morgavi, D. P., Forano, E., Martin, C. & Newbold, C. J. Microbial ecosystem and methanogenesis in ruminants. Animal 4, 1024–1036 (2010).
Guzman, C. E., Bereza-Malcolm, L. T., De Groef, B. & Franks, A. E. Presence of selected methanogens, fibrolytic bacteria, and proteobacteria in the gastrointestinal tract of neonatal dairy calves from birth to 72 hours. PLoS ONE 10, e0133048 (2015).
Friedman, N., Jami, E. & Mizrahi, I. Compositional and functional dynamics of the bovine rumen methanogenic community across different developmental stages. Environ. Microbiol. 19, 3365–3373 (2017).
Flint, H. J. The rumen microbial ecosystem — some recent developments. Trends Microbiol. 5, 483–488 (1997).
Greening, C. et al. Diverse hydrogen production and consumption pathways influence methane production in ruminants. ISME J. 13, 2617–2632 (2019).
Janssen, P. H. Influence of hydrogen on rumen methane formation and fermentation balances through microbial growth kinetics and fermentation thermodynamics. Anim. Feed. Sci. Technol. 160, 1–22 (2010).
Stams, A. J. M. & Plugge, C. M. Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat. Rev. Microbiol. 7, 568–577 (2009).
Latham, M. J. & Wolin, M. J. Fermentation of cellulose by Ruminococcus flavefaciens in the presence and absence of Methanobacterium ruminantium. Appl. Environ. Microbiol. 34, 297–301 (1977).
Thauer, R. K., Jungermann, K. & Decker, K. Energy conservation in chemotrophic anaerobic bacteria. Bacteriol. Rev. 41, 100 (1977).
Bauchop, T. & Mountfort, D. O. Cellulose fermentation by a rumen anaerobic fungus in both the absence and the presence of rumen methanogens. Appl. Environ. Microbiol. 42, 1103–1110 (1981).
Rychlik, J. L. & May, T. The effect of a methanogen, Methanobrevibacter smithii, on the growth rate, organic acid production, and specific ATP activity of three predominant ruminal cellulolytic bacteria. Curr. Microbiol. 40, 176–180 (2000).
Leahy, S. C. et al. The genome sequence of the rumen methanogen Methanobrevibacter ruminantium reveals new possibilities for controlling ruminant methane emissions. PLoS ONE 5, e8926 (2010).
Scheifinger, C. C., Linehan, B. & Wolin, M. J. H2 production by Selenomonas ruminantium in the absence and presence of methanogenic bacteria. Appl. Microbiol. 29, 480–483 (1975).
Cazier, E. A., Trably, E., Steyer, J. P. & Escudie, R. Biomass hydrolysis inhibition at high hydrogen partial pressure in solid-state anaerobic digestion. Bioresour. Technol. 190, 106–113 (2015).
Morvan, B., Bonnemoy, F., Fonty, G. & Gouet, P. Quantitative determination of H2-utilizing acetogenic and sulfate-reducing bacteria and methanogenic archaea from digestive tract of different mammals. Curr. Microbiol. 32, 129–133 (1996).
Newbold, C. J., de la Fuente, G., Belanche, A., Ramos-Morales, E. & McEwan, N. R. The role of ciliate protozoa in the rumen. Front. Microbiol. 6, 1313 (2015).
Ushida, K., Newbold, C. J. & Jouany, J.-P. Interspecies hydrogen transfer between the rumen ciliate Polyplastron multivesiculatum and Methanosarcina barkeri. J. Gen. Appl. Microbiol. 43, 129–131 (1997).
Wright, A. D. G. & Hook, S. E. 16 Manipulation of microbial ecology for sustainable animal production. Sustain. Anim. Agr. https://doi.org/10.1079/9781780640426.0254254 (2013).
Sharp, R., Ziemer, C. J., Stern, M. D. & Stahl, D. A. Taxon-specific associations between protozoal and methanogen populations in the rumen and a model rumen system. FEMS Microbiol. Ecol. 26, 71–78 (1998).
Lloyd, D. et al. Intracellular prokaryotes in rumen ciliate protozoa: detection by confocal laser scanning microscopy after in situ hybridization with fluorescent 16S rRNA probes. Eur. J. Protistol. 32, 523–531 (1996).
Ng, F. et al. An adhesin from hydrogen-utilizing rumen methanogen Methanobrevibacter ruminantium M1 binds a broad range of hydrogen-producing microorganisms. Environ. Microbiol. 18, 3010–3021 (2016).
Zhou, M., Hernandez-Sanabria, E. & Guan, L. L. Assessment of the microbial ecology of ruminal methanogens in cattle with different feed efficiencies. Appl. Environ. Microbiol. 75, 6524–6533 (2009).
Wallace, R. J. et al. The rumen microbial metagenome associated with high methane production in cattle. BMC Genomics 16, 839 (2015).
Tapio, I., Snelling, T. J., Strozzi, F. & Wallace, R. J. The ruminal microbiome associated with methane emissions from ruminant livestock. J. Anim. Sci. Biotechnol. 8, 7 (2017).
Gralka, M., Szabo, R., Stocker, R. & Cordero, O. X. Trophic interactions and the drivers of microbial community assembly. Curr. Biol. 30, R1176–R1188 (2020).
Kamke, J. et al. Rumen metagenome and metatranscriptome analyses of low methane yield sheep reveals a Sharpea-enriched microbiome characterised by lactic acid formation and utilisation. Microbiome 4, 56 (2016). This study links rumen lactic acid and hydrogen metabolism to methane emissions in sheep.
Pope, P. B. et al. Isolation of Succinivibrionaceae implicated in low methane emissions from Tammar wallabies. Science 333, 646–648 (2011).
Mi, L. et al. Comparative analysis of the microbiota between sheep rumen and rabbit cecum provides new insight into their differential methane production. Front. Microbiol. 9, 575 (2018).
Kittelmann, S. et al. Two different bacterial community types are linked with the low-methane emission trait in sheep. PLoS ONE 9, e103171 (2014).
Lee, P. C., Lee, W. G., Kwon, S., Lee, S. Y. & Chang, H. N. Succinic acid production by Anaerobiospirillum succiniciproducens: effects of the H2/CO2 supply and glucose concentration. Enzyme Microb. Technol. 24, 549–554 (1999).
Ungerfeld, E. M. & Kohn, R. A. in Ruminant Physiology: Digestion, Metabolism and Impact of Nutrition on Gene Expression, Immunology and Stress 55–85 (Wageningen Academic, 2006).
Shi, W. et al. Methane yield phenotypes linked to differential gene expression in the sheep rumen microbiome. Genome Res. 24, 1517–1525 (2014).
Morgavi, D. P., Martin, C., Jouany, J.-P. & Ranilla, M. J. Rumen protozoa and methanogenesis: not a simple cause–effect relationship. Br. J. Nutr. 107, 388–397 (2012).
Grainger, C. & Beauchemin, K. A. Can enteric methane emissions from ruminants be lowered without lowering their production? Anim. Feed. Sci. Technol. 166–167, 308–320 (2011).
Hristov, A. N. et al. An inhibitor persistently decreased enteric methane emission from dairy cows with no negative effect on milk production. Proc. Natl Acad. Sci. USA 112, 10663–10668 (2015).
Ungerfeld, E. M. Shifts in metabolic hydrogen sinks in the methanogenesis-inhibited ruminal fermentation: a meta-analysis. Front. Microbiol. 6, 37 (2015). This report analyses 54 studies on the redirection of hydrogen to potential alternative metabolic sinks when methanogenesis is inhibited.
McAllister, T. A. et al. Ruminant nutrition symposium: use of genomics and transcriptomics to identify strategies to lower ruminal methanogenesis. J. Anim. Sci. 93, 1431–1449 (2015).
Morgavi, D. P., Kelly, W. J., Janssen, P. H. & Attwood, G. T. Rumen microbial (meta)genomics and its application to ruminant production. Animal 7 (Suppl. 1), 184–201 (2013).
Patra, A., Park, T., Kim, M. & Yu, Z. Rumen methanogens and mitigation of methane emission by anti-methanogenic compounds and substances. J. Anim. Sci. Biotechnol. 8, 13 (2017). This article presents a thorough review of methanogens and methane mitigation strategies.
Kumar, S. et al. New aspects and strategies for methane mitigation from ruminants. Appl. Microbiol. Biotechnol. 98, 31–44 (2014).
Wallace, R. J., Snelling, T. J., McCartney, C. A., Tapio, I. & Strozzi, F. Application of meta-omics techniques to understand greenhouse gas emissions originating from ruminal metabolism. Genet. Sel. Evol. 49, 9 (2017).
Yang, C., Rooke, J. A., Cabeza, I. & Wallace, R. J. Nitrate and inhibition of ruminal methanogenesis: microbial ecology, obstacles, and opportunities for lowering methane emissions from ruminant livestock. Front. Microbiol. 7, 132 (2016).
Martin, C., Morgavi, D. P. & Doreau, M. Methane mitigation in ruminants: from microbe to the farm scale. Animal 4, 351–365 (2010). This article presents a detailed review of methane mitigation approaches.
Lan, W. & Yang, C. Ruminal methane production: associated microorganisms and the potential of applying hydrogen-utilizing bacteria for mitigation. Sci. Total. Environ. 654, 1270–1283 (2019).
Liu, L. et al. Nitrate decreases methane production also by increasing methane oxidation through stimulating NC10 population in ruminal culture. AMB. Express 7, 76 (2017).
Yu, Z. & Smith, G. B. Inhibition of methanogenesis by C1-and C2-polychlorinated aliphatic hydrocarbons. Environ. Toxicol. Chem. 19, 2212–2217 (2000).
Wagner, T., Wegner, C.-E., Kahnt, J., Ermler, U. & Shima, S. Phylogenetic and structural comparisons of the three types of methyl coenzyme M reductase from methanococcales and methanobacteriales. J. Bacteriol. 199, e00197-17 (2017).
Duval, S. & Kindermann, M. Use of nitrooxy organic molecules in feed for reducing enteric methnae emsions in ruminants, and/or to improve ruminant performance. International Patent Application WO 2012/084629 A1 (2012).
Burreson, B. J., Moore, R. E. & Roller, P. P. Volatile halogen compounds in the alga Asparagopsis taxiformis (Rhodophyta). J. Agric. Food Chem. 24, 856–861 (1976).
Roque, B. M. et al. Effect of the macroalgae Asparagopsis taxiformis on methane production and rumen microbiome assemblage. Anim. Microbiome 1, 3 (2019).
Machado, L. et al. Identification of bioactives from the red seaweed Asparagopsis taxiformis that promote antimethanogenic activity in vitro. J. Appl. Phycol. 28, 3117–3126 (2016).
Knight, T. et al. Chloroform decreases rumen methanogenesis and methanogen populations without altering rumen function in cattle. Anim. Feed. Sci. Technol. 166–167, 101–112 (2011).
Ungerfeld, E. M., Rust, S. R., Boone, D. R. & Liu, Y. Effects of several inhibitors on pure cultures of ruminal methanogens. J. Appl. Microbiol. 97, 520–526 (2004).
Van Wesemael, D. et al. Reducing enteric methane emissions from dairy cattle: two ways to supplement 3-nitrooxypropanol. J. Dairy Sci. 102, 1780–1787 (2019).
Machado, L., Magnusson, M., Paul, N. A., de Nys, R. & Tomkins, N. Effects of marine and freshwater macroalgae on in vitro total gas and methane production. PLoS ONE 9, e85289 (2014).
Li, X. et al. Asparagopsis taxiformis decreases enteric methane production from sheep. Anim. Prod. Sci. 58, 681 (2018).
Kelly, W. J. et al. The complete genome sequence of the rumen methanogen Methanobacterium formicicum BRM9. Stand. Genomic Sci. 9, 15 (2014).
Altermann, E., Schofield, L. R., Ronimus, R. S., Beatty, A. K. & Reilly, K. Inhibition of rumen methanogens by a novel archaeal lytic enzyme displayed on tailored bionanoparticles. Front. Microbiol. 9, 2378 (2018).
Williams, Y. J. et al. A vaccine against rumen methanogens can alter the composition of archaeal populations. Appl. Environ. Microbiol. 75, 1860–1866 (2009).
Stewart, R. D. et al. Compendium of 4,941 rumen metagenome-assembled genomes for rumen microbiome biology and enzyme discovery. Nat. Biotechnol. 37, 953–961 (2019). This important resource on rumen microbial genomes enables a better understanding of the structure and function of the rumen microbiota.
Gagen, E. J. et al. Methanogen colonisation does not significantly alter acetogen diversity in lambs isolated 17 h after birth and raised aseptically. Microb. Ecol. 64, 628–640 (2012).
Ungerfeld, E. M., Kohn, R. A., Wallace, R. J. & Newbold, C. J. A meta-analysis of fumarate effects on methane production in ruminal batch cultures. J. Anim. Sci. 85, 2556–2563 (2007).
Ungerfeld, E. M., Rust, S. R. & Burnett, R. Use of some novel alternative electron sinks to inhibit ruminal methanogenesis. Reprod. Nutr. Dev. 43, 189–202 (2003).
van Zijderveld, S. M. et al. Nitrate and sulfate: effective alternative hydrogen sinks for mitigation of ruminal methane production in sheep. J. Dairy Sci. 93, 5856–5866 (2010).
Latham, E. A., Anderson, R. C., Pinchak, W. E. & Nisbet, D. J. Insights on alterations to the rumen ecosystem by nitrate and nitrocompounds. Front. Microbiol. 7, 228 (2016).
Huisingh, J., McNeill, J. J. & Matrone, G. Sulfate reduction by a Desulfovibrio species isolated from sheep rumen. Appl. Microbiol. 28, 489–497 (1974).
Iwamoto, M., Asanuma, N. & Hino, T. Ability of Selenomonas ruminantium, Veillonella parvula, and Wolinella succinogenes to reduce nitrate and nitrite with special reference to the suppression of ruminal methanogenesis. Anaerobe 8, 209–215 (2002).
Kandylis, K. Toxicology of sulfur in ruminants: review. J. Dairy Sci. 67, 2179–2187 (1984).
Ungerfeld, E. M. A theoretical comparison between two ruminal electron sinks. Front. Microbiol. 4, 319 (2013). This important study examines and compares the nutritional and energetic implications of directing hydrogen into acetogenesis or propionate production at the expense of methanogenesis.
Foley, P. A., Kenny, D. A., Callan, J. J., Boland, T. M. & O’Mara, F. P. Effect of dl-malic acid supplementation on feed intake, methane emission, and rumen fermentation in beef cattle. J. Anim. Sci. 87, 1048–1057 (2009).
Eugène, M., Massé, D., Chiquette, J. & Benchaar, C. Meta-analysis on the effects of lipid supplementation on methane production in lactating dairy cows. Can. J. Anim. Sci. 88, 331–337 (2008).
Patra, A. K. The effect of dietary fats on methane emissions, and its other effects on digestibility, rumen fermentation and lactation performance in cattle: a meta-analysis. Livest. Sci. 155, 244–254 (2013).
Belanche, A. et al. A meta-analysis describing the effects of the essential oils blend Agolin ruminant on performance, rumen fermentation and methane emissions in dairy cows. Animals (Basel) 10, 620 (2020).
Bergen, W. G. & Bates, D. B. Ionophores: their effect on production efficiency and mode of action. J. Anim. Sci. 58, 1465–1483 (1984).
Callaway, T. R. et al. Ionophores: their use as ruminant growth promotants and impact on food safety. Curr. Issues Intest. Microbiol. 4, 43–51 (2003).
Maron, D. F., Smith, T. J. S. & Nachman, K. E. Restrictions on antimicrobial use in food animal production: an international regulatory and economic survey. Global. Health 9, 48 (2013).
Cotter, P. D., Paul Ross, R. & Hill, C. Bacteriocins — a viable alternative to antibiotics? Nat. Rev. Microbiol. 11, 95–105 (2013).
Shen, J., Liu, Z., Yu, Z. & Zhu, W. Monensin and nisin affect rumen fermentation and microbiota differently in vitro. Front. Microbiol. 8, 1111 (2017).
Lourenço, M., Ramos-Morales, E. & Wallace, R. J. The role of microbes in rumen lipolysis and biohydrogenation and their manipulation. Animal 4, 1008–1023 (2010).
Enjalbert, F., Combes, S., Zened, A. & Meynadier, A. Rumen microbiota and dietary fat: a mutual shaping. J. Appl. Microbiol. 123, 782–797 (2017).
Lovett, D. et al. Effect of forage/concentrate ratio and dietary coconut oil level on methane output and performance of finishing beef heifers. Livest. Prod. Sci. 84, 135–146 (2003).
Prabhu, R., Altman, E. & Eiteman, M. A. Lactate and acrylate metabolism by Megasphaera elsdenii under batch and steady-state conditions. Appl. Environ. Microbiol. 78, 8564–8570 (2012).
Ungerfeld, E. M. Metabolic hydrogen flows in rumen fermentation: principles and possibilities of interventions. Front. Microbiol. 11, 589 (2020).
Hristov, A. N. et al. Special topics — Mitigation of methane and nitrous oxide emissions from animal operations: I. A review of enteric methane mitigation options. J. Anim. Sci. 91, 5045–5069 (2013).
Hook, S. E., Wright, A.-D. G. & McBride, B. W. Methanogens: methane producers of the rumen and mitigation strategies. Archaea 2010, 945785 (2010).
Goopy, J. P. et al. Low-methane yield sheep have smaller rumens and shorter rumen retention time. Br. J. Nutr. 111, 578–585 (2014).
Ørskov, E. R., Ojwang, I. & Reid, G. W. A study on consistency of differences between cows in rumen outflow rate of fibrous particles and other substrates and consequences for digestibility and intake of roughages. Anim. Sci. 47, 45–51 (1988).
Herd, R. M. et al. Measures of methane production and their phenotypic relationships with dry matter intake, growth, and body composition traits in beef cattle. J. Anim. Sci. 92, 5267–5274 (2014).
Donoghue, K. A., Bird-Gardiner, T. L., Arthur, P. F., Herd, R. M. & Hegarty, R. F. Genetic parameters for methane production and relationships with production traits in Australian beef cattle. Proc. Assoc. Adv. Anim. Breed. Genet. 21, 114–117 (2015).
Breider, I. S., Wall, E. & Garnsworthy, P. C. Short communication: Heritability of methane production and genetic correlations with milk yield and body weight in Holstein-Friesian dairy cows. J. Dairy Sci. 102, 7277–7281 (2019).
New Zealand Agricultural Greenhouse Gas Research Centre (NZAGRC). Annual Report (2019) 51–59 https://www.nzagrc.org.nz/annualreport,listing,598,annual-report-2019.html (2019).
Difford, G. F. et al. Host genetics and the rumen microbiome jointly associate with methane emissions in dairy cows. PLoS Genet. 14, e1007580 (2018). This study confirms that methane production is influenced by both host genotype and rumen microbiome composition independently from each other.
Xiang, R. et al. Gene network analysis identifies rumen epithelial cell proliferation, differentiation and metabolic pathways perturbed by diet and correlated with methane production. Sci. Rep. 6, 39022 (2016).
Abecia, L., Martín-García, A. I., Martínez, G., Newbold, C. J. & Yáñez-Ruiz, D. R. Nutritional intervention in early life to manipulate rumen microbial colonization and methane output by kid goats postweaning. J. Anim. Sci. 91, 4832–4840 (2013).
Abecia, L. et al. Feeding management in early life influences microbial colonisation and fermentation in the rumen of newborn goat kids. Anim. Produc. Sci. 54, 1449–1454 (2014).
Abecia, L. et al. Analysis of the rumen microbiome and metabolome to study the effect of an antimethanogenic treatment applied in early life of kid goats. Front. Microbiol. 9, 2227 (2018).
Guyader, J. et al. Influence of rumen protozoa on methane emission in ruminants: a meta-analysis approach. Animal 8, 1816–1825 (2014).
Morgavi, D. P., Jouany, J. P. & Martin, C. Changes in methane emission and rumen fermentation parameters induced by refaunation in sheep. Aust. J. Exp. Agric. 48, 69–72 (2008).
Belanche, A., de la Fuente, G. & Newbold, C. J. Effect of progressive inoculation of fauna-free sheep with holotrich protozoa and total-fauna on rumen fermentation, microbial diversity and methane emissions. FEMS Microbiol. Ecol. 91, fiu026 (2015).
Hegarty, R. S., Bird, S. H., Vanselow, B. A. & Woodgate, R. Effects of the absence of protozoa from birth or from weaning on the growth and methane production of lambs. Br. J. Nutr. 100, 1220–1227 (2008).
Hegarty, R. S. Reducing rumen methane emissions through elimination of rumen protozoa. Aust. J. Agric. Res. 50, 1321 (1999).
Yáñez-Ruiz, D. R., Abecia, L. & Newbold, C. J. Manipulating rumen microbiome and fermentation through interventions during early life: a review. Front. Microbiol. 6, 1133 (2015). This review presents findings that relate to early-life rumen microbiome development and potential interventions in the process.
Jami, E., Israel, A., Kotser, A. & Mizrahi, I. Exploring the bovine rumen bacterial community from birth to adulthood. ISME J. 7, 1069–1079 (2013).
Zehavi, T., Probst, M. & Mizrahi, I. Insights into culturomics of the rumen microbiome. Front. Microbiol. 9, 1999 (2018).
Creevey, C. J., Kelly, W. J., Henderson, G. & Leahy, S. C. Determining the culturability of the rumen bacterial microbiome. Microb. Biotechnol. 7, 467–479 (2014).
Levy, B. & Jami, E. Exploring the prokaryotic community associated with the rumen ciliate protozoa population. Front. Microbiol. 9, 2526 (2018).
Lima, F. S. et al. Prepartum and postpartum rumen fluid microbiomes: characterization and correlation with production traits in dairy cows. Appl. Environ. Microbiol. 81, 1327–1337 (2015).
Petri, R. M. et al. Characterization of the core rumen microbiome in cattle during transition from forage to concentrate as well as during and after an acidotic challenge. PLoS ONE 8, e83424 (2013).
Wu, S., Baldwin, R. L., Li, W., Li, C. & Li, R. W. The bacterial community composition of the bovine rumen detected using pyrosequencing of 16S rRNA genes. Metagenomics 1, 1–11 (2012).
Hughes, P. & Heritage, J. Antibiotic growth-promoters in food animals. FAO Anim. Prod. Health Pap. 160, 129–152 (2004).
European Centre for Disease Prevention and Control (ECDC), European Food Safety Authority (EFSA) & European Medicines Agency (EMA). ECDC/EFSA/EMA second joint report on the integrated analysis of the consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from humans and food-producing animals. EFSA J. 15, e04872 (2017).
Van Boeckel, T. P. et al. Global trends in antimicrobial use in food animals. Proc. Natl Acad. Sci. USA 112, 5649–5654 (2015).
Landers, T. F., Cohen, B., Wittum, T. E. & Larson, E. L. A review of antibiotic use in food animals: perspective, policy, and potential. Public Health Rep. 127, 4–22 (2012).
Cuong, N. V., Padungtod, P., Thwaites, G. & Carrique-Mas, J. J. Antimicrobial usage in animal production: a review of the literature with a focus on low- and middle-income countries. Antibiotics (Basel) 7, 75 (2018).
Shterzer, N. & Mizrahi, I. The animal gut as a melting pot for horizontal gene transfer. Can. J. Microbiol. 61, 603–605 (2015).
Kav, A. B. et al. Insights into the bovine rumen plasmidome. Proc. Natl Acad. Sci. USA 109, 5452–5457 (2012).
Sabino, Y. N. V. et al. Characterization of antibiotic resistance genes in the species of the rumen microbiota. Nat. Commun. 10, 5252 (2019).
Brown Kav, A. et al. Unravelling plasmidome distribution and interaction with its hosting microbiome. Environ. Microbiol. 22, 32–44 (2020).
Auffret, M. D. et al. The rumen microbiome as a reservoir of antimicrobial resistance and pathogenicity genes is directly affected by diet in beef cattle. Microbiome 5, 159 (2017).
Noyes, N. R. et al. Characterization of the resistome in manure, soil and wastewater from dairy and beef production systems. Sci. Rep. 6, 24645 (2016).
Thomas, M. et al. Metagenomic characterization of the effect of feed additives on the gut microbiome and antibiotic resistome of feedlot cattle. Sci. Rep. 7, 12257 (2017).
Chambers, L. et al. Metagenomic analysis of antibiotic resistance genes in dairy cow feces following therapeutic administration of third generation cephalosporin. PLoS ONE 10, e0133764 (2015).
Cameron, A. & McAllister, T. A. Antimicrobial usage and resistance in beef production. J. Anim. Sci. Biotechnol. 7, 68 (2016).
Muurinen, J. et al. Influence of manure application on the environmental resistome under Finnish agricultural practice with restricted antibiotic use. Environ. Sci. Technol. 51, 5989–5999 (2017).
Tripathi, V. & Cytryn, E. Impact of anthropogenic activities on the dissemination of antibiotic resistance across ecological boundaries. Essays Biochem. 61, 11–21 (2017).
Chantziaras, I., Boyen, F., Callens, B. & Dewulf, J. Correlation between veterinary antimicrobial use and antimicrobial resistance in food-producing animals: a report on seven countries. J. Antimicrob. Chemother. 69, 827–834 (2014).
Acknowledgements
The authors acknowledge the collaborative nature of the work on the rumen microbiome by thanking the global scientific community involved in rumen microbiome research and the deciphering of its mysteries. The authors also thank E. Jami (Agricultural Research Organization, Volcani Center, Israel) and E. A. Bayer (The Weizmann Institute of Science, Israel) for critical reading of the manuscript. Research in the authors’ laboratory was supported by grants from the European Research Council (No. 640384) to I.M. and from the Israel Science Foundation (ISF No. 1947/19) to I.M. and S.M.
Author information
Authors and Affiliations
Contributions
I.M. and S.M. researched data for the article, substantially contributed to discussion of content and wrote the article. I.M., R.J.W. and S.M. conceptualized, reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Microbiology thanks S. Huws, T. McAllister and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Food and Agriculture Organization of the United Nations (FAOSTAT) enteric fermentation: http://www.fao.org/faostat/en/#data/GE
Glossary
- Core microbiome
-
A group of microorganisms commonly found within the microbiome across multiple hosts.
- Food webs
-
The interconnections among different microbial food chains.
- Electron sinks
-
In this context, microorganisms within a microbial community that accept electrons at the final step of the electron flow.
- Hydrogenotrophs
-
Methanogens that use H2 to reduce CO2 to methane.
- Methylotrophs
-
Methanogens that use a methylated compound as the input metabolite to produce methane.
- Acetoclastic methanogens
-
Methanogens that use acetate to produce methane.
- Alternative stable community states
-
Assemblages of functional groups that are locked and stabilized by metabolic feedback and result in distinct composition, function and, thus, outcome.
- Functional microbial groups
-
Groups of organisms that share similar functionality within the ecosystem.
- Horizontal gene transfer
-
(HGT). The lateral mobilization of genetic material between distinct microorganisms.
- Rumen plasmidome
-
The collective plasmid population in the rumen.
- Resistome
-
The collection of antibiotic resistance genes in a given environment.
- Biohydrogenation
-
The microbial transformation of unsaturated fatty acid to saturated fatty acid.
- Residual feed intake
-
A parameter that describes feed efficiency, measuring the difference between the actual feed intake and the predicted intake, based on an animal’s body weight, weight gain and milk composition, and is a proxy for the animal’s ability to extract energy from its feed.
Rights and permissions
About this article
Cite this article
Mizrahi, I., Wallace, R.J. & Moraïs, S. The rumen microbiome: balancing food security and environmental impacts. Nat Rev Microbiol 19, 553–566 (2021). https://doi.org/10.1038/s41579-021-00543-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-021-00543-6
This article is cited by
-
Diet and monensin influence the temporal dynamics of the rumen microbiome in stocker and finishing cattle
Journal of Animal Science and Biotechnology (2024)
-
Early-life ruminal microbiome-derived indole-3-carboxaldehyde and prostaglandin D2 are effective promoters of rumen development
Genome Biology (2024)
-
Coping with extremes: the rumen transcriptome and microbiome co-regulate plateau adaptability of Xizang goat
BMC Genomics (2024)
-
A compendium of ruminant gastrointestinal phage genomes revealed a higher proportion of lytic phages than in any other environments
Microbiome (2024)
-
Causal relationship between gut microbiota and gastrointestinal diseases: a mendelian randomization study
Journal of Translational Medicine (2024)