Abstract
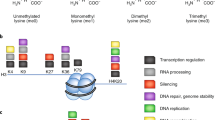
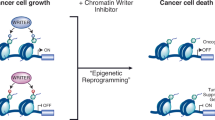
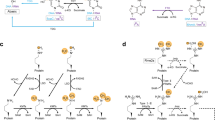
Protein lysine methylation is a crucial post-translational modification that regulates the functions of both histone and non-histone proteins. Deregulation of the enzymes or ‘writers’ of protein lysine methylation, lysine methyltransferases (KMTs), is implicated in the cause of many diseases, including cancer, mental health disorders and developmental disorders. Over the past decade, significant advances have been made in developing drugs to target KMTs that are involved in histone methylation and epigenetic regulation. The first of these inhibitors, tazemetostat, was recently approved for the treatment of epithelioid sarcoma and follicular lymphoma, and several more are in clinical and preclinical evaluation. Beyond chromatin, the many KMTs that regulate protein synthesis and other fundamental biological processes are emerging as promising new targets for drug development to treat diverse diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fischer, E. H. Phosphorylase and the origin of reversible protein phosphorylation. Biol. Chem. 391, 131–137 (2010).
Chandra, H. S. et al. Philadelphia chromosome symposium: commemoration of the 50th anniversary of the discovery of the Ph chromosome. Cancer Genet. 204, 171–179 (2011).
Wang, P., Royer, M. & Houtz, R. L. Affinity purification of ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit epsilon N-methyltransferase. Protein Expr. Purif. 6, 528–536 (1995).
Rothbart, S. B. & Baylin, S. B. Epigenetic therapy for epithelioid sarcoma. Cell 181, 211 (2020).
Cao, X. J. & Garcia, B. A. Global proteomics analysis of protein lysine methylation. Curr. Protoc. Protein Sci. https://doi.org/10.1002/cpps.16 (2016).
Husmann, D. & Gozani, O. Histone lysine methyltransferases in biology and disease. Nat. Struct. Mol. Biol. 26, 880–889 (2019).
Ambler, R. P. & Rees, M. W. Epsilon-N-methyl-lysine in bacterial flagellar protein. Nature 184, 56–57 (1959).
Murray, K. The occurrence of epsilon-N-methyl lysine in histones. Biochemistry 3, 10–15 (1964).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000). This study identfied the first histone lysine methyltransferase and established a connection between histone methylation and epigenetic mechanisms.
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Lachner, M., O’Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Cornett, E. M., Ferry, L., Defossez, P. A. & Rothbart, S. B. Lysine methylation regulators moonlighting outside the epigenome. Mol. Cell 75, 1092–1101 (2019).
Shi, X. et al. ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442, 96–99 (2006).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
Liu, S. et al. METTL13 methylation of eEF1A increases translational output to promote tumorigenesis. Cell 176, 491–504.e421 (2019). This study established a role for non-histone methylation by METTL13 in the regulation of translational elongation to promote RAS-driven cancers.
Andrews, F. H., Strahl, B. D. & Kutateladze, T. G. Insights into newly discovered marks and readers of epigenetic information. Nat. Chem. Biol. 12, 662–668 (2016).
Botuyan, M. V. et al. Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006).
Matthews, A. G. et al. RAG2 PHD finger couples histone H3 lysine 4 trimethylation with V(D)J recombination. Nature 450, 1106–1110 (2007).
Vermeulen, M. et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131, 58–69 (2007).
Kuo, A. J. et al. The BAH domain of ORC1 links H4K20me2 to DNA replication licensing and Meier-Gorlin syndrome. Nature 484, 115–119 (2012).
Wilkinson, A. W. et al. SETD3 is an actin histidine methyltransferase that prevents primary dystocia. Nature 565, 372–376 (2019).
Kwiatkowski, S. et al. SETD3 protein is the actin-specific histidine N-methyltransferase. eLife 7, e37921 (2018).
Lacoste, N., Utley, R. T., Hunter, J. M., Poirier, G. G. & Côte, J. Disruptor of telomeric silencing-1 is a chromatin-specific histone H3 methyltransferase. J. Biol. Chem. 277, 30421–30424 (2002).
Ng, H. H., Xu, R. M., Zhang, Y. & Struhl, K. Ubiquitination of histone H2B by Rad6 is required for efficient Dot1-mediated methylation of histone H3 lysine 79. J. Biol. Chem. 277, 34655–34657 (2002).
van Leeuwen, F., Gafken, P. R. & Gottschling, D. E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 109, 745–756 (2002).
Feng, Q. et al. Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr. Biol. 12, 1052–1058 (2002). This study, along with Lacoste, N. et al., Ng, H. H. et al. & van Leeuwen, F. et al. identified yeast Dot1 as the H3K79 methyltransferase.
Metzger, E. et al. KMT9 monomethylates histone H4 lysine 12 and controls proliferation of prostate cancer cells. Nat. Struct. Mol. Biol. 26, 361–371 (2019).
Falnes, P., Jakobsson, M. E., Davydova, E., Ho, A. & Małecki, J. Protein lysine methylation by seven-β-strand methyltransferases. Biochem. J. 473, 1995–2009 (2016).
Liu, J. et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m6A-dependent translation control. Nature 552, 126–131 (2017).
Biggar, K. K., Wang, Z. & Li, S. S. SnapShot: lysine methylation beyond histones. Mol. Cell 68, 1016–1016.e1 (2017).
Li, Y. et al. The target of the NSD family of histone lysine methyltransferases depends on the nature of the substrate. J. Biol. Chem. 284, 34283–34295 (2009). This study established the biochemical activity of NSD1, NSD2, NSD3 and SETD2 as H3K36 methyltransferases that have a strong preference for nucleosomal substrates.
Kuo, A. J. et al. NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol. Cell 44, 609–620 (2011).
Pfister, S. X. & Ashworth, A. Marked for death: targeting epigenetic changes in cancer. Nat. Rev. Drug Discov. 16, 241–263 (2017).
Schapira, M. Chemical inhibition of protein methyltransferases. Cell Chem. Biol. 23, 1067–1076 (2016).
Kuzmichev, A., Nishioka, K., Erdjument-Bromage, H., Tempst, P. & Reinberg, D. Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genes Dev. 16, 2893–2905 (2002).
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002).
Müller, J. et al. Histone methyltransferase activity of a Drosophila Polycomb group repressor complex. Cell 111, 197–208 (2002). This study, along with Kuzmichev, A. et al. & Cao, R. et al. (2002), provided the first descritpion of EZH2 as the H3K27 methyltransferase.
Cao, R. & Zhang, Y. SUZ12 is required for both the histone methyltransferase activity and the silencing function of the EED-EZH2 complex. Mol. Cell 15, 57–67 (2004).
Pasini, D., Bracken, A. P., Jensen, M. R., Lazzerini Denchi, E. & Helin, K. Suz12 is essential for mouse development and for EZH2 histone methyltransferase activity. EMBO J. 23, 4061–4071 (2004).
Tie, F., Stratton, C. A., Kurzhals, R. L. & Harte, P. J. The N terminus of Drosophila ESC binds directly to histone H3 and is required for E(Z)-dependent trimethylation of H3 lysine 27. Mol. Cell. Biol. 27, 2014–2026 (2007).
Margueron, R. et al. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461, 762–767 (2009).
Lee, C. H. et al. Automethylation of PRC2 promotes H3K27 methylation and is impaired in H3K27M pediatric glioma. Genes Dev. 33, 1428–1440 (2019).
Wang, X. et al. Regulation of histone methylation by automethylation of PRC2. Genes Dev. 33, 1416–1427 (2019).
Ardehali, M. B. et al. Polycomb repressive complex 2 methylates elongin A to regulate transcription. Mol. Cell 68, 872–884.e876 (2017).
Margueron, R. et al. Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol. Cell 32, 503–518 (2008).
Kaniskan, H., Martini, M. L. & Jin, J. Inhibitors of protein methyltransferases and demethylases. Chem. Rev. 118, 989–1068 (2018).
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002).
Kleer, C. G. et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc. Natl Acad. Sci. USA 100, 11606–11611 (2003).
Kim, K. H. & Roberts, C. W. Targeting EZH2 in cancer. Nat. Med. 22, 128–134 (2016).
van Haaften, G. et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat. Genet. 41, 521–523 (2009).
Morin, R. D. et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat. Genet. 42, 181–185 (2010).
Bödör, C. et al. EZH2 Y641 mutations in follicular lymphoma. Leukemia 25, 726–729 (2011).
Sneeringer, C. J. et al. Coordinated activities of wild-type plus mutant EZH2 drive tumor-associated hypertrimethylation of lysine 27 on histone H3 (H3K27) in human B-cell lymphomas. Proc. Natl Acad. Sci. USA 107, 20980–20985 (2010).
Yap, D. B. et al. Somatic mutations at EZH2 Y641 act dominantly through a mechanism of selectively altered PRC2 catalytic activity, to increase H3K27 trimethylation. Blood 117, 2451–2459 (2011).
Bödör, C. et al. EZH2 mutations are frequent and represent an early event in follicular lymphoma. Blood 122, 3165–3168 (2013).
Majer, C. R. et al. A687V EZH2 is a gain-of-function mutation found in lymphoma patients. FEBS Lett. 586, 3448–3451 (2012).
McCabe, M. T. et al. Mutation of A677 in histone methyltransferase EZH2 in human B-cell lymphoma promotes hypertrimethylation of histone H3 on lysine 27 (H3K27). Proc. Natl Acad. Sci. USA 109, 2989–2994 (2012).
Kim, K. H. et al. SWI/SNF-mutant cancers depend on catalytic and non-catalytic activity of EZH2. Nat. Med. 21, 1491–1496 (2015).
Knutson, S. K. et al. Durable tumor regression in genetically altered malignant rhabdoid tumors by inhibition of methyltransferase EZH2. Proc. Natl Acad. Sci. USA 110, 7922–7927 (2013). The first publication describing the EZH2 inhibitor EPZ-6438 (tazemetostat) that subsequently gained FDA approval.
Wilson, B. G. et al. Epigenetic antagonism between polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell 18, 316–328 (2010).
Versteege, I. et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394, 203–206 (1998).
Hasselblatt, M. et al. Nonsense mutation and inactivation of SMARCA4 (BRG1) in an atypical teratoid/rhabdoid tumor showing retained SMARCB1 (INI1) expression. Am. J. Surg. Pathol. 35, 933–935 (2011).
Modena, P. et al. SMARCB1/INI1 tumor suppressor gene is frequently inactivated in epithelioid sarcomas. Cancer Res. 65, 4012–4019 (2005).
Bitler, B. G. et al. Synthetic lethality by targeting EZH2 methyltransferase activity in ARID1A-mutated cancers. Nat. Med. 21, 231–238 (2015).
Ernst, T. et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat. Genet. 42, 722–726 (2010).
Wu, G. et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet. 44, 251–253 (2012).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013). This study described the dominant negative activity of histone oncomutations, with significant mechanistic implications for pediatric cancers.
Mohammad, F. et al. EZH2 is a potential therapeutic target for H3K27M-mutant pediatric gliomas. Nat. Med. 23, 483–492 (2017).
Piunti, A. et al. Therapeutic targeting of polycomb and BET bromodomain proteins in diffuse intrinsic pontine gliomas. Nat. Med. 23, 493–500 (2017).
Kaniskan, H., Konze, K. D. & Jin, J. Selective inhibitors of protein methyltransferases. J. Med. Chem. 58, 1596–1629 (2015).
Knutson, S. K. et al. A selective inhibitor of EZH2 blocks H3K27 methylation and kills mutant lymphoma cells. Nat. Chem. Biol. 8, 890–896 (2012).
McCabe, M. T. et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 492, 108–112 (2012). This study, along with Knutson, S. K. et al. (2012), describes the discovery of the first EZH2 inhibitors, which set the stage for tazemetostat to ultimately receive FDA approval as the first KMT inhibitory medicine.
Jiao, L. & Liu, X. Structural basis of histone H3K27 trimethylation by an active polycomb repressive complex 2. Science 350, aac4383 (2015).
Justin, N. et al. Structural basis of oncogenic histone H3K27M inhibition of human polycomb repressive complex 2. Nat. Commun. 7, 11316 (2016).
Brooun, A. et al. Polycomb repressive complex 2 structure with inhibitor reveals a mechanism of activation and drug resistance. Nat. Commun. 7, 11384 (2016).
Kasinath, V. et al. Structures of human PRC2 with its cofactors AEBP2 and JARID2. Science 359, 940–944 (2018).
Ciferri, C. et al. Molecular architecture of human polycomb repressive complex 2. eLife 1, e00005 (2012).
Vaswani, R. G. et al. Identification of (R)-N-((4-methoxy-6-methyl-2-oxo-1,2-dihydropyridin-3-yl)methyl)-2-methyl-1-(1-(1-(2,2,2-trifluoroethyl)piperidin-4-yl)ethyl)-1H-indole-3-carboxamide (CPI-1205), a potent and selective inhibitor of histone methyltransferase EZH2, suitable for phase I clinical trials for B-cell lymphomas. J. Med. Chem. 59, 9928–9941 (2016).
Kung, P. P. et al. Design and synthesis of pyridone-containing 3,4-dihydroisoquinoline-1(2H)-ones as a novel class of enhancer of zeste homolog 2 (EZH2) inhibitors. J. Med. Chem. 59, 8306–8325 (2016).
Ma, A. et al. Discovery of a first-in-class EZH2 selective degrader. Nat. Chem. Biol. 16, 214–222 (2020).
Yang, X. et al. Structure-activity relationship studies for enhancer of zeste homologue 2 (EZH2) and enhancer of zeste homologue 1 (EZH1) inhibitors. J. Med. Chem. 59, 7617–7633 (2016).
Konze, K. D. et al. An orally bioavailable chemical probe of the lysine methyltransferases EZH2 and EZH1. ACS Chem. Biol. 8, 1324–1334 (2013).
Zhao, Y. et al. EZH2 cooperates with gain-of-function p53 mutants to promote cancer growth and metastasis. EMBO J. 38, (2019).
Kim, J. et al. Polycomb- and methylation-independent roles of EZH2 as a transcription activator. Cell Rep. 25, 2808–2820 e2804 (2018).
Zhang, L. et al. Blocking immunosuppressive neutrophils deters pY696-EZH2-driven brain metastases. Sci. Transl Med. 12, eaaz5387 (2020).
Morschhauser, F. et al. Phase 2 multicenter study of tazemetostat, an EZH2 inhibitor, in patients with relapsed or refractory follicular lymphoma blood. Blood 134 (Suppl. 1), 123 (2019).
Zauderer, M. G. et al. Phase 2, multicenter study of the EZH2 inhibitor tazemetostat as monotherapy in adults with relapsed or refractory (R/R) malignant mesothelioma (MM) with BAP1 inactivation. J. Clin. Oncol. 36 (Suppl. 15), 8515 (2018).
Stacchiotti, S. et al. Safety and efficacy of tazemetostat, a first-in-class EZH2 inhibitor, in patients (pts) with epithelioid sarcoma (ES) (NCT02601950). J. Clin. Oncol. 37 (Suppl. 15), 11003 (2019).
Gounder, M. M. et al. Immunologic correlates of the abscopal effect in a SMARCB1/INI1-negative poorly differentiated chordoma after EZH2 inhibition and radiotherapy. Clin. Cancer Res. 25, 2064–2071 (2019).
Taplin, M.-E. et al. Phase Ib results of ProSTAR: CPI-1205, EZH2 inhibitor, combined with enzalutamide (E) or abiraterone/prednisone (A/P) in patients with metastatic castration-resistant prostate cancer (mCRPC). Cancer Res. 79 (Suppl. 13), CT094 (2019).
Yap, T. A. et al. Phase I study of the novel enhancer of zeste homolog 2 (EZH2) inhibitor GSK2816126 in patients with advanced hematologic and solid tumors. Clin. Cancer Res. 25, 7331–7339 (2019).
Xu, B. et al. Selective inhibition of EZH2 and EZH1 enzymatic activity by a small molecule suppresses MLL-rearranged leukemia. Blood 125, 346–357 (2015).
Honma, D. et al. Novel orally bioavailable EZH1/2 dual inhibitors with greater antitumor efficacy than an EZH2 selective inhibitor. Cancer Sci. 108, 2069–2078 (2017).
Yamagishi, M. et al. Targeting excessive EZH1 and EZH2 activities for abnormal histone methylation and transcription network in malignant lymphomas. Cell Rep. 29, 2321–2337.e2327 (2019).
Hansen, K. H. et al. A model for transmission of the H3K27me3 epigenetic mark. Nat. Cell Biol. 10, 1291–1300 (2008).
Kim, W. et al. Targeted disruption of the EZH2-EED complex inhibits EZH2-dependent cancer. Nat. Chem. Biol. 9, 643–650 (2013).
Xu, C. et al. Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc. Natl Acad. Sci. USA 107, 19266–19271 (2010).
Qi, W. et al. An allosteric PRC2 inhibitor targeting the H3K27me3 binding pocket of EED. Nat. Chem. Biol. 13, 381–388 (2017).
He, Y. et al. The EED protein-protein interaction inhibitor A-395 inactivates the PRC2 complex. Nat. Chem. Biol. 13, 389–395 (2017). This study, along with Qi, W. et al., describes the discovery of the first protein–protein interaction disrupting EED inhibitors that function as allosteric inhibitors of PRC2.
Basheer, F. et al. Contrasting requirements during disease evolution identify EZH2 as a therapeutic target in AML. J. Exp. Med. 216, 966–981 (2019).
Karantanos, T., Chistofides, A., Barhdan, K., Li, L. & Boussiotis, V. A. Regulation of T cell differentiation and function by EZH2. Front. Immunol. 7, 172 (2016).
Qiu, J., Sharma, S., Rollins, R. A. & Paul, T. A. The complex role of EZH2 in the tumor microenvironment: opportunities and challenges for immunotherapy combinations. Future Med. Chem. https://doi.org/10.4155/fmc-2020-0072 (2020).
Hamaidia, M. et al. Inhibition of EZH2 methyltransferase decreases immunoediting of mesothelioma cells by autologous macrophages through a PD-1-dependent mechanism. JCI Insight 4, e128474 (2019).
Xiao, G. et al. EZH2 negatively regulates PD-L1 expression in hepatocellular carcinoma. J. Immunother. Cancer 7, 300 (2019).
Zhou, L., Mudianto, T., Ma, X., Riley, R. & Uppaluri, R. Targeting EZH2 enhances antigen presentation, antitumor immunity, and circumvents Anti-PD-1 resistance in head and neck cancer. Clin. Cancer Res. 26, 290–300 (2020).
Goswami, S. et al. Modulation of EZH2 expression in T cells improves efficacy of anti-CTLA-4 therapy. J. Clin. Invest. 128, 3813–3818 (2018).
Zingg, D. et al. The histone methyltransferase Ezh2 controls mechanisms of adaptive resistance to tumor immunotherapy. Cell Rep. 20, 854–867 (2017).
Vlaming, H. & van Leeuwen, F. The upstreams and downstreams of H3K79 methylation by DOT1L. Chromosoma 125, 593–605 (2016).
Min, J., Feng, Q., Li, Z., Zhang, Y. & Xu, R. M. Structure of the catalytic domain of human DOT1L, a non-SET domain nucleosomal histone methyltransferase. Cell 112, 711–723 (2003).
Jones, B. et al. The histone H3K79 methyltransferase Dot1L is essential for mammalian development and heterochromatin structure. PLoS Genet. 4, e1000190 (2008).
Deshpande, A. J. et al. AF10 regulates progressive H3K79 methylation and HOX gene expression in diverse AML subtypes. Cancer Cell 26, 896–908 (2014).
Bernt, K. M. et al. MLL-rearranged leukemia is dependent on aberrant H3K79 methylation by DOT1L. Cancer Cell 20, 66–78 (2011).
Krivtsov, A. V., Hoshii, T. & Armstrong, S. A. Mixed-lineage leukemia fusions and chromatin in leukemia. Cold Spring Harb. Perspect. Med. 7, a026658 (2017).
Okada, Y. et al. hDOT1L links histone methylation to leukemogenesis. Cell 121, 167–178 (2005). This study identified a role for human DOT1L, via H3K79 methylation, in promoting MLL-driven leukaemia.
Mohan, M. et al. Linking H3K79 trimethylation to Wnt signaling through a novel Dot1-containing complex (DotCom). Genes Dev. 24, 574–589 (2010).
Chen, S. et al. The PZP domain of AF10 senses unmodified H3K27 to regulate DOT1L-mediated methylation of H3K79. Mol. Cell 60, 319–327 (2015).
Li, Y. et al. AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell 159, 558–571 (2014).
Erb, M. A. et al. Transcription control by the ENL YEATS domain in acute leukaemia. Nature 543, 270–274 (2017).
Wan, L. et al. ENL links histone acetylation to oncogenic gene expression in acute myeloid leukaemia. Nature 543, 265–269 (2017).
Godfrey, L. et al. DOT1L inhibition reveals a distinct subset of enhancers dependent on H3K79 methylation. Nat. Commun. 10, 2803 (2019).
Briggs, S. D. et al. Gene silencing: trans-histone regulatory pathway in chromatin. Nature 418, 498 (2002).
McGinty, R. K., Kim, J., Chatterjee, C., Roeder, R. G. & Muir, T. W. Chemically ubiquitylated histone H2B stimulates hDot1L-mediated intranucleosomal methylation. Nature 453, 812–816 (2008).
Altaf, M. et al. Interplay of chromatin modifiers on a short basic patch of histone H4 tail defines the boundary of telomeric heterochromatin. Mol. Cell 28, 1002–1014 (2007).
Fingerman, I. M., Li, H. C. & Briggs, S. D. A charge-based interaction between histone H4 and Dot1 is required for H3K79 methylation and telomere silencing: identification of a new trans-histone pathway. Genes Dev. 21, 2018–2029 (2007).
McGinty, R. K. et al. Structure-activity analysis of semisynthetic nucleosomes: mechanistic insights into the stimulation of Dot1L by ubiquitylated histone H2B. ACS Chem. Biol. 4, 958–968 (2009).
Anderson, C. J. et al. Structural basis for recognition of ubiquitylated nucleosome by Dot1L methyltransferase. Cell Rep. 26, 1681–1690.e1685 (2019).
Jang, S. et al. Structural basis of recognition and destabilization of the histone H2B ubiquitinated nucleosome by the DOT1L histone H3 Lys79 methyltransferase. Genes Dev. 33, 620–625 (2019).
Valencia-Sánchez, M. I. et al. Structural basis of Dot1L stimulation by histone H2B lysine 120 ubiquitination. Mol. Cell 74, 1010–1019.e6 (2019).
Worden, E. J., Hoffmann, N. A., Hicks, C. W. & Wolberger, C. Mechanism of cross-talk between H2B ubiquitination and H3 methylation by Dot1L. Cell 176, 1490–1501.e12 (2019).
Yao, T. et al. Structural basis of the crosstalk between histone H2B monoubiquitination and H3 lysine 79 methylation on nucleosome. Cell Res. 29, 330–333 (2019).
Deshpande, A. J. et al. Leukemic transformation by the MLL-AF6 fusion oncogene requires the H3K79 methyltransferase Dot1l. Blood 121, 2533–2541 (2013).
Daigle, S. R. et al. Potent inhibition of DOT1L as treatment of MLL-fusion leukemia. Blood 122, 1017–1025 (2013).
Krivtsov, A. V. et al. H3K79 methylation profiles define murine and human MLL-AF4 leukemias. Cancer Cell 14, 355–368 (2008).
Gu, Y. et al. The t(4;11) chromosome translocation of human acute leukemias fuses the ALL-1 gene, related to Drosophila trithorax, to the AF-4 gene. Cell 71, 701–708 (1992).
Tkachuk, D. C., Kohler, S. & Cleary, M. L. Involvement of a homolog of Drosophila trithorax by 11q23 chromosomal translocations in acute leukemias. Cell 71, 691–700 (1992).
Meyer, C. et al. The MLL recombinome of acute leukemias in 2017. Leukemia 32, 273–284 (2018).
Corral, J. et al. An Mll-AF9 fusion gene made by homologous recombination causes acute leukemia in chimeric mice: a method to create fusion oncogenes. Cell 85, 853–861 (1996).
Ayton, P. M. & Cleary, M. L. Transformation of myeloid progenitors by MLL oncoproteins is dependent on Hoxa7 and Hoxa9. Genes Dev. 17, 2298–2307 (2003).
Zeisig, B. B. et al. Hoxa9 and Meis1 are key targets for MLL-ENL-mediated cellular immortalization. Mol. Cell. Biol. 24, 617–628 (2004).
Brzezinka, K. et al. Functional diversity of inhibitors tackling the differentiation blockage of MLL-rearranged leukemia. J. Hematol. Oncol. 12, 66 (2019).
Nguyen, A. T., Taranova, O., He, J. & Zhang, Y. DOT1L, the H3K79 methyltransferase, is required for MLL-AF9-mediated leukemogenesis. Blood 117, 6912–6922 (2011).
Daigle, S. R. et al. Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 20, 53–65 (2011). This study provides the first example of a highly selective histone lysine methyltransferase inhibitor with in vivo efficacy in a cancer model.
Chen, L. et al. Abrogation of MLL-AF10 and CALM-AF10-mediated transformation through genetic inactivation or pharmacological inhibition of the H3K79 methyltransferase Dot1l. Leukemia 27, 813–822 (2013).
Basavapathruni, A. et al. Conformational adaptation drives potent, selective and durable inhibition of the human protein methyltransferase DOT1L. Chem. Biol. Drug Des. 80, 971–980 (2012).
Chen, C. et al. Discovery of novel Dot1L inhibitors through a structure-based fragmentation approach. ACS Med. Chem. Lett. 7, 735–740 (2016).
Scheufler, C. et al. Optimization of a fragment-based screening hit toward potent DOT1L inhibitors interacting in an induced binding pocket. ACS Med. Chem. Lett. 7, 730–734 (2016).
Shukla, N. et al. Final report of phase 1 study of the DOT1L inhibitor, pinometostat (EPZ-5676), in children with relapsed or refractory MLL-r acute leukemia. Blood 128, 2780 (2016).
Stein, E. M. et al. The DOT1L inhibitor pinometostat reduces H3K79 methylation and has modest clinical activity in adult acute leukemia. Blood 131, 2661–2669 (2018).
Klaus, C. R. et al. DOT1L inhibitor EPZ-5676 displays synergistic antiproliferative activity in combination with standard of care drugs and hypomethylating agents in MLL-rearranged leukemia cells. J. Pharmacol. Exp. Ther. 350, 646–656 (2014).
Rau, R. E. et al. DOT1L as a therapeutic target for the treatment of DNMT3A-mutant acute myeloid leukemia. Blood 128, 971–981 (2016).
Menghrajani, K. et al. A phase Ib/II study of the histone methyltransferase inhibitor pinometostat in combination with azacitidine in patients with 11q23-rearranged acute myeloid leukemia. Blood 134, 2655 (2019).
Secker, K. A. et al. Inhibition of DOT1L and PRMT5 promote synergistic anti-tumor activity in a human MLL leukemia model induced by CRISPR/Cas9. Oncogene 38, 7181–7195 (2019).
Skucha, A. et al. MLL-fusion-driven leukemia requires SETD2 to safeguard genomic integrity. Nat. Commun. 9, 1983 (2018).
Borkin, D. et al. Pharmacologic inhibition of the menin-MLL interaction blocks progression of MLL leukemia in vivo. Cancer Cell 27, 589–602 (2015).
Borkin, D. et al. Complexity of blocking bivalent protein-protein interactions: development of a highly potent inhibitor of the menin-mixed-lineage leukemia interaction. J. Med. Chem. 61, 4832–4850 (2018).
Tachibana, M., Sugimoto, K., Fukushima, T. & Shinkai, Y. Set domain-containing protein, G9a, is a novel lysine-preferring mammalian histone methyltransferase with hyperactivity and specific selectivity to lysines 9 and 27 of histone H3. J. Biol. Chem. 276, 25309–25317 (2001).
Tachibana, M. et al. G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev. 16, 1779–1791 (2002).
Tachibana, M. et al. Histone methyltransferases G9a and GLP form heteromeric complexes and are both crucial for methylation of euchromatin at H3-K9. Genes Dev. 19, 815–826 (2005).
Liu, N. et al. Recognition of H3K9 methylation by GLP is required for efficient establishment of H3K9 methylation, rapid target gene repression, and mouse viability. Genes Dev. 29, 379–393 (2015).
Collins, R. E. et al. The ankyrin repeats of G9a and GLP histone methyltransferases are mono- and dimethyllysine binding modules. Nat. Struct. Mol. Biol. 15, 245–250 (2008).
Tu, W. B. et al. MYC Interacts with the G9a histone methyltransferase to drive transcriptional repression and tumorigenesis. Cancer Cell 34, 579–595.e578 (2018).
Kato, S. et al. Gain-of-function genetic alterations of G9a drive oncogenesis. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-19-0532 (2020).
Wang, Z. et al. SETD5-coordinated chromatin reprogramming regulates adaptive resistance to targeted pancreatic cancer therapy. Cancer Cell 37, 834–849.e13 (2020).
Watson, Z. L. et al. Histone methyltransferases EHMT1 and EHMT2 (GLP/G9A) maintain PARP inhibitor resistance in high-grade serous ovarian carcinoma. Clin. Epigenet. 11, 165 (2019).
Maze, I. et al. Essential role of the histone methyltransferase G9a in cocaine-induced plasticity. Science 327, 213–216 (2010).
Anderson, E. M. et al. Knockdown of the histone di-methyltransferase G9a in nucleus accumbens shell decreases cocaine self-administration, stress-induced reinstatement, and anxiety. Neuropsychopharmacology 44, 1370–1376 (2019).
Avgustinova, A. et al. Loss of G9a preserves mutation patterns but increases chromatin accessibility, genomic instability and aggressiveness in skin tumours. Nat. Cell Biol. 20, 1400–1409 (2018).
Rowbotham, S. P. et al. H3K9 methyltransferases and demethylases control lung tumor-propagating cells and lung cancer progression. Nat. Commun. 9, 4559 (2018).
Kubicek, S. et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol. Cell 25, 473–481 (2007). This study provides the first example of a substrate-competitive, selective lysine methyltransferase inhibitor.
Dong, C. et al. G9a interacts with Snail and is critical for Snail-mediated E-cadherin repression in human breast cancer. J. Clin. Invest. 122, 1469–1486 (2012).
Chang, Y. et al. Structural basis for G9a-like protein lysine methyltransferase inhibition by BIX-01294. Nat. Struct. Mol. Biol. 16, 312–317 (2009).
Vedadi, M. et al. A chemical probe selectively inhibits G9a and GLP methyltransferase activity in cells. Nat. Chem. Biol. 7, 566–574 (2011). This study provided the first example of a selective, cellular chemical probe of G9a/GLP.
Liu, F. et al. Discovery of an in vivo chemical probe of the lysine methyltransferases G9a and GLP. J. Med. Chem. 56, 8931–8942 (2013).
Sweis, R. F. et al. Discovery and development of potent and selective inhibitors of histone methyltransferase g9a. ACS Med. Chem. Lett. 5, 205–209 (2014).
Xiong, Y. et al. Discovery of potent and selective inhibitors for G9a-like protein (GLP) lysine methyltransferase. J. Med. Chem. 60, 1876–1891 (2017).
Ferry, L. et al. Methylation of DNA ligase 1 by G9a/GLP recruits UHRF1 to replicating DNA and regulates DNA methylation. Mol. Cell 67, 550–565.e555 (2017).
Fang, J. et al. Purification and functional characterization of SET8, a nucleosomal histone H4-lysine 20-specific methyltransferase. Curr. Biol. 12, 1086–1099 (2002).
Nishioka, K. et al. PR-Set7 is a nucleosome-specific methyltransferase that modifies lysine 20 of histone H4 and is associated with silent chromatin. Mol. Cell 9, 1201–1213 (2002).
van Nuland, R. & Gozani, O. Histone H4 lysine 20 (H4K20) methylation, expanding the signaling potential of the proteome one methyl moiety at a time. Mol. Cell Proteom. 15, 755–764 (2016).
Shi, X. et al. Modulation of p53 function by SET8-mediated methylation at lysine 382. Mol. Cell 27, 636–646 (2007).
Kudithipudi, S., Dhayalan, A., Kebede, A. F. & Jeltsch, A. The SET8 H4K20 protein lysine methyltransferase has a long recognition sequence covering seven amino acid residues. Biochimie 94, 2212–2218 (2012).
McKay, D. J. et al. Interrogating the function of metazoan histones using engineered gene clusters. Dev. Cell 32, 373–386 (2015).
Milite, C. et al. The emerging role of lysine methyltransferase SETD8 in human diseases. Clin. Epigenet. 8, 102 (2016).
Couture, J. F., Collazo, E., Brunzelle, J. S. & Trievel, R. C. Structural and functional analysis of SET8, a histone H4 Lys-20 methyltransferase. Genes Dev. 19, 1455–1465 (2005).
Xiao, B. et al. Specificity and mechanism of the histone methyltransferase Pr-Set7. Genes Dev. 19, 1444–1454 (2005).
Blum, G. et al. Small-molecule inhibitors of SETD8 with cellular activity. ACS Chem. Biol. 9, 2471–2478 (2014).
Ma, A. et al. Discovery of a selective, substrate-competitive inhibitor of the lysine methyltransferase SETD8. J. Med. Chem. 57, 6822–6833 (2014).
Ma, A. et al. Structure-activity relationship studies of SETD8 inhibitors. Medchemcomm 5, 1892–1898 (2014).
Veschi, V. et al. Epigenetic siRNA and chemical screens identify SETD8 inhibition as a therapeutic strategy for p53 activation in high-risk neuroblastoma. Cancer Cell 31, 50–63 (2017).
Wu, J. et al. Downregulation of histone methyltransferase SET8 inhibits progression of hepatocellular carcinoma. Sci. Rep. 10, 4490 (2020).
Butler, K. V. et al. Structure-based design of a covalent inhibitor of the SET domain-containing protein 8 (SETD8) lysine methyltransferase. J. Med. Chem. 59, 9881–9889 (2016).
Oda, H. et al. Monomethylation of histone H4-lysine 20 is involved in chromosome structure and stability and is essential for mouse development. Mol. Cell. Biol. 29, 2278–2295 (2009).
Pannetier, M. et al. PR-SET7 and SUV4-20H regulate H4 lysine-20 methylation at imprinting control regions in the mouse. EMBO Rep. 9, 998–1005 (2008).
Schotta, G. et al. A silencing pathway to induce H3-K9 and H4-K20 trimethylation at constitutive heterochromatin. Genes Dev. 18, 1251–1262 (2004).
Schotta, G. et al. A chromatin-wide transition to H4K20 monomethylation impairs genome integrity and programmed DNA rearrangements in the mouse. Genes Dev. 22, 2048–2061 (2008).
Southall, S. M., Cronin, N. B. & Wilson, J. R. A novel route to product specificity in the Suv4-20 family of histone H4K20 methyltransferases. Nucleic Acids Res. 42, 661–671 (2014).
Wu, H. et al. Molecular basis for the regulation of the H3K4 methyltransferase activity of PRDM9. Cell Rep. 5, 13–20 (2013).
Gonzalo, S. et al. Role of the RB1 family in stabilizing histone methylation at constitutive heterochromatin. Nat. Cell Biol. 7, 420–428 (2005).
Lu, X. et al. The effect of H3K79 dimethylation and H4K20 trimethylation on nucleosome and chromatin structure. Nat. Struct. Mol. Biol. 15, 1122–1124 (2008).
Long, H. et al. H2A.Z facilitates licensing and activation of early replication origins. Nature 577, 576–581 (2020).
Sanders, S. L. et al. Methylation of histone H4 lysine 20 controls recruitment of Crb2 to sites of DNA damage. Cell 119, 603–614 (2004).
Yang, H. et al. Preferential dimethylation of histone H4 lysine 20 by Suv4-20. J. Biol. Chem. 283, 12085–12092 (2008).
Bromberg, K. D. et al. The SUV4-20 inhibitor A-196 verifies a role for epigenetics in genomic integrity. Nat. Chem. Biol. 13, 317–324 (2017).
Wang, H. et al. Purification and functional characterization of a histone H3-lysine 4-specific methyltransferase. Mol. Cell 8, 1207–1217 (2001).
Nishioka, K. et al. Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev. 16, 479–489 (2002).
Lehnertz, B. et al. p53-dependent transcription and tumor suppression are not affected in Set7/9-deficient mice. Mol. Cell 43, 673–680 (2011).
Barsyte-Lovejoy, D. et al. (R)-PFI-2 is a potent and selective inhibitor of SETD7 methyltransferase activity in cells. Proc. Natl Acad. Sci. USA 111, 12853–12858 (2014).
Hamidi, T. et al. Identification of Rpl29 as a major substrate of the lysine methyltransferase Set7/9. J. Biol. Chem. 293, 12770–12780 (2018).
Komatsu, S. et al. Overexpression of SMYD2 contributes to malignant outcome in gastric cancer. Br. J. Cancer 112, 357–364 (2015).
Komatsu, S. et al. Overexpression of SMYD2 relates to tumor cell proliferation and malignant outcome of esophageal squamous cell carcinoma. Carcinogenesis 30, 1139–1146 (2009).
Reynoird, N. et al. Coordination of stress signals by the lysine methyltransferase SMYD2 promotes pancreatic cancer. Genes Dev. 30, 772–785 (2016).
Olsen, J. B. et al. Quantitative profiling of the activity of protein lysine methyltransferase SMYD2 Using SILAC-based proteomics. Mol. Cell Proteom. 15, 892–905 (2016).
Bagislar, S. et al. Smyd2 is a Myc-regulated gene critical for MLL-AF9 induced leukemogenesis. Oncotarget 7, 66398–66415 (2016).
Cowen, S. D. et al. Design, synthesis, and biological activity of substrate competitive SMYD2 inhibitors. J. Med. Chem. 59, 11079–11097 (2016).
Ferguson, A. D. et al. Structural basis of substrate methylation and inhibition of SMYD2. Structure 19, 1262–1273 (2011).
Sweis, R. F. et al. Discovery of A-893, a new cell-active benzoxazinone inhibitor of lysine methyltransferase SMYD2. ACS Med. Chem. Lett. 6, 695–700 (2015).
Nguyen, H. et al. LLY-507, a cell-active, potent, and selective inhibitor of protein-lysine methyltransferase SMYD2. J. Biol. Chem. 290, 13641–13653 (2015).
Eggert, E. et al. Discovery and characterization of a highly potent and selective aminopyrazoline-based in vivo probe (BAY-598) for the protein lysine methyltransferase SMYD2. J. Med. Chem. 59, 4578–4600 (2016).
Huang, J. et al. Repression of p53 activity by Smyd2-mediated methylation. Nature 444, 629–632 (2006).
Thomenius, M. J. et al. Small molecule inhibitors and CRISPR/Cas9 mutagenesis demonstrate that SMYD2 and SMYD3 activity are dispensable for autonomous cancer cell proliferation. PLoS ONE 13, e0197372 (2018).
Mazur, P. K. et al. SMYD3 links lysine methylation of MAP3K2 to Ras-driven cancer. Nature 510, 283–287 (2014). This study identified a role for protein lysine methylation in integrating MAP kinase signalling in the cytoplasm to promote pancreatic and lung cancer.
Sarris, M. E., Moulos, P., Haroniti, A., Giakountis, A. & Talianidis, I. Smyd3 Is a transcriptional potentiator of multiple cancer-promoting genes and required for liver and colon cancer development. Cancer Cell 29, 354–366 (2016).
Hamamoto, R. et al. SMYD3 encodes a histone methyltransferase involved in the proliferation of cancer cells. Nat. Cell Biol. 6, 731–740 (2004).
Fabini, E. et al. Unveiling the biochemistry of the epigenetic regulator SMYD3. Biochemistry 58, 3634–3645 (2019).
Van Aller, G. S. et al. Smyd3 regulates cancer cell phenotypes and catalyzes histone H4 lysine 5 methylation. Epigenetics 7, 340–343 (2012).
Kunizaki, M. et al. The lysine 831 of vascular endothelial growth factor receptor 1 is a novel target of methylation by SMYD3. Cancer Res. 67, 10759–10765 (2007).
Van Aller, G. S. et al. Structure-based design of a novel SMYD3 inhibitor that bridges the SAM-and MEKK2-binding pockets. Structure 24, 774–781 (2016).
Mitchell, L. H. et al. Novel oxindole sulfonamides and sulfamides: EPZ031686, the first orally bioavailable small molecule SMYD3 inhibitor. ACS Med. Chem. Lett. 7, 134–138 (2016).
Huang, C. et al. Discovery of irreversible inhibitors targeting histone methyltransferase, SMYD3. ACS Med. Chem. Lett. 10, 978–984 (2019).
Bennett, R. L., Swaroop, A., Troche, C. & Licht, J. D. The role of nuclear receptor-binding SET domain family histone lysine methyltransferases in cancer. Cold Spring Harb. Perspect. Med. 7, a026708 (2017).
Yang, S. et al. Molecular basis for oncohistone H3 recognition by SETD2 methyltransferase. Genes Dev. 30, 1611–1616 (2016).
Böttcher, J. et al. Fragment-based discovery of a chemical probe for the PWWP1 domain of NSD3. Nat. Chem. Biol. 15, 822–829 (2019).
Sun, Y. et al. Histone methyltransferase SETDB1 is required for prostate cancer cell proliferation, migration and invasion. Asian J. Androl. 16, 319–324 (2014).
Wong, C. M. et al. Up-regulation of histone methyltransferase SETDB1 by multiple mechanisms in hepatocellular carcinoma promotes cancer metastasis. Hepatology 63, 474–487 (2016).
Zhu, Y., Sun, D., Jakovcevski, M. & Jiang, Y. Epigenetic mechanism of SETDB1 in brain: implications for neuropsychiatric disorders. Transl Psychiatry 10, 115 (2020).
Kim, Y., Wang, S. E. & Jiang, Y. H. Epigenetic therapy of Prader-Willi syndrome. Transl Res. 208, 105–118 (2019).
Woodcock, C. B., Yu, D., Zhang, X. & Cheng, X. Human HemK2/KMT9/N6AMT1 is an active protein methyltransferase, but does not act on DNA in vitro, in the presence of Trm112. Cell Discov. 5, 50 (2019).
Kusevic, D., Kudithipudi, S. & Jeltsch, A. Substrate specificity of the HEMK2 protein glutamine methyltransferase and identification of novel substrates. J. Biol. Chem. 291, 6124–6133 (2016).
Figaro, S., Scrima, N., Buckingham, R. H. & Heurgué-Hamard, V. HemK2 protein, encoded on human chromosome 21, methylates translation termination factor eRF1. FEBS Lett. 582, 2352–2356 (2008).
Allali-Hassani, A. et al. Discovery of a chemical probe for PRDM9. Nat. Commun. 10, 5759 (2019). This study describes the first inhibitor of PRDM9, a potential target in multiple cancers.
Houle, A. A. et al. Aberrant PRDM9 expression impacts the pan-cancer genomic landscape. Genome Res. 28, 1611–1620 (2018).
Powers, N. R. et al. The meiotic recombination activator PRDM9 trimethylates both H3K36 and H3K4 at recombination hotspots in vivo. PLoS Genet. 12, e1006146 (2016).
Mzoughi, S., Tan, Y. X., Low, D. & Guccione, E. The role of PRDMs in cancer: one family, two sides. Curr. Opin. Genet. Dev. 36, 83–91 (2016).
Roqueta-Rivera, M. et al. SETDB2 links glucocorticoid to lipid metabolism through Insig2a regulation. Cell Metab. 24, 474–484 (2016).
Sessa, A. et al. SETD5 regulates chromatin methylation state and preserves global transcriptional fidelity during brain development and neuronal wiring. Neuron 104, 271–289.e213 (2019).
Mas, Y. M. S. et al. The human mixed lineage leukemia 5 (MLL5), a sequentially and structurally divergent set domain-containing protein with no intrinsic catalytic activity. PLoS ONE 11, e0165139 (2016).
Robichaud, N., Sonenberg, N., Ruggero, D. & Schneider, R. J. Translational control in cancer. Cold Spring Harb. Perspect. Biol. 11, 254–266 (2019).
Jakobsson, M. E., Małecki, J. & Falnes, P. Regulation of eukaryotic elongation factor 1 alpha (eEF1A) by dynamic lysine methylation. RNA Biol. 15, 314–319 (2018).
Jakobsson, M. E. et al. Methylation of human eukaryotic elongation factor alpha (eEF1A) by a member of a novel protein lysine methyltransferase family modulates mRNA translation. Nucleic Acids Res. 45, 8239–8254 (2017).
Davydova, E. et al. Identification and characterization of a novel evolutionarily conserved lysine-specific methyltransferase targeting eukaryotic translation elongation factor 2 (eEF2). J. Biol. Chem. 289, 30499–30510 (2014).
Zhang, L., Hamey, J. J., Hart-Smith, G., Erce, M. A. & Wilkins, M. R. Elongation factor methyltransferase 3 – a novel eukaryotic lysine methyltransferase. Biochem. Biophys. Res. Commun. 451, 229–234 (2014).
Acknowledgements
This work was supported in part by grants from the NIH to K.P.B. (K00CA212435), O.G. (R01 CA236118) and J.J. (R01CA218600, R01CA230854, R01GM122749 and R01HD088626).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
O.G. is a co-founder of EpiCypher and Athelas Therapeutics. J.J. and H.Ü.K. are inventors of patent applications filed by the Icahn School of Medicine at Mount Sinai. J.J. is a consultant of Cullgen and Accent Therapeutics, a scientific advisory board member of Petra Pharma Corporation and an equity shareholder of Cullgen. K.P.B. is an employee at Genentech.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Chemical Probes: http://www.chemicalprobes.org
List of clinically approved kinase inhibitors: https://www.ppu.mrc.ac.uk/list-clinically-approved-kinase-inhibitors
Supplementary information
Glossary
- Reader domains
-
Modules on proteins that bind to post-translationally modified lysine residues on other proteins in a manner that is dependent on the immediate surrounding sequence and the state of methylation on the lysine.
- Writers of protein lysine methylation
-
Enzymes that catalyse the addition of one, two or three methyl moieties to the ε-nitrogen of lysine residues.
- Nucleosome
-
The fundamental building block of chromatin, which consists of ~146 base pairs of DNA wrapped around a protein core unit made of two copies each of histones H2A, H2B, H3 and H4.
- Chromatin-remodelling complexes
-
Large ATP-dependent multisubunit protein complexes that evict, load, alter or otherwise move nucleosomes on DNA to control DNA accessibility.
- Transcriptional elongation factors
-
Proteins that regulate the elongation step in gene transcription, which occurs after transcription is stably initiated and before transcription termination.
- Mixed-lineage leukaemia (MLL) genes
-
MLL1, MLL2, MLL3 and MLL4 encode four distinct lysine methyltransferases that catalyse methylation at histone H3 K4; MLL1 was originally identified as a gene involved in a recurrent chromosomal translocation in the neoplasm mixed-lineage leukaemia.
- Fusion protein
-
Chimeric proteins that result from the fusion of genes from different chromosomes during chromosomal translocations. They often have a new, non-physiologic activity that can unbalance cells and drive cancer pathogenesis.
- A-site
-
The aminoacyl site, or A-site, on the ribosome is the entry site for amino acid–tRNA molecules to bind and for proper base pairing between the mRNA codon and the tRNA anticodon during the elongation step of protein synthesis.
Rights and permissions
About this article
Cite this article
Bhat, K.P., Ümit Kaniskan, H., Jin, J. et al. Epigenetics and beyond: targeting writers of protein lysine methylation to treat disease. Nat Rev Drug Discov 20, 265–286 (2021). https://doi.org/10.1038/s41573-020-00108-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41573-020-00108-x
This article is cited by
-
Lysine methyltransferase SMYD2 enhances androgen receptor signaling to modulate CRPC cell resistance to enzalutamide
Oncogene (2024)
-
Mechanisms of action and resistance in histone methylation-targeted therapy
Nature (2024)
-
PRMT1-mediated PGK1 arginine methylation promotes colorectal cancer glycolysis and tumorigenesis
Cell Death & Disease (2024)
-
Novel methyltransferase G9a inhibitor induces ferroptosis in multiple myeloma through Nrf2/HO-1 pathway
Annals of Hematology (2024)
-
Roles of protein post-translational modifications in glucose and lipid metabolism: mechanisms and perspectives
Molecular Medicine (2023)