Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of coronavirus disease 2019 (COVID-19). Understanding of the fundamental processes underlying the versatile clinical manifestations of COVID-19 is incomplete without comprehension of how different immune cells are recruited to various compartments of virus-infected lungs, and how this recruitment differs among individuals with different levels of disease severity. As in other respiratory infections, leukocyte recruitment to the respiratory system in people with COVID-19 is orchestrated by specific leukocyte trafficking molecules, and when uncontrolled and excessive it results in various pathological complications, both in the lungs and in other organs. In the absence of experimental data from physiologically relevant animal models, our knowledge of the trafficking signals displayed by distinct vascular beds and epithelial cell layers in response to infection by SARS-CoV-2 is still incomplete. However, SARS-CoV-2 and influenza virus elicit partially conserved inflammatory responses in the different respiratory epithelial cells encountered early in infection and may trigger partially overlapping combinations of trafficking signals in nearby blood vessels. Here, we review the molecular signals orchestrating leukocyte trafficking to airway and lung compartments during primary pneumotropic influenza virus infections and discuss potential similarities to distinct courses of primary SARS-CoV-2 infections. We also discuss how an imbalance in vascular activation by leukocytes outside the airways and lungs may contribute to extrapulmonary inflammatory complications in subsets of patients with COVID-19. These multiple molecular pathways are potential targets for therapeutic interventions in patients with severe COVID-19.
Similar content being viewed by others
Introduction
The respiratory tract (nose, throat, larynx, trachea, bronchi and lungs) has a large surface area that is in direct contact with the outside environment. It comprises distinct epithelial cell layers and vascular beds, including the alveolar gas-exchange surfaces of the lung, which contain the largest vascular bed in the body1. Owing to constant exposure to the environment, there is a risk of acute lung infections, which can quickly develop into life-threatening diseases, especially when they lead to excessive leukocyte recruitment and activation1. The lung harbours different mucosal and alveolar compartments with distinct vascular beds (Figs 1,2). Because of the delicate air–blood barrier for gas exchange, resident immune cells in the lung maintain a very fine balance between protection and disease as they must constantly clear inhaled air particles and rapidly respond to pathogens through the highly coordinated recruitment of specific innate and adaptive leukocytes, which is critical for pathogen clearance.
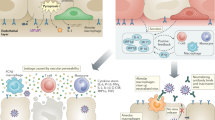
a | Shown are the major cell types (ciliated epithelial cells, goblet cells and basal cells) that form the epithelial layers of the nasal airway, lower airways and alveoli, which are targeted by respiratory viruses such as influenza virus and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). An adult human lung contains more than 3 × 108 alveoli, each covered by capillaries. The mediastinal lymph nodes are the major lung-draining lymph nodes. Some cervical lymph nodes and nasal-associated lymphoid tissue (NALT) play an additional role in priming antiviral adaptive immunity against viruses replicating in the upper airways. b | Events triggered by viral infection of the airway epithelial compartment (left) and of the alveolar epithelial compartment that constitute the lung parenchyma (right). The smooth muscle cell layers underneath the airway epithelial monolayers, neuroendocrine cells and the mucus layer on the apical aspects of these cells are omitted for clarity. Viral infection of both types of epithelial compartments triggers the release of inflammatory cytokines. These cytokines activate various epithelial-associated lymphocytes (such as innate lymphoid cells (ILCs) and resident memory T cells (TRM cells)), sentinel cells such as fibroblasts, interstitial macrophages and dendritic cells (DCs), and the pericytes associated with nearby blood vessels. In addition, some of the cytokines produced enter the circulation, reach the bone marrow and trigger generation and mobilization of innate leukocytes such as neutrophils, natural killer (NK) cells and monocytes, which are critical for viral clearance. The various blood vessels near the infected epithelial cells display arrays of leukocyte trafficking molecules (not shown), which are recognized by the circulating bone marrow-mobilized innate leukocytes. These various immune cells can then emigrate to the airway vessels and the capillaries of the virus-infected airway and alveolar compartments (Fig. 2). Virus-infected epithelial cells either die and release viral fragments or release live virus. Both fragments and particles are taken up and are processed by airway (respiratory) and alveolar DCs. On uptake and stimulation by viral particles, these different DCs leave their tissue, enter the lymphatic vessels and migrate into the draining lymph nodes (mediastinal lymph nodes). Naive T cells undergo priming by specific viral antigens in the T cell area of the lymph node, and antigen-activated CD4+ T cells then enter B cell follicles, where they provide critical signals to naive virus antigen-specific B cells, which subsequently become antibody-producing B cells (not shown). Within several days, T cells differentiate into effector T cells and egress from the lymph node, enter the blood vessels and migrate to the site of infection. These T cells are now equipped with specific receptors that allow them to emigrate through the inflamed airway blood vessels and/or the inflamed capillaries.
Leukocyte trafficking is mediated by specific adhesion molecules, chemokines and their respective receptors expressed by leukocytes. Three major vascular beds (postcapillary vessels in the respiratory tract, alveolar capillaries in the lung parenchyma and high endothelial venules (HEVs) in the draining lymph nodes) and their adjacent epithelial cells are indicated. Dashed arrows depict the low-level homeostatic emigration of circulating immune cells through the different types of blood vessel. The vessels are surrounded by basement membrane and pericytes. Other sentinel cells depicted in Fig. 1 are omitted. Top panel: in the resting state, endothelial cells of postcapillary vessels near the lower airways constitutively express integrin ligands (VCAM1 and ICAM2) and present homeostatic chemokines (CCL17, CCL20, CCL22 and CXCL16) in gradients across the vessels and epithelial cell layers. These trafficking signals mainly promote the low-level entry of leukocyte precursors (for example, precursor dendritic cells (preDCs)) and regulatory T cells (Treg cells). On viral infection, the inducible expression of trafficking molecules such as endothelial selectins (E-selectin and P-selectin), pro-inflammatory chemokines such as CCL2, CCL5, CXCL1, CXCL9 and CXCL10, and de novo transcribed ICAM1, together with VCAM1 and ICAM2, triggers massive emigration (thick arrows) of various innate immune cells (such as neutrophils and inflammatory monocytes) and natural killer (NK) cells from blood towards the infected airways. During later stages of infection, virus-specific CD8+ and CD4+ effector T cells (not shown) enter these infected compartments. Homeostatic chemokines (such as CXCL16) allow recruited CD8+ effector T cells to differentiate into epithelial-associated resident memory T cells (TRM cells). Uncontrolled viral spread or dysregulated epithelial and endothelial cell activation results in destructive leukocyte recruitment. During infection, resident innate lymphoid cells (ILCs) and TRM cells proliferate, and DCs carrying viral antigens either enter lymphatic vessels or stay in the inflamed tissue, where they present viral antigens to recruited effector T cells. Unresolved infection gives rise to inducible bronchus-associated lymphoid tissue, which recruits naive and effector T and B cells through HEVs (not shown). Middle panel: in the resting state, distinct integrin ligands, such as ICAM1, are constitutively expressed by resting endothelial cells that line the pulmonary capillaries in the lung parenchyma. Various homeostatic chemokines (such as CCL17, CCL22 and CXCL16) are secreted by nearby alveolar epithelial cells and endothelial cells that line the capillaries, and these chemokines promote the low-level entry of Treg cells and leukocyte precursors such as preDCs (not shown). In addition, patrolling monocytes crawl on all these endothelial cells using endothelial ICAM1 and remove dying endothelial cells. Alveolar macrophages are attached to the epithelial cells that constitute individual alveoli and capture airborne particles without eliciting inflammation. Just like in the respiratory tract, on viral infection, the emigration of multiple types of immune cell towards the infected alveolus is guided by enhanced expression of chemokines. These are produced primarily by individual infected epithelial cells and by virus-infected alveolar macrophages. Myeloid leukocytes may use their integrin macrophage 1 antigen (MAC1) to attach to and crawl along inflamed capillaries by as yet unidentified ligands. Bottom panel: constitutive expression of integrin ligands (not shown) on the endothelial cells lining specialized lymph node blood vessels called HEVs and the chemokines displayed by these cells, such as CCL19, CCL21, CXCL12 and CXCL13, allows the constant entry of recirculating naive T and B lymphocytes into lung-draining lymph nodes. On viral infection, in addition to naive lymphocytes, NK cells and specific myeloid leukocytes can emigrate from the HEVs, responding to signals from inflammatory chemokines such as CCL2, CXCL9 and CXCL10 displayed on these venules in an inducible manner in addition to the constitutively displayed chemokines CCL19, CXCL12 and CXCL13. For more details on individual trafficking molecules used by these different leukocytes to cross each of these vascular beds, please refer to Table 2. ADCs, alveolar DCs; LFA1, lymphocyte function-associated antigen 1; RDCs, respiratory DCs; VLA4, very late antigen 4. Adapted from ref.1, Springer Nature Limited.
The key targets for viral infections are the bronchial and alveolar epithelial cells, which are constantly exposed to air irritants and pathogens. Infected epithelial cells produce a number of inflammatory mediators, thereby orchestrating an immune response that involves multiple cell types critical for viral clearance and the establishment of antiviral immune memory. During infections with highly pathogenic viruses, such as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) or pneumotropic influenza viruses (for example, certain H1N1 and H5N1 influenza virus strains), a failure to rapidly clear these infections can lead to excessive uncontrolled inflammation, resulting in lung injury. Influenza viruses and SARS-CoV-2 use different molecular targets for entry and spread to the airways and lungs (Table 1; Box 1), and do not induce identical antiviral responses in the epithelial cells they infect2. Consequently, different influenza virus strains and SARS-CoV-2 evoke distinct inflammatory programmes during their spread in the airways and lungs. In particular, there is a difference with regard to the induction of type I and type III interferons by the epithelial cells that are initially infected. SARS-CoV-2 was reported to interfere more efficiently than influenza viruses with various cellular translation machineries implicated in antiviral immunity3,4,5. Nevertheless, given the conserved nature of antiviral responses, during the progression of SARS-CoV-2 infection, especially in patients with severe disease, most of the cytokines implicated in innate and adaptive leukocyte trafficking to sites of viral infection are likely similar to those elicited by influenza virus infections, and in particular to those induced in response to pneumotropic influenza virus strains that are able to replicate both in the lower airways and in the alveoli6,7 (Table 1). However, if SARS-CoV-2 is not efficiently cleared from the respiratory system, and in sharp contrast to influenza virus, it can spread to other organs (Table 1) using diverse mechanisms of extrapulmonary spread and resulting in a variety of clinical manifestations8.
In the absence of protective vaccines, therapeutic interventions should aim to optimize beneficial antiviral responses while restraining uncontrolled leukocyte trafficking, inflammation and injury associated with SARS-CoV-2 infections. Excessive pulmonary infiltration of monocyte-derived macrophages secreting enhanced levels of pro-inflammatory cytokines is a hallmark of severe pneumonia induced by SARS-CoV-2 (ref.9). How these leukocytes and other immune cells are recruited to various compartments of virus-infected airways and lungs at different stages of infection, which leukocyte trafficking molecules orchestrate this recruitment and how excessive accumulation of specific leukocyte subsets results in pathological complications in a subset of patients are still incompletely understood, as we lack key experimental data on leukocyte trafficking in SARS-CoV-2 infection in physiologically relevant animal models.
Here, we review our current knowledge of the molecular basis of immune cell migration and function in distinct airway and lung compartments in infections, with a specific focus on pathogenic influenza viruses, in an attempt to draw, wherever relevant, parallels to initial observations thus far reported in patients with severe coronavirus disease 2019 (COVID-19). We discuss how the intricate communication between influenza virus, epithelial cells, vascular cells, stromal cells, resident immune cells and circulating immune cells can regulate various leukocyte recruitment processes, and how parallel processes may occur during SARS-CoV-2 infections. We also highlight the involvement of specific vascular changes and the expression of leukocyte trafficking molecules in the thromboinflammatory processes associated with extrapulmonary SARS-CoV-2 complications. A discussion of classical studies on leukocyte recruitment to bacteria-infected or lipopolysaccharide-treated lungs is not included, as these are beyond the scope of this Perspective.
Trafficking signals on distinct vessels
Leukocyte trafficking from the blood to both the resting and pathogen-infected respiratory tract occurs through three major types of blood vessel: the postcapillary venules in the trachea and other blood vessels along the bronchial tree, the alveolar capillaries in the lung parenchyma, and the specialized cuboidal postcapillary venules within draining lymph nodes (LNs) termed ‘high endothelial venules’ (HEVs), which are the main portal for the entry of circulating naive and memory lymphocytes into LNs (Figs 1,2). These vessels are lined with endothelial cells, which sit at the interface between blood and tissues and contribute to haemostasis, immunity and inflammatory reactions10,11 (Fig. 1). Canonical inflammatory cytokines such as tumour necrosis factor (TNF), IL-1β and interferons, which are secreted by virus-infected epithelial and stromal cells, act directly on these various blood vessels and alveolar capillaries to induce the de novo expression of an array of leukocyte trafficking molecules on the surface of endothelial cells (Tables 1,2). These molecules function in a temporally regulated manner to recruit both innate and adaptive leukocytes to the infected airways and lungs. By contrast, endothelial cells of HEVs express distinct arrays of leukocyte trafficking molecules in a largely constitutive manner12.
To exit these different vessels, leukocytes that have penetrated the endothelial layers lining these vessels must cross the vascular basement membrane and a discontinuous pericyte layer (Fig. 2), and then follow chemotactic cues to reach their final destination (such as infected epithelial cells or different LN compartments). The basement membrane is a dense but discontinuous barrier composed of collagen IV, specialized laminin isoforms and interconnected proteoglycans13. Some of these glycoproteins are decorated with heparan sulfate moieties that serve as scaffolds on which chemokines, locally produced by inflamed blood vessels, pericytes and perivascular immune cells, become immobilized at high local concentrations14. As observed for other tissues, the basement membrane and the different interstitial tissues surrounding the distinct pulmonary vascular beds are likely remodelled during inflammation15. The proteolytic and actomyosin machineries used by individual leukocytes to cross these extravascular barriers under inflammatory settings are still poorly understood, especially in the context of viral lung infections.
Postcapillary venules
Tracheal and bronchial postcapillary venules and other blood vessels surround the bronchial tree and support leukocyte recruitment into the various airways following infection16,17. As very little is known about the transcriptional changes experienced by the endothelial cells that constitute these various blood vessels during viral infections, it is still unknown whether the main molecular changes induced in these vessels in response to pulmonary influenza virus and SARS-CoV-2 infections are similar. As in other tissues, in response to multiple inflammatory cues, these vessels are thought to mediate sequential multistep adhesive cascades initiated by leukocyte capture under shear flow, followed by rolling and firm arrest18. In the steady state, the endothelial cells of these various vessels express the cell adhesion molecules VCAM1 and ICAM2, which bind to the integrins very late antigen 4 (VLA4) and lymphocyte function-associated antigen 1 (LFA1), respectively, but they can undergo rapid transcriptional changes in response to distinct inflammatory cytokines to express additional cell adhesion molecules19 (Table 1). These include endothelial selectins and their glycoprotein ligands20, which mediate the initial capture and rolling of various blood-borne leukocytes on these vessels under shear flow (Table 2), and ICAM1, the high-affinity ligand for the integrins LFA1 and macrophage 1 antigen (MAC1)18, which together with VCAM1 mediate the arrest of rolling leukocyte on the inflamed vessels (Table 2). In response to different combinations of inflammatory cytokines (Table 1), the endothelial cells and their main perivascular counterparts, pericytes (Fig. 2), also express a large array of chemokines (for example, CXCL1, CXCL8, CCL2, CCL5 and the interferon-stimulated chemokines CXCL9, CXCL10 and CXC11), additional lipid mediators such as platelet-activating factor and leukotriene B4, and complement factors such as C5a21 (see Box 2). During infection, these endothelial cells and their associated pericytes, as well as virus-infected epithelial cells, can also produce the cytokines granulocyte colony-stimulating factor (G-CSF), granulocyte–macrophage colony-stimulating factor (GM-CSF) and IL-3, which act systemically and enhance myelopoiesis and the mobilization of newly generated myeloid leukocytes in the bone marrow22 (Fig. 1). These vessels can also rapidly mobilize Weibel–Palade bodies containing prestored von Willebrand factor, the chemokine CXCL8 (in humans) and the cell adhesion molecule P-selectin in response to various acute injury signals such as thrombin11,23. Another specialized property of postcapillary venules is the restricted expression of atypical chemokine receptor 1 (ACKR1; A. Rot, personal communication). On the basis of its functions in other tissues, this unique chemokine transcytosis receptor is likely to function in chemokine mobilization and presentation selectively in peribronchial postcapillary venules24,25,26, but its contribution to leukocyte recruitment to airways infected with influenza virus is still unknown. Other atypical chemoattractant receptors (such as the C5a receptor C5AR2 (ref.27)) may also contribute to leukocyte extravasation in influenza and other pulmonary infections.
Given that SARS-CoV-2 can initially attenuate antiviral type I and type III interferon production in infected epithelial cells4,5,28, the precise combinations of the multiple inflammatory endothelial stimuli triggered by this virus, as opposed to distinct pathogenic influenza virus strains, are likely different. Furthermore, the leukocyte trafficking signals and permeability changes induced in distinct blood vessels and postcapillary venules during different stages of viral infection and clearance (Box 3) are likely to differ between different respiratory compartments (for example, upper airways and lower airways) owing to differences in epithelial cell composition, relative infection levels and the types of cytokine produced (Fig. 1). In addition, the composition of perivascular and interstitial sentinel cells (such as pericytes, perivascular immune cells such as dendritic cells (DCs) and macrophages, fibroblasts, innate lymphoid cells (ILCs) and regulatory T cells (Treg cells)) in different virus-infected airway compartments and their individual responses to virus-triggered inflammatory cytokines may also affect the composition of leukocyte trafficking signals induced on the blood vessels near which they reside.
Alveolar capillaries
A second class of blood vessels that support leukocyte recruitment to the lung parenchyma are the alveolar capillaries29 (Fig. 2; Table 2). The endothelial lining of these alveolar capillaries is structurally distinct from that of postcapillary venules30. The transition from arterial to capillary endothelial cells is reflected in an elongation and flattening of the endothelial cell lining to improve gas exchange. Pulmonary capillaries also demonstrate lower permeability to solutes and fluids than the larger pulmonary blood vessels30. Alveolar capillaries have smaller diameters than leukocytes, a constraint which leads to a slow transit time31. These capillaries therefore contain a large number of marginated neutrophils, monocytes and naive and memory T cells that can rapidly cross the endothelium in response to chemotactic cues released within individual alveoli1. Most notably, leukocyte emigration through these narrow vessels and into the alveolar space occurs in the absence of selectin-mediated rolling29. Furthermore, the endothelial cells lining these vessels lack Weibel–Palade bodies; however, they express P-selectin and von Willebrand factor11 (Table 2).
Endothelial cells of alveolar capillaries express constitutively high levels of ICAM1 as well as ICAM2, a low-affinity ligand of LFA1. However, as opposed to postcapillary venules, they do not express VCAM1 or ACKR1 (refs19,32) (A. Rot, personal communication). The functions of these vascular ICAMs in leukocyte recruitment to influenza virus-infected alveoli are still unclear. Patrolling monocytes, a specialized subset of housekeeping monocytes that express constitutively activated LFA1, interact with vascular ICAM1 and ICAM2, which enable these cells to bind to damaged blood vessels in multiple organs33. It is therefore likely that these monocytes use ICAM1 and ICAM2 to patrol alveolar capillaries for signs of damage. The role of these ligands (and of other potential CD18 integrin ligands) in neutrophil emigration from inflamed capillaries, in general, and in influenza virus-infected mice, in particular, is still disputed34. Notably, ICAM1 is also constitutively expressed by alveolar type 1 pneumocytes19. However, the function of this pool of epithelial ICAM1 in guiding leukocytes to enter influenza virus-infected alveolar compartments remains unclear. Rather than promoting transepithelial leukocyte crossing, alveolar epithelial ICAM1 may facilitate cytotoxic T lymphocyte-mediated and natural killer (NK) cell-mediated killing of influenza virus-infected alveolar epithelial cells35.
High endothelial venules
HEVs are the third class of specialized blood vessels12 (Fig. 2). These serve as the main portal for naive T and B lymphocytes as well as central memory lymphocytes, recirculating neutrophils and ILCs36,37 to enter the draining LNs of the lung and specialized tertiary lymphoid organs along the bronchial tree38. The main draining LNs of the lungs are mediastinal LNs, which are positioned along the lower airways (Fig. 1). In addition, cervical LNs drain parts of the upper airways and are implicated in virus-specific T cell priming in mouse models of influenza virus and parainfluenza virus infections39 and potentially also in immune reactions to SARS-CoV-2. In addition to HEVs and their surrounding pericytes (Fig. 2), the two predominant stromal cell types in all draining LNs are lymphatic endothelial cells and fibroblastic reticular cells40. Lymphatic endothelial cells control lymphocyte egress and facilitate the entry of antigen-bearing DCs and soluble antigens into LNs. Fibroblastic reticular cells form a complex conduit system of microchannels and coordinate the entry and motility of lymphocytes through the LN41. Fibroblastic reticular cells also express homeostatic chemokines, including CXCL13, CCL19 and CCL21, as well as key integrin ligands, such as VCAM1 and ICAM1 (ref.42).
Another secondary lymphoid tissue in the respiratory system is the nasopharyngeal cavity (nasal)-associated lymphoid tissue. Prolonged infection in mice and humans induces the generation of additional bronchus-associated lymphoid tissue. The key leukocyte trafficking molecules expressed by the different HEVs within mediastinal LNs, nasal-associated lymphoid tissue and inducible bronchus-associated lymphoid tissue include a group of constitutively expressed glycoprotein ligands for L-selectin (also known as CD62L), which capture lymphocytes and subsets of monocytes and allow them to roll on the HEV surface in search of stop signals43,44. The LFA1 integrin ligands ICAM1 and ICAM2 are critical for lymphocyte arrest on all LN HEVs45 (Table 2). Notably, VCAM1 is particularly enriched on HEVs of human inducible bronchus-associated lymphoid tissues and may contribute to the recruitment of memory T and B cells to these lymphoid tissues during late stages of influenza virus infections46. Naive T and B lymphocytes require the chemokine receptor CCR7 to transmit critical intracellular signals that change the conformation of their LFA1 into a functional state in order to allow these cells to stop on HEVs and cross these vessels47. The main chemokine ligand for CCR7, CCL21, is displayed on both the luminal and the basolateral aspects of HEVs48. Under strong inflammatory conditions, HEVs can express or present additional adhesive and chemokine signals, which recruit subsets of regulatory NK cells and inflammatory monocytes49,50,51. During various infections, inflammatory monocytes are recruited to inflamed HEVs via specific CCR2-binding chemokines that are displayed by the endothelial cells of the HEVs50,51. As inflammatory monocytes express L-selectin and LFA1, but lack CCR7, they are normally restricted from homing to resting LNs. The CCR2-binding chemokines and additional inflammatory chemokines produced by various cells in the infected LNs (for example, CXCR3 ligands49) may be transported to HEVs through the LN conduits52. In addition to recruiting monocytes and NK cells, CXCR3 ligands displayed on inflamed HEVs can also promote the recruitment of plasmacytoid DCs with high CXCR3 expression (Table 2). Viral infections may also induce the expression of endothelial selectins (P-selectin and E-selectin) that function independently of L-selectin in recruiting these plasmacytoid DCs and NK cells to the inflamed HEVs49,53. So far, these molecular changes have been shown for HEVs within peripheral LNs but are likely shared with HEVs of lung-draining LNs.
Leukocytes recruited to infected lungs
The initial innate immune response to respiratory infections orchestrates the generation of adaptive immunity and facilitates the recruitment of various immune cell subsets. In the following subsections, we discuss the trafficking of these immune cells across distinct vascular beds and lymphatic vessels in influenza virus infections and, where known, in SARS-CoV-2 infections. We also briefly discuss the potential contributions of these leukocyte subsets to either productive or dysregulated immune responses, as well as to the resolution of viral infection-induced lung injury.
Neutrophil trafficking and vascular damage
Neutrophils, monocytes and NK cells are among the first innate immune cells to be recruited to the lung following viral infections. In influenza virus infections, neutrophils may promote clearance, or aggravate pathology, via a feedforward inflammatory circuit54,55. The cytokine G-CSF and the chemokine CXCL4, which are produced by multiple types of epithelial and endothelial cells of influenza virus-infected lungs, promote neutrophil mobilization from the bone marrow to the inflamed lung56,57. Several other neutrophil-recruiting chemokines are released by influenza virus-infected epithelial cells, alveolar macrophages and interstitial macrophages (Fig. 2), including CXCL1 (mice and humans), CXCL8 (also known as IL-8) (humans), CXCL2 and CCL3 (mice and humans)58. Neutrophil recruitment is also enhanced by platelets, which bind to and tether activated neutrophils to inflamed blood vessels59 (Fig. 3). Once they breach the endothelial lining of different blood vessels, neutrophils secrete matrix metallopeptidases and elastase, which cleave type IV collagen within the endothelial basement membranes of these various vessels60. In the interstitium, neutrophils are rapidly recruited to the bronchoalveolar space following chemokine gradients that are assumed to be generated across these various epithelial barriers during viral infections such as influenza or COVID-19 (ref.61).
a | Dysregulation of the renin–angiotensin–aldosterone system (RAAS) as a consequence of the downregulation of angiotensin-converting enzyme 2 (ACE2) by viral binding leads to decreased cleavage of angiotensin I and angiotensin II, resulting in elevated vasoconstriction and increased vascular permeability. Viral binding also results in endothelial cell damage, endothelial cell activation and thromboinflammation. b | Thromboinflammation in blood vessels is driven by the activation of endothelial cells and blood monocytes, which increase tissue factor membrane expression. Tissue factor is a key driver of thrombin generation. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)-infected or damaged endothelial cells, as well as endothelial cells stimulated by systemic or locally produced cytokines, upregulate the expression of adhesion molecules such as ICAM1 and of monocyte and neutrophil chemoattractants such as CXCL1 (not shown) and CCL2. The binding of thrombin to endothelial receptors (not shown) mobilizes vesicles containing prestored P-selectin and von Willebrand factor (vWF) (not shown), which, in turn, facilitate the recruitment of neutrophils and bind to and activate circulating platelets, respectively. The endothelial cell damage induced by the virus also exposes endothelial tissue factor, which further amplifies platelet deposition and thrombus formation. Moreover, monocytes and neutrophils are recruited to the damaged vessels by deposited platelets. Monocyte-derived tissue factor-rich microvesicles also activate the extrinsic coagulation pathway. Neutrophils recruited by the damaged endothelial cells and platelets can release neutrophil extracellular traps (NETs), which activate the intrinsic (contact activation) coagulation pathway, leading to massive fibrin deposition and blood clotting. Both neutrophils and monocytes express the integrin macrophage 1 antigen (MAC1), which allows them to bind to the damaged endothelial cells, activated platelets and deposited fibrin. COVID-19, coronavirus disease 2019. Adapted from ref.8 and ref.65, Springer Nature Limited.
Important viral clearance mechanisms of neutrophils include the production of pro-inflammatory cytokines, the phagocytosis of dying virus-infected epithelial cells and macrophages, and the release of granular content, including cationic antimicrobial peptides such as defensins and cathelicidins62. Neutrophils entrapped inside inflamed pulmonary vessels can produce neutrophil extracellular traps (NETs) in a process termed ‘NETosis’, and these are thought to capture viral particles and thereby restrict viral dissemination63. However, NETosis can also aggravate damage to the pulmonary endothelia and epithelia in severe influenza-induced pneumonia64. NETs also activate the extrinsic and intrinsic coagulation pathways65 (Fig. 3). Pro-NETotic states, reflected in elevated serum levels of myeloperoxidase (MPO)–DNA and of citrullinated histone H3, were recently reported in patients with severe COVID-19 (ref.66). In some settings, neutrophils can also secrete large amounts of CCL2, promoting monocyte recruitment67. Thus, elevated recruitment and activation of neutrophils can promote acute lung injury and acute respiratory distress syndrome, which are the main complications of severe influenza virus and SARS-CoV-2 infections68,69. These conditions are characterized by increased permeability of lung capillaries and the entry of solutes and fluids into the alveolar lumen70 (Box 3). In addition to increasing pulmonary oedema and facilitating leukocyte emigration, elevated vascular permeability may potentially facilitate SARS-CoV-2 dissemination to the circulation71.
Monocyte trafficking and pathologies
In humans, three circulating monocyte subsets have been classified on the basis of relative expression levels of CD14 and CD16 (also known as FcγRIII): classical (more than 80% of circulating monocytes), intermediate (about 15% of circulating monocytes) and non-classical (less than 5% of circulating monocytes) monocyte subsets72. The last subset, also termed ‘patrolling monocytes’33, is long-lived. Through the expression of their constitutively active LFA1, these cells bind to ICAM1 and ICAM2, which are constitutively expressed by the endothelial cells of different blood vessels, including capillaries. This binding allows patrolling monocytes to ‘crawl’ along these endothelial cells and detect signs of injury without emigrating to blood vessels (Table 2). By contrast, classical and intermediate monocytes are guided by endothelial selectins, multiple inflammatory chemokines and integrin ligands to emigrate to inflamed blood vessels within different lung compartments during different phases of influenza virus infection (Table 2). These recruited monocytes may also facilitate neutrophil extravasation across the inflamed alveolar capillaries73. Inflammatory monocytes that infiltrate the lung undergo differentiation into either monocyte-derived DCs or ‘exudate macrophages’, which release a large array of inflammatory chemokines (for example, CXCR3 ligands) as well as type I interferons74,75. A hallmark of severe influenza virus infections, which has also been reported in patients with severe COVID-19, is the CCR2-mediated exit (mobilization) of these various monocyte subsets from the bone marrow76. A fraction of these inflammatory monocytes replace the tissue-resident alveolar macrophages, which normally constitute 90% of the total lung macrophages1 and normally die during early phases of SARS-CoV-2 infection9. During influenza virus infection, alveolar macrophages and inflammatory monocytes contribute to the progressive clearance of the virus via Fc receptor-mediated antibody-derived cellular phagocytosis of infected cells65,77. However, pro-inflammatory monocyte-derived macrophages (M1 macrophages) can also contribute to pulmonary damage by secreting elevated levels of inducible nitric oxide synthase, TNF, IL-6 and matrix metallopeptidases65. Similar mechanisms may occur in patients with severe SARS-CoV-2 infection, as this is often associated with a cytokine storm78 and the excessive recruitment of IL-6-producing monocytes to infected lungs65. Systemic levels of IL-6 also appear to be directly correlated with the severity of COVID-19 (ref.2). There is also concern that macrophages may become infected with SARS-CoV-2 through antibody-dependent enhancement, a phenomenon where the uptake of virus is facilitated by non-neutralizing antibodies that bind to Fc receptors on macrophages79. However, the relevance of antibody-dependent enhancement for SARS-CoV-2 pathologies, especially during a primary infection, remains unclear80.
NK cell trafficking and functions
Circulating CD19−CD3−CD56+ NK cells are recruited to the inflamed lung early during influenza virus infection81. There is a well-documented division of labour within the NK cell compartment. CD56low NK cells express high levels of CD16 and killer cell immunoglobulin-like receptors82, conferring them with the ability to lyse infected cells via their activating receptors NKp44 and NKp46, which can directly bind to the glycoprotein haemagglutinin (HA) on the surface of influenza viruses83 (Box 1). CD56hi NK cells, which lack these receptors, perform regulatory functions by secreting cytokines and chemokines on stimulation by IL-18, which is secreted by epithelial cells, and IL-12, which is secreted by DCs84. The cytotoxic CD56low NK cell subtype and the regulatory CD56hi NK cell subtype express similar sets of chemokine receptors58 (Table 2). In addition, IL-15, produced by influenza virus-infected lung airways85, is chemotactic for most NK cells86. A potential direct contribution of regulatory NK cells to influenza virus-associated and SARS-CoV-2-associated acute respiratory distress syndrome is still unclear, but a subset of these regulatory NK cells produces CXCL1 and CXCL2 and enhances pulmonary neutrophil recruitment. Given that lung-recruited monocytes produce large amounts of the CXCR3-binding chemokines CXCL9 and CXCL10 (ref.9), NK cells responding to these chemokines can orchestrate additional monocyte and neutrophil accumulation, which can further worsen the lung injury87. Similarly to what is observed in patients infected with influenza A virus subtype H1N1, NK cell cytopenia in the blood, coupled with enhanced infiltration of NK cells into the inflamed bronchoalveolar space, appears to directly correlate with COVID-19 severity88. Although there are other similarities with influenza virus infections, the beneficial versus pathological contribution of different NK cell subsets during severe SARS-CoV-2 infections and the precise combinations of trafficking molecules used by these cells to accumulate in infected lungs await further elucidation.
Innate lymphocytes in lungs and LNs
ILCs, which include the NK-cell related ILC1, ILC2 and ILC3 subsets, are early producers of interferon-γ, IL-13 and IL-17, respectively, and have been implicated in various immune responses in lung infections and asthma89. In influenza virus infections, ILCs play versatile roles, including in viral immunosurveillance and repair of lung damage90. ILC1, ILC2 and ILC3 are primarily tissue resident both in lymphoid organs and in non-lymphoid organs, as they originate in situ from tissue progenitors91. At later stages of infection, however, haematogenous replacement of tissue ILCs has been shown to play an important role in ILC homeostasis91. For instance, the ILC1 and ILC2 subsets released from the bone marrow can migrate into inflamed lungs and also recirculate through resting and inflamed LNs91. The trafficking signals used by these different ILCs to enter influenza virus-infected lung and lung-draining LNs are still largely unknown (Table 2).
DC trafficking during influenza virus infection
The infected respiratory tract and its draining LNs are the main sites of initiation of T cell responses following the capture of viral antigens by respiratory DCs or by LN macrophages74. During influenza virus infections, and likely also during SARS-CoV-2 infection, airway and interstitial DCs play an essential role in mounting protective antiviral CD8+ T cell responses and in controlling the degree of inflammation93. The DC network in the respiratory tract is composed of several subsets that differ in phenotype, anatomical distribution and function (reviewed in refs92,93). The major resident DC subsets in the normal (non-inflamed) murine lung are respiratory conventional DCs (cDCs) and alveolar DCs. Virus-infected human and murine lungs are enriched with additional DC subsets derived from monocytes or from precursors of cDCs. Precursor DCs are recruited to the infected respiratory tract from the bone marrow in response to CCR2 chemokines released into the circulation (Fig. 1). Epithelial cells that are infected by influenza virus produce additional DC-activating cytokines such as IL-1β, GM-CSF and IL-3. By releasing various pathogen-associated molecular patterns and damage-associated molecular patterns, they facilitate DC differentiation and maturation92. The binding of natural or cross-reactive antibodies to influenza virus particles also leads to the formation of immune complexes that can drive DC maturation94. Whether such immune complexes also occur in the early stages of SARS-CoV-2 infection is questionable as cross-reactive antibodies are rare95.
The acquisition of influenza virus antigens by cDCs is achieved mainly by phagocytic engulfment of cell-free virions or of dying/dead infected epithelial cells that contain viral antigens96, and these highly conserved mechanisms are likely shared during SARS-CoV-2 infections. In mouse models, the clearance of influenza virus from the lung has been shown to depend on migratory langerin-positive CD103+CD11b− DCs, which need to upregulate CCR7 in order to egress from the bronchial tissues and enter draining LNs92,94. DCs may also use lymphatic vessel endothelial hyaluronic acid receptor 1 (LYVE1) to migrate through lymphatic vessels enriched with hyaluronan as shown in a mouse skin contact hypersensitivity model97. During early phases of influenza virus infection, migratory DCs are transported into the subcapsular sinus of draining LNs via afferent lymphatic vessels, gain access to the LN parenchyma and enter the T cell paracortex of the mediastinal LN58,98. Although DCs are critical for the priming of both CD8+ and CD4+ T cells in all known viral infections92,93, in the late stages of infections, cross-reactive memory B cells that had been generated in response to previous infections enter infected mediastinal LNs via HEVs and also participate in antigen presentation to CD4+ T cells39. Although studies are still lacking, it is very likely that in primary SARS-CoV-2 infections this pathway is limited as SARS-CoV-2 cross-reactive memory B cells are rare in naive individuals99.
T cell trafficking to sites of infection
The clearance of a primary infection with pneumotropic influenza viruses and likely of SARS-CoV-2 infection requires potent cytotoxic CD8+ T cell responses100,101. In addition, there appear to be contributions to viral clearance from subsets of CD4+ T cells (most clearly identified in patients with influenza) that acquire cytotoxic functions and are generated from naive and central memory virus-specific CD4+ T cell clones102,103. The various effector T cells generated in mediastinal and other respiratory LNs (Fig. 1) egress from these LNs through the efferent lymphatic vessels in response to signals from the sphingolipid S1P, which is constitutively expressed by these vessels104, and return to the blood. They accumulate in the airways and the lung parenchyma in response to combinations of distinct trafficking signals expressed by airway and alveolar blood vessel endothelial cells (Table 2). As these T cells downregulate their LN homing receptors (for example, CCR7 and L-selectin) and upregulate LFA1 and VLA4, and chemokine receptors recognized by ligands presented on the airway blood vessels and alveolar capillaries, these effector T cells can readily emigrate from these vessels rather than returning to lung-draining LNs (Fig.2; Table 2). Notably, influenza virus antigens can also be transferred from the lung to the spleen and bone marrow, where they are presented by various antigen-presenting cells and elicit CD4+ and CD8+ T cell priming and differentiation105. These effector T cells can also return to the circulation and home to vessels near the various infected airways. Whether SARS-CoV-2 antigens are similarly transferred to the spleen and bone marrow and elicit similar antivirus T cell priming is still unknown105. The trafficking properties of effector T cells generated in the spleen and the bone marrow, in particular their chemokine receptor repertoire, are, however, likely different from those generated in lung-draining LNs as the latter involve specific imprinting by lung DCs106.
As is the case for innate leukocytes, the entry of virus-specific effector T cells via capillaries into the alveolar space is unlikely to involve the classical multistep adhesive cascades that are typically initiated by selectin-mediated capture and rolling on inflamed postcapillary venules (Fig. 2). Rather, effector T cells bind to ICAM1 and ICAM2, which are abundantly and constitutively expressed by all alveolar capillaries and prolong the time during which these T cells are retained inside the capillaries107. Without these LFA1-dependent interactions, effector T cells are predicted to respond less efficiently to chemotactic cues displayed across the capillary linings of individual inflamed alveoli107 (Table 2). On the other hand, effector T cell entry to the bronchial lamina propria during viral infections, including influenza virus infections, likely involves E-selectin, P-selectin and VCAM1, a key VLA4 ligand expressed by both resting and inflamed postcapillary vessels19,32 (Table 2; Fig. 2). Although CD4+ and CD8+ effector T cells express the VCAM1-binding integrin VLA4 (ref.108) as well as multiple glycoprotein ligands for these selectins, including the P-selectin and E-selectin ligand PSGL1 (ref.109), it is still unclear when and how these different selectins function in capturing circulating effector T cells in the postcapillary venules around influenza virus-infected or SARS-CoV-2-infected bronchial airways. It is likely, however, that these T cells must bind to one or several of the canonical chemokine receptors, including CCR2, CCR4, CXCR3 and CCR5, as well as leukotriene B4 receptor 1 (BLT1), to successfully cross these vessels and enter the infected lamina propria106,110,111 (Table 2; Fig. 2). Subgroups of individuals carrying the CCR5Δ32 loss-of-function mutation suffered from increased lethality during the 2009 H1N1 influenza pandemic, implicating this chemokine receptor in beneficial lymphocyte migration and function in this infection. Whether this polymorphism is also a risk factor for patients with COVID-19 remains an open question. However, it has been reported that CCR5 blocking can reduce viral loads in critically ill patients with COVID-19 (ref.112). Circulating memory CD8+ T cells may use CCR5 also for recruitment into airways during secondary viral infections113. After crossing the vascular endothelial layers of these blood vessels and their basement membrane, and navigating through the collagen-rich interstitium guided by chemokines that bind to CXCR3, CXCR6 and CCR5 (ref.21), effector T cells either cross the proximal epithelial layer to reach the airway lumen or become trapped inside or below this layer114. IL-15 produced by influenza virus-infected airways is also involved in effector T cell recruitment115. A recent genome-wide association study on patients with severe COVID-19 identified single-nucleotide polymorphisms in CXCR6 that are associated with reduced expression of the key chemokine receptor CXCR6 (ref.116). Although preliminary, this study points to a potential role of CXCR6 in efficient effector T cell recruitment and protective function in SARS-CoV-2-infected airways during primary infections.
As acute viral lung infections are cleared, short-lived CD8+ effector T cells are replaced by CD127hi memory precursor T cells, which are capable of generating long-lived lung CD8+ resident memory T cells (TRM cells), primarily along the bronchial tree117. These cells are guided by the homeostatic bronchial epithelial cell-derived CXCR6 ligand CXCL16 (ref.114). Other long-lived memory cells can recirculate via lymphoid organs as central memory T cells or via other peripheral tissues as effector memory T cells. After influenza virus clearance, TRM cells enriched near the bronchial epithelia upregulate CD49a (also known as VLA1), an integrin that serves as a receptor for collagen IV, a key component of the epithelial basement membrane, and CD103, an integrin that binds to E-cadherin expressed by numerous airway epithelial cells. Moreover, these lymphocytes concomitantly downregulate LFA1 expression117. In addition, influenza virus-specific CD4+ effector T cells can differentiate into TRM cells that express elevated levels of LFA1 (ref.102), which may allow them to bind to nearby epithelial cells that constitutively express ICAM1, but it is still unclear whether these cells persist and have long-term protective properties. Notably, prior exposure to various influenza viruses has been shown to expand the pool of TRM cells to provide partial protection from heterosubtypic influenza virus strains103,117,118. Such tissue-resident SARS-CoV-2 cross-reactive CD8+ and CD4+ memory T cells might also exist in individuals previously exposed to seasonally circulating coronavirus strains119,120. The protective potential of such cross-reactive CD8+ and CD4+ T cells in primary SARS-CoV-2 infections, is, however, still unclear.
Leukocyte trafficking in lung repair
Lung recovery after viral infection has been studied in depth in mouse and ferret models of H1N1 influenza virus infection121. During infection, the collagenous assemblies in which both bronchioles and alveoli are embedded are extensively remodelled and take prolonged time to resume their original states122. The resolution of lung influenza virus infections is controlled by several key mechanisms and involves various resolving mediators, including lipoxins and protectins123. For instance, protectin D1 levels correlate inversely with influenza virus replication and immunopathology124. Peroxisome proliferator-activated receptor-γ, a transcription factor expressed on numerous immune cells and platelets and activated by various endogenous ligands, is another key resolution factor, primarily owing to its ability to downregulate nuclear factor-κB-mediated transcription125. The binding of prostaglandins to peroxisome proliferator-activated receptor-γ attenuates monocyte and neutrophil trafficking, dampens the transcription of inflammatory mediators and increases survival126. Another factor in the resolution of lungs following influenza virus infection is the atypical receptor ACKR2, which scavenges multiple inflammatory CC chemokines127. Deficiency in ACKR2 results in pulmonary dysfunction owing to accelerated recruitment of pathogenic lymphocytes to influenza virus-infected lungs128.
At later phases of influenza virus infections, subsets of CD8+ effector T cells become a major source of IL-10 near sites of epithelial infection, which dampens exaggerated innate and adaptive immune responses. These cells facilitate tissue repair along with macrophages, NK cells and ILCs, in particular ILC2 cells90. Soon after the initiation of resolution of inflammation, newly recruited Treg cells accumulate in various lung compartments, where they can attenuate inflammation via the production of transforming growth factor-β and IL-10 (ref.129). Treg cells recruited by blood vessels near terminal bronchioles also activate progenitor bronchioalveolar stem cells to differentiate into type 1 and type 2 pneumocytes130. Treg cells also attenuate fibrosis by inhibiting fibrocyte recruitment and proliferation, tightly regulated by the CXCR4–CXCL12 axis131. Treg cells express the growth factor amphiregulin, and Treg cell-specific amphiregulin deficiency impairs lung recovery in a murine model of influenza132. Amphiregulin is also produced by ILC2 cells, which are involved in recovery following influenza virus infection in mice90. As potent producers of IL-5, lung ILC2 cells may also regulate the infiltration of tissue-repairing eosinophil subsets at late stages of influenza virus infection90. Many other leukocyte subsets (NK cells, NKT cells, ILC3 cells, T helper 17 cells (TH17 cells), TH22 cells and γδ T cells) mitigate epithelial injury via IL-22 production133,134. Regulatory NKT cells also reduce the accumulation of inflammatory monocytes in influenza virus-infected lungs135. γδ T cells accumulate in the bronchoalveolar space only at late phases of influenza virus infection and are thought to contribute to lung recovery by multiple mechanisms in addition to IL-22 production136. The involvement of similar leukocyte subsets in lung repair processes during recovery from severe COVID-19 awaits confirmation by studies on patient samples and animal models. The trafficking signals that allow leukocytes involved in lung repair to migrate from the pulmonary circulation into different lung compartments also remain unknown.
Vascular pathologies beyond the lung
Most influenza viruses are restricted to the respiratory tract, owing to their requirement of a trypsin-like enzyme that is specifically expressed in the lung epithelium (Box 1). SARS-CoV-2 infectivity is considerably broader than that of influenza virus137 (Table 1). Arterial and venous endothelial cells express angiotensin-converting enzyme 2 (ACE2)138, the main entry receptor for SARS-CoV-2, and are therefore potential targets for infection. In addition, the SARS-CoV-2 entry receptor neuropilin 1 is expressed in a variety of endothelial cells and neurons across tissues139. The exceptionally broad infectivity of SARS-CoV-2 allows the virus, once disseminated into non-respiratory blood vessels, to infect other organs, as has been observed in patients with severe COVID-19 (refs8,140). In these and other subsets of patients, SARS-CoV-2 might also invade peripheral nerve terminals and gain access to the central nervous system141.
Another complication of severe SARS-CoV-2 infection is excessive coagulation and thromboinflammation8 (Fig. 3). This type of inflammation involves excessive thrombin production, the inhibition of fibrinolysis, activation of complement pathways, deposition of microthrombi and microvascular dysfunction138,142. In contrast to patients with fatal influenza virus infections, patients who die of SARS-CoV-2 infection often have widely disseminated blood clots143. Endothelial cells play essential roles in regulating vascular haemostasis, and resting endothelial cells prevent coagulation through the presentation of antithrombin III by heparin-like molecules, the synthesis of thrombomodulin and prostacyclin, and the maintenance of low levels of tissue factor144,145. The vascular pathology of COVID-19 might arise from direct ACE2 downregulation induced by viral entry into cells, resulting in a perturbed renin–angiotensin–aldosterone system146 and increased levels of the ACE2 substrate angiotensin II, a vasoconstrictor and profibrotic peptide8,147 (Fig. 3). The hypercoagulability seen in patients with severe COVID-19 appears to be worsened by platelet activation, which in turn contributes to additional neutrophil and monocyte recruitment and thromboinflammation65 (Fig. 3).
It is likely that blood vessels in the lung are the first to be affected by this SARS-CoV-2-mediated pathology. Once viral particles spread beyond the pulmonary vascular barriers, an impairment of the antihaemostatic protective properties of pulmonary vessels may result in pulmonary embolism, leading to a possibly life-threatening underoxygenation of the blood142. Infection-mediated endothelial cell injury in patients with COVID-19 is characterized also by elevated levels of von Willebrand factor and P-selectin and the presence of activated neutrophils and macrophages in multiple vascular beds, including the kidneys, heart, small intestine and liver8. Endothelial damage might also account for the high rates of cardiovascular complications in patients infected with SARS-CoV-2 (ref.8). These processes likely enhance the local stimulation of endothelial cells and induce the de novo transcription of E-selectin, ICAM1, VCAM1 and inflammatory chemokines that promote neutrophil and monocyte recruitment (Table 2). These events may result in a vicious cycle of vessel damage, inflammation, additional leukocyte recruitment and enhanced thromboinflammation (Fig. 3). The systemic stimulation and damages to the respiratory and non-respiratory vascular systems can be further amplified by a cytokine storm originating in the lungs. Lung endothelial cells are orchestrators of cytokine amplification during influenza virus infections and likely contribute also to the cytokine storm reported in severe SARS-CoV-2 infections148. The risk factors for these SARS-CoV-2 infection complications are pre-existing conditions such as hypertension (high blood pressure), high cholesterol levels, diabetes, obesity and heart diseases, which are all morbidities associated with general endothelial cell dysfunction and a high susceptibility to additional damage140,149. The propensity of such patients to develop serious complications can be attributed to the chronic inflammatory status of their vessels as well as to abnormal accumulation of inflammatory myeloid leukocytes in these extrapulmonary vessels8. It is therefore anticipated that systemic vascular and coagulation therapy might both attenuate the pathological leukocyte recruitment to extrapulmonary vessels and curb the destructive functions of these leukocytes in these vessels2. The inhibition of specific leukocyte adhesion receptors and chemoattractant receptors, particularly on neutrophils and monocytes (Fig. 3), could therefore prove therapeutically beneficial to both respiratory and non-respiratory manifestations of SARS-CoV-2 infection.
Conclusion
The COVID-19 outbreak represents the greatest worldwide infectious challenge in decades. Although featuring distinct mechanisms of infection and pathological outcomes, SARS-CoV-2 and influenza viruses likely share similar mechanisms of airway and lung inflammation and recovery5. When cellular stimulation, communication and effector functions are balanced, in both influenza virus and SARS-CoV-2 infections, viral clearance occurs with minimal damage to the infected airways and lungs. For SARS-CoV-2, a key outstanding question is when and how the lung and extrapulmonary blood vessels are targeted by SARS-CoV-2, and under what conditions the virus disseminates to other organs, either via the circulation or potentially via multiple neuronal routes141. In addition, the trafficking molecules used by leukocytes to interact with and emigrate from pulmonary blood vessels and to accumulate in extrapulmonary SARS-CoV-2-infected tissues, as well as the role of these leukocyte trafficking molecules in systemic vascular damage, deserve extensive research. Another open question is how the vascular complications of severe SARS-CoV-2 infections are affected by metabolic chronic inflammatory states and whether these states also affect infectivity. Obesity, for example, is a major co-morbidity factor in SARS-CoV-2 infection. This condition can affect numerous cell types in addition to vascular cells and can bias lung immunity to a type 17 response150. The involvement of a type 17 response in SARS-CoV-2-associated lung pathology remains an intriguing topic to investigate. Other pressing questions are the link between severe course of the disease and defective adaptive immunity (for example, in elderly patients151) or the significance of pre-existing SARS-CoV-2 cross-reactive memory T cells119. Complementary immunological, single-cell transcriptomic and histological analyses of blood and tissue samples from patients with COVID-19 and post-mortem analyses should shed more light on these important issues152. New transgenic mouse strains that allow close mimicking of the course of human SARS-CoV-2 infection and pathology might also help to dissect the involvement of specific leukocyte trafficking molecules in severe SARS-CoV-2 infections. Several known and newly developed immunomodulatory drugs (for example, dexamethasone153, coagulation inhibitors, neutralizing antibodies and antiviral drugs) are currently being tested in numerous clinical trials in patients with COVID-19. However, to date, almost no drugs that specifically target the trafficking of pathological leukocytes or their adhesive interactions with inflamed blood vessels or platelet plaques have been tested for their ability to improve clinical outcomes in patients with severe COVID-19. We propose that inhibitors of such specific leukocyte trafficking molecules (Table 2), especially integrin-blocking monoclonal antibodies and small-molecule chemokine receptor blockers, which are extensively characterized in different inflammatory models154, could be tested for the improvement of current treatments.
References
Holt, P. G., Strickland, D. H., Wikstrom, M. E. & Jahnsen, F. L. Regulation of immunological homeostasis in the respiratory tract. Nat. Rev. Immunol. 8, 142–152 (2008).
Tay, M. Z., Poh, C. M., Rénia, L., MacAry, P. A. & Ng, L. F. P. The trinity of COVID-19: immunity, inflammation and intervention. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-020-0311-8 (2020).
Lazear, H. M., Schoggins, J. W. & Diamond, M. S. Shared and distinct functions of type I and type III interferons. Immunity 50, 907–923 (2019).
Thoms, M. et al. Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2. Science https://doi.org/10.1126/science.abc8665 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045.e9 (2020).
Maines, T. R. et al. Pathogenesis of emerging avian influenza viruses in mammals and the host innate immune response. Immunol. Rev. 225, 68–84 (2008).
Tavares, L. P., Teixeira, M. M. & Garcia, C. C. The inflammatory response triggered by Influenza virus: a two edged sword. Inflamm. Res. 66, 283–302 (2017).
Gupta, A. et al. Extrapulmonary manifestations of COVID-19. Nat. Med. 26, 1017–1032 (2020).
Liao, M. et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat. Med. 26, 842–844 (2020).
Mantovani, A., Bussolino, F. & Dejana, E. Cytokine regulation of endothelial cell function. FASEB J. 6, 2591–2599 (1992).
Ochoa, C. D., Wu, S. & Stevens, T. New developments in lung endothelial heterogeneity: von Willebrand factor, P-selectin, and the Weibel-Palade body. Semin. Thrombosis Hemost. 36, 301–308 (2010).
Girard, J. P., Moussion, C. & Forster, R. HEVs, lymphatics and homeostatic immune cell trafficking in lymph nodes. Nat. Rev. Immunol. 12, 762–773 (2012).
Sorokin, L. The impact of the extracellular matrix on inflammation. Nat. Rev. Immunol. 10, 712–723 (2010).
Bao, X. et al. Endothelial heparan sulfate controls chemokine presentation in recruitment of lymphocytes and dendritic cells to lymph nodes. Immunity 33, 817–829 (2010).
Overstreet, M. G. et al. Inflammation-induced interstitial migration of effector CD4 T cells is dependent on integrin alpha. Nat Immunol. https://doi.org/10.1038/ni.2682 (2013).
Wanner, A. Circulation of the airway mucosa. J. Appl. Physiol. 67, 917–925 (1989).
Doerschuk, C. M. Leukocyte trafficking in alveoli and airway passages. Respir. Res. 1, 136–140 (2000).
Nourshargh, S. & Alon, R. Leukocyte migration into inflamed tissues. Immunity 41, 694–707 (2014).
Petrovich, E. et al. Lung ICAM-1 and ICAM-2 support spontaneous intravascular effector lymphocyte entrapment but are not required for neutrophil entrapment or emigration inside endotoxin inflamed lungs. FASEB J. 30, 1767–1778 (2016).
Müller, A. M., Cronen, C., Müller, K.-M. & Kirkpatrick, C. J. Heterogeneous expression of cell adhesion molecules by endothelial cells in ARDS. J. Pathol. 198, 270–275 (2002).
Griffith, J. W., Sokol, C. L. & Luster, A. D. Chemokines and chemokine receptors: positioning cells for host defense and immunity. Annu. Rev. Immunol. 32, 659–702 (2014).
Borriello, F. et al. Innate immune modulation by GM-CSF and IL-3 in health and disease. Int. J. Mol. Sci. 20, 834 (2019).
Pober, J. S. & Sessa, W. C. Evolving functions of endothelial cells in inflammation. Nat. Rev. Immunol. 7, 803–815 (2007).
Lee, J. S. et al. Duffy antigen facilitates movement of chemokine across the endothelium in vitro and promotes neutrophil transmigration in vitro and in vivo. J. Immunol. 170, 5244–5251 (2003).
Girbl, T. et al. Distinct compartmentalization of the chemokines CXCL1 and CXCL2 and the atypical receptor ACKR1 determine discrete stages of neutrophil diapedesis. Immunity 49, 1062–1076.e6 (2018).
Thiriot, A. et al. Differential DARC/ACKR1 expression distinguishes venular from non-venular endothelial cells in murine tissues. BMC Biol. 15, 45 (2017).
Miyabe, Y., Miyabe, C., Mani, V., Mempel, T. R. & Luster, A. D. Atypical complement receptor C5aR2 transports C5a to initiate neutrophil adhesion and inflammation. Sci. Immunol. 4, eaav5951 (2019).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science https://doi.org/10.1126/science.abc6027 (2020).
Doerschuk, C. M. Mechanisms of leukocyte sequestration in inflamed lungs. Microcirculation 8, 71–88 (2001).
Schneberger, D., Sethi, R. S. & Singh, B. in Molecular and Functional Insights Into the Pulmonary Vasculature. Advances in Anatomy Embryology and Cell Biology vol. 228 (ed. Parthasarathi, K.) 21–39 (Springer, 2018).
Doerschuk, C. M., Beyers, N., Coxson, H. O., Wiggs, B. & Hogg, J. C. Comparison of neutrophil and capillary diameters and their relation to neutrophil sequestration in the lung. J. Appl. Physiol. 74, 3040–3045 (1993).
Zarnegar, B. et al. Influenza infection in mice induces accumulation of lung mast cells through the recruitment and maturation of mast cell progenitors. Front. Immunol. 8, 1–12 (2017).
Carlin, L. M. et al. Nr4a1-dependent Ly6C(low) monocytes monitor endothelial cells and orchestrate their disposal. Cell 153, 362–375 (2013).
Reutershan, J. & Ley, K. Bench-to-bedside review: acute respiratory distress syndrome - how neutrophils migrate into the lung. Crit. Care 8, 453–461 (2004).
Dustin, M. L. & Long, E. O. Cytotoxic immunological synapses. Immunol. Rev. 235, 24–34 (2010).
Bogoslowski, A. et al. Neutrophils recirculate through lymph nodes to survey tissues for pathogens. J. Immunol. https://doi.org/10.4049/jimmunol.2000022 (2020).
Dutton, E. E. et al. Peripheral lymph nodes contain migratory and resident innate lymphoid cell populations. Sci. Immunol. 4, 31 (2019).
Foo, S. Y. & Phipps, S. Regulation of inducible BALT formation and contribution to immunity and pathology. Mucosal Immunol. 3, 537–544 (2010).
Topham, D. J., Tripp, R. A., Hamilton-Easton, A. M., Sarawar, S. R. & Doherty, P. C. Quantitative analysis of the influenza virus-specific CD4+ T cell memory in the absence of B cells and Ig. J. Immunol. 157, 2947–2952 (1996).
Krishnamurty, A. T. & Turley, S. J. Lymph node stromal cells: cartographers of the immune system. Nat. Immunol. 21, 369–380 (2020).
Bajenoff, M. et al. Stromal cell networks regulate lymphocyte entry, migration, and territoriality in lymph nodes. Immunity 25, 989–1001 (2006).
Onder, L. & Ludewig, B. A fresh view on lymph node organogenesis. Trends Immunol. 39, 775–787 (2018).
Campbell, J. J. et al. Chemokines and the arrest of lymphocytes rolling under flow conditions. Science 279, 381–384 (1998).
Luster, A. D., Alon, R. & von Andrian, U. H. Immune cell migration in inflammation: present and future therapeutic targets. Nat. Immunol. 6, 1182–1190 (2005).
Boscacci, R. T. et al. Comprehensive analysis of lymph node stroma-expressed Ig superfamily members reveals redundant and nonredundant roles for ICAM-1, ICAM-2, and VCAM-1 in lymphocyte homing. Blood 116, 915–925 (2010).
Hye, M. J., He, Y. & Sangster, M. Y. Broad dispersion and lung localization of virus-specific memory B cells induced by influenza pneumonia. Proc. Natl Acad. Sci. USA 105, 3485–3490 (2008).
Shamri, R. et al. Lymphocyte arrest requires instantaneous induction of an extended LFA-1 conformation mediated by endothelium-bound chemokines. Nat. Immunol. 6, 497–506 (2005).
Stein, J. V. et al. The CC chemokine thymus-derived chemotactic agent 4 (TCA-4, secondary lymphoid tissue chemokine, 6Ckine, exodus-2) triggers lymphocyte function-associated antigen 1-mediated arrest of rolling T lymphocytes in peripheral lymph node high endothelial venules. J. Exp. Med. 191, 61–76 (2000).
Martin-Fontecha, A. et al. Induced recruitment of NK cells to lymph nodes provides IFN-gamma for TH1 priming. Nat. Immunol. 5, 1260–1265 (2004).
Palframan, R. T. et al. Inflammatory chemokine transport and presentation in HEV: a remote control mechanism for monocyte recruitment to lymph nodes in inflamed tissues. J. Exp. Med. 194, 1361–1373 (2001).
Nakano, H. et al. Blood-derived inflammatory dendritic cells in lymph nodes stimulate acute T helper type 1 immune responses. Nat. Immunol. 10, 394–402 (2009).
Gretz, J. E., Norbury, C. C., Anderson, A. O., Proudfoot, A. E. I. & Shaw, S. Lymph-borne chemokines and other low molecular weight molecules reach high endothelial venules via specialized conduits while a functional barrier limits access to the lymphocyte microenvironments in lymph node cortex. J. Exp. Med. 192, 1425–1439 (2000).
Diacovo, T. G., Blasius, A. L., Mak, T. W., Cella, M. & Colonna, M. Adhesive mechanisms governing interferon-producing cell recruitment into lymph nodes. J. Exp. Med. 202, 687–696 (2005).
Fujisawa, H. Neutrophils play an essential role in cooperation with antibody in both protection against and recovery from pulmonary infection with influenza virus in mice. J. Virol. 82, 2772–2783 (2008).
Brandes, M., Klauschen, F., Kuchen, S. & Germain, R. N. A systems analysis identifies a feedforward inflammatory circuit leading to lethal influenza infection. Cell 154, 197–212 (2013).
Guo, L. et al. Critical role of CXCL4 in the lung pathogenesis of influenza (H1N1) respiratory infection. Mucosal Immunol. 10, 1529–1541 (2017).
Wang, H., Aloe, C., Wilson, N. & Bozinovski, S. G-CSFR antagonism reduces neutrophilic inflammation during pneumococcal and influenza respiratory infections without compromising clearance. Sci. Rep. 9, 1–12 (2019).
Grégoire, C. et al. The trafficking of natural killer cells. Immunol. Rev. 220, 169–182 (2007).
Pulavendran, S. et al. Combination therapy targeting platelet activation and virus replication protects mice against lethal influenza pneumonia. Am. J. Respir. Cell Mol. Biol. 61, 689–701 (2019).
Bradley, L. M., Douglass, M. F., Chatterjee, D., Akira, S. & Baaten, B. J. G. Matrix metalloprotease 9 mediates neutrophil migration into the airways in response to influenza virus-induced toll-like receptor signaling. PLoS Pathog. 8, e1002641 (2012).
Washburn, M. L., Crosby, R., Remlinger, K., Wang, F. & Creech, D. Therapeutically attenuating neutrophil recruitment with a CXCR2 antagonist in combination with oseltamivir ameliorates influenza-induced lung injury and disease. Open Forum Infect. Dis. https://doi.org/10.1093/ofid/ofz106c (2019).
Tate, M. D. et al. Neutrophils ameliorate lung injury and the development of severe disease during influenza infection. J. Immunol. 183, 7441–7450 (2009).
Jenne, C. N. et al. Neutrophils recruited to sites of infection protect from virus challenge by releasing neutrophil extracellular traps. Cell Host Microbe 13, 169–180 (2013).
Zhu, L. et al. High level of neutrophil extracellular traps correlates with poor prognosis of severe influenza A infection. J. Infect. Dis. 217, 428–437 (2018).
Merad, M. & Martin, J. C. Pathological inflammation in patients with COVID-19: a key role for monocytes and macrophages. Nat. Rev. Immunol. 20, 355–362 (2020).
Zuo, Y. et al. Neutrophil extracellular traps COVID-19. JCI Insight https://doi.org/10.1172/jci.insight.138999 (2020).
Soehnlein, O., Lindbom, L. & Weber, C. Mechanisms underlying neutrophil-mediated monocyte recruitment. Blood 114, 4613–4623 (2009).
Lai, C. et al. C-C motif chemokine ligand 2 (CCL2) mediates acute lung injury induced by lethal influenza H7N9 virus. Front. Microbiol. 8, 587 (2017).
Huang, C. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506 (2020).
Millar, F. R., Summers, C., Griffiths, M. J., Toshner, M. R. & Proudfoot, A. G. The pulmonary endothelium in acute respiratory distress syndrome: insights and therapeutic opportunities. Thorax 71, 462–473 (2016).
Polidoro, R. B., Hagan, R. S., de Santis Santiago, R. & Schmidt, N. W. Overview: systemic inflammatory response derived from lung injury caused by SARS-CoV-2 infection explains severe outcomes in COVID-19. Front. Immunol. 11, 1626 (2020).
Guilliams, M., Mildner, A. & Yona, S. Developmental and functional heterogeneity of monocytes. Immunity 49, 595–613 (2018).
Maus, U. A. et al. Monocytes are potent facilitators of alveolar neutrophil emigration during lung inflammation: role of the CCL2-CCR2 axis. J. Immunol. 170, 3273–3278 (2003).
Guilliams, M., Lambrecht, B. N. & Hammad, H. Division of labor between lung dendritic cells and macrophages in the defense against pulmonary infections. Mucosal Immunol. 6, 464–473 (2013).
Cao, W. et al. Rapid differentiation of monocytes into type I IFN-producing myeloid dendritic cells as an antiviral strategy against influenza virus infection. J. Immunol. 189, 2257–2265 (2012).
Lin, S. J. et al. The pathological effects of CCR2+ inflammatory monocytes are amplified by an IFNAR1-triggered chemokine feedback loop in highly pathogenic influenza infection. J. Biomed. Sci. 21, 99 (2014).
Tay, M. Z., Wiehe, K. & Pollara, J. Antibody dependent cellular phagocytosis in antiviral immune responses. Front. Immunol. 10, 332 (2019).
Moore, J. B. & June, C. H. Cytokine release syndrome in severe COVID-19. Science 368, 473–474 (2020).
Tetro, J. A. Is COVID-19 receiving ADE from other coronaviruses? Microbes Infect. 22, 72–73 (2020).
Bournazos, S., Gupta, A. & Ravetch, J. V. The role of IgG Fc receptors in antibody-dependent enhancement. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-020-00410-0 (2020).
Gazit, R. et al. Lethal influenza infection in the absence of the natural killer cell receptor gene Ncr1. Nat. Immunol. 7, 517–523 (2006).
Jost, S. & Altfeld, M. Control of human viral infections by natural killer cells. Annu. Rev. Immunol. 31, 163–194 (2013).
Arnon, T. I. et al. Recognition of viral hemagglutinins by NKp44 but not by NKp30. Eur. J. Immunol. 31, 2680–2689 (2001).
Cooper, M. A., Fehniger, T. A. & Caligiuri, M. A. The biology of human natural killer-cell subsets. Trends Immunol. 22, 633–640 (2001).
Verbist, K. C., Rose, D. L., Cole, C. J., Field, M. B. & Klonowski, K. D. IL-15 Participates in the respiratory innate immune response to influenza virus infection. PLoS ONE 7, e37539 (2012).
Allavena, P., Giardina, G., Bianchi, G. & Mantovani, A. IL-15 is chemotactic for natural killer cells and stimulates their adhesion to vascular endothelium. J. Leukoc. Biol. 61, 729–735 (1997).
Hoegl, S. et al. NK cells regulate CXCR2 + neutrophil recruitment during acute lung injury. J. Leukoc. Biol. 101, 471–480 (2017).
Giamarellos-Bourboulis, E. J. et al. Complex immune dysregulation in COVID-19 patients with severe respiratory failure. Cell Host Microbe https://doi.org/10.1016/j.chom.2020.04.009 (2020).
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015).
LA, M. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011).
Gasteiger, G., Fan, X., Dikiy, S., Lee, S. Y. & Rudensky, A. Y. Tissue residency of innate lymphoid cells in lymphoid and nonlymphoid organs. Science 350, 981–985 (2015).
Lambrecht, B. N. & Hammad, H. Lung dendritic cells in respiratory viral infection and asthma: from protection to immunopathology. Annu. Rev. Immunol. 30, 243–270 (2012).
Worbs, T., Hammerschmidt, S. I. & Förster, R. Dendritic cell migration in health and disease. Nat. Rev. Immunol. 17, 30–48 (2017).
Guilliams, M., Bruhns, P., Saeys, Y., Hammad, H. & Lambrecht, B. N. The function of Fcγ receptors in dendritic cells and macrophages. Nat. Rev. Immunol. 14, 94–108 (2014).
Huang, A. T. et al. A systematic review of antibody mediated immunity to coronaviruses: antibody kinetics, correlates of protection, and association of antibody responses with severity of disease. Preprint at medRxiv https://doi.org/10.1101/2020.04.14.20065771 (2020).
Helft, J. et al. Cross-presenting CD103+ dendritic cells are protected from influenza virus infection. J.Clin. Invest. 122, 4037–4047 (2012).
Johnson, L. A. et al. Dendritic cells enter lymph vessels by hyaluronan-mediated docking to the endothelial receptor LYVE-1. Nat. Immunol. 18, 762–770 (2017).
Legge, K. L. & Braciale, T. J. Accelerated migration of respiratory dendritic cells to the regional lymph nodes is limited to the early phase of pulmonary infection. Immunity 18, 265–277 (2003).
Zhang, Y. et al. Protective humoral immunity in SARS-CoV-2 infected pediatric patients. Cell. Mol. Immunol. 17, 768–770 (2020).
Topham, D. J., Tripp, R. A. & Doherty, P. C. CD8+ T cells clear influenza virus by perforin or Fas-dependent processes. J. Immunol. 159, 5197–5200 (1997).
Thevarajan, I. et al. Breadth of concomitant immune responses prior to patient recovery: a case report of non-severe COVID-19. Nat. Med. 26, 453–455 (2020).
Teijaro, J. R. et al. Cutting edge: tissue-retentive lung memory CD4 T cells mediate optimal protection to respiratory virus infection. J. Immunol. 187, 5510–5514 (2011).
Strutt, T. M. et al. Multipronged CD4 + T-cell effector and memory responses cooperate to provide potent immunity against respiratory virus. Immunol. Rev. 255, 149–164 (2013).
Shiow, L. R. et al. CD69 acts downstream of interferon-α/β to inhibit S1P 1 and lymphocyte egress from lymphoid organs. Nature 440, 540–544 (2006).
Wu, H. et al. Modeling of influenza-specific CD8+ T cells during the primary response indicates that the spleen is a major source of effectors. J. Immunol. 187, 4474–4482 (2011).
Mikhak, Z., Strassner, J. P. & Luster, A. D. Lung dendritic cells imprint T cell lung homing and promote lung immunity through the chemokine receptor CCR4. J. Exp. Med. 210, 1855–1869 (2013).
Galkina, E. et al. Preferential migration of effector CD8+ T cells into the interstitium of the normal lung. J. Clin. Invest. 115, 3473–3483 (2005).
Singbartl, K. et al. A CD2-green fluorescence protein-transgenic mouse reveals very late antigen-4-dependent CD8+ lymphocyte rolling in inflamed venules. J. Immunol. 166, 7520–7526 (2001).
Ley, K. & Kansas, G. S. Selectins in T-cell recruitment to non-lymphoid tissues and sites of inflammation. Nat. Rev. Immunol. 4, 325–335 (2004).
Tager, A. M. et al. Leukotriene B4 receptor BLT1 mediates early effector T cell recruitment. Nat. Immunol. 4, 982–990 (2003).
Dhume, K., Finn, C. M., Strutt, T. M., Sell, S. & McKinstry, K. K. T-bet optimizes CD4 T-cell responses against influenza through CXCR3-dependent lung trafficking but not functional programming. Mucosal Immunol. 12, 1220–1230 (2019).
Pattterson, B. et al. Disruption of the CCL5/RANTES-CCR5 pathway restores immune homeostasis and reduces plasma viral load in critical COVID-19. Preprint at medRxiv https://doi.org/10.1101/2020.05.02.20084673 (2020).
Kohlmeier, J. E. et al. The chemokine receptor CCR5 plays a key role in the early memory CD8+ T cell response to respiratory virus infections. Immunity 29, 101–113 (2008).
Wein, A. N. et al. CXCR6 regulates localization of tissue-resident memory CD8 T cells to the airways. J. Exp. Med. 216, 2748–2762 (2019).
Verbist, K. C., Cole, C. J., Field, M. B. & Klonowski, K. D. A role for IL-15 in the migration of effector CD8 T cells to the lung airways following influenza infection. J. Immunol. 186, 174–182 (2011).
Ellinghaus, D. et al. Genomewide association study of severe Covid-19 with respiratory failure. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2020283 (2020).
Ray, S. J. et al. The collagen binding alpha1beta1 integrin VLA-1 regulates CD8 T cell-mediated immune protection against heterologous influenza infection. Immunity 20, 167–179 (2004).
Chapman, T. J., Lambert, K. & Topham, D. J. Rapid reactivation of extralymphoid CD4 T cells during secondary infection. PLoS ONE 6, e20493 (2011).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell https://doi.org/10.1016/j.cell.2020.05.015 (2020).
Le Bert, N. et al. SARS-CoV-2-specific T cell immunity in cases of COVID-19 and SARS, and uninfected controls. Nature https://doi.org/10.1038/s41586-020-2550-z (2020).
Beers, M. F. & Morrisey, E. E. The three R’s of lung health and disease: repair, remodeling, and regeneration. J. Clin. Invest. 121, 2065–2073 (2011).
Talmi-Frank, D. et al. Extracellular matrix proteolysis by MT1-MMP contributes to influenza-related tissue damage and mortality. Cell Host Microbe 20, 458–470 (2016).
Russell, C. D. & Schwarze, J. The role of pro-resolution lipid mediators in infectious disease. Immunology 141, 166–173 (2014).
Morita, M. et al. The lipid mediator protectin D1 inhibits influenza virus replication and improves severe influenza. Cell 153, 112–125 (2013).
Croasdell, A. et al. PPAR γ and the innate immune system mediate the resolution of inflammation. PPAR Res. 2015, 1–20 (2015).
Cloutier, A. et al. The prostanoid 15-deoxy-Δ 12,14-prostaglandin-J2 reduces lung inflammation and protects mice against lethal influenza infection. J. Infect. Dis. 205, 621–630 (2012).
Jamieson, T. et al. The chemokine receptor D6 limits the inflammatory response in vivo. Nat. Immunol. 6, 403–411 (2005).
Tavares, L. P. et al. ACKR2 contributes to pulmonary dysfunction by shaping CCL5:CCR5-dependent recruitment of lymphocytes during influenza A infection in mice. Am. J. Physiol. Lung Cell. Mol. Physiol. 318, L655–L670 (2020).
Moser, E. K., Hufford, M. M. & Braciale, T. J. Late engagement of CD86 after influenza virus clearance promotes recovery in a FoxP3+ regulatory T cell dependent manner. PLoS Pathog. 10, e1004315 (2014).
D’Alessio, F. R. et al. CD4+CD25+Foxp3+ Tregs resolve experimental lung injury in mice and are present in humans with acute lung injury. J. Clin. Invest. 119, 2898–2913 (2009).
Garibaldi, B. T. et al. Regulatory T cells reduce acute lung injury fibroproliferation by decreasing fibrocyte recruitment. Am. J. Respir. Cell Mol. Biol. 48, 35–43 (2013).
Arpaia, N. et al. A distinct function of regulatory t cells in tissue protection. Cell 162, 1078–1089 (2015).
Kumar, P., Thakar, M. S., Ouyang, W. & Malarkannan, S. IL-22 from conventional NK cells is epithelial regenerative and inflammation protective during influenza infection. Mucosal Immunol. 6, 69–82 (2013).
Mcaleer, J. P. & Kolls, J. K. Directing traffic: IL-17 and IL-22 coordinate pulmonary immune defense. Immunol. Rev. 260, 129–144 (2014).
Kok, W. L. et al. Pivotal advance: invariant NKT cells reduce accumulation of inflammatory monocytes in the lungs and decrease immune-pathology during severe influenza A virus infection. J. Leukoc. Biol. 91, 357–368 (2012).
Nüssing, S. et al. Innate and adaptive T cells in influenza disease. Front. Med. 12, 34–47 (2018).
Puelles, V. G. et al. Multiorgan and renal tropism of SARS-CoV-2. N. Engl. J. Med. https://doi.org/10.1056/NEJMc2011400 (2020).
Varga, Z. et al. Endothelial cell infection and endotheliitis in COVID-19. Lancet 395, 1417–1418 (2020).
Cantuti-Castelvetri, L. et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and provides a possible pathway into the central nervous system. Preprint at bioRxiv https://doi.org/10.1101/2020.06.07.137802 (2020).
Ackermann, M. et al. Pulmonary vascular endothelialitis, thrombosis, and angiogenesis in Covid-19. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2015432 (2020).
Pezzini, A. & Padovani, A. Lifting the mask on neurological manifestations of COVID-19. Nat. Rev. Neurol. https://doi.org/10.1038/s41582-020-0398-3 (2020).
Bikdeli, B. et al. Pharmacological agents targeting thromboinflammation in COVID-19: review and implications for future research. Thrombosis Haemost. 120, 1004–1024 (2020).
Li, Q. et al. Early transmission dynamics in wuhan, china, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 382, 1199–1207 (2020).
Yau, J. W., Teoh, H. & Verma, S. Endothelial cell control of thrombosis. BMC Cardiovas. Disord. https://doi.org/10.1186/s12872-015-0124-z (2015).
Nawroth, P. & Kisiel W. S. D. The role of endothelium in the homeostatic balance of haemostasis. Clin. Haematol. 14, 531–546 (1985).
Mancia, G., Rea, F., Ludergnani, M., Apolone, G. & Corrao, G. Renin–angiotensin–aldosterone system blockers and the risk of covid-19. N. Engl. J. Med. https://doi.org/10.1056/nejmoa2006923 (2020).
Imai, Y., Kuba, K. & Penninger, J. M. Angiotensin-converting enzyme 2 in acute respiratory distress syndrome. Cell. Mol. Life Sci. 64, 2006–2012 (2007).
Teijaro, J. R. et al. Endothelial cells are central orchestrators of cytokine amplification during influenza virus infection. Cell 146, 980–991 (2011).
Ruan, Q., Yang, K., Wang, W., Jiang, L. & Song, J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 46, 846–848 (2020).
Kim, H. Y. et al. Interleukin-17-producing innate lymphoid cells and the NLRP3 inflammasome facilitate obesity-associated airway hyperreactivity. Nat. Med. 20, 54–61 (2014).
Moderbacher, C. R. et al. Antigen-specific adaptive immunity to SARS-CoV-2 in acute COVID-19 and associations with age and disease severity. Cell https://doi.org/10.1016/j.cell.2020.09.038 (2020).
Vabret, N. et al. Immunology of COVID-19: current state of the science. Immunity https://doi.org/10.1016/j.immuni.2020.05.002 (2020).
RECOVERY Collaborative Group Horby, P. et al. Dexamethasone in hospitalized patients with Covid-19 — preliminary report. N. Engl. J. Med. https://doi.org/10.1056/nejmoa2021436 (2020).
Schall, T. J. & Proudfoot, A. E. I. Overcoming hurdles in developing successful drugs targeting chemokine receptors. Nat. Rev. Immunol. 11, 355–363 (2011).
Karakus, U., Pohl, M. O. & Stertz, S. Breaking the convention: sialoglycan variants, coreceptors, and alternative receptors for influenza A virus entry. J. Virol. https://doi.org/10.1128/JVI.01357-19 (2019).
Chan, C. M. et al. Hemagglutinin of influenza A virus binds specifically to cell surface nucleolin and plays a role in virus internalization. Virology 494, 78–88 (2016).
Fujioka, Y. et al. A sialylated voltage-dependent Ca 2+ channel binds hemagglutinin and mediates influenza A virus entry into mammalian cells. Cell Host Microbe 23, 809–818.e5 (2018).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Ziegler, C. G. K. et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell https://doi.org/10.1016/j.cell.2020.04.035 (2020).
Heurich, A. et al. TMPRSS2 and ADAM17 Cleave ACE2 differentially and only proteolysis by TMPRSS2 augments entry driven by the severe acute respiratory syndrome coronavirus spike protein. J. Virol. 88, 1293–1307 (2014).
Taubenberger, J. K. & Morens, D. M. The pathology of influenza virus infections. Annu. Rev. Pathol. Mech. Dis. 3, 499–522 (2008).
Nicol, M. Q. & Dutia, B. M. The role of macrophages in influenza A virus infection. Future Virol. 9, 847–862 (2014).
Cline, T. D., Beck, D. & Bianchini, E. Influenza virus replication in macrophages: balancing protection and pathogenesis. J. Gen. Virol. 98, 2401–2412 (2017).
Mao, H. et al. Influenza virus directly infects human natural killer cells and induces cell apoptosis. J. Virol. 83, 9215–9222 (2009).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. https://doi.org/10.1038/s41591-020-0868-6 (2020).
Hamming, I. et al. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 203, 631–637 (2004).
Hikmet, F., Méar, L., Uhlén, M. & Lindskog, C. The protein expression profile of ACE2 in human tissues. https://doi.org/10.1101/2020.03.31.016048 (2020).
Chen, M. et al. Elevated ACE2 expression in the olfactory neuroepithelium: implications for anosmia and upper respiratory SARS-CoV-2 entry and replication. Preprint at bioRxiv https://doi.org/10.1101/2020.05.08.084996 (2020).
Chu, H. et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa410 (2020).
Rossignol, M., Gagnon, M. L. & Klagsbrun, M. Genomic organization of human neuropilin-1 and neuropilin-2 genes: Identification and distribution of splice variants and soluble isoforms. Genomics 70, 211–222 (2000).
Nogusa, S. et al. RIPK3 activates parallel pathways of MLKL-driven necroptosis and FADD-mediated apoptosis to protect against influenza a virus. Cell Host Microbe 20, 13–24 (2016).
Braun, J. et al. SARS-CoV-2-reactive T cells in healthy donors and patients with COVID-19. Nature https://doi.org/10.1038/s41586-020-2598-9 (2020).
Doherty, P. C. et al. Consequences of viral infections for lymphocyte compartmentalization and homeostasis. Semin. Immunol. 9, 365–373 (1997).
Diao, B. et al. Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). Front. Immunol. 11, 827 (2020).
Memoli, M. J. et al. Evaluation of antihemagglutinin and antineuraminidase antibodies as correlates of protection in an influenza A/H1N1 virus healthy human challenge model. mBio https://doi.org/10.1128/mBio.00417-16 (2016).
Guan, W. et al. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 382, 1708–1720 (2020).
Brochot, E. et al. Anti-spike anti-nucleocapsid and neutralizing antibodies in SARS-CoV-2 hospitalized patients and asymptomatic carriers. Preprint at medRxiv https://doi.org/10.1101/2020.05.12.20098236 (2020).
Kaneko, N. et al. Loss of Bcl-6-expressing t follicular helper cells and germinal centers in COVID-19. Cell https://doi.org/10.1016/j.cell.2020.08.025 (2020).
McGonagle, D., O’Donnell, J. S., Sharif, K., Emery, P. & Bridgewood, C. Immune mechanisms of pulmonary intravascular coagulopathy in COVID-19 pneumonia. Lancet Rheumatol. https://doi.org/10.1016/S2665-9913(20)30174-0 (2020).
Mao, L. et al. Neurologic manifestations of hospitalized patients with coronavirus disease 2019 in Wuhan, China. JAMA Neurol. 77, 683–690 (2020).
Ivanov, S. et al. Interleukin-22 reduces lung inflammation during influenza A virus infection and protects against secondary bacterial infection. J. Virol. 87, 6911–6924 (2013).
Tomankova, T., Kriegova, E. & Liu, M. Chemokine receptors and their therapeutic opportunities in diseased lung: far beyond leukocyte trafficking. Am. J. Physiol. Cell. Mol. Physiol. 308, L603–L618 (2015).
Paules, C. & Subbarao, K. Influenza. Lancet 390, 697–708 (2017).
Mansfield, K. G. Viral tropism and the pathogenesis of influenza in the mammalian host. Am. J. Pathol. 171, 1089–1092 (2007).
Stevens, J. et al. Glycan microarray analysis of the hemagglutinins from modern and pandemic influenza viruses reveals different receptor specificities. J. Mol. Biol. 355, 1143–1155 (2006).
Richard, M. et al. Influenza A viruses are transmitted via the air from the nasal respiratory epithelium of ferrets. Nat. Commun. 11, 1–11 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8 (2020).
Kandasamy, M. et al. Complement mediated signaling on pulmonary CD103+ dendritic cells is critical for their migratory function in response to influenza infection. PLoS Pathog. 9, e1003115 (2013).
Gralinski, L. E. et al. Complement activation contributes to severe acute respiratory syndrome coronavirus pathogenesis. mBio https://doi.org/10.1128/mBio.01753-18 (2018).
Risitano, A. M. et al. Complement as a target in COVID-19? Nat Rev. Immunol. https://doi.org/10.1038/s41577-020-0320-7 (2020).
Koupenova, M. et al. The role of platelets in mediating a response to human influenza infection. Nat. Commun. 10, 1–18 (2019).
Magro, C. et al. Complement associated microvascular injury and thrombosis in the pathogenesis of severe COVID-19 infection: a report of five cases. Transl. Res. https://doi.org/10.1016/j.trsl.2020.04.007 (2020).
Sun, S. et al. Treatment with anti-C5a antibody improves the outcome of H7N9 virus infection in African green monkeys. Clin. Infect. Dis. 60, 586–595 (2015).
Gao, T. et al. Highly pathogenic coronavirus N protein aggravates lung injury by MASP-2-mediated complement over-activation. Preprint at medRxiv https://doi.org/10.1101/2020.03.29.20041962 (2020).
Taveau, J. C. et al. Structure of artificial and natural VE-cadherin-based adherens junctions. Biochem. Soc. Trans. 36, 189–193 (2008).
Medford, A. R. L., Ibrahim, N. B. N. & Millar, A. B. Vascular endothelial growth factor receptor and coreceptor expression in human acute respiratory distress syndrome. J. Crit. Care 24, 236–242 (2009).
Schlegel, N. & Waschke, J. CAMP with other signaling cues converges on Rac1 to stabilize the endothelial barrier - a signaling pathway compromised in inflammation. Cell Tissue Res. 355, 587–596 (2014).
Acknowledgements
The authors thank S. Jung, I. Sagi and S. Feigelson (Weizmann Institute of Science), A. Sapoznikov (Israel Institute for Biological Research, Ness-Ziona), A. Rot (Queen Mary University of London, UK) and A. Ariel (University of Haifa, Israel) for fruitful discussions. They also thank S. Schwarzbaum for editorial assistance. This work was supported by GIF I-1470-412.13/2018, ISF 791/17, funds from the Minerva Foundation, ERA-Net E-Rate-3, and the Ministry of Health of Israel (to R.A.). This project was also funded in part by federal funds from the US National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services under CEIRS contract no. HHSN272201400005C (to D.J.T.) and P01-AI102851 (to D.J.T., D. Fowell, principal investigator) as well as 1U01HL122700 (to G. Pryhuber of the LungMAP Biorepository at the University of Rochester) and training grant GM007356 to M.S. N.G.’s research is supported by the BMBF (COVIMMUNE consortium) and the German Research Foundation grants EXC 2151, 390873048, 369799452 and 272482170. A.Z.’s research is supported by German Research Foundation grants ZA428/12-1, SFB1009A05 ZA428/18-1 and ZA428/17-1 and by IZKF grant Za2/001/18.
Author information
Authors and Affiliations
Contributions
R.A. wrote the leukocyte trafficking sections of this Perspective and coordinated the writing of other sections and boxes. D.J.T., N.G. and A.Z. wrote specific sections and revised the Perspective. D.J.T. and R.A. supervised the writing of paragraphs contributed by M.S., S.K., A.K. and E.C.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Immunology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Alon, R., Sportiello, M., Kozlovski, S. et al. Leukocyte trafficking to the lungs and beyond: lessons from influenza for COVID-19. Nat Rev Immunol 21, 49–64 (2021). https://doi.org/10.1038/s41577-020-00470-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41577-020-00470-2
This article is cited by
-
Monocyte and macrophage function in respiratory viral infections
Animal Diseases (2023)
-
d-dimer as a biomarker for COVID-19-associated pulmonary thromboembolism: a narrative review from molecular pathways to the imaging findings
The Egyptian Journal of Bronchology (2023)
-
Innate immune evasion strategies of SARS-CoV-2
Nature Reviews Microbiology (2023)
-
Amino acid catabolite markers for early prognostication of pneumonia in patients with COVID-19
Nature Communications (2023)
-
New genetic and epigenetic insights into the chemokine system: the latest discoveries aiding progression toward precision medicine
Cellular & Molecular Immunology (2023)