Abstract
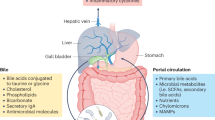
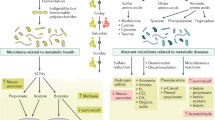
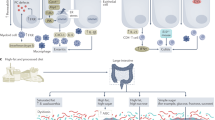
Low-grade inflammation is the hallmark of metabolic disorders such as obesity, type 2 diabetes and nonalcoholic fatty liver disease. Emerging evidence indicates that these disorders are characterized by alterations in the intestinal microbiota composition and its metabolites, which translocate from the gut across a disrupted intestinal barrier to affect various metabolic organs, such as the liver and adipose tissue, thereby contributing to metabolic inflammation. Here, we discuss some of the recently identified mechanisms that showcase the role of the intestinal microbiota and barrier dysfunction in metabolic inflammation. We propose a concept by which the gut microbiota fuels metabolic inflammation and dysregulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ridker, P. M., Cushman, M., Stampfer, M. J., Tracy, R. P. & Hennekens, C. H. Inflammation, aspirin, and the risk of cardiovascular disease in apparently healthy men. N. Engl. J. Med. 336, 973–979 (1997).
Ridker, P. M., Hennekens, C. H., Buring, J. E. & Rifai, N. C-reactive protein and other markers of inflammation in the prediction of cardiovascular disease in women. N. Engl. J. Med. 342, 836–843 (2000). In 28,263 healthy postmenopausal women, high-sensitivity C-reactive protein was the strongest predictor of the risk of cardiovascular events.
Pradhan, A. D., Manson, J. E., Rifai, N., Buring, J. E. & Ridker, P. M. C-reactive protein, interleukin 6, and risk of developing type 2 diabetes mellitus. JAMA 286, 327–334 (2001).
Hotamisligil, G. S. Inflammation, metaflammation and immunometabolic disorders. Nature 542, 177–185 (2017).
Donath, M. Y. & Shoelson, S. E. Type 2 diabetes as an inflammatory disease. Nat. Rev. Immunol. 11, 98–107 (2011).
Tilg, H. & Moschen, A. R. Insulin resistance, inflammation, and non-alcoholic fatty liver disease. Trends Endocrinol. Metab. 19, 371–379 (2008).
NCD Risk Factor Collaboration (NCD-RisC). Rising rural body-mass index is the main driver of the global obesity epidemic in adults. Nature 569, 260–264 (2019).
Pickup, J. C., Mattock, M. B., Chusney, G. D. & Burt, D. NIDDM as a disease of the innate immune system: association of acute-phase reactants and interleukin-6 with metabolic syndrome X. Diabetologia 40, 1286–1292 (1997).
Festa, A., D’Agostino, R., Jr, Tracy, R. P. & Haffner, S. M. Insulin Resistance Atherosclerosis Study. Elevated levels of acute-phase proteins and plasminogen activator inhibitor-1 predict the development of type 2 diabetes: the insulin resistance atherosclerosis study. Diabetes 51, 1131–1137 (2002).
Freeman, D. J. et al. C-reactive protein is an independent predictor of risk for the development of diabetes in the West of Scotland Coronary Prevention Study. Diabetes 51, 1596–1600 (2002).
Thorand, B. et al. C-reactive protein as a predictor for incident diabetes mellitus among middle-aged men: results from the MONICA Augsburg cohort study, 1984-1998. Arch. Intern. Med. 163, 93–99 (2003).
Tilg, H., Moschen, A. R. & Roden, M. NAFLD and diabetes mellitus. Nat. Rev. Gastroenterol. Hepatol. 14, 32–42 (2017).
Park, S. H. et al. Insulin resistance and C-reactive protein as independent risk factors for non-alcoholic fatty liver disease in non-obese Asian men. J. Gastroenterol. Hepatol. 19, 694–698 (2004).
Haukeland, J. W. et al. Systemic inflammation in nonalcoholic fatty liver disease is characterized by elevated levels of CCL2. J. Hepatol. 44, 1167–1174 (2006).
Yoneda, M. et al. High-sensitivity C-reactive protein is an independent clinical feature of nonalcoholic steatohepatitis (NASH) and also of the severity of fibrosis in NASH. J. Gastroenterol. 42, 573–582 (2007).
Chiang, C. H., Huang, C. C., Chan, W. L., Chen, J. W. & Leu, H. B. The severity of non-alcoholic fatty liver disease correlates with high sensitivity C-reactive protein value and is independently associated with increased cardiovascular risk in healthy population. Clin. Biochem. 43, 1399–1404 (2010).
Zimmermann, E. et al. C-reactive protein levels in relation to various features of non-alcoholic fatty liver disease among obese patients. J. Hepatol. 55, 660–665 (2011).
Pihlajamaki, J. et al. Serum interleukin 1 receptor antagonist as an independent marker of non-alcoholic steatohepatitis in humans. J. Hepatol. 56, 663–670 (2012).
Liang, H. et al. A low dose lipid infusion is sufficient to induce insulin resistance and a pro-inflammatory response in human subjects. PLOS ONE 13, e0195810 (2018).
Kubes, P. & Mehal, W. Z. Sterile inflammation in the liver. Gastroenterology 143, 1158–1172 (2012).
Ertunc, M. E. & Hotamisligil, G. S. Lipid signaling and lipotoxicity in metaflammation: indications for metabolic disease pathogenesis and treatment. J. Lipid Res. 57, 2099–2114 (2016).
Fu, S., Watkins, S. M. & Hotamisligil, G. S. The role of endoplasmic reticulum in hepatic lipid homeostasis and stress signaling. Cell Metab. 15, 623–634 (2012).
Hosogai, N. et al. Adipose tissue hypoxia in obesity and its impact on adipocytokine dysregulation. Diabetes 56, 901–911 (2007).
Pasarica, M. et al. Reduced adipose tissue oxygenation in human obesity: evidence for rarefaction, macrophage chemotaxis, and inflammation without an angiogenic response. Diabetes 58, 718–725 (2009).
Scherer, P. E., Williams, S., Fogliano, M., Baldini, G. & Lodish, H. F. A novel serum protein similar to C1q, produced exclusively in adipocytes. J. Biol. Chem. 270, 26746–26749 (1995). The first description and characterization of adiponectin, a key adipokine in humans.
Tilg, H. & Moschen, A. R. Adipocytokines: mediators linking adipose tissue, inflammation and immunity. Nat. Rev. Immunol. 6, 772–783 (2006).
Hotamisligil, G. S., Shargill, N. S. & Spiegelman, B. M. Adipose expression of tumor necrosis factor-alpha: direct role in obesity-linked insulin resistance. Science 259, 87–91 (1993). The first study to prove a key role for TNF in obesity and insulin resistance in rodent models.
Kern, P. A. et al. The expression of tumor necrosis factor in human adipose tissue. Regulation by obesity, weight loss, and relationship to lipoprotein lipase. J. Clin. Invest. 95, 2111–2119 (1995).
Fried, S. K., Bunkin, D. A. & Greenberg, A. S. Omental and subcutaneous adipose tissues of obese subjects release interleukin-6: depot difference and regulation by glucocorticoid. J. Clin. Endocrinol. Metab. 83, 847–850 (1998).
Moschen, A. R. et al. Anti-inflammatory effects of excessive weight loss: potent suppression of adipose interleukin 6 and tumour necrosis factor alpha expression. Gut 59, 1259–1264 (2010).
Mohamed-Ali, V. et al. Subcutaneous adipose tissue releases interleukin-6, but not tumor necrosis factor-alpha, in vivo. J. Clin. Endocrinol. Metab. 82, 4196–4200 (1997).
du Plessis, J. et al. Association of adipose tissue inflammation with histologic severity of nonalcoholic fatty liver disease. Gastroenterology 149, 635–648 (2015).
Akira, S. & Takeda, K. Toll-like receptor signalling. Nat. Rev. Immunol. 4, 499–511 (2004).
O’Neill, L. A., Golenbock, D. & Bowie, A. G. The history of Toll-like receptors — redefining innate immunity. Nat. Rev. Immunol. 13, 453–460 (2013).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Cani, P. D. Human gut microbiome: hopes, threats and promises. Gut 67, 1716–1725 (2018).
Schroeder, B. O. & Backhed, F. Signals from the gut microbiota to distant organs in physiology and disease. Nat. Med. 22, 1079–1089 (2016).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Levy, M., Kolodziejczyk, A. A., Thaiss, C. A. & Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 17, 219–232 (2017).
Devkota, S. et al. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nature 487, 104–108 (2012). This paper provides a mechanistic basis for how a Western-type diet might affect the prevalence of immune-mediated disorders such as inflammatory bowel disease.
Bischoff, S. C. et al. Intestinal permeability-a new target for disease prevention and therapy. BMC Gastroenterol. 14, 189 (2014).
Volynets, V. et al. Nutrition, intestinal permeability, and blood ethanol levels are altered in patients with nonalcoholic fatty liver disease (NAFLD). Dig. Dis. Sci. 57, 1932–1941 (2012).
Mooradian, A. D., Morley, J. E., Levine, A. S., Prigge, W. F. & Gebhard, R. L. Abnormal intestinal permeability to sugars in diabetes mellitus. Diabetologia 29, 221–224 (1986).
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Gilbert, J. A. et al. Current understanding of the human microbiome. Nat. Med. 24, 392–400 (2018).
Deschasaux, M. et al. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat. Med. 24, 1526–1531 (2018).
Backhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl Acad. Sci. U S A 101, 15718–15723 (2004).
Hooper, L. V. et al. Molecular analysis of commensal host-microbial relationships in the intestine. Science 291, 881–884 (2001).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006). This study provides evidence that the obese microbiome in rodents has an increased capacity to harvest energy from the diet and that this trait is transmissible.
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Aron-Wisnewsky, J. et al. Major microbiota dysbiosis in severe obesity: fate after bariatric surgery. Gut 68, 70–82 (2019).
Cotillard, A. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588 (2013).
Thaiss, C. A. et al. Persistent microbiome alterations modulate the rate of post-dieting weight regain. Nature 540, 544–551 (2016).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012). The first large human study from China demonstrating a gut microbiome signature in T2D.
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013). The first European study presenting evidence for compositional and functional changes in the metagenomes of women with T2D.
Loomba, R. et al. Gut microbiome-based metagenomic signature for non-invasive detection of advanced fibrosis in human nonalcoholic fatty liver disease. Cell Metab. 25, 1054–1062 (2017). The first large study in a NAFLD population demonstrating a gut microbiome signature, especially in cases associated with advanced fibrosis.
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143, 913–916 (2012).
Allin, K. H. et al. Aberrant intestinal microbiota in individuals with prediabetes. Diabetologia 61, 810–820 (2018).
Pedersen, H. K. et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535, 376–381 (2016).
Udayappan, S. D. et al. Intestinal Ralstonia pickettii augments glucose intolerance in obesity. PLOS ONE 12, e0181693 (2017).
Natividad, J. M. et al. Bilophila wadsworthia aggravates high fat diet induced metabolic dysfunctions in mice. Nat. Commun. 9, 2802 (2018).
Udayappan, S. et al. Oral treatment with Eubacterium hallii improves insulin sensitivity in db/db mice. NPJ Biofilms Microbiomes 2, 16009 (2016).
Zhao, L. et al. Gut bacteria selectively promoted by dietary fibers alleviate type 2 diabetes. Science 359, 1151–1156 (2018). A randomized clinical study demonstrating a beneficial effect of dietary fibres in subjects with T2D by modulating the gut microbiota.
Houghton, D. et al. Systematic review assessing the effectiveness of dietary intervention on gut microbiota in adults with type 2 diabetes. Diabetologia 61, 1700–1711 (2018).
Gentile, C. L. & Weir, T. L. The gut microbiota at the intersection of diet and human health. Science 362, 776–780 (2018).
Kolodziejczyk, A. A., Zheng, D., Shibolet, O. & Elinav, E. The role of the microbiome in NAFLD and NASH. EMBO Mol. Med. 11, e9302 (2018).
Boursier, J. et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology 63, 764–775 (2016).
Mouzaki, M. et al. Intestinal microbiota in patients with nonalcoholic fatty liver disease. Hepatology 58, 120–127 (2013).
Del Chierico, F. et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology 65, 451–464 (2017).
Tilg, H., Cani, P. D. & Mayer, E. A. Gut microbiome and liver diseases. Gut 65, 2035–2044 (2016).
Soderborg, T. K. et al. The gut microbiota in infants of obese mothers increases inflammation and susceptibility to NAFLD. Nat. Commun. 9, 4462 (2018).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64 (2014).
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 358, 359–365 (2017). This study provides evidence on how oral bacteria might colonize the intestinal tract and drive immune-mediated inflammatory disorders.
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012).
Fiordaliso, M. et al. Dietary oligofructose lowers triglycerides, phospholipids and cholesterol in serum and very low density lipoproteins of rats. Lipids 30, 163–167 (1995).
Li, Z. et al. Probiotics and antibodies to TNF inhibit inflammatory activity and improve nonalcoholic fatty liver disease. Hepatology 37, 343–350 (2003).
Zmora, N., Suez, J. & Elinav, E. You are what you eat: diet, health and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 16, 35–56 (2019).
Cani, P. D. et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56, 1761–1772 (2007). Landmark study demonstrating how endotoxins might affect metabolic disorders and associated inflammation.
Stenman, L. K., Holma, R. & Korpela, R. High-fat-induced intestinal permeability dysfunction associated with altered fecal bile acids. World J. Gastroenterol. 18, 923–929 (2012).
Jin, X., Yu, C. H., Lv, G. C. & Li, Y. M. Increased intestinal permeability in pathogenesis and progress of nonalcoholic steatohepatitis in rats. World J. Gastroenterol. 13, 1732–1736 (2007).
Pendyala, S., Walker, J. M. & Holt, P. R. A high-fat diet is associated with endotoxemia that originates from the gut. Gastroenterology 142, 1100–1101 (2012). This study shows that a high-fat diet or Western-style diet in healthy human subjects leads to endotoxaemia.
Teixeira, T. F. et al. Intestinal permeability parameters in obese patients are correlated with metabolic syndrome risk factors. Clin. Nutr. 31, 735–740 (2012).
Brignardello, J. et al. Pilot study: alterations of intestinal microbiota in obese humans are not associated with colonic inflammation or disturbances of barrier function. Aliment. Pharmacol. Ther. 32, 1307–1314 (2010).
Genser, L. et al. Increased jejunal permeability in human obesity is revealed by a lipid challenge and is linked to inflammation and type 2 diabetes. J. Pathol. 246, 217–230 (2018).
Laurans, L. et al. Genetic deficiency of indoleamine 2,3-dioxygenase promotes gut microbiota-mediated metabolic health. Nat. Med. 24, 1113–1120 (2018).
Luther, J. et al. Hepatic injury in nonalcoholic steatohepatitis contributes to altered intestinal permeability. Cell Mol. Gastroenterol. Hepatol. 1, 222–232 (2015).
Yuan, J. et al. Endotoxemia unrequired in the pathogenesis of pediatric nonalcoholic steatohepatitis. J. Gastroenterol. Hepatol. 29, 1292–1298 (2014).
Miele, L. et al. Increased intestinal permeability and tight junction alterations in nonalcoholic fatty liver disease. Hepatology 49, 1877–1887 (2009).
Strowski, M. Z. & Wiedenmann, B. Probiotic carbohydrates reduce intestinal permeability and inflammation in metabolic diseases. Gut 58, 1044–1045 (2009).
Giorgio, V. et al. Intestinal permeability is increased in children with non-alcoholic fatty liver disease, and correlates with liver disease severity. Dig. Liver Dis. 46, 556–560 (2014).
Damms-Machado, A. et al. Gut permeability is related to body weight, fatty liver disease, and insulin resistance in obese individuals undergoing weight reduction. Am. J. Clin. Nutr. 105, 127–135 (2017).
Peterson, L. W. & Artis, D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 14, 141–153 (2014).
Brown, E. M., Sadarangani, M. & Finlay, B. B. The role of the immune system in governing host-microbe interactions in the intestine. Nat. Immunol. 14, 660–667 (2013).
McDole, J. R. et al. Goblet cells deliver luminal antigen to CD103+ dendritic cells in the small intestine. Nature 483, 345–349 (2012).
Luck, H. et al. Regulation of obesity-related insulin resistance with gut anti-inflammatory agents. Cell Metab. 21, 527–542 (2015). This study shows that a gut-specific anti-inflammatory agent (5-aminosalicyclic acid) improves bowel inflammation and metabolic parameters by reducing intestinal permeability and endotoxaemia.
Garidou, L. et al. The gut microbiota regulates intestinal CD4 T cells expressing RORγt and controls metabolic disease. Cell Metab. 22, 100–112 (2015).
Johnson, A. M. et al. High fat diet causes depletion of intestinal eosinophils associated with intestinal permeability. PLOS ONE 10, e0122195 (2015).
Monteiro-Sepulveda, M. et al. Jejunal T cell inflammation in human obesity correlates with decreased enterocyte insulin signaling. Cell Metab. 22, 113–124 (2015).
Ma, T. Y. et al. TNF-α-induced increase in intestinal epithelial tight junction permeability requires NF-κB activation. Am. J. Physiol. Gastrointest. Liver Physiol. 286, G367–G376 (2004).
Winer, D. A., Luck, H., Tsai, S. & Winer, S. The intestinal immune system in obesity and insulin resistance. Cell Metab. 23, 413–426 (2016).
Martinez-Lopez, M. et al. Microbiota sensing by mincle-syk axis in dendritic cells regulates interleukin-17 and -22 production and promotes intestinal barrier integrity. Immunity 50, 446–461 (2019).
Sonnenberg, G. F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012).
Kruglov, A. A. et al. Nonredundant function of soluble LTα3 produced by innate lymphoid cells in intestinal homeostasis. Science 342, 1243–1246 (2013).
Stockinger, B. & Omenetti, S. The dichotomous nature of T helper 17 cells. Nat. Rev. Immunol. 17, 535–544 (2017).
Upadhyay, V. et al. Lymphotoxin regulates commensal responses to enable diet-induced obesity. Nat. Immunol. 13, 947–953 (2012).
Pamir, N., McMillen, T. S., Edgel, K. A., Kim, F. & LeBoeuf, R. C. Deficiency of lymphotoxin-α does not exacerbate high-fat diet-induced obesity but does enhance inflammation in mice. Am. J. Physiol. Endocrinol. Metab. 302, E961–E971 (2012).
Darnaud, M. et al. Enteric delivery of regenerating family member 3 alpha alters the intestinal microbiota and controls inflammation in mice with colitis. Gastroenterology 154, 1009–1023 (2018).
Fatkhullina, A. R. et al. An interleukin-23-interleukin-22 axis regulates intestinal microbial homeostasis to protect from diet-induced atherosclerosis. Immunity 49, 943–957 (2018). This paper shows that inactivation of the IL-23–IL-22 signalling pathway deteriorates atherosclerosis by affecting intestinal barrier function, dysbiosis and expansion of pathogenic bacteria.
Aden, K. et al. Epithelial IL-23R signaling licenses protective IL-22 responses in intestinal inflammation. Cell Rep. 16, 2208–2218 (2016).
Ngo, V. L. et al. A cytokine network involving IL-36γ, IL-23, and IL-22 promotes antimicrobial defense and recovery from intestinal barrier damage. Proc. Natl Acad. Sci. USA 115, E5076–E5085 (2018).
Lee, J. S. et al. Interleukin-23-independent IL-17 production regulates intestinal epithelial permeability. Immunity 43, 727–738 (2015).
Mohammed, N., Tang, L., Jahangiri, A., de Villiers, W. & Eckhardt, E. Elevated IgG levels against specific bacterial antigens in obese patients with diabetes and in mice with diet-induced obesity and glucose intolerance. Metabolism 61, 1211–1214 (2012).
Wang, X. et al. Interleukin-22 alleviates metabolic disorders and restores mucosal immunity in diabetes. Nature 514, 237–241 (2014).
Dalmas, E. et al. T cell-derived IL-22 amplifies IL-1β-driven inflammation in human adipose tissue: relevance to obesity and type 2 diabetes. Diabetes 63, 1966–1977 (2014).
Fabbrini, E. et al. Association between specific adipose tissue CD4+ T-cell populations and insulin resistance in obese individuals. Gastroenterology 145, 366–374 (2013).
Sumarac-Dumanovic, M. et al. Increased activity of interleukin-23/interleukin-17 proinflammatory axis in obese women. Int. J. Obes. 33, 151–156 (2009).
Harley, I. T. et al. IL-17 signaling accelerates the progression of nonalcoholic fatty liver disease in mice. Hepatology 59, 1830–1839 (2014).
Zuniga, L. A. et al. IL-17 regulates adipogenesis, glucose homeostasis, and obesity. J. Immunol. 185, 6947–6959 (2010).
Amar, J. et al. Intestinal mucosal adherence and translocation of commensal bacteria at the early onset of type 2 diabetes: molecular mechanisms and probiotic treatment. EMBO Mol. Med. 3, 559–572 (2011).
Everard, A. et al. Intestinal epithelial MyD88 is a sensor switching host metabolism towards obesity according to nutritional status. Nat. Commun. 5, 5648 (2014).
Vijay-Kumar, M. et al. Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 328, 228–231 (2010). In this study, it is shown that mice deficient in TLR5 develop hyperphagia and several features of metabolic syndrome, including obesity, hypertension, dyslipidaemia and insulin resistance.
Wen, H. et al. Fatty acid–induced NLRP3-ASC inflammasome activation interferes with insulin signaling. Nat. Immunol. 12, 408–415 (2011).
Stienstra, R. et al. Inflammasome is a central player in the induction of obesity and insulin resistance. Proc. Natl Acad. Sci. USA 108, 15324–15329 (2011).
Levy, M. et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell 163, 1428–1443 (2015).
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012).
Denou, E. et al. Defective NOD2 peptidoglycan sensing promotes diet-induced inflammation, dysbiosis, and insulin resistance. EMBO Mol. Med. 7, 259–274 (2015).
Ahuja, M. et al. Orai1-mediated antimicrobial secretion from pancreatic acini shapes the gut microbiome and regulates gut innate immunity. Cell Metab. 25, 635–646 (2017).
Ding, S. et al. High-fat diet: bacteria interactions promote intestinal inflammation which precedes and correlates with obesity and insulin resistance in mouse. PLOS ONE 5, e12191 (2010).
Ghoshal, S., Witta, J., Zhong, J., de Villiers, W. & Eckhardt, E. Chylomicrons promote intestinal absorption of lipopolysaccharides. J. Lipid Res. 50, 90–97 (2009).
Laugerette, F. et al. Emulsified lipids increase endotoxemia: possible role in early postprandial low-grade inflammation. J. Nutr. Biochem. 22, 53–59 (2011).
Thaiss, C. A. et al. Hyperglycemia drives intestinal barrier dysfunction and risk for enteric infection. Science 359, 1376–1383 (2018). This study demonstrates the relevance of hyperglycaemia in regulation of the intestinal barrier and associated systemic inflammation.
Sellmann, C. et al. Diets rich in fructose, fat or fructose and fat alter intestinal barrier function and lead to the development of nonalcoholic fatty liver disease over time. J. Nutr. Biochem. 26, 1183–1192 (2015).
Chassaing, B. et al. Dietary emulsifiers impact the mouse gut microbiota promoting colitis and metabolic syndrome. Nature 519, 92–96 (2015).
Elia, M., Goren, A., Behrens, R., Barber, R. W. & Neale, G. Effect of total starvation and very low calorie diets on intestinal permeability in man. Clin. Sci. 73, 205–210 (1987).
Suzuki, T. & Hara, H. Dietary fat and bile juice, but not obesity, are responsible for the increase in small intestinal permeability induced through the suppression of tight junction protein expression in LETO and OLETF rats. Nutr. Metab. 7, 19 (2010).
Tropini, C. et al. Transient osmotic perturbation causes long-term alteration to the gut microbiota. Cell 173, 1742–1754 (2018).
Cani, P. D. et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 57, 1470–1481 (2008). In this study, antibiotic therapy improved systemic metabolic dysfunction and deletion of Cd14 in ob/ob mice mimicked the effects achieved by antibiotics.
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353 (2016).
Schroeder, B. O. et al. Bifidobacteria or fiber protects against diet-induced microbiota-mediated colonic mucus deterioration. Cell Host Microbe 23, 27–40 (2018).
Li, D. et al. Amelioration of intestinal barrier dysfunction by berberine in the treatment of nonalcoholic fatty liver disease in rats. Pharmacogn. Mag. 13, 677–682 (2017).
Cani, P. D. et al. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut 58, 1091–1103 (2009).
Ghosh, S. S., Bie, J., Wang, J. & Ghosh, S. Oral supplementation with non-absorbable antibiotics or curcumin attenuates western diet-induced atherosclerosis and glucose intolerance in LDLR-/- mice-role of intestinal permeability and macrophage activation. PLOS ONE 9, e108577 (2014).
Grander, C. et al. Recovery of ethanol-induced Akkermansia muciniphila depletion ameliorates alcoholic liver disease. Gut 67, 891–901 (2017).
Plovier, H. et al. A purified membrane protein from Akkermansia muciniphila or the pasteurized bacterium improves metabolism in obese and diabetic mice. Nat. Med. 23, 107–113 (2017).
Wang, Y. et al. Lactobacillus rhamnosus GG culture supernatant ameliorates acute alcohol-induced intestinal permeability and liver injury. Am. J. Physiol. Gastrointest. Liver Physiol. 303, G32–G41 (2012).
Natividad, J. M. et al. Impaired aryl hydrocarbon receptor ligand production by the gut microbiota is a key factor in metabolic syndrome. Cell Metab. 28, 737–749 (2018).
Wang, K. et al. Parabacteroides distasonis alleviates obesity and metabolic dysfunctions via production of succinate and secondary bile acids. Cell Rep. 26, 222–235 (2019).
Krumbeck, J. A. et al. Probiotic bifidobacterium strains and galactooligosaccharides improve intestinal barrier function in obese adults but show no synergism when used together as synbiotics. Microbiome 6, 121 (2018).
Guo, C. et al. Bile acids control inflammation and metabolic disorder through inhibition of NLRP3 inflammasome. Immunity 45, 802–816 (2016).
Golden, J. M. et al. Ursodeoxycholic acid protects against intestinal barrier breakdown by promoting enterocyte migration via EGFR- and COX-2-dependent mechanisms. Am. J. Physiol. Gastrointest. Liver Physiol. 315, G259–G271 (2018).
Gadaleta, R. M. et al. Farnesoid X receptor activation inhibits inflammation and preserves the intestinal barrier in inflammatory bowel disease. Gut 60, 463–472 (2011).
Wahlstrom, A., Sayin, S. I., Marschall, H. U. & Backhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Suez, J., Zmora, N., Segal, E. & Elinav, E. The pros, cons, and many unknowns of probiotics. Nat. Med. 25, 716–729 (2019).
Wei, X. et al. Fatty acid synthase modulates intestinal barrier function through palmitoylation of mucin 2. Cell Host Microbe 11, 140–152 (2012).
Muccioli, G. G. et al. The endocannabinoid system links gut microbiota to adipogenesis. Mol. Syst. Biol. 6, 392 (2010).
Jenne, C. N. & Kubes, P. Immune surveillance by the liver. Nat. Immunol. 14, 996–1006 (2013).
Sun, L. et al. A marker of endotoxemia is associated with obesity and related metabolic disorders in apparently healthy Chinese. Diabetes Care 33, 1925–1932 (2010).
Pussinen, P. J., Havulinna, A. S., Lehto, M., Sundvall, J. & Salomaa, V. Endotoxemia is associated with an increased risk of incident diabetes. Diabetes Care 34, 392–397 (2011). This paper shows that patients with diabetes have higher systemic endotoxin activity compared with non-diabetic individuals.
Camargo, A. et al. Postprandial endotoxemia may influence the development of type 2 diabetes mellitus: from the CORDIOPREV study. Clin. Nutr. 38, 529–538 (2018).
Mehta, N. N. et al. Experimental endotoxemia induces adipose inflammation and insulin resistance in humans. Diabetes 59, 172–181 (2010).
Membrez, M. et al. Gut microbiota modulation with norfloxacin and ampicillin enhances glucose tolerance in mice. FASEB J 22, 2416–2426 (2008).
Pang, J. et al. Significant positive association of endotoxemia with histological severity in 237 patients with non-alcoholic fatty liver disease. Aliment. Pharmacol. Ther. 46, 175–182 (2017).
Luche, E. et al. Metabolic endotoxemia directly increases the proliferation of adipocyte precursors at the onset of metabolic diseases through a CD14-dependent mechanism. Mol. Metab. 2, 281–291 (2013).
Ye, D. et al. Toll-like receptor-4 mediates obesity-induced non-alcoholic steatohepatitis through activation of X-box binding protein-1 in mice. Gut 61, 1058–1067 (2012).
Amar, J. et al. Energy intake is associated with endotoxemia in apparently healthy men. Am. J. Clin. Nutr. 87, 1219–1223 (2008).
Cani, P. D. et al. Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia 50, 2374–2383 (2007).
Fabbiano, S. et al. Functional gut microbiota remodeling contributes to the caloric restriction-induced metabolic improvements. Cell Metab. 28, 907–921 (2018).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016).
Fei, N. & Zhao, L. An opportunistic pathogen isolated from the gut of an obese human causes obesity in germfree mice. ISME J. 7, 880–884 (2013).
Yang, Q., Vijayakumar, A. & Kahn, B. B. Metabolites as regulators of insulin sensitivity and metabolism. Nat. Rev. Mol. Cell Biol. 19, 654–672 (2018).
Clarke, T. B. et al. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat. Med. 16, 228–231 (2010).
Hergott, C. B. et al. Peptidoglycan from the gut microbiota governs the lifespan of circulating phagocytes at homeostasis. Blood 127, 2460–2471 (2016).
Chan, K. L. et al. Circulating NOD1 activators and hematopoietic NOD1 contribute to metabolic inflammation and insulin resistance. Cell Rep. 18, 2415–2426 (2017).
Schertzer, J. D. et al. NOD1 activators link innate immunity to insulin resistance. Diabetes 60, 2206–2215 (2011).
Cavallari, J. F. et al. Muramyl dipeptide-based postbiotics mitigate obesity-induced insulin resistance via IRF4. Cell Metab. 25, 1063–1074 (2017).
Tang, W. H. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1584 (2013).
Zhu, W. et al. Gut microbial metabolite tmao enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016). In this study, TMAO was characterized as a major gut-derived metabolite affecting platelet hyper-reactivity, suggesting a major role for the gastrointestinal tract in the pathogenesis of thrombosis.
Brown, J. M. & Hazen, S. L. The gut microbial endocrine organ: bacterially derived signals driving cardiometabolic diseases. Annu. Rev. Med. 66, 343–359 (2015).
Shan, Z. et al. Association between microbiota-dependent metabolite trimethylamine-N-oxide and type 2 diabetes. Am. J. Clin. Nutr. 106, 888–894 (2017).
Tang, W. H. et al. Increased trimethylamine N-oxide portends high mortality risk independent of glycemic control in patients with type 2 diabetes mellitus. Clin. Chem. 63, 297–306 (2017).
Chen, Y. M. et al. Associations of gut-flora-dependent metabolite trimethylamine-N-oxide, betaine and choline with non-alcoholic fatty liver disease in adults. Sci. Rep. 6, 19076 (2016).
Li, P. et al. Plasma concentration of trimethylamine-N-oxide and risk of gestational diabetes mellitus. Am. J. Clin. Nutr. 108, 603–610 (2018).
Brown, J. M. & Hazen, S. L. Microbial modulation of cardiovascular disease. Nat. Rev. Microbiol. 16, 171–181 (2018).
Tang, W. H. & Hazen, S. L. Microbiome, trimethylamine N-oxide, and cardiometabolic disease. Transl. Res. 179, 108–115 (2017).
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Perry, R. J. et al. Acetate mediates a microbiome-brain-beta-cell axis to promote metabolic syndrome. Nature 534, 213–217 (2016).
Hoyles, L. et al. Molecular phenomics and metagenomics of hepatic steatosis in non-diabetic obese women. Nat. Med. 24, 1070–1080 (2018). In this study, the authors identify phenylacetic acid, a microbial product, as a trigger of hepatic steatosis, providing further evidence for a role of the gut microbiota in this process.
Koh, A. et al. Microbially produced imidazole propionate impairs insulin signaling through mTORC1. Cell 175, 947–961 (2018). Here, imidazole propionate, a microbial metabolite derived from histidine, is shown to circulate at increased concentrations in patients with type 2 diabetes and contribute to insulin resistance.
Amar, J. et al. Blood microbiota dysbiosis is associated with the onset of cardiovascular events in a large general population: the D.E.S.I.R. study. PLOS ONE 8, e54461 (2013). In this population-based study, the authors suggest a relationship between circulating blood microbiota, atherosclerosis and cardiovascular complications.
Potgieter, M., Bester, J., Kell, D. B. & Pretorius, E. The dormant blood microbiome in chronic, inflammatory diseases. FEMS Microbiol. Rev. 39, 567–591 (2015).
Paisse, S. et al. Comprehensive description of blood microbiome from healthy donors assessed by 16 S targeted metagenomic sequencing. Transfusion 56, 1138–1147 (2016).
Schierwagen, R. et al. Circulating microbiome in blood of different circulatory compartments. Gut https://doi.org/10.1136/gutjnl-2018-316227 (2018).
Lelouvier, B. et al. Changes in blood microbiota profiles associated with liver fibrosis in obese patients: a pilot analysis. Hepatology 64, 2015–2027 (2016).
Puri, P. et al. The circulating microbiome signature and inferred functional metagenomics in alcoholic hepatitis. Hepatology 67, 1284–1302 (2018). This paper shows that heavy alcohol consumption affects intestinal barrier function and is associated with the appearance of a circulating microbiome.
Zulian, A. et al. Adipose tissue microbiota in humans: an open issue. Int. J. Obes. 40, 1643–1648 (2016).
Manfredo Vieira, S. et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science 359, 1156–1161 (2018).
Zegarra-Ruiz, D. F. et al. A diet-sensitive commensal Lactobacillus strain mediates TLR7-dependent systemic autoimmunity. Cell Host Microbe 25, 113–127 (2019).
Nakamoto, N. et al. Gut pathobionts underlie intestinal barrier dysfunction and liver T helper 17 cell immune response in primary sclerosing cholangitis. Nat. Microbiol. 4, 492–503 (2019).
Nathan, D. M. Long-term complications of diabetes mellitus. N. Engl. J. Med. 328, 1676–1685 (1993).
Diehl, A. M. & Day, C. Cause, pathogenesis, and treatment of nonalcoholic steatohepatitis. N. Engl. J. Med. 377, 2063–2072 (2017).
Chambers, J. C. et al. C-reactive protein, insulin resistance, central obesity, and coronary heart disease risk in Indian Asians from the United Kingdom compared with European whites. Circulation 104, 145–150 (2001).
Visser, M., Bouter, L. M., McQuillan, G. M., Wener, M. H. & Harris, T. B. Low-grade systemic inflammation in overweight children. Pediatrics 107, E13 (2001).
Lee, Y. S., Wollam, J. & Olefsky, J. M. An integrated view of immunometabolism. Cell 172, 22–40 (2018).
Johnson, A. R., Milner, J. J. & Makowski, L. The inflammation highway: metabolism accelerates inflammatory traffic in obesity. Immunol. Rev. 249, 218–238 (2012).
Emerging Risk Factors Collaboration, Kaptoge, S. et al. C-reactive protein concentration and risk of coronary heart disease, stroke, and mortality: an individual participant meta-analysis. Lancet 375, 132–140 (2010).
Emerging Risk Factors Collaboration, Kaptoge, S. et al. C-reactive protein, fibrinogen, and cardiovascular disease prediction. N. Engl. J. Med. 367, 1310–1320 (2012).
Nissen, S. E. et al. Statin therapy, LDL cholesterol, C-reactive protein, and coronary artery disease. N. Engl. J. Med. 352, 29–38 (2005).
Ridker, P. M. et al. C-reactive protein levels and outcomes after statin therapy. N. Engl. J. Med. 352, 20–28 (2005).
Ridker, P. M. et al. Measurement of C-reactive protein for the targeting of statin therapy in the primary prevention of acute coronary events. N. Engl. J. Med. 344, 1959–1965 (2001).
Ridker, P. M. et al. Antiinflammatory therapy with canakinumab for atherosclerotic disease. N. Engl. J. Med. 377, 1119–1131 (2017). A landmark clinical study demonstrating a key role for IL-1β in metabolic inflammation and associated cardiovascular complications.
Ridker, P. M. et al. Relationship of C-reactive protein reduction to cardiovascular event reduction following treatment with canakinumab: a secondary analysis from the CANTOS randomised controlled trial. Lancet 391, 319–328 (2018).
Ridker, P. M. et al. Modulation of the interleukin-6 signalling pathway and incidence rates of atherosclerotic events and all-cause mortality: analyses from the Canakinumab Anti-Inflammatory Thrombosis Outcomes Study (CANTOS). Eur. Heart J. 39, 3499–3507 (2018).
Brandsma, E. et al. A proinflammatory gut microbiota increases systemic inflammation and accelerates atherosclerosis. Circ. Res. 124, 94–100 (2019).
Yoshida, N. et al. Bacteroides vulgatus and Bacteroides dorei reduce gut microbial lipopolysaccharide production and inhibit atherosclerosis. Circulation 138, 2486–2498 (2018).
Leite, A. Z. et al. Detection of increased plasma interleukin-6 levels and prevalence of Prevotella copri and Bacteroides vulgatus in the feces of type 2 diabetes patients. Front. Immunol. 8, 1107 (2017).
Dewulf, E. M. et al. Insight into the prebiotic concept: lessons from an exploratory, double blind intervention study with inulin-type fructans in obese women. Gut 62, 1112–1121 (2013).
Kahn, S. E., Cooper, M. E. & Del Prato, S. Pathophysiology and treatment of type 2 diabetes: perspectives on the past, present, and future. Lancet 383, 1068–1083 (2014).
Johnson, A. M. & Olefsky, J. M. The origins and drivers of insulin resistance. Cell 152, 673–684 (2013).
Tilg, H. & Moschen, A. R. Inflammatory mechanisms in the regulation of insulin resistance. Mol. Med. 14, 222–231 (2008).
Samuel, V. T. & Shulman, G. I. Mechanisms for insulin resistance: common threads and missing links. Cell 148, 852–871 (2012).
Kiechl, S. et al. Blockade of receptor activator of nuclear factor-κB (RANKL) signaling improves hepatic insulin resistance and prevents development of diabetes mellitus. Nat. Med. 19, 358–363 (2013).
Hotamisligil, G. S. Inflammation and metabolic disorders. Nature 444, 860–867 (2006).
Larsen, C. M. et al. Interleukin-1-receptor antagonist in type 2 diabetes mellitus. N. Engl. J. Med. 356, 1517–1526 (2007).
Larsen, C. M. et al. Sustained effects of interleukin-1 receptor antagonist treatment in type 2 diabetes. Diabetes Care 32, 1663–1668 (2009).
Everett, B. M. et al. Anti-inflammatory therapy with canakinumab for the prevention and management of diabetes. J. Am. Coll. Cardiol. 71, 2392–2401 (2018).
Netea, M. G. et al. A guiding map for inflammation. Nat. Immunol. 18, 826–831 (2017).
Tilg, H. & Moschen, A. R. Evolution of inflammation in nonalcoholic fatty liver disease: the multiple parallel hits hypothesis. Hepatology 52, 1836–1846 (2010).
Mehal, W. Z. The Gordian Knot of dysbiosis, obesity and NAFLD. Nat. Rev. Gastroenterol. Hepatol. 10, 637–644 (2013).
Evans, A. S. Causation and disease: the Henle-Koch postulates revisited. Yale J. Biol. Med. 49, 175–195 (1976).
Fung, T. C., Olson, C. A. & Hsiao, E. Y. Interactions between the microbiota, immune and nervous systems in health and disease. Nat. Neurosci. 20, 145–155 (2017).
Blander, J. M., Longman, R. S., Iliev, I. D., Sonnenberg, G. F. & Artis, D. Regulation of inflammation by microbiota interactions with the host. Nat. Immunol. 18, 851–860 (2017).
Duan, Y. et al. Inflammatory links between high fat diets and diseases. Front. Immunol. 9, 2649 (2018).
Caesar, R., Tremaroli, V., Kovatcheva-Datchary, P., Cani, P. D. & Backhed, F. Crosstalk between gut microbiota and dietary lipids aggravates wat inflammation through tlr signaling. Cell Metab. 22, 658–668 (2015).
Roager, H. M. et al. Whole grain-rich diet reduces body weight and systemic low-grade inflammation without inducing major changes of the gut microbiome: a randomised cross-over trial. Gut 68, 83–93 (2019).
Mardinoglu, A. et al. An integrated understanding of the rapid metabolic benefits of a carbohydrate-restricted diet on hepatic steatosis in humans. Cell Metab. 27, 559–571 (2018).
Yan, Y. et al. Omega-3 fatty acids prevent inflammation and metabolic disorder through inhibition of NLRP3 inflammasome activation. Immunity 38, 1154–1163 (2013).
Mitchell, S. J. et al. Nicotinamide improves aspects of healthspan, but not lifespan, in mice. Cell Metab. 27, 667–676 (2018).
Agudelo, L. Z. et al. Kynurenic acid and Gpr35 regulate adipose tissue energy homeostasis and inflammation. Cell Metab. 27, 378–392 (2018).
Syed, I. et al. Palmitic acid hydroxystearic acids activate gpr40, which is involved in their beneficial effects on glucose homeostasis. Cell Metab. 27, 419–427 (2018).
Hatori, M. et al. Time-restricted feeding without reducing caloric intake prevents metabolic diseases in mice fed a high-fat diet. Cell Metab. 15, 848–860 (2012).
Clarke, S. F. et al. Exercise and associated dietary extremes impact on gut microbial diversity. Gut 63, 1913–1920 (2014).
Geng, L. et al. Exercise alleviates obesity-induced metabolic dysfunction via enhancing fgf21 sensitivity in adipose tissues. Cell Rep. 26, 2738–2752 (2019).
Seganfredo, F. B. et al. Weight-loss interventions and gut microbiota changes in overweight and obese patients: a systematic review. Obes. Rev. 18, 832–851 (2017).
Ryan, K. K. et al. FXR is a molecular target for the effects of vertical sleeve gastrectomy. Nature 509, 183–188 (2014).
Labrecque, J., Laforest, S., Michaud, A., Biertho, L. & Tchernof, A. Impact of bariatric surgery on white adipose tissue inflammation. Can. J. Diabetes 41, 407–417 (2017).
Verbeek, J. et al. Roux-en-Y gastric bypass attenuates hepatic mitochondrial dysfunction in mice with non-alcoholic steatohepatitis. Gut 64, 673–683 (2015).
de Groot, P. et al. Donor metabolic characteristics drive effects of faecal microbiota transplantation on recipient insulin sensitivity, energy expenditure and intestinal transit time. Gut https://doi.org/10.1136/gutjnl-2019-318320 (2019).
Rathinam, V. A. K., Zhao, Y. & Shao, F. Innate immunity to intracellular LPS. Nat. Immunol. 20, 527–533 (2019).
Vijayan, A., Rumbo, M., Carnoy, C. & Sirard, J. C. Compartmentalized antimicrobial defenses in response to flagellin. Trends Microbiol. 26, 423–435 (2018).
Wolf, A. J. & Underhill, D. M. Peptidoglycan recognition by the innate immune system. Nat. Rev. Immunol. 18, 243–254 (2018).
Kanneganti, T. D. The signposts and winding roads to immunity and inflammation. Nat. Rev. Immunol. 19, 81–82 (2019).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Winkler, P., Ghadimi, D., Schrezenmeir, J. & Kraehenbuhl, J. P. Molecular and cellular basis of microflora-host interactions. J. Nutr. 137, 756S–772S (2007).
Kotas, M. E. & Locksley, R. M. Why innate lymphoid cells? Immunity 48, 1081–1090 (2018).
Kumar, V. & Ahmad, A. Role of MAIT cells in the immunopathogenesis of inflammatory diseases: new players in old game. Int. Rev. Immunol. 37, 90–110 (2018).
Satoh, M. & Iwabuchi, K. Role of natural killer T cells in the development of obesity and insulin resistance: insights from recent progress. Front. Immunol. 9, 1314 (2018).
Newton, R., Priyadharshini, B. & Turka, L. A. Immunometabolism of regulatory T cells. Nat. Immunol. 17, 618–625 (2016).
Reilly, S. M. & Saltiel, A. R. Adapting to obesity with adipose tissue inflammation. Nat. Rev. Endocrinol. 13, 633–643 (2017).
Shimizu, I., Yoshida, Y., Suda, M. & Minamino, T. DNA damage response and metabolic disease. Cell Metab. 20, 967–977 (2014).
Hotamisligil, G. S. Endoplasmic reticulum stress and the inflammatory basis of metabolic disease. Cell 140, 900–917 (2010).
Grazioli, S. & Pugin, J. Mitochondrial damage-associated molecular patterns: from inflammatory signaling to human diseases. Front. Immunol. 9, 832 (2018).
Breton, J. et al. Gut commensal E. coli proteins activate host satiety pathways following nutrient-induced bacterial growth. Cell Metab. 23, 324–334 (2016).
Cani, P. D. & de Vos, W. M. Next-generation beneficial microbes: the case of Akkermansia muciniphila. Front. Microbiol. 8, 1765 (2017).
Maruvada, P., Leone, V., Kaplan, L. M. & Chang, E. B. The human microbiome and obesity: moving beyond associations. Cell Host Microbe 22, 589–599 (2017).
Marchesi, J. R. et al. The gut microbiota and host health: a new clinical frontier. Gut 65, 330–339 (2016).
Acknowledgements
The authors thank members of the Tilg and Elinav laboratories for discussions and apologize to authors whose work was not included due to space constraints. H.T. is supported by the Excellence Initiative (Competence Centres for Excellent Technologies — COMET) of the Austrian Research Promotion Agency FFG: Research Centre of Excellence in Vascular Ageing Tyrol, VASCage (K-Project No. 843536) funded by BMVIT, BMWFW, Wirtschaftsagentur Wien and Standortagentur Tirol. N.Z. is supported by a Gilead Biosciences Fellowship. T.E.A. is grateful for the support from the Austrian Science Fund (FWF, P 29379-B28), the Austrian Society of Gastroenterology and Hepatology (ÖGGH), and the European Crohn’s and Colitis Organization (ECCO). E.E. is supported by Y. and R. Ungar, the Abisch Frenkel Foundation for the Promotion of Life Sciences, the Gurwin Family Fund for Scientific Research, the Leona M. and Harry B. Helmsley Charitable Trust, the Crown Endowment Fund for Immunological Research, the estate of J. Gitlitz, the estate of L. Hershkovich, the Benoziyo Endowment Fund for the Advancement of Science, the Adelis Foundation, J.L. and V. Schwartz, A. and G. Markovitz, A. and C. Adelson, the French National Centre for Scientific Research (CNRS), D. L. Schwarz, the V. R. Schwartz Research Fellow Chair, L. Steinberg, J. N. Halpern, A. Edelheit, grants funded by the European Research Council, a Marie Curie Integration grant, the German–Israeli Foundation for Scientific Research and Development, the Israel Science Foundation, the Minerva Foundation, the Rising Tide Foundation, the Helmholtz Foundation, and the European Foundation for the Study of Diabetes. E.E. is a senior fellow of the Canadian Institute of Advanced Research (CIFAR) and an international scholar of the Bill and Melinda Gates Foundation and Howard Hughes Medical Institute (HHMI).
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, made substantial contribution to discussion of content, and wrote, reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
E.E. is a paid consultant for DayTwo and BiomX. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Immunology thanks K. Clément, B. Jabri and O. Pedersen for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- ob/ob mice
-
A mouse model of metabolic dysregulation and obesity that arises from increased appetite due to a leptin mutation (that renders these mice functionally leptin deficient).
- Metabolic endotoxaemia
-
A state that favours the translocation of microbial components (such as lipopolysaccharide) to the bloodstream, which promotes metabolic disease.
- Chylomicrons
-
Transport vesicles (so-called lipoprotein particles) for absorbed dietary lipids.
- Berberine
-
A plant-derived alkaloid of an ancient Chinese herb, Coptis chinensis.
- Low-density lipoprotein receptor-deficient mice
-
A mouse model of atherosclerosis caused by a targeted deletion of the gene encoding the low-density lipoprotein receptor (LDLR). In humans, homozygous mutations in LDLR cause familial hypercholesterolaemia, a disease characterized by pronounced hyperlipidaemia and accelerated atherosclerotic cardiovascular disease.
Rights and permissions
About this article
Cite this article
Tilg, H., Zmora, N., Adolph, T.E. et al. The intestinal microbiota fuelling metabolic inflammation. Nat Rev Immunol 20, 40–54 (2020). https://doi.org/10.1038/s41577-019-0198-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41577-019-0198-4
This article is cited by
-
Probiotics ameliorate endocrine disorders via modulating inflammatory pathways: a systematic review
Genes & Nutrition (2024)
-
Effects of dietary fibre on metabolic health and obesity
Nature Reviews Gastroenterology & Hepatology (2024)
-
Gut microbial metabolite facilitates colorectal cancer development via ferroptosis inhibition
Nature Cell Biology (2024)
-
Vitamin D3 supplementation shapes the composition of gut microbiota and improves some obesity parameters induced by high-fat diet in mice
European Journal of Nutrition (2024)
-
Decreased expression of the NLRP6 inflammasome is associated with increased intestinal permeability and inflammation in obesity with type 2 diabetes
Cellular and Molecular Life Sciences (2024)