Abstract
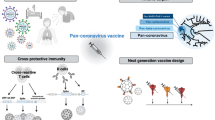
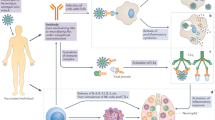
Vaccines are urgently needed to control the coronavirus disease 2019 (COVID-19) pandemic and to help the return to pre-pandemic normalcy. A great many vaccine candidates are being developed, several of which have completed late-stage clinical trials and are reporting positive results. In this Progress article, we discuss which viral elements are used in COVID-19 vaccine candidates, why they might act as good targets for the immune system and the implications for protective immunity.
Similar content being viewed by others
Introduction
As of 3 December 2020, the coronavirus disease 2019 (COVID-19) pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection, has spread to 220 countries, areas or territories with more than 63 million laboratory-confirmed cases and more than 1.4 million deaths (World Health Organization (WHO)), leading to widespread social and economic disruption. The development of a safe and effective vaccine is urgently needed to help bring an end to this pandemic.
SARS-CoV-2 is a member of the Coronaviridae family, which comprises many virulent strains that infect humans and animals, including SARS-CoV and Middle East respiratory syndrome CoV (MERS-CoV)1. To date, a number of CoV vaccines have been licensed for use in domestic animals against canine CoV, feline CoV, bovine CoV (BCoV), porcine epidemic diarrhoea virus, transmissible gastroenteritis virus (TGEV) and infectious bronchitis virus (IBV)2; however, until now, none has been licensed for use in humans. Two vaccine candidates for SARS-CoV and three for MERS-CoV are in phase I clinical trials (WHO). These prior experiences of vaccine development for animal and human CoVs have provided important insights into the development of vaccines for SARS-CoV-2 infection.
To develop a vaccine against a newly emerged virus, it is important to understand the immune correlates of protection. Although much remains to be determined regarding immune correlates of protection for SARS-CoV-2 infection, emerging data have demonstrated the importance of both humoral and cellular immunity in protection. A strong correlation has been found between vaccine-induced neutralizing antibodies (nAbs) and a reduction of viral loads in non-human primates (NHPs) after SARS-CoV-2 infection3,4,5,6. In humans, passive administration of convalescent plasma7,8,9,10, purified IgG11,12 or monoclonal antibodies13 have been reported to show benefit for the treatment and prevention of infection by SARS-CoV-2. In particular, a nAb recently received authorization by the US Food and Drug Administration for emergency use as a treatment for COVID-19 (ref.14). Moreover, analysis of a COVID-19 outbreak aboard a fishery vessel with high infection rates supported the correlation of nAbs with protection15. In addition to nAbs, T cell responses also play critical protective roles in CoV infections. The depletion of T cells in mice has been shown to impair virus clearance in SARS-CoV, MERS-CoV and SARS-CoV-2 infections16,17,18,19. In patients, virus-specific CD4+ and CD8+ T cell responses are associated with milder disease, suggesting an involvement in protective immunity against COVID-19 (refs20,21,22). Therefore, an ideal vaccine is expected to evoke both the humoral and cellular arms of the immune system. However, an important safety concern for the development of a SARS-CoV-2 vaccine or of antibody-based therapies is the potential risk of vaccine enhancement of the disease, also known as antibody-dependent enhancement (ADE) and enhanced respiratory disease (ERD)23. Antibodies that can bind to a virus without neutralizing activities can cause ADE via Fcγ receptor-mediated virus uptake, allowing subsequent replication of the virus or Fc-mediated effector functions of the antibody–virus immune complex, allowing immunopathology23,24. This effect is typically associated with flaviviruses, such as dengue virus25,26 and Zika virus27, but it has also been described in CoV infection. Cats immunized with vaccinia virus expressing a viral protein of feline infectious peritonitis virus (FIPV; a feline CoV) or passively administered with anti-FIPV antibodies showed early mortality when challenged with the live virus28,29,30,31. ADE was also observed for SARS-CoV and MERS-CoV in animal models32,33,34,35,36,37. In addition to ADE, vaccine-induced enhancement of disease can also be caused by T helper 2 (TH2) cell-biased immunopathology, leading to ERD38,39,40,41. Although some studies of SARS-CoV in animal models do not show evidence of ADE or ERD33,42,43, safety should be considered when designing vaccines for SARS-CoV-2.
With continuing cases and deaths from the COVID-19 pandemic, researchers worldwide are racing to develop COVID-19 vaccines. According to the landscape document from the WHO, COVID-19 vaccine candidates generally fall into seven strategies (Box 1), which can be divided into three broad categories44: first, protein-based vaccines that generate target antigens in vitro such as inactivated virus vaccines, virus-like particles and protein subunit vaccines; second, gene-based vaccines that deliver genes encoding viral antigens to host cells for in vivo production such as virus-vectored vaccines, DNA vaccines and mRNA vaccines; and, third, a combination of both protein-based and gene-based approaches to produce protein antigen or antigens both in vitro and in vivo, typically represented by live-attenuated virus vaccines. As of December 2020, the WHO has documented more than 214 COVID-19 vaccine candidates, with 51 of them in clinical evaluation, 13 in phase III trials and several vaccines now being authorized for use in some regions (WHO; COVID-19 Vaccine tracker). In this Progress article, we summarize and discuss the targets used in vaccine candidates, focusing on those candidates already advanced into clinical trials and with published data.
SARS-CoV-2 proteins as targets
SARS-CoV-2 contains four major structural proteins, namely spike (S), membrane (M) and envelope (E) proteins, all of which are embedded in the viral surface envelope, and nucleocapsid (N) protein, which is in the ribonucleoprotein core1,45 (Fig. 1). S proteins are responsible for recognition of the host cellular receptor to initiate virus entry. M proteins are embedded in the envelope and shape the virion envelope. E proteins are small polypeptides that are crucial for CoV infectivity. N proteins make up the helical nucleocapsid and bind along the viral RNA genome. In addition to these structural proteins, SARS-CoV-2 encodes 16 non-structural proteins (nsp1–16) and 9 accessory proteins. Several of these viral proteins could potentially serve as targets of vaccine-induced immune responses.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) contains four major structure proteins: spike (S), membrane (M) and envelope (E) proteins, which are embedded on the virion surface, and nucleocapsid (N) protein, which binds viral RNA inside the virion. The S protein trimer in its pre-fusion conformation is shown. The S protein comprises the S1 subunit (which includes the N-terminal domain (NTD) and the receptor-binding domain (RBD)) (the receptor-binding motif (RBM) within the RBD is also labelled) and the S2 subunit (which includes fusion peptide (FP), connecting region (CR), heptad repeat 1 (HR1), heptad repeat (HR2) and central helix (CH)). The SARS-CoV-2 S protein binds to its host receptor, the dimeric human angiotensin-converting enzyme 2 (hACE2), via the RBD and dissociates the S1 subunits. Cleavage at both S1–S2 and S2′ sites allows structural rearrangement of the S2 subunit required for virus–host membrane fusion. The S2-trimer in its post-fusion arrangement is shown. The RBD is an attractive vaccine target. The generation of an RBD-dimer or RBD-trimer has been shown to enhance the immunogenicity of RBD-based vaccines. A stabilized S-trimer shown with a C-terminal trimer-tag is a vaccine target. The pre-fusion S protein is generally metastable during in vitro preparations and prone to transform into its post-fusion conformation. Mutation of two residues (K986 and V987) to proline stabilizes S protein (S-2P) and prevents the pre-fusion to post-fusion structural change.
S protein
S protein is the main protein used as a target in COVID-19 vaccines. S protein consists of a membrane-distal S1 subunit and a membrane-proximal S2 subunit and exists in the virus envelope as a homotrimer. The S1 subunit determines receptor recognition via its receptor-binding domain (RBD), whereas the S2 subunit is responsible for membrane fusion, which is required for virus entry (Fig. 1). In MERS-CoV, SARS-CoV or SARS-CoV-2, the RBD is located in the C-terminal domain of the S1 subunit46,47,48. In some CoVs, the N-terminal domain (NTD) of the S1 subunit can be used for receptor binding (such as in mouse hepatitis virus) or might also be involved in virus attachment to host cells by recognizing specific sugar molecules (such as in TGEV, BCoV and IBV) or has an important role in the pre-fusion to post-fusion transition of the S protein49,50,51. The S2 subunit contains the fusion peptide (FP), connecting region (CR), heptad repeat 1 (HR1) and HR2 around a central helix as a helix-turn-helix structure (Fig. 1). Structural evidence has proposed a model for the rearrangement of SARS-CoV-2 S protein following recognition of the host cell receptor52,53. S protein can be proteolytically cleaved at both the S1–S2 and S2′ cleavage sites, releasing the structural constraints on the FP (Fig. 1). The engagement of the RBD with its cellular receptor, human angiotensin-converting enzyme 2 (hACE2), leads to the dissociation of the S1 subunit and concomitantly initiates the refolding process of the spring-loaded S2 subunit, which protrudes the FP at its end for membrane fusion. The S2 subunit in its post-fusion conformation folds as a long helical bundle with the FP inserted into the host cell membrane (Fig. 1). Theoretically, nAbs can target the S protein to inhibit virus infection at multiple stages during the virus entry process. The RBD is the major target for nAbs that interfere with viral receptor binding54,55,56. To date, most of the potent nAbs to SARS-CoV-2 target the RBD57,58,59,60,61,62,63,64,65,66,67,68,69. In addition, nAbs targeting the NTD have been reported in SARS-CoV-2 and MERS-CoV infection49,63,70,71, making it another potential target for inclusion in a vaccine. The S2 subunit is also a potential target for nAbs that interfere with the structural rearrangement of the S protein and the insertion of FP required for virus–host membrane fusion49,71,72,73. The S protein is also a target for T cell responses; studies of SARS-CoV, MERS-CoV and SARS-CoV-2 have described both CD4+ and CD8+ T cell epitopes in the S protein74,75.
To date, three COVID-19 vaccine candidates based on adenovirus expressing the full-length S protein have entered phase III clinical trials (Table 1). One, which is being developed in China, is based on human adenovirus type 5 (Ad5)76,77. The second, being developed in the UK, uses recombinant chimpanzee adenovirus, ChAdOx1 (refs78,79). The third vaccine candidate, from Russia, combines recombinant human Ad26 and Ad5 in a prime–boost vaccination regimen80. The preclinical data showed that the recombinant ChAdOx1 vaccine induced nAb production and balanced TH1 and TH2 immune responses in NHPs78 (Table 1). The vaccine prevented COVID-19-associated pneumonia in NHPs and reduced viral loads in both bronchoalveolar lavage fluid and the respiratory tract78. Phase II trial studies showed that all of these vaccine candidates induced nAb production and TH1 cell-biased responses in humans76,79,80 (Table 1).
In addition to the adenovirus-vector strategy, a DNA vaccine candidate expressing the S protein has also completed phase II clinical trials. This vaccine elicited nAbs and S protein-specific T cell responses in both mice and guinea pigs81 (Table 1). Recently, a native-like trimeric S protein was reported for a COVID-19 subunit vaccine candidate being developed in China, in which the S protein is fused to a trimer-tag, the C-terminal region of human type Iα collagen, to form a disulfide-bonded homotrimer82. This strategy stabilizes the S protein antigen in its trimeric form and increases the antigen yield82. The recombinant trimeric protein structurally mimics the native S protein presented on virus particles and could serve as a target for nAbs, a strategy that is also being pursued in the development of an HIV vaccine83. Vaccination of the S-trimer together with AS03 (a squalene-based adjuvant developed by GlaxoSmithKline) or CpG 1018 (a Toll-like receptor 9 agonist adjuvant developed by Dynavax) elicited high levels of nAbs and TH1 cell-biased responses in mice and NHPs. It could protect NHPs from challenge with SARS-CoV-2, with reduced viral loads and pathology in the lungs84. This candidate vaccine is currently under evaluation in phase I clinical trials (Table 1).
The S protein is metastable when produced as a recombinant protein and prone to transform from its pre-fusion to a post-fusion conformation, shedding the S1 subunit (Fig. 1). However, the S1 subunit is the immunodominant antigen during CoV infections due to its accessibility for immune recognition and it contains neutralizing epitopes mainly on its RBD. Strategies to stabilize the S protein in its pre-fusion conformation and enhance pre-fusion S protein expression are thought to increase the quality and quantity of vaccine-induced antibodies targeting the functionally relevant epitopes on the S1 subunit. Pallesen et al. reported two proline substitutions (2P) at the apex of the central helix and HR1 that can retain the S proteins of MERS-CoV, SARS-CoV and HKU1 in the antigenically optimal pre-fusion conformation85 (Fig. 1). The resulting antigen, S-2P, induced much greater nAb titres than wild-type S protein in mice. Learning from the previous experience with these CoVs, the S-2P design is now being used in several vaccine strategies against COVID-19. SARS-CoV-2 S-2P (comprising proline substitutions at residues K986 and V987) is used as the target antigen in three gene-based vaccine candidates (mRNA vaccines by Moderna/National Institute of Allergy and Infectious Diseases (NIAID) and BioNTech/Pfizer and a recombinant Ad26 vaccine by Janssen Pharmaceutical Companies) and a protein-based candidate (by Novavax)4,86,87,88,89,90,91,92,93,94,95,96 (Table 1). Moreover, mutation at the cleavage sites in the S protein is also believed to stabilize the pre-fusion conformation of the S protein. S-2P in the Janssen Ad26-vectored vaccine (Ad26.COV2.S) and in the Novavax protein-based vaccine (NVX-CoV2373) contains additional mutations at the S1–S2 polybasic cleavage site from RRAR to SRAG or QQAQ to render it protease resistant, which helps to further stabilize the S protein in its pre-fusion conformation96,97. All these candidates elicited protective immunity in animal models86,87,90 and have advanced into phase III clinical trials (Table 1). In particular, a preclinical study showed that Ad26 expressing S-2P elicited higher nAb titres and better protection in NHPs than other candidates based on the wild-type S protein4. The clinical studies for the two mRNA vaccines (BNT162b2 by BioNTech/Pfizer and mRNA1273 by Moderna/NIAID) and the protein subunit vaccine, NVX-CoV2373, encouragingly showed that all these vaccines elicited both high titres of nAbs beyond the levels observed in convalescent patients and substantial T cell responses88,89,91,93. Therefore, stabilization of the S protein in its pre-fusion conformation represents an effective avenue to improve vaccine efficacy. Notably, although the data are yet to be published, recent press releases from BioNTech/Pfizer and Moderna/NIAID have announced that their mRNA vaccines have more than 90% efficacy in preventing COVID-19 in phase III clinical trials98,99. The UK has just approved the BioNTech/Pfizer vaccine100.
Following their initial observations in MERS-CoV, SARS-CoV and SARS-CoV-2, Hsieh et al. generated a new variant form of the S protein ectodomain, known as HexaPro, which comprises six beneficial proline substitutions, including the two from S-2P101. These proline substitutions are located at flexible loops or N termini of helices in the FP, HR1 and CR, further constraining the structural rearrangement of the S2 subunit and stabilizing the pre-fusion S protein. HexaPro exhibits approximately 10-fold higher expression than S-2P, thus representing a promising antigen design101.
The RBD
The RBD binds to the host receptor via a receptor-binding motif (RBM) on its external subdomain in SARS-CoV, MERS-CoV or SARS-CoV-2 (refs47,50,102). The surface of the S protein is extensively shielded from antibody recognition by glycans, with the notable exception of the RBD, which explains the immunodominance of RBD epitopes103. Most SARS-CoV-2 nAbs bind to RBD and block the RBD–hACE2 interaction, thus inhibiting virus attachment57,58,59,60,61,62,63,64,65,66,67,68,69. RBD is an attractive vaccine target because it elicits high-quality, functionally relevant antibodies, while avoiding the potential risk of ADE, which is generally thought to be mediated by weak nAbs or non-nAbs24,30,104. For example, antibodies targeting an epitope (S597-603) downstream of the C terminus of the SARS-CoV RBD markedly enhanced SARS-CoV infection36,105. The RBD also contains epitopes for T cell responses, as shown in studies of SARS-CoV, MERS-CoV and SARS-CoV-2 (refs74,106,107,108). RBD-based antigens have been described in previous studies for SARS-CoV and MERS-CoV vaccine development74,109,110,111.
To date, several RBD-based vaccines for COVID-19 have entered clinical trials. Yang et al. reported an RBD-based COVID-19 vaccine candidate generated using a protein subunit strategy107 (Table 1). This vaccine induced nAbs in mice, rabbits and NHPs, and protected NHPs against challenge with SARS-CoV-2. They further showed that the immune sera, as opposed to splenic T cells, played the protective role in mice107. Consistent with this, a recent report of an RBD-based DNA vaccine also showed that the nAbs, and not the induced T cells, are the immune correlates of protection against COVID-19 in NHPs3. An RBD-based mRNA vaccine is being developed in China and is currently in phase I trials. This candidate vaccine, ARCoV, which expresses SARS-CoV RBD delivered by lipid nanoparticles, induced both nAb production and TH1 cell-biased responses in mouse and NHP models. Vaccination protected mice against challenge with a mouse-adapted SARS-CoV-2 strain106.
However, the use of RBD in vaccines is compromised by its limited immunogenicity owing to its small molecular size and possible mixed forms of multiple complexes (as monomers, dimers or trimers). Strategies to overcome these drawbacks include increasing antigen size (for example, by fusing the RBD with an Fc domain112,113,114,115,116) or by RBD multimerization (for example, by displaying multiple copies of RBD on particles117,118,119). Recently, to address these limitations, our team described a generalizable strategy to design a dimeric form of the RBD of beta-CoV antigens suitable for use against SARS-CoV-2, MERS-CoV and SARS-CoV120 (Fig. 1). The RBDs from SARS-CoV, MERS-CoV or SARS-CoV-2 spontaneously form dimers in solution120,121,122,123,124. Structural analyses showed, for MERS-CoV118 and SARS-CoV121, that both RBD protomers in a dimer stack on top of each other via the core subdomains and expose the RBM, the major site recognized by nAbs, indicating similar RBD-dimer structures for other CoVs. Structure-guided design yielded homogeneous RBD-dimers as a tandem-repeat single chain. The RBD-dimer antigen induced 10-fold to 100-fold higher nAb titres than the conventional RBD-monomer and was protective in a mouse model120. We developed a COVID-19 vaccine candidate, ZF2001, comprising the RBD-dimer as the target. This protein subunit vaccine is currently being evaluated in phase III clinical trials (Table 1). In addition to the RBD-dimer, an mRNA vaccine, BNT162b1 (BioNTech/Pfizer), was reported to express an RBD-trimer stabilized by the foldon trimerization domain (Fig. 1). The phase I/II studies of this vaccine encouragingly showed that two doses of the vaccine induced nAbs to levels higher than those in convalescent patients as well as inducing TH1 cell-biased responses94,95 (Table 1). Indeed, it is assumed that multivalent antigens would allow the crosslinking of B cell receptors for better B cell activation125.
Interestingly, a recent study described a vaccine candidate comprising multiple copies of the SARS-CoV-2 RBD displayed in arrays on nanoparticles that induced a markedly lower binding to nAb ratio than a S-2P-based vaccine119, indicating the humoral responses are focused on epitopes recognized by nAbs for the RBD-based vaccines, which are believed to have lower ADE potential.
S1-NTD and the S2 subunit
The S1-NTD also contains epitopes for CoV nAbs found in infected patients49,63,70,71,126,127 and has been considered a potential target in CoV vaccines. NTD-targeting nAbs generally do not directly block receptor binding but rather interfere with receptor binding126,127 or restrain the S protein conformational changes required for the pre-fusion to post-fusion transition49,127. It is notable that SARS-CoV-2 NTD-targeting nAbs generally exhibit lower neutralizing potency than RBD-specific nAbs49. We previously reported an NTD-based vaccine against MERS-CoV128. Vaccination with NTD protein elicited nAbs and NTD-specific T cell responses. Furthermore, it reduced lung abnormalities in a MERS-CoV challenge mouse model, although the immunogenicity and protective efficacy of the NTD protein were weaker than the RBD protein128. The inclusion of NTD in a COVID-19 vaccine would broaden the neutralizing epitopes and reduce the potential of viral escape of host immunity. Yet, so far, NTD-based vaccines against COVID-19 have not been reported.
For the S2 subunit, peptides derived from HR1 or HR2 of the S2 subunit from SARS-CoV, MERS-CoV and SARS-CoV-2 have been described that inhibit viral fusion with target cells and thereby prevent virus infection. Moreover, nAbs have been reported to target the S2 subunit of CoVs, including SARS-CoV-2 (refs49,129,130,131), suggesting the S2 subunit as a COVID-19 vaccine target. However, the membrane-proximal S2 subunit contains more extensive N-glycan shielding132,133 and is less accessible for immune recognition than the S1 subunit and is therefore less immunogenic. Rabbits immunized with SARS-CoV-2 S2 protein showed much lower nAb titres than those immunized with the S1 subunit or RBD proteins134. S2 subunit-targeting antibodies isolated from convalescent patients showed weaker neutralizing activities against SARS-CoV-2 than RBD-targeting antibodies64. These studies suggest that the S2 subunit alone may not be an effective target for humoral responses. Nevertheless, because of the relative sequence conservation of the S2 subunit between virus species, the S2 subunit is targeted by cross-reactive antibodies and CD4+ T cells recognizing both SARS-CoV-2 and other human CoVs64,131,135,136, suggesting a potential target in universal CoV vaccines.
M, E and N proteins
Unlike the S protein, CoV M and E proteins are poorly immunogenic for humoral responses, presumably owing to their small ectodomains for immune cell recognition and small molecular sizes1,137 (Fig. 1). Adoptive transfer of sera from donors immunized with a virus vector expressing M or E protein did not protect mice against SARS-CoV-2 infection16. Therefore, M and E proteins have never been explored as vaccine targets alone against SARS-CoV-2 or other CoVs. Nonetheless, the sequence identity of M or E proteins among SARS-CoV, MERS-CoV and SARS-CoV-2 is much higher than for the S protein and RBD, suggesting the potential of M and E proteins as targets for cross-reactive T cells. Indeed, several T cell epitopes have been identified in M and E proteins in previous studies of SARS-CoV and MERS-CoV immunity74. In this regard, M and E proteins may help to broaden the T cell response and improve cross-protection if included in a SARS-CoV-2 vaccine.
The N protein is the most abundant viral protein and is highly immunogenic during CoV infections139. It is a major target for antibody responses and also contains T cell epitopes140. N-specific antibodies were reported to protect mice against mouse hepatitis virus, a mouse CoV, via Fc-mediated effector functions141,142. However, anti-N immune sera did not protect against SARS-CoV-2 infection in a mouse model16. Immunization with N protein can also elicit CD4+ and CD8+ T cell responses in mice143. N-specific CD8+ T cell epitopes are known to protect chickens against IBV infection144. Venezuelan equine encephalitis virus replicon particles expressing an N-specific CD4+ T cell epitope showed complete protection against SARS-CoV infection138. These virus replicon particles also conferred partial cross-protection against MERS-CoV owing to the protein sequence conservation between viruses, resulting in a reduced viral load138. However, its potential as a CoV vaccine target was largely undermined by early studies of SARS-CoV showing that vaccines expressing N protein did not provide protection and, on the contrary, enhanced infection-induced pneumonia via increased pulmonary eosinophil infiltration and TH2 cell-biased responses39,41, causing ERD. Therefore, the inclusion of N protein in CoV vaccines is complicated by balancing viral clearance and immunopathogenesis and no N protein-based vaccine has been reported for COVID-19.
The whole virus as a target
Inactivated virus and live-attenuated virus vaccines use the whole virus as vaccine targets. They contain all structural proteins (S, N, M and E proteins) and live-attenuated virus vaccines can also generate non-structural and accessory proteins in vivo. Therefore, these vaccine candidates can induce broader antibody and T cell responses than the above-mentioned vaccines, which are based on a single protein or protein fragments.
Three vaccine candidates (all being developed in China) based on an inactivated virus strategy are currently in phase III clinical trials and another one is at the phase I/II stage (Table 1). In a preclinical study, mice immunized with an inactivated virus vaccine, PiCoVacc, elicited S protein-specific (including RBD-specific) and N-specific antibodies5. PiCoVacc and another inactivated virus vaccine, BBIBP-CorV, elicited substantial nAb production in NHPs but did not induce T cell responses5,6. Furthermore, they protected NHPs against challenge by SARS-CoV-2, suggesting an important role for humoral responses as the immune correlates of protection. Initial clinical trial studies of these two inactivated vaccine candidates showed that, similar to in NHPs, these vaccines elicited substantial nAbs in humans145,146. These inactivated vaccine candidates did not induce either TH1 or TH2 cell-associated cytokines in those vaccinated. As the uptake of inactivated vaccine components by host cells is very limited or negligible, antigen processing for peptide presentation to T cells is inefficient and may explain why these vaccines generally do not elicit T cell responses. Moreover, these inactivated vaccines all use Alum as the adjuvant, which is generally a poor stimulator of T cell responses (Table 1). Notably, although enhancement of disease has been associated with inactivated SARS-CoV vaccines in animal models38,40, such an effect has not been reported for inactivated vaccines for SARS-CoV-2 in either preclinical or clinical studies.
Live-attenuated virus vaccines have been licensed for animals against IBV, TGEV, BCoV and FIPV140. However, waning immunity has been found following IBV and BCoV vaccines140, raising concerns about the duration of immune responses elicited by SARS-CoV-2 vaccines. Indeed, it is worth noting a study showing that nAb titres in 93.3% of patients with COVID-19 declined gradually over the 3-month study period147. The relationship between impaired immunological memory and CoV proteins in the live virus needs to be further explored. The process of virus attenuation for these vaccines is usually time consuming. So far, only three live-attenuated vaccines against COVID-19 are in preclinical evaluation, with codon-deoptimization used as the attenuation strategy (WHO).
Concluding remarks
An ideal COVID-19 vaccine target would be expected to induce high titres of nAbs, reduce non-nAb production to minimize ADE potential, elicit robust TH1 cell-biased responses but low TH2 cell-biased responses to lower the ERD potential, maintain long-lasting immunological memory, and provide cross-protection between CoVs. No combination of different targets has yet been tested as a multiple-target vaccine but it might be worthwhile in the future to look at this possibility.
To date, several vaccine candidates have reached the final stages for vaccine safety and protection efficacy in large-scale clinical trials, with very recent announcements from BioNTech/Pfizer and Moderna/NIAID claiming safety and very high levels of protection efficacy for their leading mRNA vaccine candidates. It is worth noting that it is currently difficult to compare the various vaccines as there are no standardized assays for neutralization and challenge studies. The pros and cons of these vaccine candidates with the respective targets and strategies will need to be further dissected to better understand their safety, immunogenicity and protection rate. Moreover, careful longitudinal studies will be needed to ascertain the duration of any protective immunity and potential disease enhancement for each vaccine candidate148. Moreover, immunization programmes require further exploration. For instance, almost all vaccine candidates in clinical trials are delivered via intramuscular or intradermal routes. It is unclear whether nAbs in the sera reach the respiratory system, especially the lungs, to function. It is possible that mucosal vaccination via the respiratory system would be beneficial to induce immune responses at the mucosa and protect against SARS-CoV-2 transmission in situ through the respiratory tract. Accordingly, a recent report describes an intranasal ChAd vaccine that provides mostly sterilizing immunity to SARS-CoV-2 infection in a mouse model149. In general, as COVID-19 vaccines will be the first such efforts for a human CoV control strategy, more challenges remain.
References
Masters S. P. & Perlman S. in Fields virology 6th edn Ch. 28 (eds Knipe, D. M. & Howley, P. M.). 825–858 (Wolters Kluwer Health/Lippincott Williams & Wilkins, 2013).
Tizard, I. R. Vaccination against coronaviruses in domestic animals. Vaccine 38, 5123–5130 (2020).
Yu, J. et al. DNA vaccine protection against SARS-CoV-2 in rhesus macaques. Science 369, 806–811 (2020).
Mercado, N. B. et al. Single-shot Ad26 vaccine protects against SARS-CoV-2 in rhesus macaques. Nature 586, 583–588 (2020).
Gao, Q. et al. Development of an inactivated vaccine candidate for SARS-CoV-2. Science 369, 77–81 (2020).
Wang, H. et al. Development of an inactivated vaccine candidate, BBIBP-CorV, with potent protection against SARS-CoV-2. Cell 182, 713–721 (2020).
Bloch, E. M. et al. Deployment of convalescent plasma for the prevention and treatment of COVID-19. J. Clin. Invest. 130, 2757–2765 (2020).
Shen, C. et al. Treatment of 5 critically ill patients with COVID-19 with convalescent plasma. JAMA 323, 1582–1589 (2020).
Fung, M. et al. Treatment of immunocompromised COVID-19 patients with convalescent plasma. Transpl. Infect. Dis. https://doi.org/10.1111/tid.13477 (2020).
Hueso, T. et al. Convalescent plasma therapy for B-cell depleted patients with protracted COVID-19 disease. Blood 136, 2290–2295 (2020).
Liu, X., Cao, W. & Li, T. High-dose intravenous immunoglobulins in the treatment of severe acute viral pneumonia: the known mechanisms and clinical effects. Front. Immunol. 11, 1660 (2020).
Burrage, D. R., Koushesh, S. & Sofat, N. Immunomodulatory drugs in the management of SARS-CoV-2. Front. Immunol. 11, 1844 (2020).
Chen, P. et al. SARS-CoV-2 neutralizing antibody LY-CoV555 in outpatients with Covid-19. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2029849 (2020).
Mahase, E. Covid-19: FDA authorises neutralising antibody bamlanivimab for non-admitted patients. BMJ 371, m4362 (2020).
Addetia, A. et al. Neutralizing antibodies correlate with protection from SARS-CoV-2 in humans during a fishery vessel outbreak with high attack rate. J. Clin. Microbiol. 58, e02107-20 (2020).
Sun, J. et al. Generation of a broadly useful model for COVID-19 pathogenesis, vaccination, and treatment. Cell 182, 734–743 (2020).
Chen, J. et al. Cellular immune responses to severe acute respiratory syndrome coronavirus (SARS-CoV) infection in senescent BALB/c mice: CD4+ T cells are important in control of SARS-CoV infection. J. Virol. 84, 1289–1301 (2010).
Zhao, J., Zhao, J. & Perlman, S. T cell responses are required for protection from clinical disease and for virus clearance in severe acute respiratory syndrome coronavirus-infected mice. J. Virol. 84, 9318–9325 (2010).
Zhao, J. et al. Rapid generation of a mouse model for Middle East respiratory syndrome. Proc. Natl Acad. Sci. USA 111, 4970–4975 (2014).
Zhu, L. et al. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of COVID-19 and influenza patients. Immunity 53, 685–696 (2020).
Mathew, D. et al. Deep immune profiling of COVID-19 patients reveals distinct immunotypes with therapeutic implications. Science 369, eabc851 (2020).
Sekine, T. et al. Robust T cell immunity in convalescent individuals with asymptomatic or mild COVID-19. Cell 183, 158–168 (2020).
Lee, W. S., Wheatley, A. K., Kent, S. J. & DeKosky, B. J. Antibody-dependent enhancement and SARS-CoV-2 vaccines and therapies. Nat. Microbiol. 5, 1185–1191 (2020).
Rey, F. A., Stiasny, K., Vaney, M. C., Dellarole, M. & Heinz, F. X. The bright and the dark side of human antibody responses to flaviviruses: lessons for vaccine design. EMBO Rep. 19, 206–224 (2018).
Halstead, S. B. & O’Rourke, E. J. Dengue viruses and mononuclear phagocytes. I. Infection enhancement by non-neutralizing antibody. J. Exp. Med. 146, 201–217 (1977).
Halstead, S. B., O’Rourke, E. J. & Allison, A. C. Dengue viruses and mononuclear phagocytes. II. Identity of blood and tissue leukocytes supporting in vitro infection. J. Exp. Med. 146, 218–229 (1977).
Katzelnick, L. C. et al. Zika virus infection enhances future risk of severe dengue disease. Science 369, 1123–1128 (2020).
Takano, T., Kawakami, C., Yamada, S., Satoh, R. & Hohdatsu, T. Antibody-dependent enhancement occurs upon re-infection with the identical serotype virus in feline infectious peritonitis virus infection. J. Vet. Med. Sci. 70, 1315–1321 (2008).
Takano, T., Yamada, S., Doki, T. & Hohdatsu, T. Pathogenesis of oral type I feline infectious peritonitis virus (FIPV) infection: antibody-dependent enhancement infection of cats with type I FIPV via the oral route. J. Vet. Med. Sci. 81, 911–915 (2019).
Hohdatsu, T., Nakamura, M., Ishizuka, Y., Yamada, H. & Koyama, H. A study on the mechanism of antibody-dependent enhancement of feline infectious peritonitis virus infection in feline macrophages by monoclonal antibodies. Arch. Virol. 120, 207–217 (1991).
Vennema, H. et al. Early death after feline infectious peritonitis virus challenge due to recombinant vaccinia virus immunization. J. Virol. 64, 1407–1409 (1990).
Jaume, M. et al. Anti-severe acute respiratory syndrome coronavirus spike antibodies trigger infection of human immune cells via a pH- and cysteine protease-independent FcγR pathway. J. Virol. 85, 10582–10597 (2011).
Kam, Y. W. et al. Antibodies against trimeric S glycoprotein protect hamsters against SARS-CoV challenge despite their capacity to mediate FcγRII-dependent entry into B cells in vitro. Vaccine 25, 729–740 (2007).
Wan, Y. et al. Molecular mechanism for antibody-dependent enhancement of coronavirus entry. J. Virol. 94, e02015-19 (2020).
Wang, S. F. et al. Antibody-dependent SARS coronavirus infection is mediated by antibodies against spike proteins. Biochem. Biophys. Res. Commun. 451, 208–214 (2014).
Wang, Q. et al. Immunodominant SARS coronavirus epitopes in humans elicited both enhancing and neutralizing effects on infection in non-human primates. ACS Infect. Dis. 2, 361–376 (2016).
Liu, L. et al. Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight 4, e123158 (2019).
Tseng, C. T. et al. Immunization with SARS coronavirus vaccines leads to pulmonary immunopathology on challenge with the SARS virus. PLoS ONE 7, e35421 (2012).
Deming, D. et al. Vaccine efficacy in senescent mice challenged with recombinant SARS-CoV bearing epidemic and zoonotic spike variants. PLoS Med. 3, e525 (2006).
Bolles, M. et al. A double-inactivated severe acute respiratory syndrome coronavirus vaccine provides incomplete protection in mice and induces increased eosinophilic proinflammatory pulmonary response upon challenge. J. Virol. 85, 12201–12215 (2011).
Yasui, F. et al. Prior immunization with severe acute respiratory syndrome (SARS)-associated coronavirus (SARS-CoV) nucleocapsid protein causes severe pneumonia in mice infected with SARS-CoV. J. Immunol. 181, 6337–6348 (2008).
Luo, F. et al. Evaluation of antibody-dependent enhancement of SARS-CoV infection in rhesus macaques immunized with an inactivated SARS-CoV vaccine. Virol. Sin. 33, 201–204 (2018).
Qin, E. et al. Immunogenicity and protective efficacy in monkeys of purified inactivated Vero-cell SARS vaccine. Vaccine 24, 1028–1034 (2006).
Graham, B. S. Rapid COVID-19 vaccine development. Science 368, 945–946 (2020).
Srinivasan, S. et al. Structural genomics of SARS-CoV-2 indicates evolutionary conserved functional regions of viral proteins. Viruses 12, 360 (2020).
Yuan, Y. et al. Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat. Commun. 8, 15092 (2017).
Chen, Q. et al. A report of clinical diagnosis and treatment of nine cases of coronavirus disease 2019. J. Med. Virol. 92, 683–687 (2020).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Chi, X. et al. A neutralizing human antibody binds to the N-terminal domain of the Spike protein of SARS-CoV-2. Science 369, 650–655 (2020).
Lu, G., Wang, Q. & Gao, G. F. Bat-to-human: spike features determining ‘host jump’ of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends Microbiol. 23, 468–478 (2015).
Li, F. Receptor recognition mechanisms of coronaviruses: a decade of structural studies. J. Virol. 89, 1954–1964 (2015).
Cai, Y. et al. Distinct conformational states of SARS-CoV-2 spike protein. Science 369, 1586–1592 (2020).
Gao, G. F. in Combating the Threat of Pandemic Influenza: Drug Discovery Approaches (eds Torrence, P. F.) 226–246 (John Wiley & Sons, 2007).
Premkumar, L. et al. The receptor binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci. Immunol. 5, eabc8413 (2020).
Wan, J. et al. Human-IgG-neutralizing monoclonal antibodies block the SARS-CoV-2 infection. Cell Rep. 32, 107918 (2020).
Yuan, M., Liu, H., Wu, N. C. & Wilson, I. A. Recognition of the SARS-CoV-2 receptor binding domain by neutralizing antibodies. Biochem. Biophys. Res. Commun. https://doi.org/10.1016/j.bbrc.2020.10.012 (2020).
Robbiani, D. F. et al. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature 584, 437–442 (2020).
Ju, B. et al. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature 584, 115–119 (2020).
Shi, R. et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 584, 120–124 (2020).
Du, S. et al. Structurally resolved SARS-CoV-2 antibody shows high efficacy in severely infected hamsters and provides a potent cocktail pairing strategy. Cell 183, 1013–1023 (2020).
Zost, S. J. et al. Potently neutralizing and protective human antibodies against SARS-CoV-2. Nature 584, 443–449 (2020).
Wu, Y. et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 368, 1274–1278 (2020).
Brouwer, P. J. M. et al. Potent neutralizing antibodies from COVID-19 patients define multiple targets of vulnerability. Science 369, 643–650 (2020).
Wec, A. Z. et al. Broad neutralization of SARS-related viruses by human monoclonal antibodies. Science 369, 731–736 (2020).
Rogers, T. F. et al. Isolation of potent SARS-CoV-2 neutralizing antibodies and protection from disease in a small animal model. Science 369, 956–963 (2020).
Hansen, J. et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 369, 1010–1014 (2020).
Guo, Z. D. et al. Aerosol and surface distribution of severe acute respiratory syndrome coronavirus 2 in hospital wards, Wuhan, China, 2020. Emerg. Infect. Dis. 26, 1583–1591 (2020).
Barnes, C. O. et al. Structures of human antibodies bound to SARS-CoV-2 spike reveal common epitopes and recurrent features of antibodies. Cell 182, 828–842 (2020).
Cao, Y. et al. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent patients’ B cells. Cell 182, 73–84 (2020).
Liu, L. et al. Potent neutralizing antibodies against multiple epitopes on SARS-CoV-2 spike. Nature 584, 450–456 (2020).
Jiang, S., Zhang, X. & Du, L. Therapeutic antibodies and fusion inhibitors targeting the spike protein of SARS-CoV-2. Expert Opin. Ther. Targets https://doi.org/10.1080/14728222.2020.1820482 (2020).
Zhu, J. et al. Following the rule: formation of the 6-helix bundle of the fusion core from severe acute respiratory syndrome coronavirus spike protein and identification of potent peptide inhibitors. Biochem. Biophys. Res. Commun. 319, 283–288 (2004).
Sun, H. et al. Structural basis of HCoV-19 fusion core and an effective inhibition peptide against virus entry. Emerg. Microbes Infect. 9, 1238–1241 (2020).
Liu, W. J. et al. T-cell immunity of SARS-CoV: Implications for vaccine development against MERS-CoV. Antivir. Res. 137, 82–92 (2017).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501 (2020).
Zhu, F. C. et al. Immunogenicity and safety of a recombinant adenovirus type-5-vectored COVID-19 vaccine in healthy adults aged 18 years or older: a randomised, double-blind, placebo-controlled, phase 2 trial. Lancet 396, 479–488 (2020).
Zhu, F. C. et al. Safety, tolerability, and immunogenicity of a recombinant adenovirus type-5 vectored COVID-19 vaccine: a dose-escalation, open-label, non-randomised, first-in-human trial. Lancet 395, 1845–1854 (2020).
van Doremalen, N. et al. ChAdOx1 nCoV-19 vaccine prevents SARS-CoV-2 pneumonia in rhesus macaques. Nature 586, 578–582 (2020).
Folegatti, P. M. et al. Safety and immunogenicity of the ChAdOx1 nCoV-19 vaccine against SARS-CoV-2: a preliminary report of a phase 1/2, single-blind, randomised controlled trial. Lancet 396, 467–478 (2020).
Logunov, D. Y. et al. Safety and immunogenicity of an rAd26 and rAd5 vector-based heterologous prime-boost COVID-19 vaccine in two formulations: two open, non-randomised phase 1/2 studies from Russia. Lancet 396, 887–897 (2020).
Smith, T. R. F. et al. Immunogenicity of a DNA vaccine candidate for COVID-19. Nat. Commun. 11, 2601 (2020).
Ma, J. et al. Cryo-EM structure of S-Trimer, a subunit vaccine candidate for COVID-19. Preprint at bioRxiv https://doi.org/10.1101/2020.09.21.306357 (2020).
Sanders, R. W. & Moore, J. P. Native-like Env trimers as a platform for HIV-1 vaccine design. Immunol. Rev. 275, 161–182 (2017).
Liang, J. G. et al. S-Trimer, a COVID-19 subunit vaccine candidate, induces protective immunity in nonhuman primates. Preprint at bioRxiv https://doi.org/10.1101/2020.09.24.311027 (2020).
Pallesen, J. et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen. Proc. Natl Acad. Sci. USA 114, E7348–E7357 (2017).
Corbett, K. S. et al. Evaluation of the mRNA-1273 vaccine against SARS-CoV-2 in nonhuman primates. N. Engl. J. Med. 383, 1544–1555 (2020).
Corbett, K. S. et al. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness. Nature 586, 567–571 (2020).
Jackson, L. A. et al. An mRNA vaccine against SARS-CoV-2 — preliminary report. N. Engl. J. Med. 383, 1920–1931 (2020).
Walsh, E. E. et al. RNA-based COVID-19 vaccine BNT162b2 selected for a pivotal efficacy study. Preprint at medRxiv https://doi.org/10.1101/2020.08.17.20176651 (2020).
Tian, J.-H. et al. SARS-CoV-2 spike glycoprotein vaccine candidate NVX-CoV2373 elicits immunogenicity in baboons and protection in mice. Preprint at bioRxiv https://doi.org/10.1101/2020.06.29.178509 (2020).
Keech, C. et al. Phase 1-2 trial of a SARS-CoV-2 recombinant spike protein nanoparticle vaccine. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2026920 (2020).
Tostanoski, L. H. et al. Ad26 vaccine protects against SARS-CoV-2 severe clinical disease in hamsters. Nat. Med. 26, 1694–1700 (2020).
Walsh, E. E. et al. Safety and immunogenicity of two RNA-based Covid-19 vaccine candidates. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2027906 (2020).
Mulligan, M. J. et al. Phase 1/2 study of COVID-19 RNA vaccine BNT162b1 in adults. Nature 586, 589–593 (2020).
Sahin, U. et al. COVID-19 vaccine BNT162b1 elicits human antibody and TH1 T-cell responses. Nature 586, 594–599 (2020).
Bangaru, S. et al. Structural analysis of full-length SARS-CoV-2 spike protein from an advanced vaccine candidate. Science 370, 1089–1094 (2020).
Bos, R. et al. Ad26 vector-based COVID-19 vaccine encoding a prefusion-stabilized SARS-CoV-2 Spike immunogen induces potent humoral and cellular immune responses. NPJ Vaccines 5, 91 (2020).
Pfizer. Pfizer and BioNTech announce vaccine candidate against COVID-19 achieved success in first interim analysis from phase 3 study. Pfizer https://www.pfizer.com/news/press-release/press-release-detail/pfizer-and-biontech-announce-vaccine-candidate-against (2020).
Moderna. Moderna’s COVID-19 vaccine candidate meets its primary efficacy endpoint in the first interim analysis of the phase 3 COVE study. Moderna https://investors.modernatx.com/node/10316/pdf (2020).
Mahase, E. Covid-19: UK approves Pfizer and BioNTech vaccine with rollout due to start next week. BMJ 371, m4714 (2020).
Hsieh, C. L. et al. Structure-based design of prefusion-stabilized SARS-CoV-2 spikes. Science 369, 1501–1505 (2020).
Lu, G. et al. Molecular basis of binding between novel human coronavirus MERS-CoV and its receptor CD26. Nature 500, 227–231 (2013).
Grant, O. C., Montgomery, D., Ito, K. & Woods, R. J. Analysis of the SARS-CoV-2 spike protein glycan shield reveals implications for immune recognition. Sci. Rep. 10, 14991 (2020).
Weiss, R. C. & Scott, F. W. Antibody-mediated enhancement of disease in feline infectious peritonitis: comparisons with dengue hemorrhagic fever. Comp. Immunol. Microbiol. Infect. Dis. 4, 175–189 (1981).
Su, S., Du, L. & Jiang, S. Learning from the past: development of safe and effective COVID-19 vaccines. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-020-00462-y (2020).
Zhang, N. N. et al. A Thermostable mRNA vaccine against COVID-19. Cell 182, 1271–1283 (2020).
Yang, J. et al. A vaccine targeting the RBD of the S protein of SARS-CoV-2 induces protective immunity. Nature 586, 572–577 (2020).
Zhou, M. et al. Screening and identification of severe acute respiratory syndrome-associated coronavirus-specific CTL epitopes. J. Immunol. 177, 2138–2145 (2006).
Jiang, S. et al. Roadmap to developing a recombinant coronavirus S protein receptor-binding domain vaccine for severe acute respiratory syndrome. Expert Rev. Vaccines 11, 1405–1413 (2012).
Wang, C. B. Analysis of low positive rate of nucleic acid detection method used for diagnosis of novel coronavirus pneumonia [In Chinese]. Zhonghua Yi Xue Za Zhi 100, E010 (2020).
Zhou, Y., Yang, Y., Huang, J., Jiang, S. & Du, L. Advances in MERS-CoV vaccines and therapeutics based on the receptor-binding domain. Viruses 11, 60 (2019).
Gu, H. et al. Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy. Science 369, 1603–1607 (2020).
Du, L. et al. Receptor-binding domain of SARS-CoV spike protein induces long-term protective immunity in an animal model. Vaccine 25, 2832–2838 (2007).
Du, L. et al. A truncated receptor-binding domain of MERS-CoV spike protein potently inhibits MERS-CoV infection and induces strong neutralizing antibody responses: implication for developing therapeutics and vaccines. PLoS ONE 8, e81587 (2013).
Ma, C. et al. Searching for an ideal vaccine candidate among different MERS coronavirus receptor-binding fragments–the importance of immunofocusing in subunit vaccine design. Vaccine 32, 6170–6176 (2014).
He, Y. et al. Receptor-binding domain of SARS-CoV spike protein induces highly potent neutralizing antibodies: implication for developing subunit vaccine. Biochem. Biophys. Res. Commun. 324, 773–781 (2004).
Wang, C. et al. Novel chimeric virus-like particles vaccine displaying MERS-CoV receptor-binding domain induce specific humoral and cellular immune response in mice. Antivir. Res. 140, 55–61 (2017).
Kim, Y. S. et al. Chaperna-mediated assembly of ferritin-based middle east respiratory syndrome-coronavirus nanoparticles. Front. Immunol. 9, 1093 (2018).
Walls, A. C. et al. Elicitation of potent neutralizing antibody responses by designed protein nanoparticle vaccines for SARS-CoV-2. Cell 183, 1367–1382 (2020).
Dai, L. et al. A universal design of betacoronavirus vaccines against COVID-19, MERS, and SARS. Cell 182, 722–733 (2020).
Hwang, W. C. et al. Structural basis of neutralization by a human anti-severe acute respiratory syndrome spike protein antibody, 80R. J. Biol. Chem. 281, 34610–34616 (2006).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Xiao, X., Feng, Y., Chakraborti, S. & Dimitrov, D. S. Oligomerization of the SARS-CoV S glycoprotein: dimerization of the N-terminus and trimerization of the ectodomain. Biochem. Biophys. Res. Commun. 322, 93–99 (2004).
Zhang, S. et al. Structural definition of a unique neutralization epitope on the receptor-binding domain of MERS-CoV spike glycoprotein. Cell Rep. 24, 441–452 (2018).
Bachmann, M. F. & Zinkernagel, R. M. Neutralizing antiviral B cell responses. Annu. Rev. Immunol. 15, 235–270 (1997).
Wang, N. et al. Structural definition of a neutralization-sensitive epitope on the MERS-CoV S1-NTD. Cell Rep. 28, 3395–3405 (2019).
Zhou, H. et al. Structural definition of a neutralization epitope on the N-terminal domain of MERS-CoV spike glycoprotein. Nat. Commun. 10, 3068 (2019).
Jiaming, L. et al. The recombinant N-terminal domain of spike proteins is a potential vaccine against Middle East respiratory syndrome coronavirus (MERS-CoV) infection. Vaccine 35, 10–18 (2017).
Huang, Y., Yang, C., Xu, X. F., Xu, W. & Liu, S. W. Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 41, 1141–1149 (2020).
Song, G. et al. Cross-reactive serum and memory B cell responses to spike protein in SARS-CoV-2 and endemic coronavirus infection. Preprint at bioRxiv https://doi.org/10.1101/2020.09.22.308965 (2020).
Ng, K. W. et al. Preexisting and de novo humoral immunity to SARS-CoV-2 in humans. Science https://doi.org/10.1126/science.abe1107 (2020).
Watanabe, Y., Allen, J. D., Wrapp, D., McLellan, J. S. & Crispin, M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science 369, 330–333 (2020).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292 (2020).
Ravichandran, S. et al. Antibody signature induced by SARS-CoV-2 spike protein immunogens in rabbits. Sci. Transl. Med. 12, eabc3539 (2020).
Jeyanathan, M. et al. Immunological considerations for COVID-19 vaccine strategies. Nat. Rev. Immunol. 20, 615–632 (2020).
Braun, J. et al. SARS-CoV-2-reactive T cells in healthy donors and patients with COVID-19. Nature 587, 270–274 (2020).
Du, L., He, Y., Jiang, S. & Zheng, B. J. Development of subunit vaccines against severe acute respiratory syndrome. Drugs Today 44, 63–73 (2008).
Zhao, J. et al. Airway memory CD4+ T cells mediate protective immunity against emerging respiratory coronaviruses. Immunity 44, 1379–1391 (2016).
Long, Q. X. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 26, 845–848 (2020).
Sariol, A. & Perlman, S. Lessons for COVID-19 immunity from other coronavirus infections. Immunity 53, 248–263 (2020).
Nakanaga, K., Yamanouchi, K. & Fujiwara, K. Protective effect of monoclonal antibodies on lethal mouse hepatitis virus infection in mice. J. Virol. 59, 168–171 (1986).
Lecomte, J. et al. Protection from mouse hepatitis virus type 3-induced acute disease by an anti-nucleoprotein monoclonal antibody. Brief report. Arch. Virol. 97, 123–130 (1987).
Liu, S. J. et al. Immunological characterizations of the nucleocapsid protein based SARS vaccine candidates. Vaccine 24, 3100–3108 (2006).
Collisson, E. W., Pei, J., Dzielawa, J. & Seo, S. H. Cytotoxic T lymphocytes are critical in the control of infectious bronchitis virus in poultry. Dev. Comp. Immunol. 24, 187–200 (2000).
Zhang, Y. et al. Safety, tolerability, and immunogenicity of an inactivated SARS-CoV-2 vaccine in healthy adults aged 18-59 years: a randomised, double-blind, placebo-controlled, phase 1/2 clinical trial. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(20)30843-4 (2020).
Xia, S. et al. Effect of an inactivated vaccine against SARS-CoV-2 on safety and immunogenicity outcomes: interim analysis of 2 randomized clinical trials. JAMA 324, 951–960 (2020).
Wang, K. et al. Longitudinal dynamics of the neutralizing antibody response to SARS-CoV-2 infection. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa1143 (2020).
Moore, J. P. & Klasse, P. J. COVID-19 Vaccines: “Warp Speed” needs mind melds, not warped minds. J. Virol. 94, e01083-20 (2020).
Hassan, A. O. et al. A single-dose intranasal ChAd vaccine protects upper and lower respiratory tracts against SARS-CoV-2. Cell 183, 169–184 (2020).
Xia, S. et al. Safety and immunogenicity of an inactivated SARS-CoV-2 vaccine, BBIBP-CorV: a randomised, double-blind, placebo-controlled, phase 1/2 trial. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(20)30831-8 (2020).
Che, Y. et al. Randomized, double-blinded and placebo-controlled phase II trial of an inactivated SARS-CoV-2 vaccine in healthy adults. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa1703 (2020).
Ramasamy, M. N. et al. Safety and immunogenicity of ChAdOx1 nCoV-19 vaccine administered in a prime-boost regimen in young and old adults (COV002): a single-blind, randomised, controlled, phase 2/3 trial. Lancet https://doi.org/10.1016/S0140-6736(20)32466-1 (2020).
Anderson, E. J. et al. Safety and immunogenicity of SARS-CoV-2 mRNA-1273 vaccine in older adults. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2028436 (2020).
Guebre-Xabier, M. et al. NVX-CoV2373 vaccine protects cynomolgus macaque upper and lower airways against SARS-CoV-2 challenge. Vaccine 38, 7892–7896 (2020).
Acknowledgements
The authors thank Q. Wang and K. Xu for discussions and insightful input during the preparation of this article. Work in the G.F.G. laboratory is supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (CAS) (XDB29010202). L.D. is supported by Youth Innovation Promotion Association of the CAS (2018113). The work is also partially supported by the Yanqi Lake Meeting organized by the Academic Divisions of the CAS.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
L.D. and G.F.G. are listed as inventors on patent applications for a MERS-CoV RBD-dimer vaccine and on pending patent applications for RBD-dimer-based CoV vaccines. The pending patents for RBD-dimers as protein subunit vaccines for MERS-CoV and SARS-CoV-2 have been licensed to Anhui Zhifei Longcom Biopharmaceutical Co. Ltd, China.
Additional information
Peer review information
Nature Reviews Immunology thanks J. Crowe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
COVID-19 Vaccine Tracker: https://vac-lshtm.shinyapps.io/ncov_vaccine_landscape/
WHO: https://www.who.int/
Rights and permissions
About this article
Cite this article
Dai, L., Gao, G.F. Viral targets for vaccines against COVID-19. Nat Rev Immunol 21, 73–82 (2021). https://doi.org/10.1038/s41577-020-00480-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41577-020-00480-0
This article is cited by
-
Targeting the receptor binding domain and heparan sulfate binding for antiviral drug development against SARS-CoV-2 variants
Scientific Reports (2024)
-
Enhancing antibody responses by multivalent antigen display on thymus-independent DNA origami scaffolds
Nature Communications (2024)
-
Oral administration of a recombinant modified RBD antigen of SARS-CoV-2 as a possible immunostimulant for the care of COVID-19
Microbial Cell Factories (2024)
-
Nanotechnology-based theranostic and prophylactic approaches against SARS-CoV-2
Immunologic Research (2024)
-
The receptor binding domain of SARS-CoV-2 Omicron subvariants targets Siglec-9 to decrease its immunogenicity by preventing macrophage phagocytosis
Nature Immunology (2024)