Abstract
Genomic imprinting, the monoallelic and parent-of-origin-dependent expression of a subset of genes, is required for normal development, and its disruption leads to human disease. Imprinting defects can involve isolated or multilocus epigenetic changes that may have no evident genetic cause, or imprinting disruption can be traced back to alterations of cis-acting elements or trans-acting factors that control the establishment, maintenance and erasure of germline epigenetic imprints. Recent insights into the dynamics of the epigenome, including the effect of environmental factors, suggest that the developmental outcomes and heritability of imprinting disorders are influenced by interactions between the genome, the epigenome and the environment in germ cells and early embryos.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harb. Perspect. Biol. 6, a018382 (2014).
Onuchic, V. et al. Allele-specific epigenome maps reveal sequence-dependent stochastic switching at regulatory loci. Science 361, eaar3146 (2018).
Baran, Y. et al. The landscape of genomic imprinting across diverse adult human tissues. Genome Res. 25, 927–936 (2015).
Court, F. et al. Genome-wide parent-of-origin DNA methylation analysis reveals the intricacies of human imprinting and suggests a germline methylation-independent mechanism of establishment. Genome Res. 24, 554–569 (2014). A full description of allelic DMRs in humans and the first report of tissue-specific imprinting centres are presented.
Smith, Z. D. et al. DNA methylation dynamics of the human preimplantation embryo. Nature 511, 611–615 (2014). The methylation landscape in human gametes and pre-implantation embryos is described.
Peters, J. The role of genomic imprinting in biology and disease: an expanding view. Nat. Rev. Genet. 15, 517–530 (2014).
Eggermann, T. et al. Imprinting disorders: a group of congenital disorders with overlapping patterns of molecular changes affecting imprinted loci. Clin. Epigenet. 7, 123 (2015).
Horsthemke, B. Mechanisms of imprint dysregulation. Am. J. Med. Genet. 154C, 321–328 (2010).
Haig, D. Colloquium papers: Transfers and transitions: parent-offspring conflict, genomic imprinting, and the evolution of human life history. Proc. Natl Acad. Sci. USA 107 (Suppl. 1), 1731–1735 (2010).
Eggermann, K. et al. EMQN best practice guidelines for the molecular genetic testing and reporting of chromosome 11p15 imprinting disorders: Silver-Russell and Beckwith-Wiedemann syndrome. Eur. J. Hum. Genet. 24, 1377–1387 (2016).
Sanli, I. & Feil, R. Chromatin mechanisms in the developmental control of imprinted gene expression. Int. J. Biochem. Cell Biol. 67, 139–147 (2015).
Monk, D. et al. Recommendations for a nomenclature system for reporting methylation aberrations in imprinted domains. Epigenetics 13, 117–121 (2018).
Singh, P. et al. Chromosome-wide analysis of parental allele-specific chromatin and DNA methylation. Mol. Cell. Biol. 31, 1757–1770 (2011).
Riso, V. et al. ZFP57 maintains the parent-of-origin-specific expression of the imprinted genes and differentially affects non-imprinted targets in mouse embryonic stem cells. Nucleic Acids Res. 44, 8165–8178 (2016).
Kota, S. K. et al. ICR noncoding RNA expression controls imprinting and DNA replication at the Dlk1-Dio3 domain. Dev. Cell 31, 19–33 (2014).
Rougeulle, C., Glatt, H. & Lalande, M. The Angelman syndrome candidate gene, UBE3A/E6-AP, is imprinted in brain. Nat. Genet. 17, 14–15 (1997).
Vu, T. H. & Hoffman, A. R. Imprinting of the Angelman syndrome gene, UBE3A, is restricted to brain. Nat. Genet. 17, 12–13 (1997).
Travers, M. E. et al. Insights into the molecular mechanism for type 2 diabetes susceptibility at the KCNQ1 locus from temporal changes in imprinting status in human islets. Diabetes 62, 987–992 (2013).
Hanna, C. W. et al. Pervasive polymorphic imprinted methylation in the human placenta. Genome Res. 26, 756–767 (2016).
Zink, F. et al. Insights into imprinting from parent-of-origin phased methylomes and transcriptomes. Nat. Genet. 50, 1542–1552 (2018). The partial phenotypes associated with parent-of-origin-specific genotypes that influence imprinted gene expression are described.
Sanchez-Delgado, M. et al. Human oocyte-derived methylation differences persist in the placenta revealing widespread transient imprinting. PLOS Genet. 12, e1006427 (2016). Oocyte-derived methylation and polymorphic transient DMRs are demonstrated.
Tan, L., Xing, D., Chang, C. H., Li, H. & Xie, X. S. Three-dimensional genome structures of single diploid human cells. Science 361, 924–928 (2018). Allelic contacts and novel CTCF-mediated loops at the H19 – Igf2 locus are identified by single-cell Hi-C profiling.
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Kanduri, C. Long noncoding RNAs: lessons from genomic imprinting. Biochim. Biophys. Acta 1859, 102–111 (2016).
Niemczyk, M. et al. Imprinted chromatin around DIRAS3 regulates alternative splicing of GNG12-AS1, a long noncoding RNA. Am. J. Hum. Genet. 93, 224–235 (2013).
Inoue, A., Jiang, L., Lu, F., Suzuki, T. & Zhang, Y. Maternal H3K27me3 controls DNA methylation-independent imprinting. Nature 547, 419–424 (2017). Maternal H3K27me3 controls DNA-methylation-independent imprinting in early mouse embryos independent of DMRs.
Hutter, B., Helms, V. & Paulsen, M. Tandem repeats in the CpG islands of imprinted genes. Genomics 88, 323–332 (2006).
Beygo, J. et al. The molecular function and clinical phenotype of partial deletions of the IGF2/H19 imprinting control region depends on the spatial arrangement of the remaining CTCF-binding sites. Hum. Mol. Genet. 22, 544–557 (2013).
Sparago, A., Cerrato, F. & Riccio, A. Is ZFP57 binding to H19/IGF2:IG-DMR affected in Silver-Russell syndrome? Clin. Epigenet. 10, 23 (2018).
De Crescenzo, A. et al. A novel microdeletion in the IGF2/H19 imprinting centre region defines a recurrent mutation mechanism in familial Beckwith-Wiedemann syndrome. Eur. J. Med. Genet. 54, e451–e454 (2011).
Soares, M. L. et al. Targeted deletion of a 170-kb cluster of LINE-1 repeats and implications for regional control. Genome Res. https://doi.org/10.1101/gr.221366.117 (2018).
Varrault, A. et al. Zac1 regulates an imprinted gene network critically involved in the control of embryonic growth. Dev. Cell 11, 711–722 (2006).
Gabory, A. et al. H19 acts as a trans regulator of the imprinted gene network controlling growth in mice. Development 136, 3413–3421 (2009).
Stelzer, Y., Sagi, I., Yanuka, O., Eiges, R. & Benvenisty, N. The noncoding RNA IPW regulates the imprinted DLK1-DIO3 locus in an induced pluripotent stem cell model of Prader-Willi syndrome. Nat. Genet. 46, 551–557 (2014).
Girardot, M., Cavaille, J. & Feil, R. Small regulatory RNAs controlled by genomic imprinting and their contribution to human disease. Epigenetics 7, 1341–1348 (2012).
Court, F. et al. Genome-wide allelic methylation analysis reveals disease-specific susceptibility to multiple methylation defects in imprinting syndromes. Hum. Mutat. 34, 595–602 (2013).
Docherty, L. E. et al. Mutations in NLRP5 are associated with reproductive wastage and multilocus imprinting disorders in humans. Nat. Commun. 6, 8086 (2015). NLRP5 mutations in MLIDs are identified.
Azzi, S. et al. A prospective study validating a clinical scoring system and demonstrating phenotypical-genotypical correlations in Silver-Russell syndrome. J. Med. Genet. 52, 446–453 (2015).
Begemann, M. et al. Maternal variants in NLRP and other maternal effect proteins are associated with multilocus imprinting disturbance in offspring. J. Med. Genet. 55, 497–504 (2018). Deleterious variants in maternal-effect genes associated with the SCMC in individuals with MLIDs are identified.
Hackett, J. A. et al. Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine. Science 339, 448–452 (2013). Erasure of CpG methylation in mouse PGCs occurs via conversion to 5hmC that is driven by high levels of TET1 and TET2.
Seisenberger, S. et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 48, 849–862 (2012).
Guo, F. et al. The transcriptome and DNA methylome landscapes of human primordial germ cells. Cell 161, 1437–1452 (2015). Paper presents first genome-wide descriptions of the transcriptome and the DNA methylome landscapes of human PGCs.
Tang, W. W. et al. A unique gene regulatory network resets the human germline epigenome for development. Cell 161, 1453–1467 (2015).
Gkountela, S. et al. DNA demethylation dynamics in the human prenatal germline. Cell 161, 1425–1436 (2015).
Dawlaty, M. M. et al. Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev. Cell 24, 310–323 (2013).
Buiting, K. et al. Epimutations in Prader-Willi and Angelman syndromes: a molecular study of 136 patients with an imprinting defect. Am. J. Hum. Genet. 72, 571–577 (2003).
Poplinski, A., Tuttelmann, F., Kanber, D., Horsthemke, B. & Gromoll, J. Idiopathic male infertility is strongly associated with aberrant methylation of MEST and IGF2/H19 ICR1. Int. J. Androl 33, 642–649 (2010).
Hiura, H., Obata, Y., Komiyama, J., Shirai, M. & Kono, T. Oocyte growth-dependent progression of maternal imprinting in mice. Genes Cells 11, 353–361 (2006).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Gahurova, L. et al. Transcription and chromatin determinants of de novo DNA methylation timing in oocytes. Epigenetics Chromatin 10, 25 (2017).
Smallwood, S. A. et al. Dynamic CpG island methylation landscape in oocytes and preimplantation embryos. Nat. Genet. 43, 811–814 (2011).
Yu, B. et al. Genome-wide, single-cell DNA methylomics reveals increased Non-CpG methylation during human oocyte maturation. Stem Cell Rep. 9, 397–407 (2017).
Bourc’his, D., Xu, G. L., Lin, C. S., Bollman, B. & Bestor, T. H. Dnmt3L and the establishment of maternal genomic imprints. Science 294, 2536–2539 (2001).
Kaneda, M. et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 429, 900–903 (2004).
Kato, Y. et al. Role of the Dnmt3 family in de novo methylation of imprinted and repetitive sequences during male germ cell development in the mouse. Hum. Mol. Genet. 16, 2272–2280 (2007).
Shirane, K. et al. Mouse oocyte methylomes at base resolution reveal genome-wide accumulation of non-CpG methylation and role of DNA methyltransferases. PLOS Genet. 9, e1003439 (2013).
Chotalia, M. et al. Transcription is required for establishment of germline methylation marks at imprinted genes. Genes Dev. 23, 105–117 (2009).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015). Targeting of DNA methylation to transcribed gene bodies by SETD2-mediated methylation of H3K36me3 and a functional recruitment of DNMT3B is demonstrated.
Dhayalan, A. et al. The Dnmt3a PWWP domain reads histone 3 lysine 36 trimethylation and guides DNA methylation. J. Biol. Chem. 285, 26114–26120 (2010).
Ciccone, D. N. et al. KDM1B is a histone H3K4 demethylase required to establish maternal genomic imprints. Nature 461, 415–418 (2009).
Ooi, S. K. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448, 714–717 (2007).
Stewart, K. R. et al. Dynamic changes in histone modifications precede de novo DNA methylation in oocytes. Genes Dev. 29, 2449–2462 (2015).
Petrussa, L., Van de Velde, H. & De Rycke, M. Dynamic regulation of DNA methyltransferases in human oocytes and preimplantation embryos after assisted reproductive technologies. Mol. Hum. Reprod. 20, 861–874 (2014).
Henckel, A., Chebli, K., Kota, S. K., Arnaud, P. & Feil, R. Transcription and histone methylation changes correlate with imprint acquisition in male germ cells. EMBO J. 31, 606–615 (2012).
Singh, V. B. et al. Blocked transcription through KvDMR1 results in absence of methylation and gene silencing resembling Beckwith-Wiedemann syndrome. Development 144, 1820–1830 (2017).
Beygo, J. et al. A maternal deletion upstream of the imprint control region 2 in 11p15 causes loss of methylation and familial Beckwith-Wiedemann syndrome. Eur. J. Hum. Genet. 24, 1280–1286 (2016).
Valente, F. M. et al. Transcription alterations of KCNQ1 associated with imprinted methylation defects in the Beckwith-Wiedemann locus. Genet. Med. https://doi.org/10.1038/s41436-018-0416-7 (2019).
Lewis, M. W. et al. Angelman syndrome imprinting center encodes a transcriptional promoter. Proc. Natl Acad. Sci. USA 112, 6871–6875 (2015).
Smith, Z. D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Eckersley-Maslin, M. A., Alda-Catalinas, C. & Reik, W. Dynamics of the epigenetic landscape during the maternal-to-zygotic transition. Nat. Rev. Mol. Cell Biol. 19, 436–450 (2018).
Shen, L. et al. Tet3 and DNA replication mediate demethylation of both the maternal and paternal genomes in mouse zygotes. Cell Stem Cell 15, 459–471 (2014).
Hirasawa, R. et al. Maternal and zygotic Dnmt1 are necessary and sufficient for the maintenance of DNA methylation imprints during preimplantation development. Genes Dev. 22, 1607–1616 (2008).
Maenohara, S. et al. Role of UHRF1 in de novo DNA methylation in oocytes and maintenance methylation in preimplantation embryos. PLOS Genet. 13, e1007042 (2017).
Nakamura, T. et al. PGC7 binds histone H3K9me2 to protect against conversion of 5mC to 5hmC in early embryos. Nature 486, 415–419 (2012).
Nakamura, T. et al. PGC7/Stella protects against DNA demethylation in early embryogenesis. Nat. Cell Biol. 9, 64–71 (2007).
Meyer, E. et al. Germline mutation in NLRP2 (NALP2) in a familial imprinting disorder (Beckwith-Wiedemann Syndrome). PLOS Genet. 5, e1000423 (2009).
Parry, D. A. et al. Mutations causing familial biparental hydatidiform mole implicate c6orf221 as a possible regulator of genomic imprinting in the human oocyte. Am. J. Hum. Genet. 89, 451–458 (2011).
Murdoch, S. et al. Mutations in NALP7 cause recurrent hydatidiform moles and reproductive wastage in humans. Nat. Genet. 38, 300–302 (2006).
Sanchez-Delgado, M. et al. Absence of maternal methylation in biparental hydatidiform moles from women with NLRP7 maternal-effect mutations reveals widespread placenta-specific imprinting. PLOS Genet. 11, e1005644 (2015).
Soellner, L. et al. Maternal heterozygous NLRP7 variant results in recurrent reproductive failure and imprinting disturbances in the offspring. Eur. J. Hum. Genet. 25, 924–929 (2017).
Zhu, K. et al. Identification of a human subcortical maternal complex. Mol. Hum. Reprod. 21, 320–329 (2015).
Virant-Klun, I., Leicht, S., Hughes, C. & Krijgsveld, J. Identification of maturation-specific proteins by single-cell proteomics of human oocytes. Mol. Cell. Proteomics 15, 2616–2627 (2016).
Tong, Z. B. et al. Mater, a maternal effect gene required for early embryonic development in mice. Nat. Genet. 26, 267–268 (2000).
Yurttas, P. et al. Role for PADI6 and the cytoplasmic lattices in ribosomal storage in oocytes and translational control in the early mouse embryo. Development 135, 2627–2636 (2008).
Zheng, P. & Dean, J. Role of Filia, a maternal effect gene, in maintaining euploidy during cleavage-stage mouse embryogenesis. Proc. Natl Acad. Sci. USA 106, 7473–7478 (2009).
Mahadevan, S. et al. Maternally expressed NLRP2 links the subcortical maternal complex (SCMC) to fertility, embryogenesis and epigenetic reprogramming. Sci. Rep. 7, 44667 (2017). The generation of a maternal-effect mouse model for Nlrp2 is presented, which reveals disrupted imprinted methylation consistent with human MLIDs.
Sills, E. S. et al. Pathogenic variant in NLRP7 (19q13.42) associated with recurrent gestational trophoblastic disease: data from early embryo development observed during in vitro fertilization. Clin. Exp. Reprod. Med. 44, 40–46 (2017).
Li, X. et al. A maternal-zygotic effect gene, Zfp57, maintains both maternal and paternal imprints. Dev. Cell 15, 547–557 (2008).
Quenneville, S. et al. In embryonic stem cells, ZFP57/KAP1 recognize a methylated hexanucleotide to affect chromatin and DNA methylation of imprinting control regions. Mol. Cell 44, 361–372 (2011).
Coluccio, A. et al. Individual retrotransposon integrants are differentially controlled by KZFP/KAP1-dependent histone methylation, DNA methylation and TET-mediated hydroxymethylation in naive embryonic stem cells. Epigenetics Chromatin 11, 7 (2018).
Anvar, Z. et al. ZFP57 recognizes multiple and closely spaced sequence motif variants to maintain repressive epigenetic marks in mouse embryonic stem cells. Nucleic Acids Res. 44, 1118–1132 (2016).
Strogantsev, R. et al. Allele-specific binding of ZFP57 in the epigenetic regulation of imprinted and non-imprinted monoallelic expression. Genome Biol. 16, 112 (2015).
Imbeault, M., Helleboid, P. Y. & Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 543, 550–554 (2017). Human genome targets for 350 KRAB-domain-containing ZFPs are identified.
Mackay, D. J. et al. Hypomethylation of multiple imprinted loci in individuals with transient neonatal diabetes is associated with mutations in ZFP57. Nat. Genet. 40, 949–951 (2008).
Messerschmidt, D. M. et al. Trim28 is required for epigenetic stability during mouse oocyte to embryo transition. Science 335, 1499–1502 (2012).
Sharif, J. et al. The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature 450, 908–912 (2007).
Dalgaard, K. et al. Trim28 haploinsufficiency triggers bi-stable epigenetic obesity. Cell 164, 353–364 (2016).
Dagar, V. et al. Genetic variation affecting DNA methylation and the human imprinting disorder, Beckwith-Wiedemann syndrome. Clin. Epigenetics 10, 114 (2018).
Abi Habib, W. et al. 11p15 ICR1 partial deletions associated with IGF2/H19 DMR hypomethylation and Silver-Russell syndrome. Hum. Mutat. 38, 105–111 (2017).
Demars, J. et al. Analysis of the IGF2/H19 imprinting control region uncovers new genetic defects, including mutations of OCT-binding sequences, in patients with 11p15 fetal growth disorders. Hum. Mol. Genet. 19, 803–814 (2010).
Freschi, A. et al. Tissue-specific and mosaic imprinting defects underlie opposite congenital growth disorders in mice. PLOS Genet. 14, e1007243 (2018).
Hur, S. K. et al. Humanized H19/Igf2 locus reveals diverged imprinting mechanism between mouse and human and reflects Silver-Russell syndrome phenotypes. Proc. Natl Acad. Sci. USA 113, 10938–10943 (2016).
Bruno, C. et al. Germline correction of an epimutation related to Silver-Russell syndrome. Hum. Mol. Genet. 24, 3314–3321 (2015).
Berland, S. et al. Evidence for anticipation in Beckwith-Wiedemann syndrome. Eur. J. Hum. Genet. 21, 1344–1348 (2013).
Cortessis, V. K. et al. Comprehensive meta-analysis reveals association between multiple imprinting disorders and conception by assisted reproductive technology. J. Assist. Reprod. Genet. 35, 943–952 (2018). This meta-analysis demonstrates the association between imprinting disorders and ARTs.
Taniguchi, K., Kawai, T. & Hata, K. Placental development and nutritional environment. Adv. Exp. Med. Biol. 1012, 63–73 (2018).
Cox, G. F. et al. Intracytoplasmic sperm injection may increase the risk of imprinting defects. Am. J. Hum. Genet. 71, 162–164 (2002).
DeBaun, M. R., Niemitz, E. L. & Feinberg, A. P. Association of in vitro fertilization with Beckwith-Wiedemann syndrome and epigenetic alterations of LIT1 and H19. Am. J. Hum. Genet. 72, 156–160 (2003).
Maher, E. R. et al. Beckwith-Wiedemann syndrome and assisted reproduction technology (ART). J. Med. Genet. 40, 62–64 (2003).
Mussa, A. et al. Assisted reproductive techniques and risk of Beckwith-Wiedemann syndrome. Pediatrics 140, e20164311 (2017).
Tee, L. et al. Epimutation profiling in Beckwith-Wiedemann syndrome: relationship with assisted reproductive technology. Clin. Epigenet. 5, 23 (2013).
Tenorio, J. et al. Clinical and molecular analyses of Beckwith-Wiedemann syndrome: comparison between spontaneous conception and assisted reproduction techniques. Am. J. Med. Genet. 170A, 2740–2749 (2016).
Chen, Z. et al. Characterization of global loss of imprinting in fetal overgrowth syndrome induced by assisted reproduction. Proc. Natl Acad. Sci. USA 112, 4618–4623 (2015).
Canovas, S. et al. DNA methylation and gene expression changes derived from assisted reproductive technologies can be decreased by reproductive fluids. eLife 6, e23670 (2017).
de Waal, E. et al. The cumulative effect of assisted reproduction procedures on placental development and epigenetic perturbations in a mouse model. Hum. Mol. Genet. 24, 6975–6985 (2015).
Ludwig, M. et al. Increased prevalence of imprinting defects in patients with Angelman syndrome born to subfertile couples. J. Med. Genet. 42, 289–291 (2005).
Hyrapetian, M., Loucaides, E. M. & Sutcliffe, A. G. Health and disease in children born after assistive reproductive therapies (ART). J. Reprod. Immunol. 106, 21–26 (2014).
Weinerman, R., Ord, T., Bartolomei, M. S., Coutifaris, C. & Mainigi, M. The superovulated environment, independent of embryo vitrification, results in low birthweight in a mouse model. Biol. Reprod. 97, 133–142 (2017).
Dankert, D. et al. Pre- and postovulatory aging of murine oocytes affect the transcript level and poly(A) tail length of maternal effect genes. PLOS ONE 9, e108907 (2014).
Demond, H. et al. Preovulatory aging in vivo and in vitro affects maturation rates, abundance of selected proteins, histone methylation pattern and spindle integrity in murine oocytes. PLOS ONE 11, e0162722 (2016).
Lu, Y. Q., He, X. C. & Zheng, P. Decrease in expression of maternal effect gene Mater is associated with maternal ageing in mice. Mol. Hum. Reprod. 22, 252–260 (2016).
Camprubi, C. et al. Stability of genomic imprinting and gestational-age dynamic methylation in complicated pregnancies conceived following assisted reproductive technologies. Biol. Reprod. 89, 50 (2013).
Ji, M. et al. ART manipulation after controlled ovarian stimulation may not increase the risk of abnormal expression and DNA methylation at some CpG sites of H19,IGF2 and SNRPN in foetuses: a pilot study. Reprod. Biol. Endocrinol. 16, 63 (2018).
White, C. R. et al. High frequency of imprinted methylation errors in human preimplantation embryos. Sci. Rep. 5, 17311 (2015).
Conradt, E. et al. Incorporating epigenetic mechanisms to advance fetal programming theories. Dev. Psychopathol. 30, 807–824 (2018).
Han, L. et al. Embryonic defects induced by maternal obesity in mice derive from Stella insufficiency in oocytes. Nat. Genet. 50, 432–442 (2018). This study demonstrates that a maternal environmental insult may affect genomic integrity, epigenetic marking and developmental competence at the cleavage stage of offspring.
Tserga, A., Binder, A. M. & Michels, K. B. Impact of folic acid intake during pregnancy on genomic imprinting of IGF2/H19 and 1-carbon metabolism. FASEB J. 31, 5149–5158 (2017).
Motzek, A. et al. Abnormal hypermethylation at imprinting control regions in patients with S-adenosylhomocysteine hydrolase (AHCY) Deficiency. PLOS ONE 11, e0151261 (2016).
Amarasekera, M. et al. Genome-wide DNA methylation profiling identifies a folate-sensitive region of differential methylation upstream of ZFP57-imprinting regulator in humans. FASEB J. 28, 4068–4076 (2014).
Kessler, N. J., Waterland, R. A., Prentice, A. M. & Silver, M. J. Establishment of environmentally sensitive DNA methylation states in the very early human embryo. Sci. Adv. 4, eaat2624 (2018). This analysis of metastable epialleles in a rural Gambian population demonstrates epigenetic sensitivity to season of birth in early embryonic development.
Van de Pette, M. et al. Visualizing changes in Cdkn1c expression links early-life adversity to imprint mis-regulation in adults. Cell Rep. 18, 1090–1099 (2017). A mouse model demonstrates that maternal dietary change promotes transient alteration in DNA methylation and enduring misexpression of an imprinted gene.
Ivanova, E., Chen, J. H., Segonds-Pichon, A., Ozanne, S. E. & Kelsey, G. DNA methylation at differentially methylated regions of imprinted genes is resistant to developmental programming by maternal nutrition. Epigenetics 7, 1200–1210 (2012).
Heo, J. et al. Sirt1 regulates DNA methylation and differentiation potential of embryonic stem cells by antagonizing Dnmt3l. Cell Rep. 18, 1930–1945 (2017).
Kim, J. H. et al. Perinatal bisphenol A exposure promotes dose-dependent alterations of the mouse methylome. BMC Genomics 15, 30 (2014).
Kochmanski, J. J. et al. Longitudinal effects of developmental bisphenol A exposure on epigenome-wide DNA hydroxymethylation at imprinted loci in mouse blood. Environ. Health Perspect. 126, 077006 (2018).
Susiarjo, M., Sasson, I., Mesaros, C. & Bartolomei, M. S. Bisphenol a exposure disrupts genomic imprinting in the mouse. PLOS Genet. 9, e1003401 (2013).
Trapphoff, T., Heiligentag, M., El Hajj, N., Haaf, T. & Eichenlaub-Ritter, U. Chronic exposure to a low concentration of bisphenol A during follicle culture affects the epigenetic status of germinal vesicles and metaphase II oocytes. Fertil. Steril. 100, 1758–1767 (2013).
Li, Y., Duan, F., Zhou, X., Pan, H. & Li, R. Differential responses of GC1 spermatogonia cells to high and low doses of bisphenol A. Mol. Med. Rep. 18, 3034–3040 (2018).
Junge, K. M. et al. MEST mediates the impact of prenatal bisphenol A exposure on long-term body weight development. Clin. Epigenetics 10, 58 (2018).
Ding, Z. M. et al. Bisphenol AF negatively affects oocyte maturation of mouse in vitro through increasing oxidative stress and DNA damage. Chem. Biol. Interact. 278, 222–229 (2017).
Pathak, R. & Feil, R. Environmental effects on chromatin repression at imprinted genes and endogenous retroviruses. Curr. Opin. Chem. Biol. 45, 139–147 (2018).
Huang, H. S. et al. Topoisomerase inhibitors unsilence the dormant allele of Ube3a in neurons. Nature 481, 185–189 (2011).
Meng, L. et al. Towards a therapy for Angelman syndrome by targeting a long non-coding RNA. Nature 518, 409–412 (2015).
Kim, Y. et al. Targeting the histone methyltransferase G9a activates imprinted genes and improves survival of a mouse model of Prader-Willi syndrome. Nat. Med. 23, 213–222 (2017). This proof-of-principle mouse study identifies small molecules that selectively reduce H3K9me2 at a PWS imprinting centre and highlights a potential therapy for PWS.
Lei, Y. et al. Targeted DNA methylation in vivo using an engineered dCas9-MQ1 fusion protein. Nat. Commun. 8, 16026 (2017).
Cerrato, F. et al. Different mechanisms cause imprinting defects at the IGF2/H19 locus in Beckwith-Wiedemann syndrome and Wilms’ tumour. Hum. Mol. Genet. 17, 1427–1435 (2008).
Kagami, M. et al. Deletions and epimutations affecting the human 14q32.2 imprinted region in individuals with paternal and maternal upd(14)-like phenotypes. Nat. Genet. 40, 237–242 (2008).
Azzi, S. et al. Complex tissue-specific epigenotypes in Russell-Silver Syndrome associated with 11p15 ICR1 hypomethylation. Hum. Mutat. 35, 1211–1220 (2014).
Nazlican, H. et al. Somatic mosaicism in patients with Angelman syndrome and an imprinting defect. Hum. Mol. Genet. 13, 2547–2555 (2004).
Weksberg, R. et al. Discordant KCNQ1OT1 imprinting in sets of monozygotic twins discordant for Beckwith-Wiedemann syndrome. Hum. Mol. Genet. 11, 1317–1325 (2002).
Riess, A. et al. First report on concordant monozygotic twins with Silver-Russell syndrome and ICR1 hypomethylation. Eur. J. Med. Genet. 59, 1–4 (2016).
Bliek, J. et al. Lessons from BWS twins: complex maternal and paternal hypomethylation and a common source of haematopoietic stem cells. Eur. J. Hum. Genet. 17, 1625–1634 (2009).
Greenberg, M. V. et al. Transient transcription in the early embryo sets an epigenetic state that programs postnatal growth. Nat. Genet. 49, 110–118 (2017). This study demonstrates that transient transcription in pre-implantation embryo sets an epigenetic state that programmes postnatal growth.
Probst, A. V. et al. A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development. Dev. Cell 19, 625–638 (2010).
De Iaco, A. et al. DUX-family transcription factors regulate zygotic genome activation in placental mammals. Nat. Genet. 49, 941–945 (2017).
Nativio, R. et al. Disruption of genomic neighbourhood at the imprinted IGF2-H19 locus in Beckwith-Wiedemann syndrome and Silver-Russell syndrome. Hum. Mol. Genet. 20, 1363–1374 (2011).
Acknowledgements
The authors thank F. Cerrato, M. V. Cubellis and A. Sparago for careful reading of the manuscript. The authors apologize to all authors of studies that could not be cited owing to the concise nature of this Review. Individual authors thank the following funders for research support: Ministry of Education, Universities and Research, Research Projects of National Interest (MIUR PRIN) 2015 (JHLY35), Telethon-Italia (GGP15131 and AIRC IG18671) (A.R.); Deutsche Forschungsgemeinschaft (DFG) (EG110/15-1) (T.E.); Spanish Ministry of Economy and Competitiveness (MINECO; BFU2014-53093-R and BFU2017-85571-R) co-funded with the European Union Regional Development Fund (FEDER) (D.M.). E.R.M. acknowledges support from National Institute for Health Research (NIHR) Cambridge Biomedical Research Centre and an NIHR Senior Investigator Award. The University of Cambridge has received salary support in respect of E.R.M. from the National Health Service (NHS) in the East of England through the Clinical Academic Reserve. The views expressed are those of the authors and not necessarily those of the NIHR, NHS or Department of Health.
Reviewer information
Nature Reviews Genetics thanks Y.-H. Jiang and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Genomic imprinting
-
The epigenetic marking of a gene on the basis of parental origin that results in monoallelic expression.
- Epialleles
-
Epigenetic profiles that are maintained in somatic tissues, resulting in interindividual variation.
- Imprinting disorders
-
Diseases associated with disruption of imprinted gene expression that can be caused by genetic or epigenetic defects.
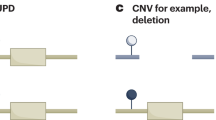
- Uniparental disomy
-
(UPD). A genetic defect characterized by the presence of two copies of a chromosome or part of the chromosome derived from only one parent.
- Epimutations
-
With reference to imprinting disorders, epigenetic changes that affect the regulation of imprinted loci. An epimutation is primary if there is no detectable genetic cause and secondary if it is associated with a genetic cause.
- cis-acting elements
-
DNA sequences that regulate the expression of a gene that is present on the same chromosome.
- trans-acting factors
-
Proteins that regulate the expression of a gene.
- Epigenetic reprogramming
-
The erasure of pre-existing epigenetic marks that enable subsequent remodelling of chromatin.
- Epigenome
-
The chromatin modifications influencing genome function but not involving the underlying DNA sequence that can be propagated through cell division.
- Imprinting centre
-
Also known as imprinting control region. A functional definition of germline differentially methylated regions (gDMRs) that have been shown through either genetic targeting in mice or mutations in humans to regulate imprinted gene expression. Not all gDMRs have been shown to be imprinting centres.
- Germline differentially methylated region
-
(gDMR). A region of differential DNA methylation between the parental alleles in somatic cells that originates from the gametes. gDMRs that survive embryonic reprogramming are generally associated with imprinted genes.
- Loss of methylation
-
(LOM). With reference to imprinting disorders, loss of differential imprinting centre methylation detected in individuals that causes deregulation of imprinted genes in the domain controlled by the imprinting centre.
- Gain of methylation
-
(GOM). With reference to imprinting disorders, gain of methylation on the unmethylated allele of the imprinting centre. It is detected in people and causes deregulation of imprinted genes in the domain controlled by the imprinting centre.
- Secondary DMRs
-
Also known as somatic differentially methylated regions (DMRs). Regions of differential DNA methylation between parental alleles that do not originate in the germ line. They are regulated in a hierarchical fashion by a nearby imprinting centre region.
- Multilocus imprinting disturbances
-
(MLIDs). Methylation anomalies at imprinted differentially methylated regions in individuals with imprinting disorders in addition to those that are normally associated with disease.
- Zygote
-
A fertilized ovum before the first cell division that represents the earliest stage of embryonic development. The genome of the zygote is a combination of the DNA in each gamete.
- Primordial germ cells
-
(PGCs). Stem cell-like cells found in the gonadal ridge of developing embryos that develop into gametes following sex-specific epigenetic reprogramming and meiosis.
- Blastocyst
-
The developmental stage of a mammalian embryo just before implantation, consisting of an inner cell mass, which will form the embryo, and a cavity with an outer layer called trophoblast, which gives rise to the placenta.
- Genome activation
-
The initiation of gene expression in the developing embryo. The initial burst of expression is termed zygotic genome activation and is regulated by pioneer transcription factors during the oocyte-to-embryo transition. Initiation of expression in cleavage embryos is referred to as embryonic genome activation.
- Pronucleus
-
The haploid nucleus from a male or female gamete before the genetic material fuse at syngamy.
- Maternal-effect gene
-
A gene coding for an oocyte-derived transcript or protein that is required for the early development of the embryo.
- Hydatidiform mole
-
A benign gestational trophoblastic disease developing during pregnancy and resulting from abnormal fertilization. It is characterized by trophoblastic proliferation and little or no embryonic tissue. It is commonly sporadic and contains only sperm DNA. Occasionally, it can be biparental, recurrent and familial, following an autosomal recessive mode of inheritance.
- Penetrance
-
The proportion of individuals in a population with a specific genotype who show an associated phenotypic trait. Incomplete penetrance occurs when not all individuals carrying a dominant deleterious genetic variant express the associated clinical phenotype.
- Subcortical maternal complex
-
(SCMC). A large multiprotein complex comprising NLRP5, OOEP, TLE6, PADI6 and KHDC3L that localizes to the outermost regions of the cytoplasm in oocytes and is excluded from regions of cell-to-cell contact in cleavage embryos.
- Endogenous retroviruses
-
Also known as retrotransposons. Repetitive genetic elements present in the genome that, similarly to retroviruses, use the activity of reverse transcriptase to move from one locus to another.
- Protamines
-
Basic proteins that largely replace histones in the nucleus of mature sperm for more condensed DNA packaging.
- Haploinsufficiency
-
A situation in which one-half of the normal level of a gene product, usually as a consequence of a loss-of-function mutation, is not sufficient for normal function.
- Anticipation
-
A phenomenon whereby the symptoms of a genetic disorder become apparent at an earlier age or with greater severity in succeeding generations.
- Assisted reproductive technologies
-
(ART). Techniques used to achieve pregnancy during the treatment of infertility. ARTs cover a wide spectrum of treatments including the use of fertility drugs, intrauterine insemination and in vitro fertilization and/or intracytoplasmic sperm injection.
Rights and permissions
About this article
Cite this article
Monk, D., Mackay, D.J.G., Eggermann, T. et al. Genomic imprinting disorders: lessons on how genome, epigenome and environment interact. Nat Rev Genet 20, 235–248 (2019). https://doi.org/10.1038/s41576-018-0092-0
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-018-0092-0
This article is cited by
-
Genetically transitional disease: conceptual understanding and applicability to rheumatic disease
Nature Reviews Rheumatology (2024)
-
Detection of DNA methylation signatures through the lens of genomic imprinting
Scientific Reports (2024)
-
Risk assessment of assisted reproductive technology and parental age at childbirth for the development of uniparental disomy-mediated imprinting disorders caused by aneuploid gametes
Clinical Epigenetics (2023)
-
Widespread allele-specific topological domains in the human genome are not confined to imprinted gene clusters
Genome Biology (2023)
-
Molecular characterisation of 36 multilocus imprinting disturbance (MLID) patients: a comprehensive approach
Clinical Epigenetics (2023)