Abstract
Most rare diseases still lack approved treatments despite major advances in research providing the tools to understand their molecular basis, as well as legislation providing regulatory and economic incentives to catalyse the development of specific therapies. Addressing this translational gap is a multifaceted challenge, for which a key aspect is the selection of the optimal therapeutic modality for translating advances in rare disease knowledge into potential medicines, known as orphan drugs. With this in mind, we discuss here the technological basis and rare disease applicability of the main therapeutic modalities, including small molecules, monoclonal antibodies, protein replacement therapies, oligonucleotides and gene and cell therapies, as well as drug repurposing. For each modality, we consider its strengths and limitations as a platform for rare disease therapy development and describe clinical progress so far in developing drugs based on it. We also discuss selected overarching topics in the development of therapies for rare diseases, such as approval statistics, engagement of patients in the process, regulatory pathways and digital tools.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
08 January 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41573-019-0059-7
References
Global Genes. Rare diseases. RARE facts. Global Genes https://globalgenes.org/rare-facts/ (2019).
Tambuyzer, E. Rare diseases, orphan drugs and their regulation: questions and misconceptions. Nat. Rev. Drug Discov. 9, 921–929 (2010).
Farnaes, L. et al. Rapid whole-genome sequencing decreases infant morbidity and cost of hospitalization. NPJ Genom. Med. 3, 10 (2018).
Gurovich, Y. et al. Identifying facial phenotypes of genetic disorders using deep learning. Nat. Med. 25, 60–64 (2019).
Plowright, A. T. et al. Heart regeneration: opportunities and challenges for drug discovery with novel chemical and therapeutic methods or agents. Angew. Chem. Int. Ed. 53, 4056–4075 (2014).
Valeur, E. et al. New modalities for challenging targets in drug discovery. Angew. Chem. Int. Ed. Engl. 56, 10294–10323 (2017).
Scannell, J. W. et al. Diagnosing the decline in pharmaceutical R&D efficiency. Nat. Rev. Drug Discov. 11, 191 (2012).
Rodgers, G. et al. Glimmers in illuminating the druggable genome. Nat. Rev. Drug Discov. 17, 301–302 (2018).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Macarron, R. et al. Impact of high-throughput screening in biomedical research. Nat. Rev. Drug Discov. 10, 188–195 (2011).
Lipinski, C. A. et al. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 23, 3–25 (1997).
Gerry, C. J. & Schreiber, S. L. Chemical probes and drug leads from advances in synthetic planning and methodology. Nat. Rev. Drug Discov. 17, 333–352 (2018).
Shultz, M. D. Two decades under the influence of the rule of five and the changing properties of approved oral drugs. J. Med. Chem. 62, 1701–1714 (2018).
Plenge, R. M. Disciplined approach to drug discovery and early development. Sci. Transl Med. 8, 349ps15 (2016).
Scott, A. How CRISPR is transforming drug discovery. Nature 555, S10–S11 (2018).
Takahashi, T. Organoids for drug discovery and personalized medicine. Annu. Rev. Pharmacol. Toxicol. 59, 447–462 (2019).
Elitt, M. S., Barbar, L. & Tesar, P. J. Drug screening for human genetic diseases using iPSC models. Hum. Mol. Genet. 27(R2), R89–R98 (2018).
Ebert, A. D. et al. Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 457, 277–280 (2009).
Groen, E. J. N., Talbot, K. & Gillingwater, T. H. Advances in therapy for spinal muscular atrophy: promises and challenges. Nat. Rev. Neurol. 14, 214–224 (2018).
Artegiani, B. & Clevers, H. Use and application of 3D-organoid technology. Hum. Mol. Genet. 27(R2), R99–R107 (2018).
Strynatka, K. A. et al. How surrogate and chemical genetics in model organisms can suggest therapies for human genetic diseases. Genetics 208, 833–851 (2018).
Van Goor, F. et al. Correction of the F508del-CFTR protein processing defect in vitro by the investigational drug VX-809. Proc. Natl Acad. Sci. USA 108, 18843–18848 (2011).
Van Goor, F. et al. Rescue of CF airway epithelial cell function in vitro by a CFTR potentiator, VX-770. Proc. Natl Acad. Sci. USA 106, 18825–18830 (2009).
US Food and Drug Administration. FDA expands approved use of Kalydeco to treat additional mutations of cystic fibrosis (FDA, 2017).
Keating, D. et al. VX-445-tezacaftor-ivacaftor in patients with cystic fibrosis and one or two Phe508del alleles. N. Engl. J. Med. 379, 1612–1620 (2018).
US Food and Drug Administration. FDA approves new breakthrough therapy for cystic fibrosis (FDA, 2019).
Strug, L. J. et al. Recent advances in developing therapeutics for cystic fibrosis. Hum. Mol. Genet. 27(R2), R173–R186 (2018).
US Food and Drug Administration. Orkambi prescribing information. FDA https://www.accessdata.fda.gov/drugsatfda_docs/label/2015/206038Orig1s000lbl.pdf (2018).
Platt, F. M. Emptying the stores: lysosomal diseases and therapeutic strategies. Nat. Rev. Drug Discov. 17, 133–150 (2018).
Keeling, K. M. et al. Therapeutics based on stop codon readthrough. Annu. Rev. Genomics Hum. Genet. 15, 371–394 (2014).
Guiraud, S. & Davies, K. E. Pharmacological advances for treatment in Duchenne muscular dystrophy. Curr. Opin. Pharmacol. 34, 36–48 (2017).
Welch, E. M. et al. PTC124 targets genetic disorders caused by nonsense mutations. Nature 447, 87–91 (2007).
Squire, S. et al. Prevention of pathology in mdx mice by expression of utrophin: analysis using an inducible transgenic expression system. Hum. Mol. Genet. 11, 3333–3344 (2002).
Goldstein, G. Overview of the development of Orthoclone OKT3: monoclonal antibody for therapeutic use in transplantation. Transplant Proc. 19 (2 Suppl. 1), 1–6 (1987).
Carter, P. J. & Lazar, G. A. Next generation antibody drugs: pursuit of the high-hanging fruit. Nat. Rev. Drug Discov. 17, 197–223 (2018).
Grilo, A. L. & Mantalaris, A. The increasingly human and profitable monoclonal antibody market. Trends Biotechnol. 37, 9–16 (2018).
Kohler, G. & Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 256, 495–497 (1975).
Potter, M. The early history of plasma cell tumors in mice, 1954-1976. Adv. Cancer Res. 98, 17–51 (2007).
Clackson, T. et al. Making antibody fragments using phage display libraries. Nature 352, 624–628 (1991).
Chames, P. et al. Therapeutic antibodies: successes, limitations and hopes for the future. Br. J. Pharmacol. 157, 220–233 (2009).
Smith, K. et al. Rapid generation of fully human monoclonal antibodies specific to a vaccinating antigen. Nat. Protoc. 4, 372–384 (2009).
Ecker, D. M., Jones, S. D. & Levine, H. L. The therapeutic monoclonal antibody market. mAbs 7, 9–14 (2015).
Sedykh, S. E. et al. Bispecific antibodies: design, therapy, perspectives. Drug Des. Devel. Ther. 12, 195–208 (2018).
Thakur, A. & Lum, L. G. “NextGen” biologics: bispecific antibodies and emerging clinical results. Exp. Opin. Biol. Ther. 16, 675–688 (2016).
Rath, T. et al. Fc-fusion proteins and FcRn: structural insights for longer-lasting and more effective therapeutics. Crit. Rev. Biotechnol. 35, 235–254 (2015).
Nasiri, H. et al. Antibody-drug conjugates: promising and efficient tools for targeted cancer therapy. J. Cell Physiol. 233, 6441–6457 (2018).
Lambert, J. M. & Berkenblit, A. Antibody–drug conjugates for cancer treatment. Annu. Rev. Med. 69, 191–207 (2018).
Fleischmann, R. M. et al. Anakinra, a recombinant human interleukin-1 receptor antagonist (r-metHuIL-1ra), in patients with rheumatoid arthritis: A large, international, multicenter, placebo-controlled trial. Arthritis Rheum. 48, 927–934 (2003).
De Benedetti, F. et al. Canakinumab for the treatment of autoinflammatory recurrent fever syndromes. N. Engl. J. Med. 378, 1908–1919 (2018).
Hoffman, H. M. et al. Efficacy and safety of rilonacept (interleukin-1 Trap) in patients with cryopyrin-associated periodic syndromes: results from two sequential placebo-controlled studies. Arthritis Rheum. 58, 2443–2452 (2008).
Mahlangu, J. et al. Emicizumab prophylaxis in patients who have hemophilia A without inhibitors. N. Engl. J. Med. 379, 811–822 (2018).
Scully, M. et al. Caplacizumab treatment for acquired thrombotic thrombocytopenic purpura. N. Engl. J. Med. 380, 335–346 (2019).
Frenzel, A. et al. Designing human antibodies by phage display. Transfus. Med. Hemother. 44, 312–318 (2017).
Shukla, A. A. et al. Evolving trends in mAb production processes. Bioeng. Transl. Med. 2, 58–69 (2017).
Peters, R. & Harris, T. Advances and innovations in haemophilia treatment. Nat. Rev. Drug Discov. 17, 493–508 (2018).
Beck, M. Treatment strategies for lysosomal storage disorders. Dev. Med. Child Neurol. 60, 13–18 (2018).
LiverTox: Clinical and Research Information on Drug-Induced Liver Injury [Internet]. Enzyme Replacement Therapy. National Institute of Diabetes and Digestive and Kidney Diseases https://www.ncbi.nlm.nih.gov/books/NBK548796/ (2016).
Jurecka, A. & Tylki-Szyman´ska, A. Enzyme replacement therapy: lessons learned and emerging questions. Expert Opin. Orphan Drugs 3, 293–305 (2015).
Gadek, J. E. et al. Replacement therapy of alpha 1-antitrypsin deficiency. Reversal of protease-antiprotease imbalance within the alveolar structures of PiZ subjects. J. Clin. Invest. 68, 1158–1165 (1981).
Wewers, M. D. et al. Replacement therapy for alpha 1-antitrypsin deficiency associated with emphysema. N. Engl. J. Med. 316, 1055–1062 (1987).
Desnick, R. J. & Schuchman, E. H. Enzyme replacement therapy for lysosomal diseases: lessons from 20 years of experience and remaining challenges. Annu. Rev. Genomics Hum. Genet. 13, 307–335 (2012).
Brady, R. O. et al. Replacement therapy for inherited enzyme deficiency. Use of purified glucocerebrosidase in Gaucher’s disease. N. Engl. J. Med. 291, 989–993 (1974).
Chien, Y. H., Hwu, W. L. & Lee, N. C. Pompe disease: early diagnosis and early treatment make a difference. Pediatr. Neonatol. 54, 219–227 (2013).
Grabowski, G. A., Golembo, M. & Shaaltiel, Y. Taliglucerase alfa: an enzyme replacement therapy using plant cell expression technology. Mol. Genet. Metab. 112, 1–8 (2014).
Gaffke, L. et al. How close are we to therapies for Sanfilippo disease? Metab. Brain Dis. 33, 1–10 (2018).
Gonzalez, E. A. & Baldo, G. Gene therapy for lysosomal storage disorders: recent advances and limitations. J. Inborn Errors Metab. Screen. 5, 2326409816689786 (2017).
Chotirmall, S. H. et al. Alpha-1 proteinase inhibitors for the treatment of alpha-1 antitrypsin deficiency: safety, tolerability, and patient outcomes. Ther. Clin. Risk Manag. 11, 143–151 (2015).
Leadiant Biosciences. Adagen (pegademase bovine). Leadiant https://leadiant.com/products/adagen/ (2019).
US Food and Drug Administration. FDA approves a new treatment for PKU, a rare and serious genetic disease (FDA,2018).
Nicolino, M. Alglucosidase alfa: first available treatment for Pompe disease. Therapy 4, 271–277 (2007).
van Gelder, C. et al. A higher dose of enzyme therapy in patients with classic infantile Pompe disease seems to improve ventilator-free survival and motor function. BMC Musculoskelet. Disord. 14, 19 (2013).
Kirkegaard, T. Emerging therapies and therapeutic concepts for lysosomal storage diseases. Expert Opin. Orphan Drugs 1, 385–404 (2013).
Harmatz, P. Enzyme replacement therapies and immunogenicity in lysosomal storage diseases: is there a pattern? Clin. Ther. 37, 2130–2134 (2015).
Khvorova, A. & Watts, J. K. The chemical evolution of oligonucleotide therapies of clinical utility. Nat. Biotechnol. 35, 238 (2017).
Setten, R. L., Rossi, J. J. & Han, S. P. The current state and future directions of RNAi-based therapeutics. Nat. Rev. Drug Discov. 18, 421–446 (2019).
Zatsepin, T. S., Kotelevtsev, Y. V. & Koteliansky, V. Lipid nanoparticles for targeted siRNA delivery - going from bench to bedside. Int. J. Nanomed. 11, 3077–3086 (2016).
Havens, M. A., Duelli, D. M. & Hastings, M. L. Targeting RNA splicing for disease therapy. Wiley Interdiscip. Rev. RNA 4, 247–266 (2013).
Arechavala-Gomeza, V. et al. Antisense oligonucleotide-mediated exon skipping for Duchenne muscular dystrophy: progress and challenges. Curr. Gene Ther. 12, 152–160 (2012).
Dias, N. & Stein, C. A. Antisense oligonucleotides: basic concepts and mechanisms. Mol. Cancer Ther. 1, 347–355 (2002).
US Food and Drug Administration. Kynamro prescribing information. FDA https://www.accessdata.fda.gov/drugsatfda_docs/label/2013/203568s000lbl.pdf (2013).
Rinaldi, C. & Wood, M. J. A. Antisense oligonucleotides: the next frontier for treatment of neurological disorders. Nat. Rev. Neurol. 14, 9–21 (2018).
Adams, D. et al. Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis. N. Engl. J. Med. 379, 11–21 (2018).
Benson, M. D. et al. Inotersen treatment for patients with hereditary transthyretin amyloidosis. N. Engl. J. Med. 379, 22–31 (2018).
US Food and Drug Administration. Tegsedi prescribing information. FDA https://www.accessdata.fda.gov/drugsatfda_docs/label/2018/211172lbl.pdf (2018).
US Food and Drug Administration. FDA approves first drug for spinal muscular (FDA, 2016).
US Food and Drug Administration. SPINRAZA (nusinersen) injection for intrathecal use. FDA https://www.accessdata.fda.gov/drugsatfda_docs/label/2016/209531lbl.pdf (2016).
Field, M. J. & Boat, T. F. (eds) Rare Diseases and Orphan Products (National Academies Press, 2010).
van Roon-Mom, W. M. C., Roos, R. A. C. & de Bot, S. T. Dose-dependent lowering of mutant huntingtin using antisense oligonucleotides in Huntington disease patients. Nucleic Acid Ther. 28, 59–62 (2018).
Deverman, B. E. et al. Gene therapy for neurological disorders: progress and prospects. Nat. Rev. Drug Discov. 17, 641–659 (2018).
Srivastava, A. In vivo tissue-tropism of adeno-associated viral vectors. Curr. Opin. Virol. 21, 75–80 (2016).
Nance, M. E. & Duan, D. Perspective on adeno-associated virus capsid modification for Duchenne muscular dystrophy gene therapy. Hum. Gene Ther. 26, 786–800 (2015).
Asokan, A. Reengineered AAV vectors: old dog, new tricks. Discov. Med. 9, 399–403 (2010).
Journal of Gene Medicine. Gene therapy clinical trials worldwide. Wiley http://www.abedia.com/wiley/vectors.php (2018).
Chandler, R. J., Sands, M. S. & Venditti, C. P. Recombinant adeno-associated viral integration and genotoxicity: insights from animal models. Hum. Gene Ther. 28, 314–322 (2017).
Berns, K. I. et al. Adeno-associated virus type 2 and hepatocellular carcinoma? Hum. Gene Ther. 26, 779–781 (2015).
Colella, P., Ronzitti, G. & Mingozzi, F. Emerging issues in AAV-mediated in vivo gene therapy. Mol. Ther. Methods Clin. Dev. 8, 87–104 (2018).
Kohn, D. B. Historical perspective on the current renaissance for hematopoietic stem cell gene therapy. Hematol. Oncol. Clin. North Am. 31, 721–735 (2017).
Yu, S. F. et al. Self-inactivating retroviral vectors designed for transfer of whole genes into mammalian cells. Proc. Natl Acad. Sci. USA 83, 3194–3198 (1986).
Zufferey, R. et al. Self-inactivating lentivirus vector for safe and efficient in vivo gene delivery. J. Virol. 72, 9873–9880 (1998).
Martin, U. Therapeutic application of pluripotent stem cells: challenges and risks. Front. Med. 4, 229 (2017).
Di Foggia, V. et al. Induced pluripotent stem cell therapies for degenerative disease of the outer retina: disease modeling and cell replacement. J. Ocul. Pharmacol. Therap. 32, 240–252 (2016).
Yin, H., Kauffman, K. J. & Anderson, D. G. Delivery technologies for genome editing. Nat. Rev. Drug Discov. 16, 387–399 (2017).
Telen, M. J., Malik, P. & Vercellotti, G. M. Therapeutic strategies for sickle cell disease: towards a multi-agent approach. Nat. Rev. Drug Discov. 18, 139–158 (2019).
Mendell, J. R. et al. Single-dose gene-replacement therapy for spinal muscular atrophy. N. Engl. J. Med. 377, 1713–1722 (2017).
Chapin, J. C. & Monahan, P. E. Gene therapy for hemophilia: progress to date. BioDrugs 32, 9–25 (2018).
Yla-Herttuala, S. Endgame: glybera finally recommended for approval as the first gene therapy drug in the European union. Mol. Ther. 20, 1831–1832 (2012).
US Food and Drug Administration. FDA approves novel gene therapy to treat patients with a rare form of inherited vision loss (FDA, 2017).
European Commission. Union Register of medicinal products for human use. European Commission http://ec.europa.eu/health/documents/community-register/html/h1331.htm (2018).
De Ravin, S. S. et al. Lentiviral hematopoietic stem cell gene therapy for X-linked severe combined immunodeficiency. Sci. Transl Med. 8, 335ra57 (2016).
Aiuti, A. et al. Lentiviral hematopoietic stem cell gene therapy in patients with Wiskott-Aldrich syndrome. Science 341, 1233151 (2013).
Thompson, A. A. et al. Gene therapy in patients with transfusion-dependent beta-thalassemia. N. Engl. J. Med. 378, 1479–1493 (2018).
Aiuti, A., Roncarolo, M. G. & Naldini, L. Gene therapy for ADA-SCID, the first marketing approval of an ex vivo gene therapy in Europe: paving the road for the next generation of advanced therapy medicinal products. EMBO Mol. Med. 9, 737–740 (2017).
Cartier, N. et al. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science 326, 818–823 (2009).
Biffi, A. et al. Lentiviral hematopoietic stem cell gene therapy benefits metachromatic leukodystrophy. Science 341, 1233158 (2013).
Eichler, F. et al. Hematopoietic stem-cell gene therapy for cerebral adrenoleukodystrophy. N. Engl. J. Med. 377, 1630–1638 (2017).
US Food and Drug Administration. Orphan drug designation for Kymriah (Tisagenlecleucel) for the treatment of acute lymphoblastic leukemia (FDA, 2017).
US Food and Drug Administration. FDA approves CAR-T cell therapy to treat adults with certain types of large B-cell lymphoma (FDA, 2017).
Tang, J. et al. The global landscape of cancer cell therapy. Nat. Rev. Drug Discov. 17, 465–466 (2018).
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Moser, R. J. & Hirsch, M. L. AAV vectorization of DSB-mediated gene editing technologies. Curr. Gene Ther. 16, 207–219 (2016).
Kaiser, J. New gene-editing treatment might help treat a rare disorder, hints first human test. Science https://doi.org/10.1126/science.aav3226 (2018).
Kulkarni, J. A., Cullis, P. R. & van der Meel, R. Lipid nanoparticles enabling gene therapies: from concepts to clinical utility. Nucleic Acid Ther. 28, 146–157 (2018).
Nienhuis, A. W. Development of gene therapy for blood disorders: an update. Blood 122, 1556–1564 (2013).
Alliance for Regenerative Medicine. Quarterly data report Q3. ARM http://alliancerm.org/wp-content/uploads/2018/10/ARM_Q3_2018_Web-1.pdf (2018).
Oprea, T. I. et al. Associating drugs, targets and clinical outcomes into an integrated network affords a new platform for computer-aided drug repurposing. Mol. Inform. 30, 100–111 (2011).
Oprea, T. I. et al. Drug repurposing from an academic perspective. Drug Discov. Today Ther. Strateg. 8, 61–69 (2011).
Ursu, O. et al. DrugCentral 2018: an update. Nucleic Acids Res. 47(D1), D963–D970 (2019).
Pushpakom, S. et al. Drug repurposing: progress, challenges and recommendations. Nat. Rev. Drug Discov. 8, 41–58 (2019).
Tenenbaum, J. D. Translational bioinformatics: past, present, and future. Genomics Proteomics Bioinformatics 14, 31–41 (2016).
Butte, A. J. & Chen, R. Finding disease-related genomic experiments within an international repository: first steps in translational bioinformatics. AMIA Annu. Symp. Proc. 2006, 106–110 (2006).
Himmelstein, D. S. & Baranzini, S. E. Heterogeneous network edge prediction: a data integration approach to prioritize disease-associated genes. PLOS Comput. Biol. 11, e1004259 (2015).
Ghofrani, H. A., Osterloh, I. H. & Grimminger, F. Sildenafil: from angina to erectile dysfunction to pulmonary hypertension and beyond. Nat. Rev. Drug Discov. 5, 689–702 (2006).
Galie, N. et al. Sildenafil citrate therapy for pulmonary arterial hypertension. N. Engl. J. Med. 353, 2148–2157 (2005).
Colvis, C. M. & Austin, C. P. The NIH-industry new therapeutic uses pilot program: demonstrating the power of crowdsourcing. Assay Drug Dev. Technol. 13, 297–298 (2015).
Markey, K. A. et al. Assessing the efficacy and safety of an 11beta-hydroxysteroid dehydrogenase type 1 inhibitor (AZD4017) in the idiopathic intracranial hypertension drug trial, IIH:DT: clinical methods and design for a phase II randomized controlled trial. JMIR Res. Protoc. 6, e181 (2017).
Huang, R. et al. The NCGC pharmaceutical collection: a comprehensive resource of clinically approved drugs enabling repurposing and chemical genomics. Sci. Transl Med. 3, 80ps16 (2011).
Corsello, S. M. et al. The drug repurposing hub: a next-generation drug library and information resource. Nat. Med. 23, 405–408 (2017).
Mariz, S. et al. Worldwide collaboration for orphan drug designation. Nat. Rev. Drug Discov. 15, 440 (2016).
Gammie, T., Lu, C. Y. & Babar, Z. U. Access to orphan drugs: a comprehensive review of legislations, regulations and policies in 35 countries. PLOS ONE 10, e0140002 (2015).
European Medicines Agency. Patients’ and consumers’ working party (EMA, 2019).
Spencer, D. et al. Integrating rare disease patients into pre-clinical therapy development; finding our way with patient input (BioPontis Alliance, 2016).
BioPontis Alliance for Rare Diseases. Resources (BioPontis Alliance, 2019).
US Food and Drug Administration. Guidance for industry patient-reported outcome measures: use in medical product development to support labeling claims (FDA, 2009).
Contesse, M. G. et al. The case for the use of patient and caregiver perception of change assessments in rare disease clinical trials: a methodologic overview. Adv. Ther. 36, 997–1010 (2019).
Bloom, D. et al. The rules of engagement: CTTI recommendations for successful collaborations between sponsors and patient groups around clinical trials. Ther. Innov. Regul. Sci. 52, 206–213 (2018).
BioPontis Alliance for Rare Diseases. Translational research readiness tool developed with rare disease patients organizations (BioPontis Alliance, 2017).
Jayasundara, K. et al. Estimating the clinical cost of drug development for orphan versus non-orphan drugs. Orphanet J. Rare Dis. 14, 12 (2019).
Brooks, P. J., Tagle, D. A. & Groft, S. Expanding rare disease drug trials based on shared molecular etiology. Nat. Biotechnol. 32, 515–518 (2014).
Moffat, J. G. et al. Opportunities and challenges in phenotypic drug discovery: an industry perspective. Nat. Rev. Drug Discov. 16, 531–543 (2017).
Berg, A. et al. A phenotypic screen for corrector discovery using a surface liquid readout in F508del primary airway epithelia. Pediatr. Pulmonol. 50, S77–S107 (2015).
Philippakis, A. A. et al. The matchmaker exchange: a platform for rare disease gene discovery. Hum. Mutat 36, 915–921 (2015).
Nguengang Wakap, S. et al. Estimating cumulative point prevalence of rare diseases: analysis of the Orphanet database. Eur. J. Hum. Genet. https://doi.org/10.1038/s41431-019-0508-0 (2019).
Haendel, M. et al. How many rare diseases are there? Nat. Rev. Drug Discov. https://doi.org/10.1038/d41573-019-00180-y (2019).
Oprea, T. I. et al. Unexplored therapeutic opportunities in the human genome. Nat. Rev. Drug Discov. 17, 317–332 (2018).
Nguyen, D. T. et al. Pharos: collating protein information to shed light on the druggable genome. Nucleic Acids Res. 45(D1), D995–D1002 (2017).
Jia, J. et al. eRAM: encyclopedia of rare disease annotations for precision medicine. Nucleic Acids Res. 46(D1), D937–D943 (2018).
Porta, M. A Dictionary of Epidemiology 193–194 (Oxford Univ. Press, 2014).
Kempf, L., Goldsmith, J. C. & Temple, R. Challenges of developing and conducting clinical trials in rare disorders. Am. J. Med. Genet. A 176, 773–783 (2018).
US Food and Drug Administration. Rare diseases: natural history studies for drug development (FDA, 2019).
Gavin, P. The importance of natural histories for rare diseases. Expert Opin. Orphan Drugs 3, 855–857 (2015).
Temple, R. Historically controlled trials. FDA https://www.fda.gov/media/97835/download (2016).
US Food and Drug Administration. Guidance for industry (FDA, 2019).
US Food and Drug Administration. Rare diseases: natural history studies for drug development guidance for industry (FDA, 2019).
Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin. Pharmacol. Ther. 69, 89–95 (2001).
Strimbu, K. & Tavel, J. A. What are biomarkers? Curr. Opin. HIV AIDS 5, 463–466 (2010).
Federal Communications Commission. Ingestibles, wearables and embeddables (FCC, 2018).
Deiters, W., Burmann, A. & Meister, S. Strategies for digitalizing the hospital of the future [German]. Urologe A 57, 1031–1039 (2018).
Sawicki, G. S. et al. Sustained benefit from ivacaftor demonstrated by combining clinical trial and cystic fibrosis patient registry data. Am. J. Respir. Crit. Care Med. 192, 836–842 (2015).
Weidemann, F. et al. Usefulness of an implantable loop recorder to detect clinically relevant arrhythmias in patients with advanced Fabry cardiomyopathy. Am. J. Cardiol. 118, 264–274 (2016).
Menotti, F. et al. Amount and intensity of daily living activities in Charcot-Marie-Tooth 1A patients. Brain Behav. 4, 14–20 (2014).
Pande, A. et al. Machine learning to improve energy expenditure estimation in children with disabilities: a pilot study in Duchenne muscular dystrophy. JMIR Rehabil. Assist. Technol. 3, e7 (2016).
Hay, C. R. M. et al. The haemtrack home therapy reporting system: design, implementation, strengths and weaknesses: A report from UK Haemophilia Centre Doctors Organisation. Haemophilia 23, 728–735 (2017).
Calvo-Lerma, J. et al. Innovative approach for self-management and social welfare of children with cystic fibrosis in Europe: development, validation and implementation of an mHealth tool (MyCyFAPP). BMJ Open 7, e014931 (2017).
Slade, A. et al. Patient reported outcome measures in rare diseases: a narrative review. Orphanet J. Rare Dis. 13, 61 (2018).
Benjamin, K. et al. Patient-reported outcome and observer-reported outcome assessment in rare disease clinical trials: an ISPOR COA emerging good practices task force report. Value Health 20, 838–855 (2017).
Lechtzin, N. et al. Home monitoring of patients with cystic fibrosis to identify and treat acute pulmonary exacerbations. eICE study results. Am. J. Respir. Crit. Care Med. 196, 1144–1151 (2017).
Cox, G. F. The art and science of choosing efficacy endpoints for rare disease clinical trials. Am. J. Med. Genet. A 176, 759–772 (2018).
Groft, S. C. & Posada de la Paz, M. Preparing for the future of rare diseases. Adv. Exp. Med. Biol. 1031, 641–648 (2017).
Noah, B. et al. Impact of remote patient monitoring on clinical outcomes: an updated meta-analysis of randomized controlled trials. NPJ Digit. Med. 1, 20172 (2018).
Anselmo, A. C., Gokarn, Y. & Mitragotri, S. Non-invasive delivery strategies for biologics. Nat. Rev. Drug Discov. 18, 19–40 (2019).
Acknowledgements
The authors acknowledge M. Lanthier and K. Miller, who helped formulate FDA data, and M. Gomar Mengod, who helped formulate EMA data for the figures presented. The authors acknowledge work done by the combined teams of C. J. Mungall (Lawrence Berkeley National Laboratory), M. A. Haendel (Oregon Health Sciences University and Oregon State University), P. J. Robinson (Jackson Laboratories) and D.-T. Nguyen (US National Institutes of Health National Center for Advancing Translational Sciences) for Fig. 1 and J. Holmes and S. L. Mathias (University of New Mexico) for the figure related to the rare disease proteome in Box 2. The authors also sincerely acknowledge the help of F. Sasinowsky for the sections on ASOs, natural history and patient engagement, and of E. Powers for the gene and cell therapy section. The authors thank D. Spencer for rereading and commenting on the article. NIH grants U24 CA224370, U24 TR002278, U01 CA239108, UL1 TR001449 and P30 CA118100 provided funding to T.I.O.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Disclaimer
The views expressed in this article are the personal views of the authors and may not be understood or quoted as being made on behalf of or reflecting the position of regulatory agencies or organizations with which they are employed/affiliated.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Drug repurposing hub: https://clue.io/repurposing
E-Rare. E-Rare-3 call for proposals 2016: www.erare.eu/joint-call/e-rare-3-call-proposals-2016-jtc-2016-clinical-research-new-therapeutic-uses-already-0
EURORDIS Rare Diseases Europe: https://www.eurordis.org
European Lead Factory: https://www.europeanleadfactory.eu/
European Medicines Agency. Committee for Orphan Medicinal Products: https://www.ema.europa.eu/en/committees/committee-orphan-medicinal-products-comp
European Medicines Agency. Orphan designation overview: https://www.ema.europa.eu/en/human-regulatory/overview/orphan-designation-overview
European Medicines Agency. Rare diseases, orphan medicines. Getting the facts straight: https://www.ema.europa.eu/en/documents/other/rare-diseases-orphan-medicines-getting-facts-straight_en.pdf
FDA Rare Diseases Program: https://www.fda.gov/about-fda/center-drug-evaluation-and-research-cder/rare-diseases-program
Genetic and Rare Diseases Information Center of the US National Institutes of Health: https://rarediseases.info.nih.gov/
IRDiRC. Datamining and repurposing: http://www.irdirc.org/activities/task-forces/data-miningrepurposing
Mondo disease ontology: https://mondo.monarchinitiative.org/
National Institutes of Health Bridging Interventional Development Gaps Program: https://ncats.nih.gov/bridgs
National Institutes of Health Drug Record. Livertox: enzyme replacement therapy: https://www.ncbi.nlm.nih.gov/books/NBK548796/
NORD: https://rarediseases.org/
Orphanet, an online database of rare diseases and orphan drugs: http://www.orpha.net
Pharos: https://pharos.nih.gov/
Rare Diseases Registry (RaDaR) Program: https://rarediseases.info.nih.gov/radar
Supplementary information
Glossary
- Lipinski’s rule of five
-
These guidelines identify several physicochemical properties to be considered for small molecules that are intended for oral delivery: molecular mass 500 Da or less; five or fewer hydrogen-bond donors; fewer than 10 hydrogen-bond acceptors; and calculated octanol–water partition coefficient (a surrogate for the ability of a molecule to cross biological membranes) of 5 or less.
- CFTR
-
Gene encoding the cystic fibrosis transmembrane conductance regulator protein, an ion channel in the membrane of cells that produce mucus, sweat, saliva, tears and digestive enzymes. Mutations in CFTR that affect the production, processing or function of the protein underlie cystic fibrosis.
- ‘Umbrella trial’
-
A clinical trial design in which a single drug is evaluated in more than one disease simultaneously.
- Nanobody
-
A type of single-domain antibody fragment.
- Good manufacturing practice
-
A system for ensuring that products are consistently produced and controlled according to defined quality standards.
- Pegylation
-
Attaching polyethylene glycol chains to therapeutics, particularly proteins, can improve characteristics such as immunogenicity and pharmacokinetics. For example, pegylation has been used to extend the half-life of factor VIII replacement therapies for haemophilia.
- Intrathecal injection
-
Delivery of a substance directly to the spinal fluid (intrathecal space) through a drug delivery system comprising a pump and a catheter.
- Haematopoietic stem cells
-
(HSCs). Cells that can replenish all blood cell types. HSCs derived from bone marrow have been used for many years to treat cancer; patients receive a myeloablative conditioning regimen to remove diseased cells before transplantation, with the transplanted HSCs then reconstituting the haematopoietic system. A similar strategy can also be used to treat inherited blood disorders.
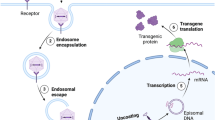
- Adeno-associated virus (AAV) vectors
-
AAV vectors are based on wild-type AAV, which has a single-stranded circular genome of roughly 4.7 kilobases. The AAV genome contains two open reading frames bounded by inverted terminal repeats into which a transgene of up to approximately five kilobases can be inserted.
- Capsid
-
A protein shell that originally encloses the viral genome.
- Fast track pathway
-
This can expedite the review of products to treat serious conditions. The process allows sponsors to have more frequent meetings and communications with the FDA to address appropriate data collection and design of clinical trials. It also allows a sponsor to be eligible for priority review and a rolling review of the application.
- Accelerated approval
-
This allows a product for a serious condition to be approved on the basis of a surrogate end point or an intermediate clinical end point. Confirmatory postmarketing trials will be needed to verify this benefit.
- Priority review
-
This is a designation that allows the FDA to act on a marketing authorization application in 6 months (compared with 10 months for standard reviews). To be eligible for priority review, the intended medicine should offer significant advancements in safety and efficacy of treatment, diagnosis or prevention of a serious condition.
- Breakthrough therapy deignation
-
This FDA designation can expedite development of drugs for which preliminary clinical evidence indicates that they may offer substantial advantages over existing treatment options for patients with serious or life-threatening diseases. Designated drugs are eligible for the expedited processing that fast track designation offers, as well as intensive guidance on efficient development from the FDA.
- Regenerative medicine advanced therapy designation
-
This FDA designation is similar to the breakthrough threapy designation and is available for cell therapies, therapeutic tissue engineering products, human cell and tissue products and combination products if the product is intended to treat serious or life-threatening diseases.
- Conditional marketing authorization
-
This European Medicines Agency pathway is similar to the accelerated approval process in the United States. Applicants may be granted a conditional marketing authorization for medicines for which the benefit of immediate availability outweighs the risk of less comprehensive clinical data than normally required.
- Approval under exceptional circumstances
-
In exceptional cases, a reduced data set is acceptable by the European Medicines Agency for candidate drugs for a rare indication with a high medical need if it is difficult to obtain sufficient data to fulfil the requirements of a full dossier for marketing authorization in a reasonable time frame. Annual review of clinical data obtained after such approval is required, with the potential to maintain or withdraw the authorization.
- Accelerated assessment
-
The evaluation of a marketing authorization application under the centralized procedure in the European Union can take up to 210 days. On request, the time frame can be reduced to 150 days if the applicant provides sufficient justification that the medicinal product is expected to be of major public health interest, particularly in cases of therapeutic innovation.
- Priority Medicines (PRIME) scheme
-
A scheme in the European Union that provides early and enhanced scientific and regulatory support for medicines that may offer a major therapeutic advantage over existing treatments, or benefit patients without treatment options.
Rights and permissions
About this article
Cite this article
Tambuyzer, E., Vandendriessche, B., Austin, C.P. et al. Therapies for rare diseases: therapeutic modalities, progress and challenges ahead. Nat Rev Drug Discov 19, 93–111 (2020). https://doi.org/10.1038/s41573-019-0049-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41573-019-0049-9
This article is cited by
-
The Role of Pharmacogenomics in Rare Diseases
Drug Safety (2024)
-
Determining Commonalities in the Experiences of Patients with Rare Diseases: A Qualitative Analysis of US Food and Drug Administration Patient Engagement Sessions
The Patient - Patient-Centered Outcomes Research (2024)
-
Systematic and quantitative analysis of stop codon readthrough in Rett syndrome nonsense mutations
Journal of Molecular Medicine (2024)
-
High-content screening identifies a small molecule that restores AP-4-dependent protein trafficking in neuronal models of AP-4-associated hereditary spastic paraplegia
Nature Communications (2024)
-
New orphan disease therapies from the proteome of industrial plasma processing waste- a treatment for aceruloplasminemia
Communications Biology (2024)