Abstract
Circular RNAs (circRNAs) are covalently closed, endogenous biomolecules in eukaryotes with tissue-specific and cell-specific expression patterns, whose biogenesis is regulated by specific cis-acting elements and trans-acting factors. Some circRNAs are abundant and evolutionarily conserved, and many circRNAs exert important biological functions by acting as microRNA or protein inhibitors (‘sponges’), by regulating protein function or by being translated themselves. Furthermore, circRNAs have been implicated in diseases such as diabetes mellitus, neurological disorders, cardiovascular diseases and cancer. Although the circular nature of these transcripts makes their detection, quantification and functional characterization challenging, recent advances in high-throughput RNA sequencing and circRNA-specific computational tools have driven the development of state-of-the-art approaches for their identification, and novel approaches to functional characterization are emerging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J. & Kleinschmidt, A. K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl Acad. Sci. USA 73, 3852–3856 (1976).
Hsu, M. T. & Coca-Prados, M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature 280, 339–340 (1979).
Cocquerelle, C., Mascrez, B., Hetuin, D. & Bailleul, B. Mis-splicing yields circular RNA molecules. FASEB J 7, 155–160 (1993).
Capel, B. et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73, 1019–1030 (1993).
Wang, P. L. et al. Circular RNA is expressed across the eukaryotic tree of life. PLOS ONE 9, e90859 (2014).
Ivanov, A. et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 10, 170–177 (2015).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L. & Brown, P. O. Cell-type specific features of circular RNA expression. PLOS Genet. 9, e1003777 (2013).
Westholm, J. O. et al. Genome-wide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 9, 1966–1980 (2014).
Maass, P. G. et al. A map of human circular RNAs in clinically relevant tissues. J. Mol. Med. 95, 1179–1189 (2017).
Xia, S. et al. Comprehensive characterization of tissue-specific circular RNAs in the human and mouse genomes. Brief. Bioinform. 18, 984–992 (2017).
Guo, J. U., Agarwal, V., Guo, H. & Bartel, D. P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 15, 409 (2014).
Zhang, X. O. et al. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 26, 1277–1287 (2016).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013). This article identifies circRNAs as a large class of post-transcriptional regulators that compete with other RNAs for binding by miRNAs and RBPs.
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLOS ONE 7, e30733 (2012). This article reveals that the production of circRNA is a general feature of gene expression in human cells.
Li, Z. et al. Exon–intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 22, 256–264 (2015).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Mol. Cell 51, 792–806 (2013).
Veno, M. T. et al. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol. 16, 245 (2015).
Rybak-Wolf, A. et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol. Cell 58, 870–885 (2015). This article provides a circRNA brain expression atlas and shows that circRNA expression correlates negatively with the expression of ADAR1.
Enuka, Y. et al. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 44, 1370–1383 (2016).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013). This study is the first to functionally characterize naturally expressed circRNAs.
Izuogu, O. G. et al. Analysis of human ES cell differentiation establishes that the dominant isoforms of the lncRNAs RMST and FIRRE are circular. BMC Genomics 19, 276 (2018).
Szabo, L. et al. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol. 16, 126 (2015).
You, X. et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 18, 603–610 (2015).
van Rossum, D., Verheijen, B. M. & Pasterkamp, R. J. Circular RNAs: novel regulators of neuronal development. Front. Mol. Neurosci. 9, 74 (2016).
Bachmayr-Heyda, A. et al. Correlation of circular RNA abundance with proliferation — exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci. Rep. 5, 8057 (2015). This study is the first to report a global reduction in circRNA abundance in cancer relative to normal tissues and presents a model for how circRNAs may accumulate in non-proliferating cells.
Moldovan, L.-I. et al. High-throughput RNA sequencing from paired lesional- and non-lesional skin reveals major alterations in the psoriasis circRNAome. Preprint at bioRxiv https://doi.org/10.1101/581066 (2019).
Fang, Y. et al. Screening of circular RNAs and validation of circANKRD36 associated with inflammation in patients with type 2 diabetes mellitus. Int. J. Mol. Med. 42, 1865–1874 (2018).
Hanan, M., Soreq, H. & Kadener, S. CircRNAs in the brain. RNA Biol. 14, 1028–1034 (2017).
Holdt, L. M. et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 7, 12429 (2016).
Kristensen, L. S., Hansen, T. B., Veno, M. T. & Kjems, J. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene 37, 555–565 (2018).
Li, H. et al. Comprehensive circular RNA profiles in plasma reveals that circular RNAs can be used as novel biomarkers for systemic lupus erythematosus. Clin. Chim. Acta 480, 17–25 (2018).
Aufiero, S., Reckman, Y.J., Pinto, Y.M. & Creemers, E.E. Circular RNAs open a new chapter in cardiovascular biology. Nat. Rev. Cardiol. 16, 503–514 (2019).
Vo, J. N. et al. The landscape of circular RNA in. Cancer. Cell 176, 869–881.e813 (2019).
Cortes-Lopez, M. et al. Global accumulation of circRNAs during aging in Caenorhabditis elegans. BMC Genomics 19, 8 (2018).
Gruner, H., Cortes-Lopez, M., Cooper, D. A., Bauer, M. & Miura, P. CircRNA accumulation in the aging mouse brain. Sci. Rep. 6, 38907 (2016).
Fu, X. D. & Ares, M. Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet. 15, 689–701 (2014).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 56, 55–66 (2014).
Starke, S. et al. Exon circularization requires canonical splice signals. Cell Rep. 10, 103–111 (2015).
Liang, D. et al. The output of protein-coding genes shifts to circular RNAs when the pre-mRNA processing machinery is limiting. Mol. Cell 68, 940–954.e943 (2017).
Kramer, M. C. et al. Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev. 29, 2168–2182 (2015).
Kelly, S., Greenman, C., Cook, P. R. & Papantonis, A. Exon skipping is correlated with exon circularization. J. Mol. Biol. 427, 2414–2417 (2015).
Zhang, X. O. et al. Complementary sequence-mediated exon circularization. Cell 159, 134–147 (2014). This article shows that exon circularization is often dependent on flanking intronic complementary sequences.
Conn, S. J. et al. The RNA binding protein quaking regulates formation of circRNAs. Cell 160, 1125–1134 (2015).
Errichelli, L. et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat. Commun. 8, 14741 (2017).
Eisenberg, E. & Levanon, E. Y. A-to-I RNA editing — immune protector and transcriptome diversifier. Nat. Rev. Genet. 19, 473–490 (2018).
Koh, H. R., Xing, L., Kleiman, L. & Myong, S. Repetitive RNA unwinding by RNA helicase A facilitates RNA annealing. Nucleic Acids Res. 42, 8556–8564 (2014).
Aktas, T. et al. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 544, 115–119 (2017). This study shows that DHX9 suppresses RNA processing defects, including exon circularization, originating from the Alu invasion of the human genome.
Wen, X. et al. NF90 exerts antiviral activity through regulation of PKR phosphorylation and stress granules in infected cells. J. Immunol. 192, 3753–3764 (2014).
Li, X. et al. Coordinated circRNA biogenesis and function with NF90/NF110 in viral infection. Mol. Cell 67, 214–227 (2017).
Eger, N., Schoppe, L., Schuster, S., Laufs, U. & Boeckel, J. N. Circular RNA splicing. Adv. Exp. Med. Biol. 1087, 41–52 (2018).
Ferreira, H. J. et al. Circular RNA CpG island hypermethylation-associated silencing in human cancer. Oncotarget 9, 29208–29219 (2018).
Kristensen, L. S., Okholm, T. L. H., Veno, M. T. & Kjems, J. Circular RNAs are abundantly expressed and upregulated during human epidermal stem cell differentiation. RNA Biol. 15, 280–291 (2018).
Shukla, S. et al. CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing. Nature 479, 74–79 (2011).
Bentley, D. L. Coupling mRNA processing with transcription in time and space. Nat. Rev. Genet. 15, 163–175 (2014).
Rinaldi, L. et al. Dnmt3a and Dnmt3b associate with enhancers to regulate human epidermal stem cell homeostasis. Cell Stem Cell 19, 491–501 (2016).
Chen, N. et al. A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol. 19, 218 (2018).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Zhang, Y. et al. The biogenesis of nascent circular RNAs. Cell Rep. 15, 611–624 (2016).
Huang, C., Liang, D., Tatomer, D. C. & Wilusz, J. E. A length-dependent evolutionarily conserved pathway controls nuclear export of circular RNAs. Genes Dev. 32, 639–644 (2018). This study provides the first data on how circRNAs are exported from the nucleus to the cytoplasm.
Park, O. H. et al. Endoribonucleolytic cleavage of m(6)A-containing RNAs by RNase P/MRP complex. Mol. Cell 74, 494–507 (2019).
Liu, C. X. et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell 177, 865–880 (2019). This study indicates that circRNAs are important players in innate immunity as they form duplexes of 16–26 bp, which bind and regulate PKR activity.
Hansen, T. B. et al. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 30, 4414–4422 (2011).
Kleaveland, B., Shi, C. Y., Stefano, J. & Bartel, D. P. A network of noncoding regulatory RNAs acts in the mammalian brain. Cell 174, 350–362 (2018).
Hammond, S. M., Boettcher, S., Caudy, A. A., Kobayashi, R. & Hannon, G. J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 293, 1146–1150 (2001).
Lasda, E. & Parker, R. Circular RNAs co-precipitate with extracellular vesicles: a possible mechanism for circRNA clearance. PLOS ONE 11, e0148407 (2016).
Dou, Y. et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci. Rep. 6, 37982 (2016).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 25, 981–984 (2015).
Preusser, C. et al. Selective release of circRNAs in platelet-derived extracellular vesicles. J. Extracell. Vesicles 7, 1424473 (2018).
Glazar, P., Papavasileiou, P. & Rajewsky, N. circBase: a database for circular RNAs. RNA 20, 1666–1670 (2014).
Szabo, L. & Salzman, J. Detecting circular RNAs: bioinformatic and experimental challenges. Nat. Rev. Genet. 17, 679–692 (2016).
Rigatti, R., Jia, J. H., Samani, N. J. & Eperon, I. C. Exon repetition: a major pathway for processing mRNA of some genes is allele-specific. Nucleic Acids Res. 32, 441–446 (2004).
Chuang, T.J. et al. Integrative transcriptome sequencing reveals extensive alternative trans-splicing and cis-backsplicing in human cells. Nucleic Acids Res. 46, 3671-3691 (2018).
Dahl, M. et al. Enzyme-free digital counting of endogenous circular RNA molecules in B-cell malignancies. Lab. Invest. 98, 1657–1669 (2018).
Jeck, W. R. & Sharpless, N. E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 32, 453–461 (2014).
Li, X., Yang, L. & Chen, L. L. The biogenesis, functions, and challenges of circular RNAs. Mol. Cell 71, 428–442 (2018).
Hansen, T. B. Improved circRNA identification by combining prediction algorithms. Front. Cell Dev. Biol. 6, 20 (2018).
Gao, Y., Zhang, J. & Zhao, F. Circular RNA identification based on multiple seed matching. Brief. Bioinform. 19, 803-810 (2017).
Hansen, T. B., Veno, M. T., Damgaard, C. K. & Kjems, J. Comparison of circular RNA prediction tools. Nucleic Acids Res. 44, e58 (2016).
Chen, X. et al. PRMT5 circular RNA promotes metastasis of urothelial carcinoma of the bladder through sponging miR-30c to induce epithelial–mesenchymal transition. Clin. Cancer Res. 24, 6319–6330 (2018).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009).
Chen, D.-F., Zhang, L.-J., Tan, K. & Jing, Q. Application of droplet digital PCR in quantitative detection of the cell-free circulating circRNAs. Biotechnol. Biotechnol. Equip. 32, 116-123 (2018).
Li, T. et al. Plasma circular RNA profiling of patients with gastric cancer and their droplet digital RT-PCR detection. J. Mol. Med. 96, 85–96 (2018).
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 26, 317–325 (2008).
Baker, A. M. et al. Robust RNA-based in situ mutation detection delineates colorectal cancer subclonal evolution. Nat. Commun. 8, 1998 (2017).
Erben, L., He, M. X., Laeremans, A., Park, E. & Buonanno, A. A novel ultrasensitive in situ hybridization approach to detect short sequences and splice variants with cellular resolution. Mol. Neurobiol. 55, 6169–6181 (2018).
Granados-Riveron, J. T. & Aquino-Jarquin, G. CRISPR–Cas13 precision transcriptome engineering in cancer. Cancer Res. 78, 4107–4113 (2018).
Abudayyeh, O. O. et al. RNA targeting with CRISPR-Cas13. Nature 550, 280–284 (2017).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Piwecka, M. et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357 (2017). This study presents the first circRNA knockout mouse model, which indicates that interactions between miRNA and circRNA are important for normal brain function.
Zheng, Q. et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 7, 11215 (2016).
Dudekula, D. B. et al. CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol. 13, 34–42 (2016).
Abdelmohsen, K. et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by circPABPN1. RNA Biol. 14, 361–369 (2017).
Zeng, Y. et al. A circular RNA binds to and activates AKT phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics 7, 3842–3855 (2017).
Du, W. W. et al. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 44, 2846–2858 (2016).
Du, W. W. et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 24, 357–370 (2017).
Legnini, I. et al. circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 66, 22–37 (2017).
Pamudurti, N. R. et al. Translation of circRNAs. Mol. Cell 66, 9–21.e27 (2017).
Yang, Y. et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res 27, 626–641 (2017).
Yang, Y. et al. Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J. Natl Cancer Inst. 110, 304–315 (2018).
Zhang, M. et al. A novel protein encoded by the circular form of the SHPRH gene suppresses glioma tumorigenesis. Oncogene 37, 1805–1814 (2018).
Zhang, M. et al. A peptide encoded by circular form of LINC-PINT suppresses oncogenic transcriptional elongation in glioblastoma. Nat. Commun. 9, 4475 (2018).
Stagsted, L.V., Nielsen, K.M., Daugaard, I. & Hansen, T. B. Noncoding AUG circRNAs constitute an abundant and conserved subclass of circles. Life Sci. Alliance 2, e201900398 (2019).
Denzler, R., Agarwal, V., Stefano, J., Bartel, D. P. & Stoffel, M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol. Cell 54, 766–776 (2014).
Thomson, D. W. & Dinger, M. E. Endogenous microRNA sponges: evidence and controversy. Nat. Rev. Genet. 17, 272–283 (2016).
Weng, W. et al. Circular RNA ciRS-7 — a promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin. Cancer Res. 23, 3918-3928 (2017).
Yu, L. et al. The circular RNA Cdr1as act as an oncogene in hepatocellular carcinoma through targeting miR-7 expression. PLOS ONE 11, e0158347 (2016).
Yu, C. Y. et al. The circular RNA circBIRC6 participates in the molecular circuitry controlling human pluripotency. Nat. Commun. 8, 1149 (2017).
Barbollat-Boutrand, L. et al. MicroRNA-23b-3p regulates human keratinocyte differentiation through repression of TGIF1 and activation of the TGF-ss-SMAD2 signalling pathway. Exp. Dermatol. 26, 51–57 (2017).
Hsiao, K.Y. et al. Non-coding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 77, 2339-2350 (2017).
Verduci, L. et al. The oncogenic role of circPVT1 in head and neck squamous cell carcinoma is mediated through the mutant p53/YAP/TEAD transcription-competent complex. Genome Biol. 18, 237 (2017).
Xia, P. et al. A circular RNA protects dormant hematopoietic stem cells from DNA sensor cGAS-mediated exhaustion. Immunity 48, 688–701.e687 (2018).
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS-STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Sato, T. et al. Interferon regulatory factor-2 protects quiescent hematopoietic stem cells from type I interferon-dependent exhaustion. Nat. Med. 15, 696–700 (2009).
Essers, M. A. et al. IFNα activates dormant haematopoietic stem cells in vivo. Nature 458, 904–908 (2009).
Schlee, M. & Hartmann, G. Discriminating self from non-self in nucleic acid sensing. Nat. Rev. Immunol. 16, 566–580 (2016).
Chen, Y. G. et al. Sensing self and foreign circular RNAs by intron identity. Mol. Cell 67, 228–238 (2017). This study indicates that circRNAs play important roles in immunobiology and that self–non-self discrimination depends on the introns that flank circRNAs.
Wesselhoeft, R. A. et al. RNA circularization diminishes immunogenicity and can extend translation duration in vivo. Mol. Cell 74, 508–520 (2019).
Abe, N. et al. Rolling circle translation of circular RNA in living human cells. Sci. Rep. 5, 16435 (2015).
Meyer, K. D. et al. 5′ UTR m(6)A promotes cap-independent translation. Cell 163, 999–1010 (2015).
Zhou, J. et al. Dynamic m(6)A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015).
Chen, C. Y. & Sarnow, P. Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Science 268, 415–417 (1995).
Perriman, R. & Ares, M. Jr. Circular mRNA can direct translation of extremely long repeating-sequence proteins in vivo. RNA 4, 1047–1054 (1998).
Wang, Y. & Wang, Z. Efficient backsplicing produces translatable circular mRNAs. RNA 21, 172–179 (2015).
Chen, X. et al. circRNADb: a comprehensive database for human circular RNAs with protein-coding annotations. Sci. Rep. 6, 34985 (2016).
Ng, S. Y., Bogu, G. K., Soh, B. S. & Stanton, L. W. The long noncoding RNA RMST interacts with SOX2 to regulate neurogenesis. Mol. Cell 51, 349–359 (2013).
Ng, S. Y., Johnson, R. & Stanton, L. W. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J. 31, 522–533 (2012).
Barra, J. & Leucci, E. Probing long non-coding RNA–protein interactions. Front. Mol. Biosci. 4, 45 (2017).
Du, W. W. et al. Identifying and characterizing circRNA–protein interaction. Theranostics 7, 4183–4191 (2017).
Schneider, T. et al. circRNA–protein complexes: IMP3 protein component defines subfamily of circRNPs. Sci. Rep. 6, 31313 (2016).
Bramsen, J. B. et al. A large-scale chemical modification screen identifies design rules to generate siRNAs with high activity, high stability and low toxicity. Nucleic Acids Res. 37, 2867–2881 (2009).
Bramsen, J. B. et al. Improved silencing properties using small internally segmented interfering RNAs. Nucleic Acids Res. 35, 5886–5897 (2007).
de Bruyns, A., Geiling, B. & Dankort, D. Construction of modular lentiviral vectors for effective gene expression and knockdown. Methods Mol. Biol. 1448, 3–21 (2016).
Herrera-Carrillo, E., Harwig, A. & Berkhout, B. Influence of the loop size and nucleotide composition on AgoshRNA biogenesis and activity. RNA Biol. 14, 1559–1569 (2017).
Liu, Y. P., Schopman, N. C. & Berkhout, B. Dicer-independent processing of short hairpin RNAs. Nucleic Acids Res. 41, 3723–3733 (2013).
Barrett, S. P., Parker, K. R., Horn, C., Mata, M. & Salzman, J. ciRS-7 exonic sequence is embedded in a long non-coding RNA locus. PLOS Genet. 13, e1007114 (2017).
Barrett, S. P. & Salzman, J. Circular RNAs: analysis, expression and potential functions. Development 143, 1838–1847 (2016).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 28, 2233–2247 (2014).
Schmidt, C. A., Noto, J. J., Filonov, G. S. & Matera, A. G. A method for expressing and imaging abundant, stable, circular RNAs in vivo using tRNA splicing. Methods Enzymol. 572, 215–236 (2016).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Zhu, Q., Shah, S., Dries, R., Cai, L. & Yuan, G.C. Identification of spatially associated subpopulations by combining scRNAseq and sequential fluorescence in situ hybridization data. Nat. Biotechnol. 36, 1183–1190 (2018).
Fan, X. et al. Single-cell RNA-seq transcriptome analysis of linear and circular RNAs in mouse preimplantation embryos. Genome Biol. 16, 148 (2015).
Verboom, K. et al. SMARTer single cell total RNA sequencing. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz535 (2019).
Amaria, R. N. et al. Neoadjuvant immune checkpoint blockade in high-risk resectable melanoma. Nat. Med. 24, 1649–1654 (2018).
Blank, C.U. et al. Neoadjuvant versus adjuvant ipilimumab plus nivolumab in macroscopic stage III melanoma. Nat. Med. 24, 1655–1661 (2018).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Li, R. C. et al. CiRS-7 promotes growth and metastasis of esophageal squamous cell carcinoma via regulation of miR-7/HOXB13. Cell Death Dis. 9, 838 (2018).
Barbagallo, D. et al. circSMARCA5 inhibits migration of glioblastoma multiforme cells by regulating a molecular axis involving splicing factors SRSF1/SRSF3/PTB. Int. J. Mol. Sci. 19, E480 (2018).
Begum, S., Yiu, A., Stebbing, J. & Castellano, L. Novel tumour suppressive protein encoded by circular RNA, circ-SHPRH, in glioblastomas. Oncogene 37, 4055–4057 (2018).
Zeng, K. et al. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 9, 417 (2018).
Okholm, T. L. H. et al. Circular RNA expression is abundant and correlated to aggressiveness in early-stage bladder cancer. NPJ Genom. Med. 2, 36 (2017).
Li, Y. et al. CircHIPK3 sponges miR-558 to suppress heparanase expression in bladder cancer cells. EMBO Rep. 18, 1646–1659 (2017).
Smid, M. et al. The circular RNome of primary breast cancer. Genome Res. 29, 356–366 (2019).
Bahn, J. H. et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 61, 221–230 (2015).
Memczak, S., Papavasileiou, P., Peters, O. & Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLOS ONE 10, e0141214 (2015).
Stoll, L. et al. Circular RNAs as novel regulators of beta-cell functions in normal and disease conditions. Mol. Metab. 9, 69–83 (2018).
Xu, H., Guo, S., Li, W. & Yu, P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci. Rep. 5, 12453 (2015).
Wang, K. et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 37, 2602–2611 (2016).
Miao, Q. et al. RNA-seq of circular RNAs identified circPTPN22 as a potential new activity indicator in systemic lupus erythematosus. Lupus 28, 520–528 (2019).
Lukiw, W. J. Circular RNA (circRNA) in Alzheimer’s disease (AD). Front. Genet. 4, 307 (2013).
Shi, Z. et al. The circular RNA ciRS-7 promotes APP and BACE1 degradation in an NF-kappaB-dependent manner. FEBS J. 284, 1096–1109 (2017).
Kumar, L. et al. Functional characterization of novel circular RNA molecule, circzip-2 and its synthesizing gene zip-2 in C. elegans model of Parkinson’s disease. Mol. Neurobiol. 55, 6914–6926 (2018).
Tagawa, T. et al. Discovery of Kaposi’s sarcoma herpesvirus-encoded circular RNAs and a human antiviral circular RNA. Proc. Natl Acad. Sci. USA (2018).
Acknowledgements
This work was supported by a grant to L.S.K. from the Carlsberg Foundation (CF16-0087).
Author information
Authors and Affiliations
Contributions
L.S.K. researched data for the article. All of the authors wrote and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
L.S.K. is on the advisory board of BioXpedia A/S, which provides services using some of the commercially available techniques mentioned in this article. All of the other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks J. E. Wilusz and L. Yang for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Viroids
-
Small, circular, single-stranded RNA molecules (246–401 nucleotides) that are uncoated and do not encode proteins. They are pathogenic to higher plants.
- Alternative splicing
-
A mechanism by which different forms of a mature RNA can be generated from the same primary RNA by the use of different splice sites.
- Alu elements
-
The most abundant primate-specific DNA transposable elements. These are highly repetitive and composed of ~300 bases.
- Innate immune system
-
The first-line host defence to confine and combat infection.
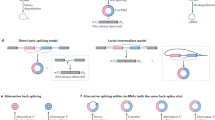
- Lariat formation
-
Splicing intermediates formed when the 5′ end of the intron being removed is joined to the branch-point adenosine with a 2′,5′-phosphodiester linkage, creating a lasso-shaped molecule.
- Debranching
-
The hydrolysis of 2′,5′-phosphodiester bonds in intron lariats by the lariat debranching enzyme, encoded by DBR1. This hydrolysis converts the intron lariat into a linearized intron, which is subsequently degraded.
- Backsplice junction (BSJ) region
-
The only region of a circular RNA (circRNA) that is distinct from the corresponding linear RNA at the primary sequence level. It is generated through the backsplicing event that generates the circRNA and is composed of a canonical 5′ splice site sequence joined to an upstream 3′ splice site sequence.
- Droplet digital PCR
-
A quantitative PCR method that uses microfluidics (oil–water separation) to amplify individual nucleic acids within individual droplets in the same tube. By measuring the fluorescence signal in each droplet, the copy number of the target molecule can be determined.
- Long-term haematopoietic stem cells
-
Haematopoietic stem cells that are defined by specific surface markers and can self-renew infinitely and differentiate to all cell types within the blood and immune system.
- Pattern recognition receptor
-
Host receptors that recognize molecules typical for pathogens. Upon recognition of pathogen-associated patterns, the innate immune system is activated.
- Internal ribosome entry sites
-
Structural RNA elements that enable the initiation of a cap-independent translation.
- Argonaute-crosslinking and immunoprecipitation
-
A method to identify and map microRNAs bound to AGO proteins and the target transcripts associated with them.
- Polysome profiling
-
A technique to study the translatome based on a sucrose-gradient separation of untranslated and translated RNA transcripts; translated RNA transcripts are associated with polysomes.
- Ribosome footprinting
-
A technique to measure translation by the high-throughput sequencing of ribosome-protected RNA fragments, which determines the position of ribosomes at codon resolution.
- Locked nucleic acids
-
A modified RNA nucleotide in which the ribose moiety is modified with a methylene bridge connecting the 2′ oxygen and 4′ carbon. It has an increased affinity for its complementary nucleotide relative to traditional DNA or RNA oligonucleotides.
- Unlocked nucleic acids
-
An acyclic RNA nucleotide that lacks the C2′–C3′ bond of the ribose moiety found in traditional RNA. It has a decreased affinity for its complementary nucleotide relative to traditional DNA or RNA oligonucleotides.
- Passenger disabled siRNA
-
A small interfering RNA (siRNA) in which an intact antisense strand is complemented with a fragmented sense strand. These siRNAs, which are known as small internally segmented interfering RNAs, eliminate off-target effects by only allowing the functional incorporation of the antisense strand into the RNA-induced silencing complex (RISC).
- AgoshRNAs
-
Short hairpin RNAs that are characterized by a relatively short base-paired stem, which allows them to avoid cleavage by Dicer. Instead, they are processed by the slicer activity of Ago2, which creates a single guide RNA strand that targets a specific RNA for degradation and has less off-target effects as no passenger strand is created.
Rights and permissions
About this article
Cite this article
Kristensen, L.S., Andersen, M.S., Stagsted, L.V.W. et al. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20, 675–691 (2019). https://doi.org/10.1038/s41576-019-0158-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-019-0158-7
This article is cited by
-
A novel peptide PDHK1-241aa encoded by circPDHK1 promotes ccRCC progression via interacting with PPP1CA to inhibit AKT dephosphorylation and activate the AKT-mTOR signaling pathway
Molecular Cancer (2024)
-
Hypoxia-induced circRNAs encoded by PPARA are highly expressed in human cardiomyocytes and are potential clinical biomarkers of acute myocardial infarction
European Journal of Medical Research (2024)
-
Circ_0000376 regulates miR-577/HK2/LDHA signaling pathway to promote the growth, invasion and glycolysis of osteosarcoma
Journal of Orthopaedic Surgery and Research (2024)
-
EIF4A3-mediated biogenesis of circSTX6 promotes bladder cancer metastasis and cisplatin resistance
Journal of Experimental & Clinical Cancer Research (2024)
-
Role of a novel circRNA-CGNL1 in regulating pancreatic cancer progression via NUDT4–HDAC4–RUNX2–GAMT-mediated apoptosis
Molecular Cancer (2024)