Abstract
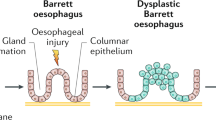
Barrett oesophagus, in which a metaplastic columnar mucosa that can predispose individuals to cancer development lines a portion of the distal oesophagus, is the only known precursor of oesophageal adenocarcinoma, the incidence of which has increased profoundly over the past several decades. Most evidence suggests that Barrett oesophagus develops from progenitor cells at the oesophagogastric junction that proliferate and undergo epithelial–mesenchymal transition as part of a wound-healing process that replaces oesophageal squamous epithelium damaged by gastroesophageal reflux disease (GERD). GERD also seems to induce reprogramming of key transcription factors in the progenitor cells, resulting in the development of the specialized intestinal metaplasia that is characteristic of Barrett oesophagus, probably through an intermediate step of metaplasia to cardiac mucosa. Genome-wide association studies suggest that patients with GERD who develop Barrett oesophagus might have an inherited predisposition to oesophageal metaplasia and that there is a shared genetic susceptibility to Barrett oesophagus and to several of its risk factors (such as GERD, obesity and cigarette smoking). In this Review, we discuss the mechanisms, pathophysiology, genetic predisposition and cells of origin of Barrett oesophagus, and opine on the clinical implications and future research directions.
Key points
-
Barrett oesophagus is defined conceptually as the condition in which a metaplastic mucosa that can predispose to cancer development lines a portion of the distal oesophagus.
-
Patients with Barrett oesophagus can have an inherited predisposition to its development, involving germline susceptibility to reflux oesophagitis, to risk factors such as gastroesophageal reflux disease (GERD) and obesity, and to tissue-specific gene expression that might favour oesophageal metaplasia.
-
Barrett oesophagus seems to develop when oesophageal squamous cells destroyed by GERD are replaced by columnar progenitor cells at the oesophagogastric junction that proliferate as part of a wound-healing process.
-
Potential cells of origin for Barrett metaplasia include basal cells of oesophageal squamous epithelium, oesophageal submucosal gland cells, transitional basal cells, residual embryonic cells, gastric gland cells and cells of compact mucous glands.
-
GERD induces the reprogramming of key transcription factors in progenitor cells to produce the specialized intestinal metaplasia with goblet cells, which is characteristic of Barrett oesophagus, probably through an intermediate step of metaplasia to cardiac mucosa.
-
Short and long segments of the specialized intestinal metaplasia seem to develop through the same pathophysiological mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Spechler, S. J. & Souza, R. F. Barrett’s esophagus. N. Engl. J. Med. 371, 836–845 (2014).
Cook, M. B. & Thrift, A. P. Epidemiology of Barrett’s esophagus and esophageal adenocarcinoma: implications for screening and surveillance. Gastrointest. Endosc. Clin. North. Am. 31, 1–26 (2021).
Sawas, T. et al. Identification of prognostic phenotypes of esophageal adenocarcinoma in 2 independent cohorts. Gastroenterology 155, 1720–1728.e4 (2018).
Curtius, K., Rubenstein, J. H., Chak, A. & Inadomi, J. M. Computational modelling suggests that Barrett’s oesophagus may be the precursor of all oesophageal adenocarcinomas. Gut 70, 435–1440 (2021).
Nowicki-Osuch, K. et al. Molecular phenotyping reveals the identity of Barrett’s esophagus and its malignant transition. Science 373, 760–767 (2021).
Ek, W. E. et al. Germline genetic contributions to risk for esophageal adenocarcinoma, Barrett’s esophagus, and gastroesophageal reflux. J. Natl Cancer Inst. 105, 1711–1718 (2013).
Palles, C. et al. Polymorphisms near TBX5 and GDF7 are associated with increased risk for Barrett’s esophagus. Gastroenterology 148, 367–378 (2015).
Levine, D. M. et al. A genome-wide association study identifies new susceptibility loci for esophageal adenocarcinoma and Barrett’s esophagus. Nat. Genet. 45, 1487–1493 (2013).
Gharahkhani, P. et al. Genome-wide association studies in oesophageal adenocarcinoma and Barrett’s oesophagus: a large-scale meta-analysis. Lancet Oncol. 17, 1363–1373 (2016).
Dong, J. et al. Sex-specific genetic associations for barrett’s esophagus and esophageal adenocarcinoma. Gastroenterology 159, 2065–2076.e1 (2020).
Sun, X. et al. Genomic regions associated with susceptibility to Barrett’s esophagus and esophageal adenocarcinoma in African Americans: the cross BETRNet admixture study. PLoS ONE 12, e0184962 (2017).
Su, Z. et al. Common variants at the MHC locus and at chromosome 16q24.1 predispose to Barrett’s esophagus. Nat. Genet. 44, 1131–1136 (2012).
Spechler, S. J. Cardiac mucosa: the heart of the problem. Gut 64, 1673–1674 (2015).
Noffsinger, A. E. in Fenoglio-Preiser’s Gastrointestinal Pathology 136–223 (Wolters Kluwer, 2017).
Chandrasoma, P. Pathophysiology of Barrett’s esophagus. Semin. Thorac. Cardiovasc. Surg. 9, 270–278 (1997).
Chandrasoma, P., Makarewicz, K., Wickramasinghe, K., Ma, Y. & Demeester, T. A proposal for a new validated histological definition of the gastroesophageal junction. Hum. Pathol. 37, 40–47 (2006).
Chandrasoma, P., Wijetunge, S., Demeester, S. R., Hagen, J. & Demeester, T. R. The histologic squamo-oxyntic gap: an accurate and reproducible diagnostic marker of gastroesophageal reflux disease. Am. J. Surg. Pathol. 34, 1574–1581 (2010).
Park, Y. S., Park, H. J., Kang, G. H., Kim, C. J. & Chi, J. G. Histology of gastroesophageal junction in fetal and pediatric autopsy. Arch. Pathol. Lab. Med. 127, 451–455 (2003).
Kilgore, S. P. et al. The gastric cardia: fact or fiction? Am. J. Gastroenterol. 95, 921–924 (2000).
Dunn, L. J., Burt, A. D., Hayes, N. & Griffin, S. M. Columnar metaplasia in the esophageal remnant after esophagectomy: a common occurrence and a valuable insight into the development of Barrett esophagus. Ann. Surg. 264, 1016–1021 (2016).
Dias Pereira, A. & Chaves, P. Columnar-lined oesophagus without intestinal metaplasia: results from a cohort with a mean follow-up of 7 years. Aliment. Pharmacol. Ther. 36, 282–289 (2012).
Ellison, E., Hassall, E. & Dimmick, J. E. Mucin histochemistry of the developing gastroesophageal junction. Pediatr. Pathol. Lab. Med. 16, 195–206 (1996).
Hahn, H. P. et al. Intestinal differentiation in metaplastic, nongoblet columnar epithelium in the esophagus. Am. J. Surg. Pathol. 33, 1006–1015 (2009).
Liu, W., Hahn, H., Odze, R. D. & Goyal, R. K. Metaplastic esophageal columnar epithelium without goblet cells shows DNA content abnormalities similar to goblet cell-containing epithelium. Am. J. Gastroenterol. 104, 816–824 (2009).
Robertson, E. V. et al. Central obesity in asymptomatic volunteers is associated with increased intrasphincteric acid reflux and lengthening of the cardiac mucosa. Gastroenterology 145, 730–739 (2013).
Robertson, E. V. et al. Hiatus hernia in healthy volunteers is associated with intrasphincteric reflux and cardiac mucosal lengthening without traditional reflux. Gut 66, 1208–1215 (2017).
Odze, R. et al. Histologic study of the esophagogastric junction of organ donors reveals novel glandular structures in normal esophageal and gastric mucosae. Clin. Transl. Gastroenterol. 12, e00346 (2021).
Paull, A. et al. The histologic spectrum of Barrett’s esophagus. N. Engl. J. Med. 295, 476–480 (1976).
Chen, W. et al. Significance of Paneth cell metaplasia in Barrett esophagus: a morphologic and clinicopathologic study. Am. J. Clin. Pathol. 143, 665–671 (2015).
Evans, J. A. et al. Clonal transitions and phenotypic evolution in Barrett’s esophagus. Gastroenterology 162, 1197–1209 (2022).
McClave, S. A., Boyce, H. W. Jr. & Gottfried, M. R. Early diagnosis of columnar-lined esophagus: a new endoscopic diagnostic criterion. Gastrointest. Endosc. 33, 413–416 (1987).
Ishimura, N. et al. Questionnaire-based survey conducted in 2011 concerning endoscopic management of Barrett’s esophagus in East Asian countries. Digestion 86, 136–146 (2012).
Sharma, P., Morales, T. G. & Sampliner, R. E. Short segment Barrett’s esophagus — the need for standardization of the definition and of endoscopic criteria. Am. J. Gastroenterol. 93, 1033–1036 (1998).
Shaheen, N. J., Falk, G. W., Iyer, P. G. & Gerson, L. B. ACG clinical guideline: diagnosis and management of Barrett’s esophagus. Am. J. Gastroenterol. 111, 30–50 (2016).
Fitzgerald, R. C. et al. British Society of Gastroenterology guidelines on the diagnosis and management of Barrett’s oesophagus. Gut 63, 7–42 (2014).
Weusten, B. et al. Endoscopic management of Barrett’s esophagus: European Society of Gastrointestinal Endoscopy (ESGE) Position Statement. Endoscopy 49, 191–198 (2017).
Merlo, L. M., Kosoff, R. E., Gardiner, K. L. & Maley, C. C. An in vitro co-culture model of esophageal cells identifies ascorbic acid as a modulator of cell competition. BMC Cancer 11, 461 (2011).
Spechler, S. J., Sharma, P., Souza, R. F., Inadomi, J. M. & Shaheen, N. J. American Gastroenterological Association medical position statement on the management of Barrett’s esophagus. Gastroenterology 140, 1084–1091 (2011).
Iascone, C., DeMeester, T. R., Little, A. G. & Skinner, D. B. Barrett’s esophagus. Functional assessment, proposed pathogenesis, and surgical therapy. Arch. Surg. 118, 543–549 (1983).
Zaninotto, G., DeMeester, T. R., Bremner, C. G., Smyrk, T. C. & Cheng, S. C. Esophageal function in patients with reflux-induced strictures and its relevance to surgical treatment. Ann. Thorac. Surg. 47, 362–370 (1989).
Gray, M. R., Donnelly, R. J. & Kingsnorth, A. N. Role of salivary epidermal growth factor in the pathogenesis of Barrett’s columnar lined oesophagus. Br. J. Surg. 78, 1461–1466 (1991).
Johnson, D. A., Winters, C., Spurling, T. J., Chobanian, S. J. & Cattau, E. L. Jr. Esophageal acid sensitivity in Barrett’s esophagus. J. Clin. Gastroenterol. 9, 23–27 (1987).
Spechler, S. J. The columnar lined oesophagus: a riddle wrapped in a mystery inside an enigma. Gut 41, 710–711 (1997).
Spechler, S. J., Zeroogian, J. M., Antonioli, D. A., Wang, H. H. & Goyal, R. K. Prevalence of metaplasia at the gastro-oesophageal junction. Lancet 344, 1533–1536 (1994).
Johnston, M. H., Hammond, A. S., Laskin, W. & Jones, D. M. The prevalence and clinical characteristics of short segments of specialized intestinal metaplasia in the distal esophagus on routine endoscopy. Am. J. Gastroenterol. 91, 1507–1511 (1996).
Nandurkar, S., Talley, N. J., Martin, C. J., Ng, T. H. & Adams, S. Short segment Barrett’s oesophagus: prevalence, diagnosis and associations. Gut 40, 710–715 (1997).
Chalasani, N., Wo, J. M., Hunter, J. G. & Waring, J. P. Significance of intestinal metaplasia in different areas of esophagus including esophagogastric junction. Dig. Dis. Sci. 42, 603–607 (1997).
Kahrilas, P. J. et al. The acid pocket: a target for treatment in reflux disease? Am. J. Gastroenterol. 108, 1058–1064 (2013).
Fletcher, J., Wirz, A., Henry, E. & McColl, K. E. Studies of acid exposure immediately above the gastro-oesophageal squamocolumnar junction: evidence of short segment reflux. Gut 53, 168–173 (2004).
Pandolfino, J. E. et al. Comparison of esophageal acid exposure at 1 cm and 6 cm above the squamocolumnar junction using the Bravo pH monitoring system. Dis. Esophagus 19, 177–182 (2006).
Iijima, K. et al. Dietary nitrate generates potentially mutagenic concentrations of nitric oxide at the gastroesophageal junction. Gastroenterology 122, 1248–1257 (2002).
Spechler, S. J. Are we underestimating acid reflux? Gut 53, 162–163 (2004).
Tosh, D. & Slack, J. M. How cells change their phenotype. Nat. Rev. Mol. Cell Biol. 3, 187–194 (2002).
Wang, D. H. The Esophageal squamous epithelial cell-still a reasonable candidate for the Barrett’s esophagus cell of origin? Cell. Mol. Gastroenterol. Hepatol. 4, 157–160 (2017).
Que, J., Garman, K. S., Souza, R. F. & Spechler, S. J. Pathogenesis and cells of origin of Barrett’s esophagus. Gastroenterology 157, 349–364.e1 (2019).
Willet, S. G. et al. Regenerative proliferation of differentiated cells by mTORC1-dependent paligenosis. EMBO J. 37, e98311 (2018).
Jin, R. U. & Mills, J. C. Are gastric and esophageal metaplasia relatives? The case for Barrett’s stemming from SPEM. Dig. Dis. Sci. 63, 2028–2041 (2018).
Brown, J. W., Cho, C. J. & Mills, J. C. Paligenosis: cellular remodeling during tissue repair. Annu. Rev. Physiol. 84, 461–483 (2022).
Zhang, W. & Wang, D. H. Origins of metaplasia in Barrett’s esophagus: is this an esophageal stem or progenitor cell disease? Dig. Dis. Sci. 63, 2005–2012 (2018).
Agoston, A. T. et al. Columnar-lined esophagus develops via wound repair in a surgical model of reflux esophagitis. Cell. Mol. Gastroenterol. Hepatol. 6, 389–404 (2018).
Asanuma, K. et al. In oesophageal squamous cells, nitric oxide causes S-nitrosylation of Akt and blocks SOX2 (sex determining region Y-Box 2) expression. Gut 65, 1416–1426 (2016).
Minacapelli, C. D. et al. Barrett’s metaplasia develops from cellular reprograming of esophageal squamous epithelium due to gastroesophageal reflux. Am. J. Physiol. Gastrointest. Liver Physiol. 312, G615–G622 (2017).
Wang, D. H. et al. Aberrant epithelial-mesenchymal Hedgehog signaling characterizes Barrett’s metaplasia. Gastroenterology 138, 1810–1822 (2010).
Wang, D. H. et al. Hedgehog signaling regulates FOXA2 in esophageal embryogenesis and Barrett’s metaplasia. J. Clin. Invest. 124, 3767–3780 (2014).
Roudebush, C., Catala-Valentin, A., Andl, T., Le Bras, G. F. & Andl, C. D. Activin A-mediated epithelial de-differentiation contributes to injury repair in an in vitro gastrointestinal reflux model. Cytokine 123, 154782 (2019).
Zhou, G. et al. Acid and bile salt up-regulate BMP4 expression in human esophageal epithelium cells. Scand. J. Gastroenterol. 44, 926–932 (2009).
Glickman, J. N. et al. Multilayered epithelium in mucosal biopsy specimens from the gastroesophageal junction region is a histologic marker of gastroesophageal reflux disease. Am. J. Surg. Pathol. 33, 818–825 (2009).
Chen, X. et al. Multilayered epithelium in a rat model and human Barrett’s esophagus: similar expression patterns of transcription factors and differentiation markers. BMC Gastroenterol. 8, 1 (2008).
Shields, H. M. et al. Prospective evaluation of multilayered epithelium in Barrett’s esophagus. Am. J. Gastroenterol. 96, 3268–3273 (2001).
Abdulnour-Nakhoul, S. et al. Characterization of esophageal submucosal glands in pig tissue and cultures. Dig. Dis. Sci. 52, 3054–3065 (2007).
Zhang, X. et al. The microscopic anatomy of the esophagus including the individual layers, specialized tissues, and unique components and their responses to injury. Ann. NY Acad. Sci. 1434, 304–318 (2018).
Leedham, S. J. et al. Individual crypt genetic heterogeneity and the origin of metaplastic glandular epithelium in human Barrett’s oesophagus. Gut 57, 1041–1048 (2008).
Coad, R. A. et al. On the histogenesis of Barrett’s oesophagus and its associated squamous islands: a three-dimensional study of their morphological relationship with native oesophageal gland ducts. J. Pathol. 206, 388–394 (2005).
Garman, K. S. Origin of Barrett’s epithelium: esophageal submucosal glands. Cell. Mol. Gastroenterol. Hepatol. 4, 153–156 (2017).
Owen, R. P. et al. Single cell RNA-seq reveals profound transcriptional similarity between Barrett’s oesophagus and oesophageal submucosal glands. Nat. Commun. 9, 4261 (2018).
Gillen, P., Keeling, P., Byrne, P. J., West, A. B. & Hennessy, T. P. Experimental columnar metaplasia in the canine oesophagus. Br. J. Surg. 75, 113–115 (1988).
Li, H. et al. Mechanisms of columnar metaplasia and squamous regeneration in experimental Barrett’s esophagus. Surgery 115, 176–181 (1994).
Krüger, L. et al. Ductular and proliferative response of esophageal submucosal glands in a porcine model of esophageal injury and repair. Am. J. Physiol. Gastrointest. Liver Physiol. 313, G180–G191 (2017).
Van Nieuwenhove, Y. & Willems, G. Gastroesophageal reflux triggers proliferative activity of the submucosal glands in the canine esophagus. Dis. Esophagus 11, 89–93 (1998).
von Furstenberg, R. J. et al. Porcine esophageal submucosal gland culture model shows capacity for proliferation and differentiation. Cell. Mol. Gastroenterol. Hepatol. 4, 385–404 (2017).
Sayin, S. I., Baumeister, T., Wang, T. C. & Quante, M. Origins of metaplasia in the esophagus: is this a GE junction stem cell disease? Dig. Dis. Sci. 63, 2013–2021 (2018).
Quante, M. et al. Bile acid and inflammation activate gastric cardia stem cells in a mouse model of Barrett-like metaplasia. Cancer Cell 21, 36–51 (2012).
Lee, Y. et al. Gastrin stimulates a cholecystokinin-2-receptor-expressing cardia progenitor cell and promotes progression of Barrett’s-like esophagus. Oncotarget 8, 203–214 (2017).
Wang, X. et al. Residual embryonic cells as precursors of a Barrett’s-like metaplasia. Cell 145, 1023–1035 (2011).
Jiang, M. et al. Transitional basal cells at the squamous-columnar junction generate Barrett’s oesophagus. Nature 550, 529–533 (2017).
Vercauteren Drubbel, A. et al. Reactivation of the Hedgehog pathway in esophageal progenitors turns on an embryonic-like program to initiate columnar metaplasia. Cell Stem Cell 28, 1411–1427 (2021).
Sarosi, G. et al. Bone marrow progenitor cells contribute to esophageal regeneration and metaplasia in a rat model of Barrett’s esophagus. Dis. Esophagus 21, 43–50 (2008).
Hutchinson, L. et al. Human Barrett’s adenocarcinoma of the esophagus, associated myofibroblasts, and endothelium can arise from bone marrow-derived cells after allogeneic stem cell transplant. Stem Cell Dev. 20, 11–17 (2011).
Gurtner, G. C., Werner, S., Barrandon, Y. & Longaker, M. T. Wound repair and regeneration. Nature 453, 314–321 (2008).
Armstrong, D. et al. The endoscopic assessment of esophagitis: a progress report on observer agreement. Gastroenterology 111, 85–92 (1996).
Mills, J. C. & Sansom, O. J. Reserve stem cells: differentiated cells reprogram to fuel repair, metaplasia, and neoplasia in the adult gastrointestinal tract. Sci. Signal. 8, re8 (2015).
Slack, J. M. Metaplasia and transdifferentiation: from pure biology to the clinic. Nat. Rev. Mol. Cell Biol. 8, 369–378 (2007).
Oberg, S., Johansson, J., Wenner, J. & Walther, B. Metaplastic columnar mucosa in the cervical esophagus after esophagectomy. Ann. Surg. 235, 338–345 (2002).
Vega, K. J. et al. Identification of the putative intestinal stem cell marker doublecortin and CaM kinase-like-1 in Barrett’s esophagus and esophageal adenocarcinoma. J. Gastroenterol. Hepatol. 27, 773–780 (2012).
Yang, J. et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 21, 341–352 (2020).
Zhang, Q. et al. Acidic bile salts induce epithelial to mesenchymal transition via VEGF signaling in non-neoplastic Barrett’s cells. Gastroenterology 156, 130–144.e110 (2019).
Kunze, B. et al. Notch signaling mediates differentiation in Barrett’s esophagus and promotes progression to adenocarcinoma. Gastroenterology 159, 575–590 (2020).
Kalluri, R. & Weinberg, R. A. The basics of epithelial-mesenchymal transition. J. Clin. Invest. 119, 1420–1428 (2009).
Peters, Y. et al. Barrett oesophagus. Nat. Rev. Dis. Primers 5, 35 (2019).
Cameron, A. J. et al. Gastroesophageal reflux disease in monozygotic and dizygotic twins. Gastroenterology 122, 55–59 (2002).
Mohammed, I., Cherkas, L. F., Riley, S. A., Spector, T. D. & Trudgill, N. J. Genetic influences in gastro-oesophageal reflux disease: a twin study. Gut 52, 1085–1089 (2003).
An, J. et al. Gastroesophageal reflux GWAS identifies risk loci that also associate with subsequent severe esophageal diseases. Nat. Commun. 10, 4219 (2019).
Dai, J. Y. et al. A newly identified susceptibility locus near FOXP1 modifies the association of gastroesophageal reflux with Barrett’s esophagus. Cancer Epidemiol. Biomark. Prev. 24, 1739–1747 (2015).
Thrift, A. P. et al. Obesity and risk of esophageal adenocarcinoma and Barrett’s esophagus: a Mendelian randomization study. J. Natl Cancer Inst. 106, dju252 (2014).
Böhmer, A. C. et al. Shared genetic etiology of obesity-related traits and Barrett’s esophagus/adenocarcinoma: insights from genome-wide association studies. Cancer Epidemiol. Biomark. Prev. 29, 427–433 (2020).
Singh, S. et al. Central adiposity is associated with increased risk of esophageal inflammation, metaplasia, and adenocarcinoma: a systematic review and meta-analysis. Clin. Gastroenterol. Hepatol. 11, 1399–1412.e7 (2013).
Dighe, S. G. et al. Germline variation in the insulin-like growth factor pathway and risk of Barrett’s esophagus and esophageal adenocarcinoma. Carcinogenesis 42, 369–377 (2021).
Consortium, G. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Schröder, J. et al. Identification of loci of functional relevance to Barrett’s esophagus and esophageal adenocarcinoma: cross-referencing of expression quantitative trait loci data from disease-relevant tissues with genetic association data. PLoS ONE 14, e0227072 (2019).
Jin, E. H. et al. A novel susceptibility locus near GRIK2 associated with erosive esophagitis in a Korean cohort. Clin. Transl. Gastroenterol. 11, e00145 (2020).
Wu, C. S. et al. Glutamate receptor, ionotropic, kainate 2 silencing by DNA hypermethylation possesses tumor suppressor function in gastric cancer. Int. J. Cancer 126, 2542–2552 (2010).
Brassai, A., Suvanjeiev, R. G., Bán, E. G. & Lakatos, M. Role of synaptic and nonsynaptic glutamate receptors in ischaemia induced neurotoxicity. Brain Res. Bull. 112, 1–6 (2015).
Browning, K. N. & Travagli, R. A. Plasticity of vagal brainstem circuits in the control of gastric function. Neurogastroenterol. Motil. Soc. 22, 1154–1163 (2010).
Tack, J. & Pandolfino, J. E. Pathophysiology of gastroesophageal reflux disease. Gastroenterology 154, 277–288 (2018).
Mittal, R. K., Holloway, R. H., Penagini, R., Blackshaw, L. A. & Dent, J. Transient lower esophageal sphincter relaxation. Gastroenterology 109, 601–610 (1995).
Buas, M. F. et al. Germline variation in inflammation-related pathways and risk of Barrett’s oesophagus and oesophageal adenocarcinoma. Gut 66, 1739–1747 (2017).
Feagins, L. A. et al. Mechanisms of oxidant production in esophageal squamous cell and Barrett’s cell lines. Am. J. Physiol. Gastrointest. Liver Physiol. 294, G411–G417 (2008).
Banerjee, B. et al. Clinical study of ursodeoxycholic acid in Barrett’s esophagus patients. Cancer Prev. Res. 9, 528–533 (2016).
Battle, A., Brown, C. D., Engelhardt, B. E. & Montgomery, S. B. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Zhang, H. Y. et al. In non-neoplastic Barrett’s epithelial cells, acid exerts early antiproliferative effects through activation of the Chk2 pathway. Cancer Res. 67, 8580–8587 (2007).
Huo, X. et al. Aspirin prevents NF-κB activation and CDX2 expression stimulated by acid and bile salts in oesophageal squamous cells of patients with Barrett’s oesophagus. Gut 67, 606–615 (2018).
Souza, R. F. et al. Differences in ERK activation in squamous mucosa in patients who have gastroesophageal reflux disease with and without Barrett’s esophagus. Am. J. Gastroenterol. 100, 551–559 (2005).
Yang, L. et al. Inflammation and intestinal metaplasia of the distal esophagus are associated with alterations in the microbiome. Gastroenterology 137, 588–597 (2009).
Verbeek, R. E. et al. Toll-like receptor 4 activation in Barrett’s esophagus results in a strong increase in COX-2 expression. J. Gastroenterol. 49, 1121–1134 (2014).
Jakszyn, P. & Gonzalez, C. A. Nitrosamine and related food intake and gastric and oesophageal cancer risk: a systematic review of the epidemiological evidence. World J. Gastroenterol. 12, 4296–4303 (2006).
Endo, H. et al. Exogenous luminal nitric oxide exposure accelerates columnar transformation of rat esophagus. Int. J. Cancer 127, 2009–2019 (2010).
Souza, R. F. et al. Gastroesophageal reflux might cause esophagitis through a cytokine-mediated mechanism rather than caustic acid injury. Gastroenterology 137, 1776–1784 (2009).
Dunbar, K. B. et al. Association of acute gastroesophageal reflux disease with esophageal histologic changes. JAMA 315, 2104–2112 (2016).
Lind, A., Koenderman, L., Kusters, J. G. & Siersema, P. D. Squamous tissue lymphocytes in the esophagus of controls and patients with reflux esophagitis and Barrett’s esophagus are characterized by a non-inflammatory phenotype. PLoS ONE 9, e106261 (2014).
Sen, M. et al. Flow based single cell analysis of the immune landscape distinguishes Barrett’s esophagus from adjacent normal tissue. Oncotarget 10, 3592–3604 (2019).
Kong, J. et al. Immature myeloid progenitors promote disease progression in a mouse model of Barrett’s-like metaplasia. Oncotarget 6, 32980–33005 (2015).
Moons, L. M. et al. Barrett’s oesophagus is characterized by a predominantly humoral inflammatory response. J. Pathol. 207, 269–276 (2005).
Somja, J. et al. Dendritic cells in Barrett’s esophagus carcinogenesis: an inadequate microenvironment for antitumor immunity? Am. J. Pathol. 182, 2168–2179 (2013).
Jeong, Y. et al. Identification and genetic manipulation of human and mouse oesophageal stem cells. Gut 65, 1077–1086 (2016).
Dobson, H., Pignatelli, M., Hopwood, D. & D’Arrigo, C. Cell adhesion molecules in oesophageal epithelium. Gut 35, 1343–1347 (1994).
Ali, I. et al. Dickkopf homologs in squamous mucosa of esophagitis patients are overexpressed compared with Barrett’s patients and healthy controls. Am. J. Gastroenterol. 101, 1437–1448 (2006).
Zhang, H. Y. et al. Differences in activity and phosphorylation of MAPK enzymes in esophageal squamous cells of GERD patients with and without Barrett’s esophagus. Am. J. Physiol. Gastrointest. Liver Physiol. 295, G470–G478 (2008).
Qu, Z. et al. Proteomic quantification and site-mapping of S-nitrosylated proteins using isobaric iodoTMT reagents. J. Proteome Res. 13, 3200–3211 (2014).
Gu, Z., Nakamura, T. & Lipton, S. A. Redox reactions induced by nitrosative stress mediate protein misfolding and mitochondrial dysfunction in neurodegenerative diseases. Mol. Neurobiol. 41, 55–72 (2010).
Macfarlane, S., Furrie, E., Macfarlane, G. T. & Dillon, J. F. Microbial colonization of the upper gastrointestinal tract in patients with Barrett’s esophagus. Clin. Infect. Dis. 45, 29–38 (2007).
Sharma, P. et al. The development and validation of an endoscopic grading system for Barrett’s esophagus: the Prague C & M criteria. Gastroenterology 131, 1392–1399 (2006).
Spechler, S. J. Intestinal metaplasia at the gastroesophageal junction. Gastroenterology 126, 567–575 (2004).
Jung, K. W. et al. Epidemiology and natural history of intestinal metaplasia of the gastroesophageal junction and Barrett’s esophagus: a population-based study. Am. J. Gastroenterol. 106, 1447–1455 (2011).
Itskoviz, D. et al. Risk of neoplastic progression among patients with an irregular Z line on long-term follow-up. Dig. Dis. Sci. 63, 1513–1517 (2018).
Thota, P. N. et al. Low risk of high-grade dysplasia or esophageal adenocarcinoma among patients with Barrett’s esophagus less than 1cm (irregular Z line) within 5 years of index endoscopy. Gastroenterology 152, 987–992 (2017).
Pohl, H. et al. Length of Barrett’s oesophagus and cancer risk: implications from a large sample of patients with early oesophageal adenocarcinoma. Gut 65, 196–201 (2016).
Barrie, J., Yanni, F., Sherif, M., Dube, A. K. & Tamhankar, A. P. Length of Barrett’s esophagus in the presence of low-grade dysplasia, high-grade dysplasia, and adenocarcinoma. Surg. Endosc. 35, 4756–4762 (2021).
Wani, S. et al. An analysis of the GIQuIC nationwide quality registry reveals unnecessary surveillance endoscopies in patients with normal and irregular Z-Lines. Am. J. Gastroenterol. 115, 1869–1878 (2020).
Evans, J. A. et al. The role of endoscopy in Barrett’s esophagus and other premalignant conditions of the esophagus. Gastrointest. Endosc. 76, 1087–1094 (2012).
Whiteman, D. C. et al. Australian clinical practice guidelines for the diagnosis and management of Barrett’s esophagus and early esophageal adenocarcinoma. J. Gastroenterol. Hepatol. 30, 804–820 (2015).
Virchow, R. Ueber metaplasie: vortrag, gehalten auf dem internationalen medicinischen Congress in Kopenhagen. Virchows Arch. 97, 21 (1884).
Spechler, S. J. Screening for Barrett’s oesophagus: are we looking for the right thing? Gut 70, 1426–1427 (2021).
Krishnamoorthi, R. et al. Factors associated with progression of Barrett’s esophagus: a systematic review and meta-analysis. Clin. Gastroenterol. Hepatol. 16, 1046–1055.e8 (2018).
Acknowledgements
The authors acknowledge funding from the US National Institutes of Health (R01-DK124185 to R.F.S. and S.J.S.) and the Baylor Scott & White Research Institute.
Author information
Authors and Affiliations
Contributions
Both authors contributed equally to all aspects of the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.F.S. has served as a consultant for Interpace Diagnostics, Castle Biosciences, Ironwood Pharmaceuticals, Phathom Pharmaceuticals, IsoThrive, CDx Diagnostics and AstraZeneca and receives research support from Phathom Pharmaceuticals and Sanofi. S.J.S. has served as a consultant for Interpace Diagnostics, Castle Biosciences, Phathom Pharmaceuticals, IsoThrive, Takeda Pharmaceuticals and Ironwood Pharmaceuticals and receives royalties as an author for UpToDate.
Peer review
Peer review information
Nature Reviews Gastroenterology & Hepatology thanks Gary Falk, Michael Quante and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Souza, R.F., Spechler, S.J. Mechanisms and pathophysiology of Barrett oesophagus. Nat Rev Gastroenterol Hepatol 19, 605–620 (2022). https://doi.org/10.1038/s41575-022-00622-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41575-022-00622-w
This article is cited by
-
Decoding spatiotemporal transcriptional dynamics and epithelial fibroblast crosstalk during gastroesophageal junction development through single cell analysis
Nature Communications (2024)
-
Pathophysiology of gastro-oesophageal reflux disease: implications for diagnosis and management
Nature Reviews Gastroenterology & Hepatology (2024)