Abstract
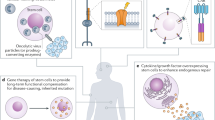
New fundamental discoveries in stem cell biology have yielded potentially transformative regenerative therapeutics. However, widespread implementation of stem-cell-derived therapeutics remains sporadic. Barriers that impede the development of these therapeutics can be linked to our incomplete understanding of how the regulatory networks that encode stem cell fate govern the development of the complex tissues and organs that are ultimately required for restorative function. Bioengineering tools, strategies and design principles represent core components of the stem cell bioengineering toolbox. Applied to the different layers of complexity present in stem-cell-derived systems — from gene regulatory networks in single stem cells to the systemic interactions of stem-cell-derived organs and tissues — stem cell bioengineering can address existing challenges and advance regenerative medicine and cellular therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Till, J. E. & McCulloch, E. A. A direct measurement of the radiation sensitivity of normal mouse bone marrow cells. Radiat. Res. 14, 213–222 (1961).
Henig, I. & Zuckerman, T. Hematopoietic stem cell transplantation — 50 years of evolution and future perspectives. Rambam Maimonides Med. J. 5, e0028 (2014).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03167203 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03178149 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02302157 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03163511?term=ViaCyte&rank=1 (2018).
Mavilio, F. et al. Correction of junctional epidermolysis bullosa by transplantation of genetically modified epidermal stem cells. Nat. Med. 12, 1397–1402 (2006).
Hirsch, T. et al. Regeneration of the entire human epidermis using transgenic stem cells. Nature 551, 327–332 (2017).
Blanpain, C. & Simons, B. D. Unravelling stem cell dynamics by lineage tracing. Nat. Rev. Mol. Cell Biol. 14, 489–502 (2013).
Dekkers, J. F. et al. A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nat. Med. 19, 939–945 (2013).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Dekkers, J. F. et al. Characterizing responses to CFTR-modulating drugs using rectal organoids derived from subjects with cystic fibrosis. Sci. Transl Med. 8, 344ra84 (2016).
Lindemans, C. A. et al. Interleukin 22 promotes intestinal-stem-cell-mediated epithelial regeneration. Nature 528, 560–564 (2015).
Wells, J. M. & Watt, F. M. Diverse mechanisms for endogenous regeneration and repair in mammalian organs. Nature 557, 322–328 (2018).
Matsumura, H. et al. Hair follicle aging is driven by transepidermal elimination of stem cells via COL17A1 proteolysis. Science 351, aad4395 (2016).
Watanabe, M. et al. Type XVII collagen coordinates proliferation in the interfollicular epidermis. eLife 6, a015206 (2017).
Jaenisch, R. & Bird, A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 33, S245–S254 (2003).
Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447, 425–432 (2007).
Alon, U. An Introduction to Systems Biology: Design Principles of Biological Circuits 60, 63–64 (Chapman and Hall, 2007).
Morrison, S. J. & Spradling, A. C. Stem cells and niches: mechanisms that promote stem cell maintenance throughout life. Cell 132, 598–611 (2008).
Yin, X. et al. Engineering stem cell organoids. Cell Stem Cell 18, 25–38 (2016).
López-Onieva, L., Fernández-Miñán, A. & González-Reyes, A. Jak/Stat signalling in niche support cells regulates dpp transcription to control germline stem cell maintenance in the Drosophila ovary. Development 135, 533–540 (2008).
Yamashita, Y. M., Mahowald, A. P., Perlin, J. R. & Fuller, M. T. Asymmetric inheritance of mother versus daughter centrosome in stem cell division. Science 315, 518–521 (2007).
Ohlstein, B. & Spradling, A. Multipotent Drosophila intestinal stem cells specify daughter cell fates by differential notch signaling. Science 315, 988–992 (2007).
Kretzschmar, K. & Clevers, H. Organoids: modeling development and the stem cell niche in a dish. Dev. Cell 38, 590–600 (2016).
Crane, G. M., Jeffery, E. & Morrison, S. J. Adult haematopoietic stem cell niches. Nat. Rev. Immunol. 17, 573–590 (2017).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Self-renewing diploid Axin2+ cells fuel homeostatic renewal of the liver. Nature 524, 180–185 (2015).
Nabhan, A., Brownfield, D. G., Harbury, P. B., Krasnow, M. A. & Desai, T. J. Single-cell Wnt signaling niches maintain stemness of alveolar type 2 cells. Science 137, 1–12 (2018).
Page, M. E., Lombard, P., Ng, F., Göttgens, B. & Jensen, K. B. The epidermis comprises autonomous compartments maintained by distinct stem cell populations. Cell Stem Cell 13, 471–482 (2013).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Pilz, G. A. et al. Live imaging of neurogenesis in the adult mouse hippocampus. Science 359, 658–662 (2018).
Raspopovic, J., Marcon, L., Russo, L. & Sharpe, J. Modeling digits. Digit patterning is controlled by a Bmp-Sox9 Wnt Turing network modulated by morphogen gradients. Science 345, 566–570 (2014).
Sick, S., Reinker, S., Timmer, J. & Schlake, T. WNT and DKK determine hair follicle spacing through a reaction-diffusion mechanism. Science 314, 1447–1450 (2006).
Economou, A. D. et al. Periodic stripe formation by a Turing mechanism operating at growth zones in the mammalian palate. Nat. Genet. 44, 348–U163 (2012).
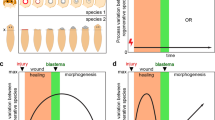
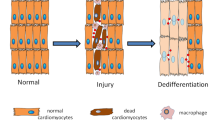
Donati, G. et al. Wounding induces dedifferentiation of epidermal Gata6+ cells and acquisition of stem cell properties. Nat. Cell Biol. 19, 603–613 (2017).
Tata, P. R. et al. Dedifferentiation of committed epithelial cells into stem cells in vivo. Nature 503, 218–223 (2013).
van Es, J. H. et al. Dll1+ secretory progenitor cells revert to stem cells upon crypt damage. Nat. Cell Biol. 14, 1099–1104 (2012).
Green, J. B. A. & Sharpe, J. Positional information and reaction-diffusion: two big ideas in developmental biology combine. Development 142, 1203–1211 (2015).
Tewary, M. et al. A stepwise model of reaction-diffusion and positional-information governs self-organized human peri-gastrulation-like patterning. Development 144, 4298–4312 (2017). This study employs an in vitro model of a stem-cell-derived, developmentally-relevant, fate-patterning system and demonstrates that reaction diffusion and positional information can work in concert to give rise to emergent complexity in stem cell systems.
Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
Turing, A. M. The chemical basis of morphogenesis. Phil. Trans. R. Soc. Lond. B Biol. Sci. 237, 37–72 (1952).
Wolpert, L. Positional information & pattern formation. Phil. Trans. Soc. R. Lond. B Biol. Sci. 295, 441–450 (1981).
Wolpert, L. Positional information and the spatial pattern of cellular differentiation. J. Theor. Biol. 25, 1–47 (1969).
Cooke, J. & Zeeman, E. C. A clock and wavefront model for control of the number of repeated structures during animal morphogenesis. J. Theor. Biol. 58, 455–476 (1976).
Thavandiran, N. et al. Design and formulation of functional pluripotent stem cell-derived cardiac microtissues. Proc. Nat. Acad. Sci. USA 110, E4698–E4707 (2013).
Droujinine, I. A. & Perrimon, N. Interorgan communication pathways in physiology: focus on Drosophila. Annu. Rev. Genet. 50, 539–570 (2016).
Evans, A. N. & Rooney, B. F. Personnel Psychological Vol. 62 (eds Craig, S. B. & Clark, A. P.) 633–636 (Wiley, 2009).
Way, J. C., Collins, J. J., Keasling, J. D. & Silver, P. A. Integrating biological redesign: where synthetic biology came from and where it needs to go. Cell 157, 151–161 (2014).
Antebi, Y. E. et al. Combinatorial signal perception in the BMP pathway. Cell 170, 1184–1196 (2017).
Mirams, G. R. et al. Chaste: an open source C + + library for computational physiology and biology. PLOS Comput. Biol. 9, e1002970 (2013).
Nerurkar, N. L., Mahadevan, L. & Tabin, C. J. BMP signaling controls buckling forces to modulate looping morphogenesis of the gut. Proc. Nat. Acad. Sci. USA 114, 2277–2282 (2017).
Cosentino, C. & Bates, D. Feedback Control in Systems Biology (CRC Press, 2011).
Bleris, L. et al. Synthetic incoherent feedforward circuits show adaptation to the amount of their genetic template. Molecular Systems Biology 7, 519 (2011) .
Freeman, M. Feedback control of intercellular signalling in development. Nature 408, 313–319 (2000). This review explores the role of positive and negative feedback loops in the dynamic regulation of developmental signalling.
Doyle, J. & Csete, M. Motifs, control, and stability. PLOS Biol. 3, e392 (2005).
Hussein, S. M. I. et al. Genome-wide characterization of the routes to pluripotency. Nature 516, 198–206 (2014).
Guo, F. et al. Single-cell multi-omics sequencing of mouse early embryos and embryonic stem cells. Cell Res. 27, 967–988 (2017).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Yan, L. et al. Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Kumar, P., Tan, Y. & Cahan, P. Understanding development and stem cells using single cell-based analyses of gene expression. Development 144, 17–32 (2017).
Zhou, F. et al. Tracing haematopoietic stem cell formation at single-cell resolution. Nature 533, 487–492 (2016).
Shakiba, N. et al. CD24 tracks divergent pluripotent states in mouse and human cells. Nat. Commun. 6, 7329 (2015). This study employs mass spectrometry analysis of the surface proteome of reprogramming cells to identify a key marker that can distinguish different reprogramming and PSC states in both murine and human systems.
O’Malley, J. et al. High-resolution analysis with novel cell-surface markers identifies routes to iPS cells. Nature 499, 88–91 (2013).
Ng, S. W. K. et al. A 17 gene stemness score for rapid determination of risk in acute leukaemia. Nature 540, 433–437 (2016). This study utilizes computational and statistical analysis to identify a 17-gene signature that is predictive of clinical survival following leukaemia.
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Yordanov, B. et al. A method to identify and analyze biological programs through automated reasoning. NPJ Syst. Biol. Appl. 2, 16010 (2016).
Chai, L. E. et al. A review on the computational approaches for gene regulatory network construction. Comput. Biol. Med. 48, 55–65 (2014).
Qiao, W. et al. Intercellular network structure and regulatory motifs in the human hematopoietic system. Mol. Syst. Biol. 10, 741–741 (2014). This study combines genomic and phenotypic data with high-content experiments to build a directional cell–cell communication network between 12 cell types in human umbilical cord blood.
Koike-Yusa, H., Li, Y., Tan, E. P., Velasco-Herrera, M. D. C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Golipour, A. et al. A late transition in somatic cell reprogramming requires regulators distinct from the pluripotency network. Cell Stem Cell 11, 769–782 (2012).
Nazareth, E. J. P., Rahman, N., Yin, T. & Zandstra, P. W. A. Multi-lineage screen reveals mTORC1 inhibition enhances human pluripotent stem cell mesendoderm and blood progenitor production. Stem Cell Rep. 6, 679–691 (2016).
Dunn, S. J., Martello, G., Yordanov, B., Emmott, S. & Smith, A. G. Defining an essential transcription factor program for naïve pluripotency. Science 344, 1156–1160 (2014). This paper reports the development of a data-constrained, computational approach to derive a simple GRN that functionally captures the mouse PSC state.
Moignard, V. et al. Decoding the regulatory network of early blood development from single-cell gene expression measurements. Nat. Biotechnol. 33, 269–276 (2015).
Kinoshita, A. Y. et al. Modeling signaling-dependent pluripotency with Boolean logic to predict cell fate transitions. Mol. Syst. Biol. 14, e7952 (2018).
Cahan, P. et al. CellNet: network biology applied to stem cell engineering. Cell 158, 903–915 (2014).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Gerrits, A. et al. Cellular barcoding tool for clonal analysis in the hematopoietic system. Blood 115, 2610–2618 (2010).
McKenna, A. et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016).
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017).
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2016). This paper describes a synthetic system to record lineage information and event histories in the genome of ESCs in a manner that is read out at the single level in situ.
Schroeder, T. Long-term single-cell imaging of mammalian stem cells. Nat. Methods 8, S30–S35 (2011). This perspective piece provides an overview of the utility of continuous long-term live imaging for providing key insights on stem cell function.
Prochazka, L., Benenson, Y. & Zandstra, P. W. Synthetic gene circuits and cellular decision-making in human pluripotent stem cells. Curr. Opin. Syst. Biol. 5, 93–103 (2017). This review explores the intersection of synthetic biology and stem cell engineering towards the development of decision-making gene circuits to control cell fate.
Teague, B. P., Guye, P. & Weiss, R. Synthetic morphogenesis. Cold Spring Harb. Perspect. Biol. 8, a023929 (2016).
Kitada, T., DiAndreth, B., Teague, B. & Weiss, R. Programming gene and engineered-cell therapies with synthetic biology. Science 359, eaad1067 (2018).
Mathur, M., Xiang, J. S. & Smolke, C. D. Mammalian synthetic biology for studying the cell. J. Cell Biol. 216, 73–82 (2017).
Cameron, D. E., Bashor, C. J. & Collins, J. J. A brief history of synthetic biology. Nat. Rev. Microbiol. 12, 381–390 (2014).
Lienert, F., Lohmueller, J. J., Garg, A. & Silver, P. A. Synthetic biology in mammalian cells: next generation research tools and therapeutics. Nat. Rev. Mol. Cell Biol. 15, 95–107 (2014).
Khalil, A. S. & Collins, J. J. Synthetic biology: applications come of age. Nat. Rev. Genet. 11, 367–379 (2010).
Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307–1311 (2011).
Kiani, S. et al. CRISPR transcriptional repression devices and layered circuits in mammalian cells. Nat. Methods 11, 723–726 (2014).
Prochazka, L., Angelici, B., Haefliger, B. & Benenson, Y. Highly modular bow-tie gene circuits with programmable dynamic behaviour. Nat. Commun. 5, 4729 (2014).
Weinberg, B. H. et al. Large-scale design of robust genetic circuits with multiple inputs and outputs for mammalian cells. Nat. Biotechnol. 35, 453–462 (2017).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell 164, 780–791 (2016).
Roybal, K. T. et al. Engineering T cells with customized therapeutic response programs using synthetic notch receptors. Cell 167, 419–432 (2016).
Saxena, P. et al. A programmable synthetic lineage-control network that differentiates human IPSCs into glucose-sensitive insulin-secreting beta-like cells. Nat. Commun. 7, 11247 (2016).
Del Vecchio, D., Dy, A. J. & Qian, Y. Control theory meets synthetic biology. J. R. Soc. Interface 13, 20160380 (2016).
Del Vecchio, D., Abdallah, H., Qian, Y. & Collins, J. J. A. Blueprint for a synthetic genetic feedback controller to reprogram cell fate. Cell Syst. 4, 109–120 (2017).
Davies, J. Using synthetic biology to explore principles of development. Development 144, 1146–1158 (2017).
Shakiba, N. & Zandstra, P. W. Engineering cell fitness: lessons for regenerative medicine. Curr. Opin. Biotechnol. 47, 7–15 (2017).
Jackson, H. J., Rafiq, S. & Brentjens, R. J. Driving CAR T cells forward. Nat. Rev. Clin. Oncol. 13, 370–383 (2016).
Li, P. et al. Morphogen gradient reconstitution reveals Hedgehog pathway design principles. Science 360, 543–548 (2018).
Sun, C. & Bernards, R. Feedback and redundancy in receptor tyrosine kinase signaling: relevance to cancer therapies. Trends Biochem. Sci. 39, 465–474 (2014).
Dublanche, Y., Michalodimitrakis, K., Kümmerer, N., Foglierini, M. & Serrano, L. Noise in transcription negative feedback loops: simulation and experimental analysis. Mol. Syst. Biol. 2, 41 (2006).
Austin, D. W. et al. Gene network shaping of inherent noise spectra. Nature 439, 608–611 (2006).
Becskei, A. & Serrano, L. Engineering stability in gene networks by autoregulation. Nature 405, 590–593 (2000).
Briat, C., Zechner, C. & Khammash, M. Design of a synthetic integral feedback circuit: dynamic analysis and DNA implementation. ACS Synth. Biol. 5, 1108–1116 (2016).
Briat, C., Gupta, A. & Khammash, M. Antithetic integral feedback ensures robust perfect adaptation in noisy biomolecular networks. Cell Syst 2, 15–26 (2016).
Raj, A. & van Oudenaarden, A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 216–226 (2008).
Maamar, H. & Dubnau, D. Bistability in the Bacillus subtilis K state (competence) system requires a positive feedback loop. Mol. Microbiol. 56, 615–624 (2005).
Xiong, W. & Ferrell, J. E. A positive-feedback-based bistable ‘memory module’ that governs a cell fate decision. Nature 426, 460–465 (2003).
Isaacs, F. J., Hasty, J., Cantor, C. R. & Collins, J. J. Prediction and measurement of an autoregulatory genetic module. Proc. Natl Acad. Sci. USA 100, 7714–7719 (2003).
Süel, G. M., Garcia-Ojalvo, J., Liberman, L. M. & Elowitz, M. B. An excitable gene regulatory circuit induces transient cellular differentiation. Nature 440, 545–550 (2006).
Kramer, B. P. & Fussenegger, M. Hysteresis in a synthetic mammalian gene network. Proc. Natl Acad. Sci. USA 102, 9517–9522 (2005).
Losick, R. & Desplan, C. Stochasticity and cell fate. Science 320, 65–68 (2008).
Wernet, M. F. et al. Stochastic spineless expression creates the retinal mosaic for colour vision. Nature 440, 174–180 (2006).
Johnston, R. J. & Desplan, C. Stochastic neuronal cell fate choices. Curr. Opin. Neurobiol. 18, 20–27 (2008).
Abranches, E. et al. Stochastic NANOG fluctuations allow mouse embryonic stem cells to explore pluripotency. Development 141, 2770–2779 (2014).
Guye, P. et al. Genetically engineering self-organization of human pluripotent stem cells into a liver bud-like tissue using Gata6. Nat. Commun. 7, 10243–10212 (2016).
Watt, F. M., Jordan, P. W. & O’Neill, C. H. Cell shape controls terminal differentiation of human epidermal keratinocytes. Proc. Natl Acad. Sci. USA 85, 5576–5580 (1988).
McBeath, R., Pirone, D. M., Nelson, C. M., Bhadriraju, K. & Chen, C. S. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 6, 483–495 (2004).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–U212 (2011).
Chen, S. et al. Interrogating cellular fate decisions with high-throughput arrays of multiplexed cellular communities. Nat. Commun. 7, 10309 (2016).
Peerani, R., Onishi, K., Mahdavi, A., Kumacheva, E. & Zandstra, P. W. Manipulation of signaling thresholds in ‘engineered stem cell niches’ identifies design criteria for pluripotent stem cell screens. PLOS ONE 4, e6438 (2009).
Onishi, K., Tonge, P. D., Nagy, A. & Zandstra, P. W. Microenvironment-mediated reversion of epiblast stem cells by reactivation of repressed JAK-STAT signaling. Integr. Biol. 4, 1367–1376 (2012).
Onishi, K., Tonge, P. D., Nagy, A. & Zandstra, P. W. Local BMP-SMAD1 signaling increases LIF receptor-dependent STAT3 responsiveness and primed-to-naive mouse pluripotent stem cell conversion frequency. Stem Cell Rep. 3, 156–168 (2014).
Peerani, R. et al. Niche-mediated control of human embryonic stem cell self-renewal and differentiation. EMBO J. 26, 4744–4755 (2007).
Bauwens, C. L. et al. Control of human embryonic stem cell colony and aggregate size heterogeneity influences differentiation trajectories. Stem Cells 26, 2300–2310 (2008).
Lee, L. H. et al. Micropatterning of human embryonic stem cells dissects the mesoderm and endoderm lineages. Stem Cell Res. 2, 155–162 (2009).
Nazareth, E. J. P. et al. High-throughput fingerprinting of human pluripotent stem cell fate responses and lineage bias. Nat. Methods 10, 1225–1231 (2013).
Nemashkalo, A., Ruzo, A., Heemskerk, I. & Warmflash, A. Morphogen and community effects determine cell fates in response to BMP4 signaling in human embryonic stem cells. Development 144, 3042–3053 (2017).
Flaim, C. J., Chien, S. & Bhatia, S. N. An extracellular matrix microarray for probing cellular differentiation. Nat. Methods 2, 119–125 (2005).
Shukla, S. et al. Progenitor T-cell differentiation from hematopoietic stem cells using Delta-like-4 and VCAM-1. Nat. Methods 14, 531–538 (2017).
Gilbert, P. M. et al. Substrate elasticity regulates skeletal muscle stem cell self-renewal in culture. Science 329, 1078–1081 (2010).
Trappmann, B. et al. Extracellular-matrix tethering regulates stem-cell fate. Nat. Mater. 11, 642–649 (2012).
Chaudhuri, O. et al. Hydrogels with tunable stress relaxation regulate stem cell fate and activity. Nat. Mater. 15, 326–334 (2016).
Madl, C. M. et al. Maintenance of neural progenitor cell stemness in 3D hydrogels requires matrix remodelling. Nat. Mater. 16, 1233–1242 (2017).
Kirouac, D. et al. Cell-cell interaction networks regulate blood stem and progenitor cell fate. Mol. Syst. Biol. 5, 293 (2009).
Kirouac, D. C. et al. Dynamic interaction networks in a hierarchically organized tissue. Mol. Syst. Biol. 6, 417 (2010).
Csaszar, E. et al. Rapid expansion of human hematopoietic stem cells by automated control of inhibitory feedback signaling. Cell Stem Cell 10, 218–229 (2012). This paper reports an integrated computational and experimental strategy that enables tunable reduction in the global levels of paracrine signalling factors in an automated closed-system for the improved expansion of HSCs.
Lipsitz, Y. Y., Timmins, N. E. & Zandstra, P. W. Quality cell therapy manufacturing by design. Nat. Biotechnol. 34, 393–400 (2016).
Goyal, S., Kim, S., Chen, I. S. Y. & Chou, T. Mechanisms of blood homeostasis: lineage tracking and a neutral model of cell populations in rhesus macaques. BMC Biol. 13, 85 (2015).
Lipsitz, Y. Y., Bedford, P., Davies, A. H., Timmins, N. E. & Zandstra, P. W. Achieving efficient manufacturing and quality assurance through synthetic cell therapy design. Cell Stem Cell 20, 13–17 (2017).
Todhunter, M. E. et al. Programmed synthesis of three-dimensional tissues. Nat. Methods 12, 975–981 (2015).
Leng, L., McAllister, A., Zhang, B., Radisic, M. & Günther, A. Mosaic hydrogels: one-step formation of multiscale soft materials. Adv. Mater. Weinheim 24, 3650–3658 (2012).
Kang, H. W. et al. A 3D bioprinting system to produce human-scale tissue constructs with structural integrity. Nat. Biotechnol. 34, 312–319 (2016).
Young, M. et al. A TRACER 3D co-culture tumour model for head and neck cancer. Biomaterials 164, 54–69 (2018).
Rodenhizer, D. et al. A three-dimensional engineered tumour for spatial snapshot analysis of cell metabolism and phenotype in hypoxic gradients. Nat. Mater. 15, 227–234 (2016).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Lancaster, M. A. et al. Guided self-organization and cortical plate formation in human brain organoids. Nat. Biotechnol. 35, 659–666 (2017).
Shao, Y. et al. A pluripotent stem cell-based model for post-implantation human amniotic sac development. Nat. Commun. 8, 208 (2017).
Shao, Y. et al. Self-organized amniogenesis by human pluripotent stem cells in a biomimetic implantation-like niche. Nat. Mater. 16, 419–425 (2017).
Cerchiari, A. E. et al. A strategy for tissue self-organization that is robust to cellular heterogeneity and plasticity. Proc. Nat. Acad. Sci. USA 112, 2287–2292 (2015).
Hughes, A. J. et al. Engineered tissue folding by mechanical compaction of the mesenchyme. Dev. Cell 44, 165–178 (2017).
Gjorevski, N. et al. Designer matrices for intestinal stem cell and organoid culture. Nature 539, 560–564 (2016).
Arora, N. et al. A process engineering approach to increase organoid yield. Development 144, 1128–1136 (2017).
Czerniecki, S. M. et al. High-throughput screening enhances kidney organoid differentiation from human pluripotent stem cells and enables automated multidimensional phenotyping. Cell Stem Cell 22, 929–940 (2018).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E. D. & Brivanlou, A. H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Morgani, S. M., Metzger, J. J., Nichols, J., Siggia, E. D. & Hadjantonakis, A. K. Micropattern differentiation of mouse pluripotent stem cells recapitulates embryo regionalized cell fate patterning. eLife 7, 1040 (2018).
Blin, G., Picart, C., Thery, M. & Puceat, M. Geometrical confinement guides Brachyury self-patterning in embryonic stem cells. bioRxiv https://doi.org/10.1101/138354 (2017).
Thery, M. Micropatterning as a tool to decipher cell morphogenesis and functions. J. Cell Sci. 123, 4201–4213 (2010).
Azioune, A., Storch, M., Bornens, M., Thery, M. & Piel, M. Simple and rapid process for single cell micro-patterning. Lab. Chip 9, 1640–1642 (2009).
Etoc, F. et al. A balance between secreted inhibitors and edge sensing controls gastruloid self-organization. Dev. Cell 39, 302–315 (2016).
Kunche, S., Yan, H., Calof, A. L., Lowengrub, J. S. & Lander, A. D. Feedback, lineages and self-organizing morphogenesis. PLOS Comput. Biol. 12, e1004814 (2016).
Rubenstein, M., Sai, Y., Chuong, C. M. & Shen, W. M. Regenerative patterning in swarm robots: mutual benefits of research in robotics & stem cell biology. Int. Dev, J. Biol. 53, 869–881 (2009).
Rubenstein, M., Cornejo, A. & Nagpal, R. Robotics. Programmable self-assembly in a thousand-robot swarm. Science 345, 795–799 (2014).
Finerty, J. C. Parabiosis in physiological studies. Physiol. Rev. 32, 277–302 (1952).
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005).
Loffredo, F. S. et al. Growth differentiation factor 11 is a circulating factor that reverses age-related cardiac hypertrophy. Cell 153, 828–839 (2013).
Katsimpardi, L. et al. Vascular and neurogenic rejuvenation of the aging mouse brain by young systemic factors. Science 344, 630–634 (2014).
Seok, J. et al. Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc. Nat. Acad. Sci. USA 110, 3507–3512 (2013).
Benam, K. H. et al. Engineered in vitro -disease models. Annu. Rev. Pathol. 10, 195–262 (2015).
Pammolli, F., Magazzini, L. & Riccaboni, M. The productivity crisis in pharmaceutical R&D. Nat. Rev. Drug Discov. 10, 428–438 (2011).
Prantil-Baun, R. et al. Physiologically based pharmacokinetic and pharmacodynamic analysis enabled by microfluidically linked organs-on-chips. Annu. Rev. Pharmacol. Toxicol. 58, 37–64 (2018).
Skardal, A. et al. Multi-tissue interactions in an integrated three-tissue organ-on a-chip platform. Sci. Rep. 7, 8837 (2017).
Huh, D. et al. Microfabrication of human organs-on-chips. Nat. Protoc. 8, 2135–2157 (2013).
Danhof, M. et al. Mechanism-based pharmacokinetic-pharmacodynamic modeling: biophase distribution, receptor theory, and dynamical systems analysis. Annu. Rev. Pharmacol. Toxicol. 47, 357–400 (2007).
Edington, C. D. et al. Interconnected microphysiological systems for quantitative biology and pharmacology studies. Sci. Rep. 8, 4530 (2018).
Kasendra, M. et al. Development of a primary human small intestine-on a-chip using biopsy-derived organoids. Sci. Rep. 8, 2 (2018).
Tsamandouras, N. et al. Integrated gut and liver microphysiological systems for quantitative in vitro pharmacokinetic studies. AAPS J. 19, 1499–1512 (2017).
Acknowledgements
The authors thank C. Bauwens for her insightful feedback on this review. The authors especially thank D. Lauffenburger (Massachusetts Institute of Technology), a strong proponent of the cue–signal–response paradigm and with whom P.W.Z. has had many helpful discussions on the engineering approach to biology over the years. The authors apologize to their colleagues whose important work could not be included because of space constraints. The authors are funded by the Canadian Institutes for Health Research and Medicine by Design, a Canada First Research Excellence Programme at the University of Toronto. P.W.Z. is the Canada Research Chair in Stem Cell Engineering.
Reviewer information
Nature Reviews Genetics thanks D. V. Schaffer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Cometing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Cell therapies
-
Clinical treatments that introduce living cellular material into a patient. They may engraft in the body, leading to long-term replacement of damaged or missing tissue, or stimulate endogenous repair and promote endogenous viability.
- Embryonic stem cell
-
(ESC). A type of pluripotent stem cell, derived from the inner cell mass of the developing embryo, that is responsible for giving rise to all of the cells in the developing fetus but not the extra-embryonic tissues.
- Organoid
-
A minimal and miniaturized organ that is developed from a suspension of stem cells in vitro. These stem cells undergo division and self-organization to give rise to a 3D structure that mimics the anatomy of organs in the body. Thus, organoids can serve as models for understanding organ development and for modelling disease states.
- Cell fate
-
A cell’s identity based on its expression of genetic, proteomic and epigenetic markers but also in terms of its functional abilities. Cell fate determines a cell’s self-renewal ability, proliferative ability, differentiation potential, survival and motility.
- Autocrine
-
A form of cellular signalling in which secreted chemicals bind to receptors on the same cell. By contrast, juxtacrine and paracrine signalling induce responses in neighbouring cells, either through direct contact (juxtacrine) or secreted chemicals (paracrine).
- Extracellular matrix
-
(ECM). A collection of extracellular molecules, including proteins, proteoglycans and polysaccharides, that supports the growth of nearby cells by providing biomechanical and biochemical cues. It enables cell adhesion and cell–cell communication.
- Gene regulatory networks
-
(GRNs). A set of genes and their direct and indirect regulatory interactions with one another. GRNs are akin to decision-making computational circuits that serve to process input signals and generate robust outputs in cell behaviour.
- Network motifs
-
Interaction patterns that recur more frequently than in randomized networks — for example, negative autoregulation (or ‘autorepression’) and the feedforward loop.
- Niches
-
The in vivo microenvironments in which stem cells reside that regulate their homeostasis and fate choices.
- Morphogenesis
-
The process by which developing organisms acquire their structure and shape.
- Bayesian networks
-
Probabilistic models that relate the dependencies of the expression of a set of genes on one another through a directed graph.
- Boolean networks
-
Models of gene regulatory networks that can predict gene expression outcomes given the initial state of genes in the network as well as the derivation of steady-state gene expression status.
- Artificial neural networks
-
Networks composed of nodes, which can be genes, that process and transmit information. The output of each node is a nonlinear function of a sum of its regulatory inputs.
- Ordinary differential equations
-
A mathematical framework capturing gene expression dynamics as a function of the presence of regulators and the rate of change of mRNA and/or protein concentration due to production and degradation.
- Reverse engineering
-
The process of analysing a system to uncover underlying design rules to create representations of the system at higher levels of abstraction (inverse of forward engineering).
- Forward engineering
-
The iterative process by which a system is designed, prototyped, tested and further optimized from a model (the classical engineering design process).
- Micropatterning
-
Technology that enables transfer of miniature ‘islands’ of extracellular matrix proteins to enforce control of the shape and size of adherent cells either as single cells or cell colonies.
- Stemness
-
The characteristic of a cell that makes it a stem cell. That is, the ability to self-renew and differentiate to specify to different cell types.
- Bioreactors
-
Vessels in which biological species, such as stem cells and their progeny, are grown, maintained and manipulated in a controlled environment (pH, oxygen and media change) for cell manufacturing pipelines.
- Bioprinting
-
Utilization of printing techniques ranging from inkjet printers to 3D printers to combine cells, biomaterials, extracellular matrix, growth factors, etc. to fabricate complex tissue surrogates in vitro.
- Fate patterning
-
A process during embryogenesis in which cell fates are allocated or ‘patterned’ as a function of space and time.
- Morphogens
-
Signalling molecules, typically soluble chemicals, for which the asymmetric distribution in a developing tissue gives rise to fate patterning and morphogenesis.
Rights and permissions
About this article
Cite this article
Tewary, M., Shakiba, N. & Zandstra, P.W. Stem cell bioengineering: building from stem cell biology. Nat Rev Genet 19, 595–614 (2018). https://doi.org/10.1038/s41576-018-0040-z
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-018-0040-z
This article is cited by
-
Single-Cell RNA Sequencing: Technological Progress and Biomedical Application in Cancer Research
Molecular Biotechnology (2023)
-
Robust and tunable signal processing in mammalian cells via engineered covalent modification cycles
Nature Communications (2022)
-
Stretch-boosted cell-mediated vascularization
Nature Biomedical Engineering (2021)
-
Endogenous suppression of WNT signalling in human embryonic stem cells leads to low differentiation propensity towards definitive endoderm
Scientific Reports (2021)
-
Microtechnology-based methods for organoid models
Microsystems & Nanoengineering (2020)