Abstract
Assembly and publication of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) genome in January 2020 enabled the immediate development of tests to detect the new virus. This began the largest global testing programme in history, in which hundreds of millions of individuals have been tested to date. The unprecedented scale of testing has driven innovation in the strategies, technologies and concepts that govern testing in public health. This Review describes the changing role of testing during the COVID-19 pandemic, including the use of genomic surveillance to track SARS-CoV-2 transmission around the world, the use of contact tracing to contain disease outbreaks and testing for the presence of the virus circulating in the environment. Despite these efforts, widespread community transmission has become entrenched in many countries and has required the testing of populations to identify and isolate infected individuals, many of whom are asymptomatic. The diagnostic and epidemiological principles that underpin such population-scale testing are also considered, as are the high-throughput and point-of-care technologies that make testing feasible on a massive scale.
Similar content being viewed by others
Introduction
On 7 January 2020, a new coronavirus was detected by metatranscriptomic sequencing of lung fluid from a patient with pneumonia-like symptoms in Wuhan, China1,2. On 10 January 2020, the assembled reference genome of this new coronavirus, termed severe acute respiratory syndrome (SARS) coronavirus 2 (SARS-CoV-2), was published, and within 2 weeks the first diagnostic tests to detect the virus were issued3,4. To date, hundreds of millions of individuals have been tested for the presence of SARS-CoV-2 (ref.5), leading to widespread public awareness of and debate regarding diagnostic concepts and technologies.
Testing for the presence of SARS-CoV-2 is typically performed for one of two reasons. First, a symptomatic patient might be tested to inform their clinical treatment. Such diagnostic testing is focused on therapeutic care and is typically performed in a well-controlled clinical setting, and the test results are usually interpreted alongside the patient’s history and symptoms. Second, testing might be performed to identify infectious individuals in a population, who are then isolated to prevent the onward infection of others. Such screening is focused on public health outcomes and aims to reduce viral transmission through a population. Individuals who are not symptomatic might be tested, and testing might need to be performed on a massive scale. These two different uses of testing have different requirements and priorities, and a test that is useful in clinical diagnosis can be ill-suited for population-scale screening.
The observation that countries with high testing rates were able to effectively control SARS-CoV-2 transmission during the initial stages of the pandemic6 supported the proposal that screening could help to limit viral transmission. Accordingly, widespread testing rapidly gained traction as an intervention that might avoid both the immediate economic costs of lockdown and the societal costs of social distancing measures. Accordingly, many countries implemented population-scale testing to monitor and reduce viral transmission. However, despite the promise of this approach, obtaining reliable test results on such a massive scale is difficult and unprecedented, and has been achieved with differing outcomes and success7. Nonetheless, the COVID-19 pandemic has transformed the role of population-scale screening. With ongoing investment and innovation, large-scale testing is likely to become a common feature of public health.
This Review focuses on the use of testing as a tool to screen populations for individuals infected with SARS-CoV-2. We discuss the changing role of such testing in the COVID-19 pandemic: the initial use of genomic epidemiology to monitor the spread of viral strains around the world; the containment of disease outbreaks using contact tracing; and the massive screening programmes that aim to suppress community transmission (Table 1; Fig. 1). We also consider the testing strategies and technologies used to address the challenges of population-scale testing. We finally discuss findings from large-scale testing programmes that have measured the exposure of entire populations to SARS-CoV-2, and how these studies have evaluated and informed our response to the pandemic.
a | Shedding of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) RNA genome (yellow shading) typically increases rapidly following infection and peaks at the presentation of symptoms (although many patients can be asymptomatic) before gradually declining. Levels of viral proteins (red shading) also rapidly increase, albeit within a narrower window. In response to infection, the body produces IgM (green shading) and IgG (blue shading) antibodies, which persist for weeks to months. b | The SARS-CoV-2 RNA genome can be detected by molecular assays, such as reverse transcription–quantitative polymerase chain reaction (RT–qPCR), before the development of symptoms. Serological tests can detect reactive IgG and IgM, which indicate past infection with SARS-CoV-2. c | Assay sensitivity is dependent on both technical performance aspects and viral load. A test might not detect viral RNA even when an individual is infectious (false negative, red shaded area) or, alternatively, might detect persistent viral RNA after an individual is no longer infectious (false positive), which demonstrates that test positivity correlates poorly with infectivity. Owing to the rapid increase in viral shedding, only a narrow window exists wherein a more sensitive assay (test B) will outperform a less sensitive assay (test A). Note that this figure illustrates a generalized COVID-19 infectious course, and in practice the relative duration of detectability and analyte abundance differ considerably between individuals. NAAT, nucleic acid antigen testing; NGS, next-generation sequencing.
Genomic epidemiology of SARS-CoV-2
During replication and transmission, viral genomes accumulate mutations that can be used to define distinct variants. Whole-genome sequencing can identify these mutations and trace the transmission of variants between individuals, populations and countries around the world. This genomic epidemiology approach was used to monitor past outbreaks of Ebola virus and Zika virus infection8,9,10 and has been widely used to reconstruct the spread of SARS-CoV-2. One of these SARS-CoV-2 strains, termed ‘B.1’, was initially transmitted to Italy, where it caused an outbreak in Lombardy before further circulating across Europe and onwards to the USA, where it seeded an outbreak in New York City11. This chain of transmission around the globe was reconstructed by the tracking of four mutations in the SARS-CoV-2 B.1 strain. The SARS-CoV-2 viral genome quickly diversified into different strains following the initial zoonotic transmission of the ancestral virus. During its initial spread, thousands of viral genomes were rapidly sequenced by research laboratories worldwide and shared in open-access databases such as the EpiCoV database from GISAID12 and the Our World in Data COVID-19 dataset. These sequences, along with metadata (including location, date and method of sampling) and tools such as Nextstrain13, could then be used to track the spread of SARS-CoV-2 strains14.
Genomic epidemiology has also been widely used to reconstruct the routes by which SARS-CoV-2 is introduced into and spreads within countries15,16,17,18,19. These studies show that SARS-CoV-2 is typically introduced into a country several times20. Subsequent transmission is stochastic, wherein a small number of infected individuals are responsible for the majority of secondary infections in ‘superspreader’ events21,22. Genomic epidemiology studies also reveal the routes of transmission into and across a country, and their results have informed public health containment measures such as regional lockdowns, quarantines and travel restrictions23.
Despite the usefulness of genomic epidemiology, the data should be interpreted with caution. Although infection with different strains can rule out a chain of transmission between two individuals, infections by the same strain do not necessarily prove a direct link in the chain of transmission24. Moreover, compared with other pathogenic viruses, the SARS-CoV-2 genome accumulates mutations slowly and the amount of viral sampling and sequencing has differed widely between countries. Low strain diversity, rapid disease spread and non-representative sampling can confound the results of genomic epidemiology studies, and our understanding of SARS-CoV-2 spread and evolution was especially unclear during the early stages of the COVID-19 pandemic11,25.
Testing of individuals for SARS-CoV-2
The RNA genome of SARS-CoV-2 can be directly detected by nucleic acid assays such as reverse transcription–quantitative polymerase chain reaction (RT–qPCR) tests, whereas the presence of viral proteins can be detected by an antigen test (Box 1). The detection of these analytes requires that a sample contains sufficient viral genome copies or levels of viral proteins that exceed the limit of detection for a given assay (Fig. 2). However, the abundance of these viral analytes varies dramatically between different anatomical locations and different stages of infection26. Thus, the ability to detect SARS-CoV-2 is ultimately framed by the biology of the virus. Indeed, the sensitivity of tests for detecting SARS-CoV-2 can be as dependent on the time and site sampled as it is on the technical performance of the assay27.
a,b | In a population with a low prevalence (5% here) of COVID-19 cases, even a highly sensitive and specific test (part a) returns many false-positive results. c,d | By contrast, a high infection prevalence (25% here) increases the likelihood that a positive result is true, despite the test performance remaining unchanged. The positive predictive value (PPV) of a test describes the probability that an individual who tests positive is actually infected35, and thus depends on both the specificity of the test and the prevalence of infection. At low prevalence values, the proportion of false-positive results is increased and the PPV is reduced. Even a highly specific test returns mostly false-positive results (and therefore has a low PPV) when the prevalence of infection is low. Adapted from ref.134, Springer Nature Limited.
The viral load of SARS-CoV-2 changes dramatically during the course of infection28,29. Viral genome abundance rapidly increases following infection, and patients can become infectious and begin shedding viral particles 3–5 days before the development of COVID-19 symptoms30. During these early stages of infection, a nasopharyngeal sample (which requires a long swab to sample the back of the nasal cavity) generally provides the highest reported viral RNA abundance31,32. However, other, less invasive options, including oral swabs, nasal swabs or saliva tests, also have detectable levels of viral analytes and can reduce exposure of the health-care workers performing the sampling33.
Viral load typically declines gradually following the onset of symptoms. During the later stages of infection, samples from the lower respiratory tract (such as sputum, bronchoalveolar lavage fluid or tracheal secretions) can yield high viral titres. Even after the cessation of all symptoms, persistent viral RNA can be detected for several weeks. However, efforts to isolate replication-competent virus from this persistent RNA have been unsuccessful, and these recovered individuals are generally considered to be non-infectious34.
The response of an individual to the test result is a key factor in the effectiveness of testing programmes, and the interpretation of test results should consider both the probability of infection and any pre-existing risk factors35 (Box 2; Fig. 2). Whereas a positive test result indicates the presence of infection and the need for isolation to prevent further transmission of the virus, a negative test result does not rule out the possibility of it being a false-negative finding36. If individuals are falsely reassured by receiving a negative test result, they might ignore other protection strategies and increase the risk of onward transmission of the virus37.
Following viral infection, the body mounts a humoral immune response that includes the production of antibodies that target SARS-CoV-2 antigens38,39,40. These antibodies directly bind to the spike protein and thereby neutralize viral action, as well as recruiting further immune responses to clear the virus. The levels of reactive (that is, anti-SARS-CoV-2) IgM antibodies typically increase within a few days of the viral infection and these antibodies persist for weeks, whereas reactive IgG antibodies appear in the middle and later stages of the infection and can remain circulating in the blood for several months41,42. Serological tests can detect reactive antibodies produced in response to viral exposure. As these tests do not directly detect the virus but instead detect the body’s response to the virus, they can indicate whether an individual was previously infected with SARS-CoV-2 (ref.43). Serological tests can also indicate the strength and direction of an individual’s immune response. However, a positive serological test result does not necessarily indicate a protective immune response, and other anti-SARS-CoV-2 immunoglobulins (for example, IgA and IgE) are often not detected by available serological tests.
Contact tracing
The introduction and initial outbreak of SARS-CoV-2 within a country can be contained by testing and contact tracing, wherein all infected individuals and their secondary contacts are identified and isolated. If these secondary exposed individuals are quarantined before they can infect others, further onward transmission of the virus is prevented44.
Large-scale contact tracing programmes have been instrumental in halting SARS-CoV-2 transmission in countries such as Vietnam, Taiwan and Japan6,45. Within days of detecting its first case on 20 January 2020, South Korea quickly scaled up the nation’s existing network of contact tracers, who had been used in the SARS-CoV outbreak in 2003. These contact tracers used patient interviews as well as medical, mobile phone and credit card transaction records to identify thousands of people linked to an outbreak in Itaewo, a popular nightclub district in Seoul. Subsequent testing identified and isolated hundreds of infected individuals, many of whom were separated by multiple cycles of transmission from the original Itaewo outbreak46.
Although simple in concept, contract tracing is often difficult to put into practice because the effectiveness of this approach depends on identifying contacts before they can infect others47,48. Given that individuals can become infectious within days of initial exposure to SARS-CoV-2 and before they develop symptoms, only a narrow window is available within which to identify individuals before they infect others. Therefore, rapid testing is of primary importance for effective contact tracing, and contacted individuals are recommended to pre-emptively isolate themselves while awaiting their test results.
Contact tracing is also laborious and time-consuming, and becomes increasingly difficult in the context of ongoing viral transmission. As more cases are diagnosed, the number of secondary contacts scales non-linearly to increasingly large numbers of people, who must in turn be identified, tested and isolated49. This increasing case burden results in delays that further decrease the effectiveness of contact tracing. The use of digital contact tracing using mobile apps and other technologies can help to automate contact tracing approaches and ease this burden, but requires widespread adoption and adherence50,51. Once the case burden exceeds a country’s contact tracing capacity, most individuals with secondary cases are contacted too late and tracing has little further effect on viral transmission.
Population-scale testing
Reducing community transmission
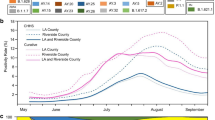
Despite efforts to contain SARS-CoV-2, community transmission has become entrenched in many countries, which has required a shift from contact tracing to the population-scale testing required to detect the large fraction of presymptomatic, mildly symptomatic and asymptomatic individuals who can unwittingly transmit the virus52,53. On 31 October 2020, Slovakia became the first country to test its entire population in an attempt to reduce viral transmission54,55,56. This massive undertaking required thousands of health-care workers to test almost all citizens using a rapid antigen test, following which individuals who tested positive and their close contacts were advised to isolate themselves. Although the effects of this massive testing effort are difficult to distinguish from those of concurrent interventions, a marked reduction in SARS-CoV-2 infections was observed following the effort57. However, this reduction was limited to regions with high viral prevalence, and testing seemingly had little effect in regions with a lower viral prevalence58.
The success of population-scale testing can be precisely measured as a reduction in the reproductive number, which indicates the extent of viral transmission through a population (Box 2). However, the amount of testing that must be performed to markedly reduce the reproductive number is difficult to evaluate. The fraction of tests that return a positive result can provide some indication as to the adequacy of the testing programme: a low test positivity rate indicates both low viral prevalence and sufficient surveillance capacity, whereas a high test positivity rate suggests that testing is inadequate and that many infected individuals are going undetected59 (Box 2).
For many countries, adequate surveillance mandates a massive amount of testing, requiring huge resources and a collaborative effort across a laboratory enterprise that includes commercial, clinical, government and research organizations57. Large coordinated systems are required for safe sample acquisition, transport and custody. In addition, test results must be returned sufficiently rapidly to enable individuals who test positive to isolate themselves before they infect others. Indeed, the speed and frequency of testing is considered as important as test sensitivity in effectively reducing viral transmission58. Such massive testing efforts can quickly strain supply chains and infrastructure, and many countries have faced shortages in key reagents60 that left them unable to implement effective, widespread testing. As a result, testing of health-care workers and symptomatic individuals is often prioritized over testing of asymptomatic individuals. Competition for reagents has often reflected global health inequities, and wide disparities in testing rates are observed between countries61. However, containing SARS-CoV-2 transmission is a global challenge, and the global responsibility for sustainable testing must be recognized.
Monitoring of viral prevalence
Population-scale testing also provides critical viral prevalence data that have informed our response to the pandemic. On 21 February 2020, the town of Vo’ in Italy reported the first death from COVID-19 and quickly entered a complete lockdown for 2 weeks. Researchers were able to test almost every town resident and to quantify the effect of lockdown on reducing the viral reproduction number (Box 2), thereby providing early validation of the effectiveness of such interventions used to combat the pandemic62.
Population-scale testing enables the rate of viral spread through a population to be measured, aids in the identification of regional hotspots and high-risk subpopulations, and facilitates the calculation of incidence and mortality rates. However, as with any measurement process, such testing is subject to the errors and biases inherent in the assay and sampling processes. Low-sensitivity tests can underestimate viral prevalence, whereas low-specificity tests can overestimate prevalence. For example, an early study conducted in California, USA, was criticized for providing an inflated measure of SARS-CoV-2 prevalence owing to the use of low-specificity serological tests63.
Serological assays that detect reactive antibodies can indicate whether an individual was previously infected with SARS-CoV-2. These assays, which are sufficiently cheap to be widely deployed, have been used in some of the largest studies of SARS-CoV-2 spread and immunization64,65. Repeated rounds of serological sampling across large, representative populations in the UK conducted as part of the real-time assessment of community transmission (REACT) study provided ongoing estimates of viral exposure during successive pandemic waves66. The REACT investigators measured viral spread through subpopulations in the UK and their data were used to determine the need for regional lockdowns.
Iceland has proved ideal for the study of COVID-19 because more than half of its citizens have been tested for SARS-CoV-2 antibodies, and the clinical outcomes of individuals with a positive test result have been closely monitored67. However, despite extensive testing, almost half of infected people in Iceland were not detected, thereby illustrating the limitations of and gaps in even the most widespread population-scale testing schemes64. A similar seroprevalence study in Spain found that population-scale testing detected just 9% of infected people during the first pandemic wave, and almost one million infected people were estimated to have gone undetected65.
Strategies to scale up testing
Population-scale testing has relied heavily on RT–qPCR, which is performed in large, high-throughput, centralized laboratories with automated equipment by skilled personnel (Box 1). These centralized laboratories achieve sensitive and reliable results owing to validated and regulated oversight of testing procedures. However, samples need to be transported to the testing site, which can often extend the testing turnaround time to several days.
The pooling of several samples enables multiple individuals to be simultaneously tested. Chinese authorities tested more than seven million people in the city of Qingdao in 3 days using a pooling approach, wherein ten samples were combined and tested with a single reaction68. If the pool returns a negative test result, then all constituent individual samples are also considered to have a negative result. Only if the pooled test returns a positive result are the constituent samples tested individually to ultimately identify the positive sample or samples. A further innovation divides the samples among overlapping pools, such that any single sample is present in a unique combination of pools69,70. If the unique combination of pools returns a positive result, the individual sample can be identified without requiring an additional round of retesting71.
Pooling of patient samples, viral transport media or extracted RNA enables more individuals to be tested with use of fewer reagents. However, the advantages of pooling are limited in the context of a high test positivity rate. If positive samples are common, the majority of pools will return positive results, and most samples still need to be tested individually. In practice, this shortcoming can be mitigated by non-random pooling of samples from shared households or groups, which increases the likelihood that positive samples will be clustered within a small number of pools72. Nevertheless, the additional rounds of testing mandated by pooling can incur reporting delays, and the dilution of positive samples within large pools can reduce test sensitivity73.
Point-of-care (POC) tests can be performed on-site, such as within a local clinic, workplace or even a patient’s home. These tests are typically antigen-based lateral flow assays that can be easily shipped to the site and are sufficiently simple to be performed and interpreted without specialized training or equipment (Box 1). These features enable POC tests to be easily and widely distributed to the population without incurring the challenges and delays associated with sample collection and reporting74. Decentralization can improve access to testing, enable earlier and more frequent testing, and reduce exposure of health-care workers. Given these advantages, POC tests are widely considered an attractive approach to achieve a massive expansion in testing75, and a diverse range of POC tests have been developed that incorporate molecular, antigen-based and serological technologies76.
In November 2020, an initial pilot scheme aimed to screen up to half a million people in Liverpool, UK, using rapid, on-site antigen tests74. This scheme offered routine and repeated testing to all residents, regardless of symptoms, in a broad effort to achieve population-scale test coverage and reduce viral transmission. This community-led testing scheme identified more than one-third of the infected but mildly symptomatic or asymptomatic individuals in Liverpool77. However, despite this achievement, the sensitivity of the antigen tests used in this scheme was markedly lower than that reported in the original test validation studies, and the tests were thought to miss almost one-third of infectious individuals78.
Monitoring of emergent strains
The mutations that accumulate in the SARS-CoV-2 genome can alter the viral phenotype and confer a selective advantage that gives rise to new strains. Genomic epidemiology showed that one strain, distinguished by a non-synonymous D614G mutation in the spike protein, first emerged in Europe before expanding to become the predominant strain worldwide79 owing to a selective fitness advantage conferred by the mutation that increased viral transmissibility80.
The integration of genome sequencing within population-scale testing can enable monitoring of the viral strains circulating within a population. Numerous countries have mandated that a proportion of positive samples is subjected to whole-genome sequencing, thereby providing ongoing surveillance of emerging and circulating variants. This sequencing information can identify emergent SARS-CoV-2 variants with differing transmission or pathogenicity, with resistance to antiviral treatment or that are at risk of vaccine escape81. In late December 2020, a new SARS-CoV-2 strain known as B.1.1.7 rapidly increased in prevalence throughout the UK, apparently outcompeting existing variants and prompting the rapid imposition of restrictions on travel to other countries82. Additional variants that might increase transmissibility and pathogenicity or reduce the efficacy of vaccines have similarly arisen in South Africa (B.1.351)83 and Brazil (P.1)84. As the effect of these variants on the viral phenotype has become apparent, authorities have recognized that global testing will be increasingly needed to monitor the emergence and circulation of new variant SARS-CoV-2 strains.
Variant diversification identified by genomic surveillance is also important to assess the influence of new mutations on the ongoing performance of molecular diagnostic tests85,86. The emergent B.1.1.7 strain harbours a large number of mutations that might prevent the binding of some primers to the spike gene and thereby reduce the sensitivity of RT–qPCR tests87 (Box 1). In response, numerous variant-specific primers have been developed, illustrating that strain diversification will require ongoing updates and validation of testing reagents.
Environmental testing
In the early stages of the COVID-19 pandemic, the routes of SARS-CoV-2 transmission were unclear, and viral particles that land on objects and surfaces (fomites) were considered to pose a risk of indirect transmission88. Testing of hospital and public surfaces close to infected patients detected the presence of viral RNA, and countries such as China implemented testing of frozen food imports89,90. However, ongoing environmental testing and epidemiological studies suggest that this mode of transmission is uncommon, and that viral transmission is instead primarily caused by contaminated respiratory droplets and aerosols91.
Nevertheless, although viral transmission via fomites is rare, the detection of SARS-CoV-2 in environmental samples can be used to infer its circulation within a community. SARS-CoV-2 can infect the gastrointestinal tract and can be shed at high concentrations in stool, and might be subsequently discharged into wastewater92. SARS-CoV-2 RNA can be sensitively detected in untreated municipal wastewater, including settled solids and sludge93,94,95. Accordingly, many municipalities have started to test sewage for the presence of SARS-CoV-2, similarly to the testing of primary sewage for poliovirus, which has been successfully used to monitor communities for decades96.
Retrospective studies show that increases in the concentration of SARS-CoV-2 RNA in raw wastewater are correlated with increases in reported COVID-19 cases83,97. For example, the detection of SARS-CoV-2 in wastewater from New Haven, Connecticut, USA, tracked rates of hospital admissions during early stages of the COVID-19 pandemic98. The detection of SARS-CoV-2 RNA in wastewater occurred days before the rise in the numbers of clinical COVID-19 cases at local hospitals and provided a valuable predictor of viral presence98.
Environmental testing has the advantage of detecting SARS-CoV-2 across a wide catchment area that encompasses many individuals. This cost-effective approach can provide community data even in the absence of large-scale testing and is unbiased with regard to people who lack access to health care. Environmental testing can also be used to closely monitor closed residential settings, such as colleges, universities, nursing homes and prisons99. Despite its potential, wastewater testing faces technical challenges in achieving sufficient sensitivity, in part because variables such as sampling location, type and volume can all affect detection. Rigorous validation and longitudinal monitoring is required for reliable, actionable wastewater testing100.
Ensuring the integrity of testing
The speed and scale of the COVID-19 pandemic has created unprecedented challenges to ensure the accuracy and reliability of testing. The implementation of population-scale screening is a massive undertaking that requires coordinated input from governmental, commercial and academic organizations. Numerous large programmes, such as the RADx initiative and the X-Prize Challenge, have been established to accelerate population-scale testing101,102.
The unique requirements of the pandemic have also called for modified regulatory regimes that encourage the development of rapid, cheap tests that are well suited to population-scale testing26. To expedite access to SARS-CoV-2 testing during the pandemic, many regulatory agencies have responded with expedited authorization of testing devices. For example, the FDA issues Emergency Use Authorizations (EUAs) if devices have a reasonable expectation of being effective (and various other criteria are met)103. Although EUAs provide welcome flexibility for commercial and laboratory developers, this approach also increases the risk that these tests could perform unexpectedly when deployed at scale in the field. For example, several poorly performing antigen tests were initially issued EUAs that were ultimately revoked, which required the FDA to independently evaluate these poor-quality tests and subsequently update its performance requirements and advice104,105.
Reference standards can improve quality and standardization across large and diverse testing schemes106. The rapid manufacture of synthetic SARS-CoV-2-positive controls has enabled molecular assays to be rapidly evaluated even in the context of limited access to patient samples. These positive controls also enable the early detection of defective reagents, such as faulty primers in RT–qPCR tests107,108. Inactivated viral fractions and serum samples have also been used to develop and evaluate the performance of serological tests109. Indeed, the proliferation of positive reference controls has been so widespread as to fuel concerns about contamination of testing and environmental surveillance schemes110,111.
The FDA has issued SARS-CoV-2 reference sample panels for the standardized evaluation of diagnostic assays109, and numerous other organizations, such as FIND, have launched standardized evaluation programmes112. Notably, findings from these standardized evaluations often differ markedly from test performance statistics published in the EUA101,113. The disparity might, in part, be due to the initial validation of test performance in hospitalized patients, who typically have high viral loads and undergo sampling in a well-controlled setting. This approach can render unreliable the estimation of test performance during population-scale screening, where many tested individuals might be presymptomatic or asymptomatic and have low viral or antibody loads114. This situation highlights that test validation is an ongoing process and that proficiency testing schemes are required to continually evaluate the performance of individual laboratories115.
Surveillance testing
SARS-CoV-2 will probably continue to circulate within human populations for many years, with ongoing regional or seasonal outbreaks29. Surveillance testing aims to identify these outbreaks early, often by the detection of a symptomatic index case, and to quickly contain their spread through contact tracing and isolation. Genomic epidemiology can distinguish between novel introductions and persistent local circulation, and can trace the source of outbreaks116, whereas environmental testing provides ongoing monitoring of large catchment areas.
Routine testing could also become more common in various aspects of life, with testing of closed communities and populations (such as prisons, schools and universities) increasingly common115,117,118. Given the role of cross-border transport in the rapid spread of SARS-CoV-2, testing might become a standard feature of international travel, with testing of incoming travellers mandatory in many countries that aim to limit reintroduction of SARS-CoV-2 or new variant strains119,120. Global surveillance of circulating SARS-CoV-2 variants using genome sequencing will also become increasingly important to match circulating variants with vaccines, and to monitor vaccine escape and the emergence of antiviral resistance7.
Clinical diagnostic testing for SARS-CoV-2 (alongside other respiratory pathogens, such as influenza viruses) is expected to become part of the standard evaluation of patients admitted to hospital with associated symptoms. Currently, the presence of SARS-CoV-2 is sufficient to direct clinical treatment; however, genome sequencing might be required to distinguish between different variant strains if they necessitate differing clinical or containment responses. Although the considerable innovation and investment in testing is likely to also benefit the detection of other infectious diseases and endemic diseases such as cancer, the disruption to national screening programmes as a result of the pandemic is anticipated to increase the case burden of these diseases for many years to come121.
Domestic and wild animal populations can also be infected with SARS-CoV-2 and could act as reservoirs for the virus. Spillover viral transmission between human and animal populations can result in changes in selection pressure that accelerate viral adaptation and evolution. Testing has found repeated, independent infections of domestic mink populations in the Netherlands and Denmark, where SARS-CoV-2 accumulated additional mutations before being reintroduced into human populations122,123,124. This observation has prompted culling of domestic mink populations as a result of fears that they could act as reservoirs for new SARS-CoV-2 strains. In addition, pressure is growing for increased testing of both wild and domestic animal populations to monitor viral prevalence and diversity.
Population-scale testing has assumed a central role in our efforts to combat the COVID-19 pandemic and will be similarly central to our preparation for future pandemics. Notably, countries such as South Korea and Taiwan that had been affected by SARS outbreaks were able to quickly scale up existing contact tracing and testing schemes, which proved key in containing COVID-19 outbreaks45,125. Many other countries are likely to establish similar national preparation plans that maintain their investment in testing infrastructure and supplies of reagents. The establishment of international genomic surveillance systems to monitor the emergence of novel SARS-CoV-2 strains and the zoonotic transmission of new viruses will enable prompt and appropriate responses, including the development and manufacture of diagnostic tests, and the rapid imposition of travel restrictions and testing requirements7. Although some of these preparations are specific to coronaviruses, which have been the source of multiple pandemics in the past two decades126, the lessons learned are also applicable to response planning for other viruses with different transmission dynamics, such as influenza viruses and Ebola virus9.
Conclusions
Population-scale testing has emerged as a major intervention strategy to manage the COVID-19 pandemic. As community transmission became entrenched within countries, the utility of population-scale testing has been intensely debated. Although widespread testing has been proposed as a solution to suppress and even stop the pandemic, the limitations of and gaps in population-scale testing have also become apparent, and both the cost and the effectiveness of mass testing on reducing viral transmission have been intensely debated.
In the early stages of the pandemic, the main limitations on widespread testing were technical, including insufficient reagents and an unprepared laboratory testing infrastructure. These practical obstacles to the implementation of testing on a massive scale favoured the use of established technologies, which explains why the vast majority of testing for SARS-CoV-2 has been performed by conventional RT–qPCR tests in centralized laboratories. These RT–qPCR tests were developed almost immediately after publication of the reference genome, were quickly scaled up through an existing infrastructure and have generally proved both accurate and reliable. However, with the pandemic ongoing, a new generation of rapid and convenient POC tests have been developed.
With sufficiently broad and frequent community testing, identification and isolation of most infectious individuals was considered possible, thereby arresting the pandemic127. However, even the most extensive testing schemes have been found to miss many infectious individuals and to provide only partial population coverage64,65. The rapid infectious course of SARS-CoV-2 and the possibility of its transmission by presymptomatic or asymptomatic individuals blunts the impact of population-scale testing, as many infectious individuals escape detection128. In France, for example, models based on health records suggest that most infected people were not tested or quarantined following the first pandemic wave in early 2020, despite the implementation of a nationwide surveillance programme129.
The benefits of widespread testing have been widely debated, with researchers arguing that screening of large populations with low infection rates is expensive and ineffective, as most positive results are false positives130. Furthermore, the effectiveness of population-scale testing becomes increasingly marginal as viral prevalence falls, and testing alone seems insufficient to eliminate viral transmission. Therefore, researchers have argued that testing should be reserved for symptomatic individuals, traced contacts, closed populations or high-risk groups with an increased probability of infection, where this approach has proven effective, and used in coordination with other interventions, such as targeted regional lockdowns or travel restrictions131,132,133.
Human populations have experienced many pandemics throughout history; however, this has been the first pandemic in which widespread and routine testing has been available. We have been able to monitor viral transmission and measure viral spread in successive pandemic waves of SARS-CoV-2 infection throughout human populations worldwide. The COVID-19 pandemic has already required unprecedented testing of millions of people, and this information has been critical to understanding viral transmission, informing effective responses and planning for future pandemics. Although population-scale testing has not provided a simple or single technical solution to the pandemic, it has served as an invaluable guide to our response.
References
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Sheridan, C. Coronavirus and the race to distribute reliable diagnostics. Nat. Biotechnol. 38, 382–384 (2020).
World Health Organization. Molecular assays to diagnose COVID-19: summary table of available protocols. WHO https://www.who.int/publications/m/item/molecular-assays-to-diagnose-covid-19-summary-table-of-available-protocols (2020).
Hasell, J. et al. A cross-country database of COVID-19 testing. Sci. Data 7, 345 (2020).
Wang, C. J., Ng, C. Y. & Brook, R. H. Response to COVID-19 in Taiwan: big data analytics, new technology, and proactive testing. JAMA 323, 1341–1342 (2020).
The COVID-19 testing debacle. Nat. Biotechnol. 38, 653 (2020).
Gardy, J. L. & Loman, N. J. Towards a genomics-informed, real-time, global pathogen surveillance system. Nat. Rev. Genet. 19, 9–20 (2018).
Gire, S. K. et al. Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345, 1369–1372 (2014).
Faria, N. R. et al. Establishment and cryptic transmission of Zika virus in Brazil and the Americas. Nature 546, 406–410 (2017).
Worobey, M. et al. The emergence of SARS-CoV-2 in Europe and North America. Science 370, 564–570 (2020). This work uses genomic epidemiology to describe the initial international spread of SARS-CoV-2 variants.
Elbe, S. & Buckland-Merrett, G. Data, disease and diplomacy: GISAID’s innovative contribution to global health. Glob. Chall. 1, 33–46 (2017).
Hadfield, J. et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123 (2018).
Forster, P., Forster, L., Renfrew, C. & Forster, M. Phylogenetic network analysis of SARS-CoV-2 genomes. Proc. Natl Acad. Sci. USA 117, 9241–9243 (2020).
Rockett, R. J. et al. Revealing COVID-19 transmission in Australia by SARS-CoV-2 genome sequencing and agent-based modeling. Nat. Med. 26, 1398–1404 (2020).
Miller, D. et al. Full genome viral sequences inform patterns of SARS-CoV-2 spread into and within Israel. Nat. Commun. 11, 5518 (2020).
Lu, J. et al. Genomic epidemiology of SARS-CoV-2 in Guangdong province, China. Cell 181, 997–1003.e9 (2020).
Fauver, J. R. et al. Coast-to-coast spread of SARS-CoV-2 during the early epidemic in the United States. Cell 181, 990–996.e5 (2020).
Seemann, T. et al. Tracking the COVID-19 pandemic in Australia using genomics. Nat. Commun. 11, 4376 (2020).
Deng, X. et al. Genomic surveillance reveals multiple introductions of SARS-CoV-2 into northern California. Science 369, 582–587 (2020).
Althouse, B. M. et al. Superspreading events in the transmission dynamics of SARS-CoV-2: opportunities for interventions and control. PLoS Biol. 18, e3000897 (2020).
Lemieux, J. E. et al. Phylogenetic analysis of SARS-CoV-2 in Boston highlights the impact of superspreading events. Science 371, eabe3261 (2020).
Oude Munnink, B. B. et al. Rapid SARS-CoV-2 whole-genome sequencing and analysis for informed public health decision-making in the Netherlands. Nat. Med. 26, 1405–1410 (2020).
Villabona-Arenas, C. J., Hanage, W. P. & Tully, D. C. Phylogenetic interpretation during outbreaks requires caution. Nat. Microbiol. 5, 876–877 (2020).
Bedford, T. et al. Cryptic transmission of SARS-CoV-2 in Washington State. Science 370, 571–575 (2020).
Wölfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature 581, 465–469 (2020).
Mina, M. J., Parker, R. & Larremore, D. B. Rethinking COVID-19 test sensitivity — a strategy for containment. N. Engl. J. Med. 383, e120 (2020).
Li, Q. et al. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 382, 1199–1207 (2020). This report determines the initial SARS-CoV-2 transmission dynamics, including infectious and incubation periods and basic reproductive number.
Cevik, M. et al. SARS-CoV-2, SARS-CoV, and MERS-CoV viral load dynamics, duration of viral shedding, and infectiousness: a systematic review and meta-analysis. Lancet Microbe 2, e13–e22 (2021).
Kissler, S. M., Tedijanto, C., Goldstein, E., Grad, Y. H. & Lipsitch, M. Projecting the transmission dynamics of SARS-CoV-2 through the postpandemic period. Science 368, 860–868 (2020).
Pan, Y., Zhang, D., Yang, P., Poon, L. L. M. & Wang, Q. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect. Dis. 20, 411–412 (2020).
Wang, W. et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 323, 1843–1844 (2020).
Vogels, C. B. F. et al. SalivaDirect: a simplified and flexible platform to enhance SARS-CoV-2 testing capacity. Med 2, 263–280.e6 (2020).
van Kampen, J. J. A. et al. Duration and key determinants of infectious virus shedding in hospitalized patients with coronavirus disease-2019 (COVID-19). Nat. Commun. 12, 1–6 (2021).
Stites, E. C. & Wilen, C. B. The interpretation of SARS-CoV-2 diagnostic tests. Med 1, 78–89 (2020).
Sethuraman, N., Jeremiah, S. S. & Ryo, A. Interpreting diagnostic tests for SARS-CoV-2. JAMA 323, 2249–2251 (2020).
McCartney, M., Sullivan, F. & Heneghan, C. Information and rational decision-making: explanations to patients and citizens about personal risk of COVID-19. BMJ Evid. Based Med. https://doi.org/10.1136/bmjebm-2020-111541 (2020).
Rodda, L. B. et al. Functional SARS-CoV-2-specific immune memory persists after mild COVID-19. Cell 184, 169–183.e17 (2020).
Tay, M. Z., Poh, C. M., Rénia, L., MacAry, P. A. & Ng, L. F. P. The trinity of COVID-19: immunity, inflammation and intervention. Nat. Rev. Immunol. 20, 363–374 (2020).
Wajnberg, A. et al. Humoral response and PCR positivity in patients with COVID-19 in the New York city region, USA: an observational study. Lancet Microbe 1, e283–e289 (2020).
Long, Q. X. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 26, 845–848 (2020).
Ng, D. L. et al. SARS-CoV-2 seroprevalence and neutralizing activity in donor and patient blood. Nat. Commun. 11, 4698 (2020).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 26, 1033–1036 (2020).
Kraemer, M. U. G. et al. The effect of human mobility and control measures on the COVID-19 epidemic in China. Science 368, 493–497 (2020).
Wu, W. K., Liou, J. M., Hsu, C. C., Lin, Y. H. & Wu, M. S. Pandemic preparedness in Taiwan. Nat. Biotechnol. 38, 932–933 (2020).
Kang, C. R. et al. Coronavirus disease exposure and spread from nightclubs, South Korea. Emerg. Infect. Dis. 26, 2499–2501 (2020).
Kretzschmar, M. E. et al. Impact of delays on effectiveness of contact tracing strategies for COVID-19: a modelling study. Lancet Public Health 5, e452–e459 (2020).
Kucharski, A. J. et al. Effectiveness of isolation, testing, contact tracing, and physical distancing on reducing transmission of SARS-CoV-2 in different settings: a mathematical modelling study. Lancet Infect. Dis. 20, 1151–1160 (2020).
Hellewell, J. et al. Feasibility of controlling COVID-19 outbreaks by isolation of cases and contacts. Lancet Glob. Health 8, e488–e496 (2020).
Ferretti, L. et al. Quantifying SARS-CoV-2 transmission suggests epidemic control with digital contact tracing. Science 368, eabb6936 (2020).
The app credibility gap. Nat. Biotechnol. 38, 768 (2020).
Long, Q. X. et al. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat. Med. 26, 1200–1204 (2020).
Li, R. et al. Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 368, 489–493 (2020).
Holt, E. Slovakia to test all adults for SARS-CoV-2. Lancet 396, 1386–1387 (2020).
Pavelka, M. et al. The impact of population-wide rapid antigen testing on SARS-CoV-2 prevalence in Slovakia. Science https://doi.org/10.1126/science.abf9648 (2021).
Frnda, J. & Durica, M. On pilot massive COVID-19 testing by antigen tests in Europe. case study: Slovakia. Infect. Dis. Rep. 13, 45–57 (2021).
Vandenberg, O., Martiny, D., Rochas, O., van Belkum, A. & Kozlakidis, Z. Considerations for diagnostic COVID-19 tests. Nat. Rev. Microbiol. 19, 171–183 (2020).
Larremore, D. B. et al. Test sensitivity is secondary to frequency and turnaround time for COVID-19 surveillance. Sci. Adv. 7, eabd5393 (2021).
Russell, T. W. et al. Reconstructing the early global dynamics of under-ascertained COVID-19 cases and infections. BMC Med. 18, 332 (2020).
Guan, D. et al. Global supply-chain effects of COVID-19 control measures. Nat. Hum. Behav. 4, 577–587 (2020).
Songok, E. A locally sustainable approach to COVID-19 testing in Africa. Lancet Microbe 1, e197 (2020).
Lavezzo, E. et al. Suppression of a SARS-CoV-2 outbreak in the Italian municipality of Vo’. Nature 584, 425–429 (2020). Large-scale testing enabled the measurement of SARS-CoV-2 transmission within an entire town and allowed the impact of interventionists’ lockdown measures on the reproductive number to be determined.
Vogel, G. Antibody surveys suggesting vast undercount of coronavirus infections may be unreliable. Science https://doi.org/10.1126/science.abc3831 (2020).
Gudbjartsson, D. F. et al. Humoral immune response to SARS-CoV-2 in Iceland. N. Engl. J. Med. 383, 1724–1734 (2020). This population-scale serological study measured the exposure of the Icelandic population to SARS-CoV-2 during the first pandemic wave and assessed the effectiveness of large-scale testing.
Pollán, M. et al. Prevalence of SARS-CoV-2 in Spain (ENE-COVID): a nationwide, population-based seroepidemiological study. Lancet 396, 535–544 (2020).
Ward, H. et al. Antibody prevalence for SARS-CoV-2 in England following the first peak of the pandemic. Nat. Commun. 12, 905 (2021).
Gudbjartsson, D. F. et al. Spread of SARS-CoV-2 in the Icelandic population. N. Engl. J. Med. 382, 2302–2315 (2020).
Xing, Y., Wong, G. W. K., Ni, W., Hu, X. & Xing, Q. Rapid response to an outbreak in Qingdao, China. N. Engl. J. Med. 383, e129 (2020).
Ghosh, S. et al. Tapestry: a single-round smart pooling technique for COVID-19 testing. Preprint at medRxiv https://doi.org/10.1101/2020.04.23.20077727 (2020).
Mutesa, L. et al. A pooled testing strategy for identifying SARS-CoV-2 at low prevalence. Nature 589, 276–280 (2020).
Shental, N. et al. Efficient high-throughput SARS-CoV-2 testing to detect asymptomatic carriers. Sci. Adv. 6, 5961–5972 (2020).
Barak, N. et al. Lessons from applied large-scale pooling of 133,816 SARS-CoV-2 RT-PCR tests. Sci. Transl Med. https://doi.org/10.1126/scitranslmed.abf2823 (2021).
Bateman, A. C., Mueller, S., Guenther, K. & Shult, P. Assessing the dilution effect of specimen pooling on the sensitivity of SARS-CoV-2 PCR tests. J. Med. Virol. 93, 1568–1572 (2020).
Iacobucci, G. Covid-19: Mass population testing is rolled out in Liverpool. BMJ 371, m4268 (2020).
Peeling, R. W., Olliaro, P. L., Boeras, D. I. & Fongwen, N. Scaling up COVID-19 rapid antigen tests: promises and challenges. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(21)00048-7 (2021).
Kevadiya, B. D. et al. Diagnostics for SARS-CoV-2 infections. Nat. Mater. https://doi.org/10.1038/s41563-020-00906-z (2021).
GOV.UK. Research and analysis. Liverpool COVID-19 community testing pilot: interim evaluation report summary. gov.uk https://www.gov.uk/government/publications/liverpool-covid-19-community-testing-pilot-interim-evaluation-report-summary/liverpool-covid-19-community-testing-pilot-interim-evaluation-report-summary (2021).
Wise, J. Covid-19: Lateral flow tests miss over half of cases, Liverpool pilot data show. BMJ 371, m4848 (2020).
Korber, B. et al. Tracking changes in SARS-CoV-2 Spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182, 812–827.e19 (2020).
Plante, J. A. et al. Spike mutation D614G alters SARS-CoV-2 fitness. Nature https://doi.org/10.1038/s41586-020-2895-3 (2020).
Thomson, E. C. et al. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell 184, 1171–1187.e20 (2021).
Davies, N. G. et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372, eabg3055 (2021). This study describes the emergence of SARS-CoV-2 variants with higher estimated reproductive numbers than other variants.
Tegally, H. et al. Emergence and rapid spread of a new severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) lineage with multiple spike mutations in South Africa. Nature https://doi.org/10.1038/s41586-021-03402-9 (2021).
Voloch, C. M. et al. Genomic characterization of a novel SARS-CoV-2 lineage from Rio de Janeiro, Brazil. J. Virol. https://doi.org/10.1128/JVI.00119-21 (2021).
Artesi, M. et al. A recurrent mutation at position 26340 of SARS-CoV-2 is associated with failure of the E Gene quantitative reverse transcription-PCR utilized in a commercial dual-target diagnostic assay. J. Clin. Microbiol. 58, e01598–20 (2020).
Ziegler, K. et al. SARS-CoV-2 samples may escape detection because of a single point mutation in the N gene. Eur. Surveill. 25, 2001650 (2020).
Rambaut, A. et al. Preliminary genomic characterisation of an emergent SARS-CoV-2 lineage in the UK defined by a novel set of spike mutations. Virological https://virological.org/t/preliminary-genomic-characterisation-of-an-emergent-sars-cov-2-lineage-in-the-uk-defined-by-a-novel-set-of-spike-mutations/563 (2020).
van Doremalen, N. et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 382, 1564–1567 (2020).
Piana, A. et al. Monitoring COVID-19 transmission risks by quantitative real-time PCR tracing of droplets in hospital and living environments. mSphere 6, e01070-20 (2021).
Pang, X. et al. Cold-chain food contamination as the possible origin of COVID-19 resurgence in Beijing. Natl Sci. Rev. https://doi.org/10.1093/nsr/nwaa264 (2020).
Zhang, R., Li, Y., Zhang, A. L., Wang, Y. & Molina, M. J. Identifying airborne transmission as the dominant route for the spread of COVID-19. Proc. Natl Acad. Sci. USA 117, 14857–14863 (2020).
Wu, Y. et al. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol. Hepatol. 5, 434–435 (2020).
Lodder, W. & de Roda Husman, A. M. SARS-CoV-2 in wastewater: potential health risk, but also data source. Lancet Gastroenterol. Hepatol. 5, 533–534 (2020).
Graham, K. E. et al. SARS-CoV-2 RNA in wastewater settled solids is associated with COVID-19 cases in a large urban sewershed. Environ. Sci. Technol. 55, 488–498 (2020).
Ahmed, W. et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 728, 138764 (2020).
Berchenko, Y. et al. Estimation of polio infection prevalence from environmental surveillance data. Sci. Transl Med. 9, eaaf6786 (2017).
Medema, G., Heijnen, L., Elsinga, G., Italiaander, R. & Brouwer, A. Presence of SARS-coronavirus-2 RNA in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environ. Sci. Technol. Lett. 7, 511–516 (2020).
Peccia, J. et al. Measurement of SARS-CoV-2 RNA in wastewater tracks community infection dynamics. Nat. Biotechnol. 38, 1164–1167 (2020). This group shows that detection of the presence of SARS-CoV-2 RNA in wastewater correlated with hospital COVID-19 admissions, thereby demonstrating that this approach can be used to monitor the presence of SARS-CoV-2 within a community catchment area.
Betancourt, W. Q. et al. COVID-19 containment on a college campus via wastewater-based epidemiology, targeted clinical testing and an intervention. Sci. Total Environ. 779, 146408 (2021).
O’Reilly, K. M., Allen, D. J., Fine, P. & Asghar, H. The challenges of informative wastewater sampling for SARS-CoV-2 must be met: lessons from polio eradication. Lancet Microbe 1, E189–E190 (2020).
MacKay, M. J. et al. The COVID-19 XPRIZE and the need for scalable, fast, and widespread testing. Nat. Biotechnol. 38, 1021–1024 (2020).
Tromberg, B. J. et al. Rapid scaling up of COVID-19 diagnostic testing in the US — the NIH RADx initiative. N. Engl. J. Med. 383, 1071–1077 (2020).
Shuren, J. & Stenzel, T. COVID-19 molecular diagnostic testing — lessons learned. N. Engl. J. Med. 383, e97 (2020).
Kalokairinou, L., Zettler, P. J., Nagappan, A., Kyweluk, M. A. & Wexler, A. The promise of direct-to-consumer COVID-19 testing: ethical and regulatory issues. J. Law Biosci. 7, lsaa069 (2020).
Abbasi, J. The promise and peril of antibody testing for COVID-19. JAMA 323, 1881–1883 (2020).
Page, M., Almond, N., Rose, N. J. & Schneider, C. K. Diagnostics and the coronavirus: don’t let the standards slip. Nat. Biotechnol. 38, 673–674 (2020).
Nalla, A. K. et al. Comparative performance of SARS-CoV-2 detection assays using seven different primer-probe sets and one assay kit. J. Clin. Microbiol. 58, e00557-20 (2020).
Vogels, C. B. F. et al. Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT–qPCR primer–probe sets. Nat. Microbiol. 5, 1299–1305 (2020).
US Food & Drug Administration. SARS-CoV-2 reference panel comparative data. FDA https://www.fda.gov/medical-devices/coronavirus-covid-19-and-medical-devices/sars-cov-2-reference-panel-comparative-data (2020).
Wang, C. Y. T., Buckley, C., Bletchly, C., Harris, P. & Whiley, D. Contamination of SARS-CoV-2 RT-PCR probes at the oligonucleotide manufacturer. Pathology 52, 814–816 (2020).
Mögling, R. et al. Delayed laboratory response to COVID-19 caused by molecular diagnostic contamination. Emerg. Infect. Dis. 26, 1944–1946 (2020).
FIND. SARS-CoV-2 diagnostic pipeline. FIND https://www.finddx.org/covid-19/pipeline/ (2021).
Fitzpatrick, M. C., Pandey, A., Wells, C. R., Sah, P. & Galvani, A. P. Buyer beware: inflated claims of sensitivity for rapid COVID-19 tests. Lancet 397, 24–25 (2021).
Takahashi, S., Greenhouse, B. & Rodríguez-Barraquer, I. Are seroprevalence estimates for severe acute respiratory syndrome coronavirus 2 biased? J. Infect. Dis. 222, 1772–1775 (2020).
Zeichhardt, H. & Kammel, M. INSTAND report on extra external quality assessment scheme group no. 340 virus genome detection SARS-CoV-2. INSTAND https://www.instand-ev.de/System/rv-files/340%20EN%20SARS-CoV-2%20Genome%20EQAS%20April%202020%2020200502j.pdf (2020).
Meredith, L. W. et al. Rapid implementation of SARS-CoV-2 sequencing to investigate cases of health-care associated COVID-19: a prospective genomic surveillance study. Lancet Infect. Dis. 20, 1263–1272 (2020).
Rafiei, Y. & Mello, M. M. The missing piece — SARS-CoV-2 testing and school reopening. N. Engl. J. Med. 383, e126 (2020).
Paltiel, A. D., Zheng, A. & Walensky, R. P. Assessment of SARS-CoV-2 screening strategies to permit the safe reopening of college campuses in the US. JAMA Netw. Open 3, e2016818 (2020).
Chinazzi, M. et al. The effect of travel restrictions on the spread of the 2019 novel coronavirus (COVID-19) outbreak. Science 368, 395–400 (2020).
Wells, C. R. et al. Impact of international travel and border control measures on the global spread of the novel 2019 coronavirus outbreak. Proc. Natl Acad. Sci. USA 117, 7504–7509 (2020).
Maringe, C. et al. The impact of the COVID-19 pandemic on cancer deaths due to delays in diagnosis in England, UK: a national, population-based, modelling study. Lancet Oncol. 21, 1023–1034 (2020).
Oude Munnink, B. B. et al. Transmission of SARS-CoV-2 on mink farms between humans and mink and back to humans. Science 371, 172–177 (2020). This group uses whole-genome sequencing to show SARS-CoV-2 transmission between minks and humans.
Oreshkova, N. et al. SARS-CoV-2 infection in farmed minks, the Netherlands, April and May 2020. Euro Surveill. 25, 2001005 (2020).
van Dorp, L. et al. No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2. Nat. Commun. 11, 5986 (2020).
Dighe, A. et al. Response to COVID-19 in South Korea and implications for lifting stringent interventions. BMC Med. 18, 321 (2020).
Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Taipale, J., Romer, P. & Linnarsson, S. Population-scale testing can suppress the spread of COVID-19. Preprint at medRxiv https://doi.org/10.1101/2020.04.27.20078329 (2020).
Gandhi, M., Yokoe, D. S. & Havlir, D. V. Asymptomatic transmission, the Achilles’ heel of current strategies to control COVID-19. N. Engl. J. Med. 382, 2158–2160 (2020).
Pullano, G. et al. Underdetection of COVID-19 cases in France threatens epidemic control. Nature 590, 134–139 (2020).
Raffle, A. E., Pollock, A. M. & Harding-Edgar, L. COVID-19 mass testing programmes. BMJ 370, m3262 (2020).
Pollock, A. M. & Lancaster, J. Asymptomatic transmission of COVID-19. BMJ 371, m4851 (2020).
Skittrall, J. P. et al. Diagnostic tool or screening programme? Asymptomatic testing for SARS-CoV-2 needs clear goals and protocols. Lancet Region. HealthEur. 1, 100002 (2021).
Flaxman, S. et al. Estimating the effects of non-pharmaceutical interventions on COVID-19 in Europe. Nature 584, 257–261 (2020).
Frasier, S. L. Coronavirus antibody tests have a mathematical pitfall. Sci. Am. 323, 12–13 (2020).
Corman, V. M. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eur. Surveill. 25, 2000045 (2020).
Liu, R. et al. Positive rate of RT-PCR detection of SARS-CoV-2 infection in 4880 cases from one hospital in Wuhan, China, from Jan to Feb 2020. Clin. Chim. Acta 505, 172–175 (2020).
Wikramaratna, P., Paton, R., Ghafari, M. & Lourenco, J. Estimating false-negative detection rate of SARS-CoV-2 by RT-PCR. Euro Surveill. 25, 2000568 (2020).
Schmid-Burgk, J. et al. LAMP-Seq: population-scale COVID-19 diagnostics using combinatorial barcoding. Preprint at bioRxiv https://doi.org/10.1101/2020.04.06.025635 (2020).
Yángüez, E. et al. HiDRA-seq: high-throughput SARS-CoV-2 detection by RNA barcoding and amplicon sequencing. Preprint at bioRxiv https://doi.org/10.1101/2020.06.02.130484 (2020).
Bloom, J. S. et al. Swab-Seq: a high-throughput platform for massively scaled up SARS-CoV-2 testing. Preprint at medRxiv https://doi.org/10.1101/2020.08.04.20167874 (2021).
Notomi, T. et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, e63 (2000).
Dao Thi, V. L. et al. A colorimetric RT-LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Sci. Transl Med. 12, eabc7075 (2020).
Piepenburg, O., Williams, C. H., Stemple, D. L. & Armes, N. A. DNA detection using recombination proteins. PLoS Biol. 4, 1115–1121 (2006).
El-Tholoth, M., Bau, H. H. & Song, J. A single and two-stage, closed-tube, molecular test for the 2019 novel coronavirus (COVID-19) at home, clinic, and points of entry. Preprint at https://doi.org/10.26434/chemrxiv.11860137.v1 (2020).
Broughton, J. P. et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 38, 870–874 (2020).
Lambert-Niclot, S. et al. Evaluation of a rapid diagnostic assay for detection of SARS-CoV-2 antigen in nasopharyngeal swabs. J. Clin. Microbiol. 58, e00977-20 (2020).
Scohy, A. et al. Low performance of rapid antigen detection test as frontline testing for COVID-19 diagnosis. J. Clin. Virol. 129, 104455 (2020).
Mak, G. C. et al. Evaluation of rapid antigen test for detection of SARS-CoV-2 virus. J. Clin. Virol. 129, 104500 (2020).
Whitman, J. D. et al. Evaluation of SARS-CoV-2 serology assays reveals a range of test performance. Nat. Biotechnol. 38, 1174–1183 (2020).
Petherick, A. Developing antibody tests for SARS-CoV-2. Lancet 395, 1101–1102 (2020).
Tang, M. S. et al. Clinical performance of two SARS-CoV-2 serologic assays. Clin. Chem. 66, 1055–1062 (2020).
Delamater, P. L., Street, E. J., Leslie, T. F., Yang, Y. T. & Jacobsen, K. H. Complexity of the basic reproduction number (R0). Emerg. Infect. Dis. 25, 1–4 (2019).
Li, Y. et al. The temporal association of introducing and lifting non-pharmaceutical interventions with the time-varying reproduction number (R) of SARS-CoV-2: a modelling study across 131 countries. Lancet Infect. Dis. 21, 193–202 (2020).
Zhang, J. et al. Changes in contact patterns shape the dynamics of the COVID-19 outbreak in China. Science 368, 1481–1486 (2020).
Kucharski, A. J. et al. Early dynamics of transmission and control of COVID-19: a mathematical modelling study. Lancet Infect. Dis. 20, 553–558 (2020).
World Health Organization. Considerations for implementing and adjusting public health and social measures in the context of COVID-19. WHO https://www.who.int/publications/i/item/considerations-in-adjusting-public-health-and-social-measures-in-the-context-of-covid-19-interim-guidance (2020).
Arnaout, R. et al. SARS-CoV2 testing: the limit of detection matters. Preprint at bioRxiv https://doi.org/10.1101/2020.06.02.131144 (2020).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks Z. Kozlakidis, R. S. Vincenzo, J. Hay and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
GISAID: https://www.gisaid.org/
Nextstrain: https://nextstrain.org/
Our World in Data COVID-19 dataset: https://ourworldindata.org/coronavirus-testing
Glossary
- Lockdown
-
The closure of non-essential businesses and activities, along with requiring individuals to remain within their homes, in an effort to reduce population mobility and contact and thereby reduce viral transmission.
- Social distancing
-
The maintenance of a sufficient distance between individuals to prevent airborne transmission of a virus.
- Superspreader
-
A single individual who infects many other individuals.
- Prevalence
-
The fraction of individuals within a population who are affected. This value can indicate the a priori probability that a randomly selected individual from a given population is infected, and is a key determinant of the utility of a screening test.
- Fomites
-
Objects (such as clothes, utensils and furniture) that are likely to be contaminated with infectious material.
Rights and permissions
About this article
Cite this article
Mercer, T.R., Salit, M. Testing at scale during the COVID-19 pandemic. Nat Rev Genet 22, 415–426 (2021). https://doi.org/10.1038/s41576-021-00360-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-021-00360-w
This article is cited by
-
Implementing a pilot study of COVID-19 self-testing in high-risk populations and remote locations: results and lessons learnt
BMC Public Health (2024)
-
Biporous silica nanostructure-induced nanovortex in microfluidics for nucleic acid enrichment, isolation, and PCR-free detection
Nature Communications (2024)
-
Metallic nanoplatforms for COVID-19 diagnostics: versatile applications in the pandemic and post-pandemic era
Journal of Nanobiotechnology (2023)
-
HaploCoV: unsupervised classification and rapid detection of novel emerging variants of SARS-CoV-2
Communications Biology (2023)
-
Effects of public-health measures for zeroing out different SARS-CoV-2 variants
Nature Communications (2023)