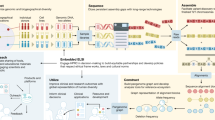
Abstract
Since the early days of the genome era, the scientific community has relied on a single ‘reference’ genome for each species, which is used as the basis for a wide range of genetic analyses, including studies of variation within and across species. As sequencing costs have dropped, thousands of new genomes have been sequenced, and scientists have come to realize that a single reference genome is inadequate for many purposes. By sampling a diverse set of individuals, one can begin to assemble a pan-genome: a collection of all the DNA sequences that occur in a species. Here we review efforts to create pan-genomes for a range of species, from bacteria to humans, and we further consider the computational methods that have been proposed in order to capture, interpret and compare pan-genome data. As scientists continue to survey and catalogue the genomic variation across human populations and begin to assemble a human pan-genome, these efforts will increase our power to connect variation to human diversity, disease and beyond.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
National Human Genome Reserach Institute. Human Genome Project FAQ. NIH https://www.genome.gov/human-genome-project/Completion-FAQ (2019).
Rouli, L., Merhej, V., Fournier, P. E. & Raoult, D. The bacterial pangenome as a new tool for analysing pathogenic bacteria. New Microbes New Infect. 7, 72–85 (2015).
Pallen, M. J. & Wren, B. W. Bacterial pathogenomics. Nature 449, 835–842 (2007).
Tettelin, H. et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial ‘pan-genome’. Proc. Natl Acad. Sci. USA 102, 13950–13955 (2005). The first work on pan-genomes in bacteria, this paper coined the term ‘pan-genome’ and the associated concepts of the ‘core’ and ‘dispensable’ genomes.
Ali, A. et al. Pan-genome analysis of human gastric pathogen H. pylori: comparative genomics and pathogenomics approaches to identify regions associated with pathogenicity and prediction of potential core therapeutic targets. Biomed. Res. Int. 2015, 139580 (2015).
Ali, A. et al. Campylobacter fetus subspecies: comparative genomics and prediction of potential virulence targets. Gene 508, 145–156 (2012).
Imperi, F. et al. The genomics of Acinetobacter baumannii: insights into genome plasticity, antimicrobial resistance and pathogenicity. IUBMB Life 63, 1068–1074 (2011).
Rasko, D. A. et al. The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates. J. Bacteriol. 190, 6881–6893 (2008).
Salipante, S. J. et al. Large-scale genomic sequencing of extraintestinal pathogenic Escherichia coli strains. Genome Res. 25, 119–128 (2015).
Trost, E. et al. Pangenomic study of Corynebacterium diphtheriae that provides insights into the genomic diversity of pathogenic isolates from cases of classical diphtheria, endocarditis, and pneumonia. J. Bacteriol. 194, 3199–3215 (2012).
Medini, D., Donati, C., Tettelin, H., Masignani, V. & Rappuoli, R. The microbial pan-genome. Curr. Opin. Genet. Dev. 15, 589–594 (2005).
1000 Genomes Project Consortium et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Sedlazeck, F. J., Lee, H., Darby, C. A. & Schatz, M. C. Piercing the dark matter: bioinformatics of long-range sequencing and mapping. Nat. Rev. Genet. 19, 329–346 (2018).
Miga, K. H. et al. Centromere reference models for human chromosomes X and Y satellite arrays. Genome Res. 24, 697–707 (2014).
Kehr, B. et al. Diversity in non-repetitive human sequences not found in the reference genome. Nat. Genet. 49, 588–593 (2017).
Jonsson, H. et al. Whole genome characterization of sequence diversity of 15,220 Icelanders. Sci. Data 4, 170115 (2017).
Maretty, L. et al. Sequencing and de novo assembly of 150 genomes from Denmark as a population reference. Nature 548, 87–91 (2017).
Eisfeldt, J., Martensson, G., Ameur, A., Nilsson, D. & Lindstrand, A. Discovery of novel sequences in 1,000 Swedish genomes. Mol. Biol. Evol. https://doi.org/10.1093/molbev/msz176 (2019).
Jacobs, G. S. et al. Multiple deeply divergent Denisovan ancestries in Papuans. Cell 177, 1010–1021.e1032 (2019).
Bai, H. et al. Whole-genome sequencing of 175 Mongolians uncovers population-specific genetic architecture and gene flow throughout North and East Asia. Nat. Genet. 50, 1696–1704 (2018).
Choudhury, A. et al. Whole-genome sequencing for an enhanced understanding of genetic variation among South Africans. Nat. Commun. 8, 2062 (2017).
Gurdasani, D. et al. The African Genome Variation Project shapes medical genetics in Africa. Nature 517, 327–332 (2015).
Mathias, R. A. et al. A continuum of admixture in the Western Hemisphere revealed by the African Diaspora genome. Nat. Commun. 7, 12522 (2016).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Telenti, A. et al. Deep sequencing of 10,000 human genomes. Proc. Natl Acad. Sci. USA 113, 11901–11906 (2016).
1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Sudmant, P. H. et al. An integrated map of structural variation in 2,504 human genomes. Nature 526, 75–81 (2015).
Sherman, R. M. et al. Assembly of a pan-genome from deep sequencing of 910 humans of African descent. Nat. Genet. 51, 30–35 (2019). This study reports over 300 Mb of novel sequence detected from the examination of African-ancestry individuals, demonstrating that a considerable amount of sequence is missing from the human reference genome.
Hall, S. S. Revolution postponed. Sci. Am. 303, 60–67 (2010).
Wade, N. A decade later, genetic map yields few new cures. N. Y. Times 12 (12 Jun 2010).
Zou, Y. et al. 1,520 reference genomes from cultivated human gut bacteria enable functional microbiome analyses. Nat. Biotechnol. 37, 179–185 (2019).
Francis, W. R. & Worheide, G. Similar ratios of introns to intergenic sequence across animal genomes. Genome Biol. Evol. 9, 1582–1598 (2017).
Piovesan, A. et al. Human protein-coding genes and gene feature statistics in 2019. BMC Res. Notes 12, 315 (2019).
Zhao, Q. et al. Pan-genome analysis highlights the extent of genomic variation in cultivated and wild rice. Nat. Genet. 50, 278–284 (2018).
Schatz, M. C. et al. Whole genome de novo assemblies of three divergent strains of rice, Oryza sativa, document novel gene space of aus and indica. Genome Biol. 15, 506 (2014).
Sun, C. et al. RPAN: rice pan-genome browser for approximately 3000 rice genomes. Nucleic Acids Res. 45, 597–605 (2017).
Gao, L. et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor. Nat. Genet. 51, 1044–1051 (2019).
Li, Y. H. et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat. Biotechnol. 32, 1045–1052 (2014).
Golicz, A. A. et al. The pangenome of an agronomically important crop plant Brassica oleracea. Nat. Commun. 7, 13390 (2016).
Hubner, S. et al. Sunflower pan-genome analysis shows that hybridization altered gene content and disease resistance. Nat. Plants 5, 54–62 (2019).
Tao, Y., Zhao, X., Mace, E., Henry, R. & Jordan, D. Exploring and exploiting pan-genomics for crop improvement. Mol. Plant 12, 156–169 (2019).
Shahbandeh, M. Rice — statistics & facts. Statistica https://www.statista.com/topics/1443/rice/ (2017).
Alonge, M. et al. RaGOO: fast and accurate reference-guided scaffolding of draft genomes. Genome Biol. 20, 224 (2019).
Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Morgante, M., De Paoli, E. & Radovic, S. Transposable elements and the plant pan-genomes. Curr. Opin. Plant Biol. 10, 149–155 (2007).
Hirsch, C. N. et al. Insights into the maize pan-genome and pan-transcriptome. Plant Cell 26, 121–135 (2014).
Hansey, C. N. et al. Maize (Zea mays L.) genome diversity as revealed by RNA-sequencing. PLoS One 7, e33071 (2012).
Ma, Y., Liu, M., Stiller, J. & Liu, C. A pan-transcriptome analysis shows that disease resistance genes have undergone more selection pressure during barley domestication. BMC Genomics 20, 12 (2019).
Gan, X. et al. Multiple reference genomes and transcriptomes for Arabidopsis thaliana. Nature 477, 419–423 (2011).
Cao, J. et al. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat. Genet. 43, 956–963 (2011).
Ganguly, P. NHGRI funds centers for advancing the reference sequence of the human genome. NIH https://www.genome.gov/news/news-release/NIH-funds-centers-for-advancing-sequence-of-human-genome-reference (2019).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Hamosh, A. et al. Online Mendelian inheritance in man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 30, 52–55 (2002).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Hach, F. et al. mrsFAST-Ultra: a compact, SNP-aware mapper for high performance sequencing applications. Nucleic Acids Res. 42, W494–W500 (2014).
Tithi, S. S., Heath, L. S. & Zhang, L. in 7th International Conference on Bioinformatics and Computational Biology (BICoB) (eds Saeed, F. & Haspel, N.) 187–192 (International Society for Computers and Their Applications, 2015).
Wenger, A. M. et al. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat. Biotechnol. 37, 1155–1162 (2019).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Pendleton, M. et al. Assembly and diploid architecture of an individual human genome via single-molecule technologies. Nat. Methods 12, 780–786 (2015).
Chaisson, M. J. et al. Resolving the complexity of the human genome using single-molecule sequencing. Nature 517, 608–611 (2015).
Lappalainen, I. et al. DbVar and DGVa: public archives for genomic structural variation. Nucleic Acids Res. 41, D936–D941 (2013).
MacDonald, J. R., Ziman, R., Yuen, R. K., Feuk, L. & Scherer, S. W. The database of genomic variants: a curated collection of structural variation in the human genome. Nucleic Acids Res. 42, D986–D992 (2014).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Preprint at bioRxiv https://doi.org/10.1101/563866 (2019).
Salzberg, S. L. Next-generation genome annotation: we still struggle to get it right. Genome Biol. 20, 92 (2019).
Pertea, M. et al. CHESS: a new human gene catalog curated from thousands of large-scale RNA sequencing experiments reveals extensive transcriptional noise. Genome Biol. 19, 208 (2018).
Audano, P. A. et al. Characterizing the major structural variant alleles of the human genome. Cell 176, 663–675.e619 (2019). The authors examined 15 PacBio-sequenced genomes to produce the largest long-read structural variant callset to date, and so discovered over 6 Mb of sequence per individual, on average, that was absent from the reference.
Duan, Z. et al. HUPAN: a pan-genome analysis pipeline for human genomes. Genome Biol. 20, 149 (2019). This study presents a pan-genome for a collection of Chinese individuals, as well as a proposed method to examine collections of human pan-genome data, provided that de novo assemblies can be performed on each individual genome.
Hehir-Kwa, J. Y. et al. A high-quality human reference panel reveals the complexity and distribution of genomic structural variants. Nat. Commun. 7, 12989 (2016).
Levy-Sakin, M. et al. Genome maps across 26 human populations reveal population-specific patterns of structural variation. Nat. Commun. 10, 1025 (2019).
Nagasaki, M. et al. Rare variant discovery by deep whole-genome sequencing of 1,070 Japanese individuals. Nat. Commun. 6, 8018 (2015).
Wong, K. H. Y., Levy-Sakin, M. & Kwok, P. Y. De novo human genome assemblies reveal spectrum of alternative haplotypes in diverse populations. Nat. Commun. 9, 3040 (2018).
Faber-Hammond, J. J. & Brown, K. H. Anchored pseudo-de novo assembly of human genomes identifies extensive sequence variation from unmapped sequence reads. Hum. Genet. 135, 727–740 (2016).
Boomsma, D. I. et al. The genome of the Netherlands: design, and project goals. Eur. J. Hum. Genet. 22, 221–227 (2014).
Genome of the Netherlands Consortium. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat. Genet. 46, 818–825 (2014).
Li, R. et al. Building the sequence map of the human pan-genome. Nat. Biotechnol. 28, 57–63 (2010). This study produces some of the first full assemblies of the human genomes of diverse populations. Asian and African genome assemblies are produced, and, based on the assemblies, the researchers estimate that a full human pan-genome might contain between 19 and 40 Mb of DNA missing from the reference.
Miga, K. H. Centromeric satellite DNAs: hidden sequence variation in the human population. Genes 10, 352 (2019).
Ameur, A. et al. De novo assembly of two Swedish genomes reveals missing segments from the human GRCh38 reference and improves variant calling of population-scale sequencing data. Genes 9, 486 (2018).
Huddleston, J. et al. Discovery and genotyping of structural variation from long-read haploid genome sequence data. Genome Res. 27, 677–685 (2017).
Shi, L. et al. Long-read sequencing and de novo assembly of a Chinese genome. Nat. Commun. 7, 12065 (2016).
Barra, V. & Fachinetti, D. The dark side of centromeres: types, causes and consequences of structural abnormalities implicating centromeric DNA. Nat. Commun. 9, 4340 (2018).
Church, D. M. et al. Modernizing reference genome assemblies. PLOS Biol. 9, e1001091 (2011).
Church, D. M. et al. Extending reference assembly models. Genome Biol. 16, 13 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Garrison, E. et al. Variation graph toolkit improves read mapping by representing genetic variation in the reference. Nat. Biotechnol. 36, 875–879 (2018). Vg is one of the leading methods to build and map reads to a variation graph, able to store a human pan-genome graph with ~180 Mb of variant sequences in under 4 Gb, with an index of ~63 Gb. Read alignment from a human genome to the variant graph can be performed in under an hour, although index and graph building are more time-consuming.
Rakocevic, G. et al. Fast and accurate genomic analyses using genome graphs. Nat. Genet. 51, 354–362 (2019).
Jain, C., Dilthey, A., Misra, S., Zhang, H. & Aluru, S. Accelerating sequence alignment to graphs. Preprint at bioRxiv https://doi.org/10.1101/651638 (2019).
Rautiainen, M., Mäkinen, V. & Marschall, T. Bit-parallel sequence-to-graph alignment. Bioinformatics 35, 3599–3607 (2019).
Iqbal, Z., Caccamo, M., Turner, I., Flicek, P. & McVean, G. De novo assembly and genotyping of variants using colored de Bruijn graphs. Nat. Genet. 44, 226–232 (2012).
Muggli, M. D. et al. Succinct colored de Bruijn graphs. Bioinformatics 33, 3181–3187 (2017).
Holley, G., Wittler, R. & Stoye, J. Bloom filter trie: an alignment-free and reference-free data structure for pan-genome storage. Algorithms Mol. Biol. 11, 3 (2016).
Hickey, G. et al. Genotyping structural variants in pangenome graphs using the vg toolkit. Preprint at bioRxiv https://doi.org/10.1101/654566 (2019).
Siren, J., Garrison, E., Novak, A. M., Paten, B. & Durbin, R. Haplotype-aware graph indexes. Bioinformatics https://doi.org/10.1093/bioinformatics/btz575 (2019).
Durbin, R. Efficient haplotype matching and storage using the positional Burrows–Wheeler transform (PBWT). Bioinformatics 30, 1266–1272 (2014).
Novak, A. M., Garrison, E. & Paten, B. A graph extension of the positional Burrows–Wheeler transform and its applications. Algorithms Mol. Biol. 12, 18 (2017).
Computational Pan-Genomics Consortium. Computational pan-genomics: status, promises and challenges. Brief. Bioinform. 19, 118–135 (2018).
Paten, B., Novak, A. M., Eizenga, J. M. & Garrison, E. Genome graphs and the evolution of genome inference. Genome Res. 27, 665–676 (2017).
Pritt, J., Chen, N. C. & Langmead, B. FORGe: prioritizing variants for graph genomes. Genome Biol. 19, 220 (2018).
Grytten, I., Rand, K. D., Nederbragt, A. J. & Sandve, G. K. Assessing graph-based read mappers against a novel baseline approach highlights strengths and weaknesses of the current generation of methods. Preprint at bioRxiv https://doi.org/10.1101/538066 (2019).
Kuhnle, A. et al. in Research in Computational Molecular Biology Vol. 11467 (ed. Cowen, L. J.) 158–173 (Springer, 2019).
Liu, Q., Shi, L. & Wang, K. Ethnicity-specific reference genome assembly by long-read sequencing. J. Mol. Genet. Med. 12, 1–3 (2018).
Graves-Lindsay, T. Reference genome improvement. National Human Genome Research Institute https://www.genome.wustl.edu/items/reference-genome-improvement/ (2018).
Sone, J. et al. Long-read sequencing identifies GGC repeat expansions in NOTCH2NLC associated with neuronal intranuclear inclusion disease. Nat. Genet. 51, 1215–1221 (2019). This research discovers a repeat expansion associated with disease by using long-read sequencing of affected families; the result highlights the limitations of approaches based on short reads to reference alignment and demonstrates that consideration of harder-to-detect variants can lead to clinically relevant discoveries.
Ballouz, S., Dobin, A. & Gillis, J. A. Is it time to change the reference genome? Genome Biol. 20, 159 (2019).
Gagie, T., Navarro, G. & Prezza, N. in Proceedings of the Twenty-Ninth Annual ACM–SIAM Symposium on Discrete Algorithms (ed. Czumaj, A.) 1459–1477 (Society for Industrial and Applied Mathematics, 2018).
Miga, K. H. et al. Telomere-to-telomere assembly a complete human X chromosome. Preprint at bioRxiv https://doi.org/10.1101/735928 (2019).
The International Human Genome Sequencing Consortium et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Schneider, V. A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Acknowledgements
This work was supported in part by the National Institutes of Health under grants R01-HL129239, R01-HG006677 and R35-GM130151.
Author information
Authors and Affiliations
Contributions
R.M.S. researched data for the article. Both authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks B. Paten and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Reference genomes
-
A reference genome is a genome sequence that is used as the representative for the species — typically, the most polished and complete sequence available for the species.
- Long-read sequencing
-
Sequencing reads on the order of 5–10 kb (Pacific Biosciences) or longer, in some cases up to 1–2 Mb in length (Oxford Nanopore Technologies). Long reads are more expensive to generate and have higher error than short reads (100–250 bp in length).
- Core genome
-
The genes or sequence shared between all individuals of a species (or other grouping).
- Dispensable genome
-
The genes or sequence not shared between all individuals of a species (or other grouping). Everything that is not a part of the core genome is part of the dispensable genome, and vice versa.
- Singleton
-
A sequence found only in a single individual in the study population or group.
- Transcriptome
-
The sequences of only the exon regions, typically inferred by sequencing RNA transcripts rather than DNA directly.
- Alignment
-
The process of computationally lining up sequencing reads to a genome (typically a reference) in order to determine where they are likely to have originated from in the genome.
- Assembly
-
The process of overlapping sequencing reads from many copies of a genome in order to piece together short sequences into longer sequences. Assembly is often performed for a whole genome, particularly when no reference is available for alignment, but it can be performed locally, as well as on regions or subsets of reads.
- Haplotype
-
A sequence on one of the two homologous chromosomes of an organism’s diploid genome. In humans, haplotypes are considered in contrast to using a single sequence to represent that sequence on both homologous copies of a chromosome.
- Admixed
-
An individual with genetic ancestry from multiple distinct populations.
Rights and permissions
About this article
Cite this article
Sherman, R.M., Salzberg, S.L. Pan-genomics in the human genome era. Nat Rev Genet 21, 243–254 (2020). https://doi.org/10.1038/s41576-020-0210-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-020-0210-7
This article is cited by
-
A comprehensive benchmark of graph-based genetic variant genotyping algorithms on plant genomes for creating an accurate ensemble pipeline
Genome Biology (2024)
-
Three near-complete genome assemblies reveal substantial centromere dynamics from diploid to tetraploid in Brachypodium genus
Genome Biology (2024)
-
A sequence-aware merger of genomic structural variations at population scale
Nature Communications (2024)
-
A review of the pangenome: how it affects our understanding of genomic variation, selection and breeding in domestic animals?
Journal of Animal Science and Biotechnology (2023)
-
Comparing methods for constructing and representing human pangenome graphs
Genome Biology (2023)