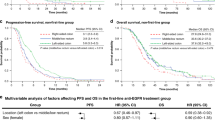
Abstract
The number of molecularly stratified treatment options available to patients with colorectal cancer (CRC) is increasing, with a parallel rise in the use of biomarkers to guide prognostication and treatment decision-making. The increase in both the number of biomarkers and their use has resulted in a progressively complex situation, evident both from the extensive interactions between biomarkers and from their sometimes complex associations with patient prognosis and treatment benefit. Current and emerging biomarkers also reflect the genomic complexity of CRC, and include a wide range of aberrations such as point mutations, amplifications, fusions and hypermutator phenotypes, in addition to global gene expression subtypes. In this Review, we provide an overview of current and emerging clinically relevant biomarkers and their role in the management of patients with CRC, illustrating the intricacies of biomarker interactions and the growing treatment opportunities created by the availability of comprehensive molecular profiling.
Key points
-
The expanded use of biomarkers to guide the treatment of patients with colorectal cancer has revealed a level of complexity arising from interactions between different biomarkers.
-
An improved understanding of the causes of primary resistance might increase response rates among patients receiving targeted therapies and enable more-effective drug combinations, exemplified by mutations in the MAPK signalling pathway for EGFR-targeted and/or BRAF-targeted therapies.
-
Immune checkpoint inhibition has provided the largest contribution to the increased use of molecularly guided therapies, and biomarkers that complement patient stratification by microsatellite instability status are likely to provide further benefit.
-
Biomarkers that indicate a poor prognosis have motivated the search for more effective therapies for specific molecular subgroups; these biomarkers typically have a limited prevalence, but their accumulation could expand the eligibility for, and benefit from, targeted treatment.
-
Some colorectal cancers harbour more than one molecular target and treatment sequencing in relation to both standard and targeted therapies is a growing challenge.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Linnekamp, J. F., Wang, X., Medema, J. P. & Vermeulen, L. Colorectal cancer heterogeneity and targeted therapy: a case for molecular disease subtypes. Cancer Res. 75, 245–249 (2015).
Dienstmann, R., Salazar, R. & Tabernero, J. Personalizing colon cancer adjuvant therapy: selecting optimal treatments for individual patients. J. Clin. Oncol. 33, 1787–1796 (2015).
Dienstmann, R. et al. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat. Rev. Cancer 17, 79–92 (2017).
Schmoll, H. J. et al. ESMO consensus guidelines for management of patients with colon and rectal cancer. A personalized approach to clinical decision making. Ann. Oncol. 23, 2479–2516 (2012).
Grothey, A. et al. Duration of adjuvant chemotherapy for stage III colon cancer. N. Engl. J. Med. 378, 1177–1188 (2018).
Tabernero, J. et al. Ramucirumab versus placebo in combination with second-line FOLFIRI in patients with metastatic colorectal carcinoma that progressed during or after first-line therapy with bevacizumab, oxaliplatin, and a fluoropyrimidine (RAISE): a randomised, double-blind, multicentre, phase 3 study. Lancet Oncol. 16, 499–508 (2015).
Grothey, A. et al. Regorafenib monotherapy for previously treated metastatic colorectal cancer (CORRECT): an international, multicentre, randomised, placebo-controlled, phase 3 trial. Lancet 381, 303–312 (2013).
Cremolini, C. et al. First-line chemotherapy for mCRC-a review and evidence-based algorithm. Nat. Rev. Clin. Oncol. 12, 607–619 (2015).
Tabernero, J. et al. Analysis of angiogenesis biomarkers for ramucirumab efficacy in patients with metastatic colorectal cancer from RAISE, a global, randomized, double-blind, phase III study. Ann. Oncol. 29, 602–609 (2018).
Toledo, R. A. et al. Exome sequencing of plasma DNA portrays the mutation landscape of colorectal cancer and discovers mutated VEGFR2 receptors as modulators of anti-angiogenic therapies. Clin. Cancer Res. 24, 3550–3559 (2018).
Van Cutsem, E. et al. ESMO consensus guidelines for the management of patients with metastatic colorectal cancer. Ann. Oncol. 27, 1386–1422 (2016).
Van Cutsem, E. et al. Fluorouracil, leucovorin, and irinotecan plus cetuximab treatment and RAS mutations in colorectal cancer. J. Clin. Oncol. 33, 692–700 (2015).
Hammond, W. A., Swaika, A. & Mody, K. Pharmacologic resistance in colorectal cancer: a review. Ther. Adv. Med. Oncol. 8, 57–84 (2016).
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486, 532–536 (2012).
Siravegna, G. et al. Clonal evolution and resistance to EGFR blockade in the blood of colorectal cancer patients. Nat. Med. 21, 795–801 (2015). This reference describes the tracking of the evolution of resistance-bearing subclones in liquid biopsy samples during treatment with anti-EGFR antibodies, thus providing a molecular explanation for the efficacy of therapy rechallenge.
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015). Prospective trial showing that MMR status predicts clinical benefit from the ICI pembrolizumab in patients with treatment-refractory metastatic cancers.
Diaz, L. A. et al. Pembrolizumab therapy for microsatellite instability high (MSI-H) colorectal cancer (CRC) and non-CRC. J. Clin. Oncol. 35, (3071 (2017).
Overman, M. J. et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): an open-label, multicentre, phase 2 study. Lancet Oncol. 18, 1182–1191 (2017).
Gong, J., Wang, C., Lee, P. P., Chu, P. & Fakih, M. Response to PD-1 blockade in microsatellite stable metastastic colorectal cancer harboring a POLE mutation. J. Natl Compr. Canc. Netw. 15, 142–147 (2017).
Hong, D. S. et al. Phase IB study of vemurafenib in combination with irinotecan and cetuximab in patients with metastatic colorectal cancer with BRAF V600E mutation. Cancer Discov. 6, 1352–1365 (2016).
Kopetz, S. et al. Randomized trial of irinotecan and cetuximab with or without vemurafenib in BRAF-mutant metastatic colorectal cancer (SWOG 1406). J. Clin. Oncol. 35, 520 (2017). Meeting abstract describing initial data from a prospective randomized trial indicating prolonged PFS in patients with BRAF V600E-mutated and RAS wild-type metastatic CRCs by addition of vemurafenib to the combination of irinotecan plus cetuximab.
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015). The international CRC subtyping consortium combined several gene expression-based classification frameworks for CRC into the four consensus molecular subtypes, based on analysis of almost 4,000 primary tumours.
van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933–945 (2015).
Siena, S. et al. Rechallenge with EGFR inhibitors in patients with metastatic colorectal cancer: effect on outcomes [abstract P-320]. Ann. Oncol. 28 (Suppl. 3), mdx261.317 (2017).
Marquart, J., Chen, E. Y. & Prasad, V. Estimation of the percentage of US patients with cancer who benefit from genome-driven oncology. JAMA Oncol. 4, 1093–1098 (2018).
Ballman, K. V. Biomarker: predictive or prognostic? J. Clin. Oncol. 33, 3968–3971 (2015).
Labianca, R. et al. Early colon cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 24, vi64–vi72 (2013).
Duffy, M. J. et al. Tumor markers in colorectal cancer, gastric cancer and gastrointestinal stromal cancers: European Group on Tumor Markers (EGTM) 2013 guidelines update. Int. J. Cancer 134, 2513–2522 (2013).
Sepulveda, A. R. et al. Molecular biomarkers for the evaluation of colorectal cancer: guideline from the American Society for Clinical Pathology, College of American Pathologists, Association for Molecular Pathology, and the American Society of Clinical Oncology. J. Clin. Oncol. 35, 1453–1486 (2017).
Thibodeau, S. N., Bren, G. & Schaid, D. Microsatellite instability in cancer of the proximal colon. Science 260, 816–819 (1993).
Lothe, R. A. et al. Genomic instability in colorectal cancer: relationship to clinicopathological variables and family history. Cancer Res. 53, 5849–5852 (1993).
Popat, S., Hubner, R. & Houlston, R. S. Systematic review of microsatellite instability and colorectal cancer prognosis. J. Clin. Oncol. 23, 609–618 (2005).
Guastadisegni, C., Colafranceschi, M., Ottini, L. & Dogliotti, E. Microsatellite instability as a marker of prognosis and response to therapy: a meta-analysis of colorectal cancer survival data. Eur. J. Cancer 46, 2788–2798 (2010).
Sargent, D. J. et al. Defective mismatch repair as a predictive marker for lack of efficacy of fluorouracil-based adjuvant therapy in colon cancer. J. Clin. Oncol. 28, 3219–3226 (2010).
Hutchins, G. et al. Value of mismatch repair, KRAS, and BRAF mutations in predicting recurrence and benefits from chemotherapy in colorectal cancer. J. Clin. Oncol. 29, 1261–1270 (2011).
Myeroff, L. L. et al. A transforming growth factor beta receptor type II gene mutation common in colon and gastric but rare in endometrial cancers with microsatellite instability. Cancer Res. 55, 5545–5547 (1995).
Rampino, N. et al. Somatic frameshift mutations in the BAX gene in colon cancers of the microsatellite mutator phenotype. Science 275, 967–969 (1997).
Souza, R. F. et al. Microsatellite instability in the insulin-like growth factor II receptor gene in gastrointestinal tumours. Nat. Genet. 14, 255–257 (1996).
Thorstensen, L. et al. WNT1 inducible signaling pathway protein 3, WISP-3, a novel target gene in colorectal carcinomas with microsatellite instability. Gastroenterology 121, 1275–1280 (2001).
Røyrvik, E. C., Ahlquist, T., Rognes, T. & Lothe, R. A. Slip slidin’ away: a duodecennial review of targeted genes in mismatch repair deficient colorectal cancer. Crit. Rev. Oncog. 13, 229–257 (2007).
Dolcetti, R. et al. High prevalence of activated intraepithelial cytotoxic T lymphocytes and increased neoplastic cell apoptosis in colorectal carcinomas with microsatellite instability. Am. J. Pathol. 154, 1805–1813 (1999).
Jass, J. R. et al. Morphology of sporadic colorectal cancer with DNA replication errors. Gut 42, 673–679 (1998).
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Pages, F. et al. In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J. Clin. Oncol. 27, 5944–5951 (2009).
Pages, F. et al. International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet 391, 2128–2139 (2018). Retrospective international multi-centre study showing that the immunoscore has prognostic value beyond that of clinicopathological prognostic factors and MSI status in patients with stage I–III colon cancers.
Grady, W. M. & Carethers, J. M. Genomic and epigenetic instability in colorectal cancer pathogenesis. Gastroenterology 135, 1079–1099 (2008).
Ward, R. et al. Microsatellite instability and the clinicopathological features of sporadic colorectal cancer. Gut 48, 821–829 (2001).
Gryfe, R. et al. Tumor microsatellite instability and clinical outcome in young patients with colorectal cancer. N. Engl. J. Med. 342, 69–77 (2000).
Samowitz, W. S. et al. Microsatellite instability in sporadic colon cancer is associated with an improved prognosis at the population level. Cancer Epidemiol. Biomarkers Prev. 10, 917–923 (2001).
Roth, A. D. et al. Integrated analysis of molecular and clinical prognostic factors in stage II/III colon cancer. J. Natl Cancer Inst. 104, 1635–1646 (2012).
Vilar, E. & Gruber, S. B. Microsatellite instability in colorectal cancer-the stable evidence. Nat. Rev. Clin. Oncol. 7, 162 (2010).
Dienstmann, R. et al. Prediction of overall survival in stage II and III colon cancer beyond TNM system: a retrospective, pooled biomarker study. Ann. Oncol. 28, 1023–1031 (2017).
Breivik, J. et al. Different genetic pathways to proximal and distal colorectal cancer influenced by sex-related factors. Int. J. Cancer 74, 664–669 (1997).
Sinicrope, F. A. et al. Prognostic impact of deficient DNA mismatch repair in patients with stage III colon cancer from a randomized trial of FOLFOX-based adjuvant chemotherapy. J. Clin. Oncol. 31, 3664–3672 (2013).
Merok, M. A. et al. Microsatellite instability has a positive prognostic impact on stage II colorectal cancer after complete resection: results from a large, consecutive Norwegian series. Ann. Oncol. 24, 1274–1282 (2013).
Klingbiel, D. et al. Prognosis of stage II and III colon cancer treated with adjuvant 5-fluorouracil or FOLFIRI in relation to microsatellite status: results of the PETACC-3 trial. Ann. Oncol. 26, 126–132 (2015).
Sinicrope, F. A. et al. DNA mismatch repair status and colon cancer recurrence and survival in clinical trials of 5-fluorouracil-based adjuvant therapy. J. Natl Cancer Inst. 103, 863–875 (2011).
Benatti, P. et al. Microsatellite instability and colorectal cancer prognosis. Clin. Cancer Res. 11, 8332–8340 (2005).
Jo, W. S. & Carethers, J. M. Chemotherapeutic implications in microsatellite unstable colorectal cancer. Cancer Biomark. 2, 51–60 (2006).
Carethers, J. M. et al. Use of 5-fluorouracil and survival in patients with microsatellite-unstable colorectal cancer. Gastroenterology 126, 394–401 (2004).
Lanza, G. et al. Immunohistochemical test for MLH1 and MSH2 expression predicts clinical outcome in stage II and III colorectal cancer patients. J. Clin. Oncol. 24, 2359–2367 (2006).
Jover, R. et al. The efficacy of adjuvant chemotherapy with 5-fluorouracil in colorectal cancer depends on the mismatch repair status. Eur. J. Cancer 45, 365–373 (2009).
Ribic, C. M. et al. Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N. Engl. J. Med. 349, 247–257 (2003). Retrospective analysis of pooled data from randomized trials involving 5-FU-based adjuvant chemotherapies in patients with stage II or III colon cancer, indicating a significant difference in the effects of treatment in patients with MSI-H and MSS tumours.
Andre, T. et al. Oxaliplatin, fluorouracil, and leucovorin as adjuvant treatment for colon cancer. N. Engl. J. Med. 350, 2343–2351 (2004).
Flejou, J. F. et al. Effect of adding oxaliplatin to adjuvant 5-fluorouracil/leucovorin (5FU/LV) in patients with defective mismatch repair (dMMR) colon cancer stage II and III included in the MOSIAC study. J. Clin. Oncol. 31, 3524 (2013).
Tougeron, D. et al. Efficacy of adjuvant chemotherapy in colon cancer with microsatellite instability: a large multicenter AGEO study. J. Natl Cancer Inst. 108, djv438 (2016).
Zaanan, A. et al. Defective mismatch repair status as a prognostic biomarker of disease-free survival in stage III colon cancer patients treated with adjuvant FOLFOX chemotherapy. Clin. Cancer Res. 17, 7470–7478 (2011).
Gavin, P. G. et al. Mutation profiling and microsatellite instability in stage II and III colon cancer: an assessment of their prognostic and oxaliplatin predictive value. Clin. Cancer Res. 18, 6531–6541 (2012).
Sinicrope, F. et al. Overall survival result and outcomes by KRAS, BRAF, and DNA mismatch repair in relation to primary tumor site in colon cancers from a randomized trial of adjuvant chemotherapy: NCCTG (Alliance) N0147. J. Clin. Oncol. 32, 3525–3525 (2014).
Saltz, L. B. et al. Irinotecan fluorouracil plus leucovorin is not superior to fluorouracil plus leucovorin alone as adjuvant treatment for stage III colon cancer: results of CALGB 89803. J. Clin. Oncol. 25, 3456–3461 (2007).
Ychou, M. et al. A phase III randomised trial of LV5FU2 + irinotecan versus LV5FU2 alone in adjuvant high-risk colon cancer (FNCLCC Accord02/FFCD9802). Ann. Oncol. 20, 674–680 (2009).
Van Cutsem, E. et al. Randomized phase III trial comparing biweekly infusional fluorouracil/leucovorin alone or with irinotecan in the adjuvant treatment of stage III colon cancer: PETACC-3. J. Clin. Oncol. 27, 3117–3125 (2009).
Bertagnolli, M. M. et al. Microsatellite instability predicts improved response to adjuvant therapy with irinotecan, fluorouracil, and leucovorin in stage III colon cancer: Cancer and Leukemia Group B Protocol 89803. J. Clin. Oncol. 27, 1814–1821 (2009).
Schutte, M. et al. Molecular dissection of colorectal cancer in pre-clinical models identifies biomarkers predicting sensitivity to EGFR inhibitors. Nat. Commun. 8, 14262 (2017).
Campbell, B. B. et al. Comprehensive analysis of hypermutation in human cancer. Cell 171, 1042–1056 (2017).
Angelova, M. et al. Characterization of the immunophenotypes and antigenomes of colorectal cancers reveals distinct tumor escape mechanisms and novel targets for immunotherapy. Genome Biol. 16, 64 (2015).
Willis, J. et al. Impact of microsatellite instability (MSI) on tumor clonal evolution in metastatic colorectal cancer (mCRC). J. Clin. Oncol. 36, 616 (2018).
Peltomaki, P. & Vasen, H. Mutations associated with HNPCC predisposition — update of ICG-HNPCC/INSiGHT mutation database. Dis. Markers 20, 269–276 (2004).
Weisenberger, D. J. et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat. Genet. 38, 787–793 (2006).
Cohen, R. et al. Clinical and molecular characterisation of hereditary and sporadic metastatic colorectal cancers harbouring microsatellite instability/DNA mismatch repair deficiency. Eur. J. Cancer 86, 266–274 (2017).
Haraldsdottir, S. et al. Patients with colorectal cancer associated with Lynch syndrome and MLH1 promoter hypermethylation have similar prognoses. Genet. Med. 18, 863–868 (2016).
Sinicrope, F. A. et al. Molecular markers identify subtypes of stage III colon cancer associated with patient outcomes. Gastroenterology 148, 88–99 (2015).
Sato, K. et al. Fusion kinases identified by genomic analyses of sporadic microsatellite instability-high colorectal cancers. Clin. Cancer Res. 25, 378–389 (2019).
Benson, A. B. et al. NCCN clinical practice guidelines in oncology: colon cancer. Version 4. 2018. NCCN https://www.nccn.org/professionals/physician_gls/pdf/colon.pdf (2018).
Dalerba, P. et al. CDX2 as a prognostic biomarker in stage II and stage III colon cancer. N. Engl. J. Med. 374, 211–222 (2016).
Bruun, J. et al. Prognostic, predictive and pharmacogenomic assessments of CDX2 refine stratification of colorectal cancer. Mol. Oncol. 12, 1639–1655 (2018).
Wang, J. et al. Prevalence of recurrent oncogenic fusion in mismatch repair-deficient colorectal carcinoma with hypermethylated MLH1 and wild-type BRAF and KRAS. Mod. Pathol. 32, 1053–1064 (2019).
Kloosterman, W. P. et al. A systematic analysis of oncogenic gene fusions in primary colon cancer. Cancer Res. 77, 3814–3822 (2017).
Sveen, A. et al. Multilevel genomics of colorectal cancers with microsatellite instability — clinical impact of JAK1 mutations and consensus molecular subtype 1. Genome Med. 9, 46 (2017).
Mlecnik, B. et al. Integrative analyses of colorectal cancer show immunoscore is a stronger predictor of patient survival than microsatellite instability. Immunity 44, 698–711 (2016).
Galon, J. et al. Cancer classification using the Immunoscore: a worldwide task force. J. Transl Med. 10, 205 (2012).
Venderbosch, S. et al. Mismatch repair status and BRAF mutation status in metastatic colorectal cancer patients: a pooled analysis of the CAIRO, CAIRO2, COIN, and FOCUS studies. Clin. Cancer Res. 20, 5322–5330 (2014). Retrospective analysis of pooled data from randomized clinical trials showing that dMMR cancers are associated with inferior PFS and OS compared with MMR-proficient cancers in patients with metastatic CRC, and that this effect is likely driven by enrichment with BRAF V600E mutations.
Koopman, M. et al. Deficient mismatch repair system in patients with sporadic advanced colorectal cancer. Br. J. Cancer 100, 266–273 (2009).
Smith, C. G. et al. Somatic profiling of the epidermal growth factor receptor pathway in tumors from patients with advanced colorectal cancer treated with chemotherapy +/- cetuximab. Clin. Cancer Res. 19, 4104–4113 (2013).
Halama, N. et al. Localization and density of immune cells in the invasive margin of human colorectal cancer liver metastases are prognostic for response to chemotherapy. Cancer Res. 71, 5670–5677 (2011).
Mlecnik, B. et al. Comprehensive intrametastatic immune quantification and major impact of immunoscore on survival. J. Natl Cancer Inst. 110, djx123 (2018).
Tanis, E. et al. Prognostic impact of immune response in resectable colorectal liver metastases treated by surgery alone or surgery with perioperative FOLFOX in the randomised EORTC study 40983. Eur. J. Cancer 51, 2708–2717 (2015).
Pugh, S. A., Harrison, R. J., Primrose, J. N. & Khakoo, S. I. T cells but not NK cells are associated with a favourable outcome for resected colorectal liver metastases. BMC Cancer 14, 180 (2014).
Halama, N. et al. The localization and density of immune cells in primary tumors of human metastatic colorectal cancer shows an association with response to chemotherapy. Cancer Immun. 9, 1 (2009).
Halama, N. et al. Hepatic metastases of colorectal cancer are rather homogeneous but differ from primary lesions in terms of immune cell infiltration. Oncoimmunology 2, e24116 (2013).
Tran, B. et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer 117, 4623–4632 (2011).
Kim, C. G. et al. Effects of microsatellite instability on recurrence patterns and outcomes in colorectal cancers. Br. J. Cancer 115, 25–33 (2016).
Heinemann, V. et al. Somatic DNA mutations, tumor mutational burden (TMB), and MSI status: association with efficacy in patients (pts) with metastatic colorectal cancer (mCRC) of FIRE-3 (AIO KRK-0306). J. Clin. Oncol. 36, 3591 (2018).
Margonis, G. A. et al. Microsatellite instability in resectable colorectal liver metastasis: an international multi-institutional analysis. J. Clin. Oncol. 36, 220 (2018).
Price, T. J. et al. Outcomes for metastatic colorectal cancer (mCRC) based on microsatellite instability. J. Clin. Oncol. 36, 759 (2018).
Jin, Z. et al. Outcome of mismatch repair-deficient metastatic colorectal cancer: the Mayo Clinic experience. Oncologist 23, 1083–1091 (2018).
Sjo, O. H. et al. Peritoneal carcinomatosis of colon cancer origin: highest incidence in women and in patients with right-sided tumors. J. Surg. Oncol. 104, 792–797 (2011).
Sorbye, H. et al. High BRAF mutation frequency and marked survival differences in subgroups according to KRAS/BRAF mutation status and tumor tissue availability in a prospective population-based metastatic colorectal cancer cohort. PLOS ONE 10, e0131046 (2015).
Goldstein, J. et al. Multicenter retrospective analysis of metastatic colorectal cancer (CRC) with high-level microsatellite instability (MSI-H). Ann. Oncol. 25, (1032–1038 (2014).
Venook, A. P. et al. Impact of primary (1°) tumor location on overall survival (OS) and progression-free survival (PFS) in patients (pts) with metastatic colorectal cancer (mCRC): analysis of CALGB/SWOG 80405 (Alliance). J. Clin. Oncol. 34, 3504–3504 (2016).
Holch, J. W., Ricard, I., Stintzing, S., Modest, D. P. & Heinemann, V. The relevance of primary tumor location in patients with metastatic colorectal cancer: a meta-analysis of first-line clinical trials. Eur. J. Cancer 70, 87–98 (2017).
Loupakis, F. et al. Primary tumor location as a prognostic factor in metastatic colorectal cancer. J. Natl Cancer Inst. 107, dju427 (2015).
Cremolini, C. et al. Primary tumor sidedness and benefit from FOLFOXIRI plus bevacizumab as initial therapy for metastatic colorectal cancer. Retrospective analysis of the TRIBE trial by GONO. Ann. Oncol. 29, 1528–1534 (2018).
Loree, J. et al. Classifying colorectal cancer by tumor location rather than sidedness highlights a continuum in mutation profiles and consensus molecular subtypes. Clin. Cancer Res. 24, 1062–1072 (2018).
Steinert, G. et al. Immune escape and survival mechanisms in circulating tumor cells of colorectal cancer. Cancer Res. 74, 1694–11704 (2014).
US Food & Drug Administration. FDA approves first cancer treatment for any solid tumor with a specific genetic feature. FDA.gov https://www.fda.gov/newsevents/newsroom/pressannouncements/ucm560167.htm (2017).
Le, D. T. et al. Mismatch-repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02563002 (2018).
Diaz, L. A. et al. Phase 3, open-label, randomized study of first-line pembrolizumab (pembro) versus investigator-choice chemotherapy for mismatch repair-deficient (dMMR) or microsatellite instability-high (MSI-H) metastatic colorectal carcinoma (mCRC): KEYNOTE-177. J. Clin. Oncol. 35 (Suppl. 4), TPS3618 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02997228 (2019).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02912559 (2019).
Overman, M. J. et al. Durable clinical benefit with nivolumab plus ipilimumab in DNA mismatch repair-deficient/microsatellite instability-high metastatic colorectal cancer. J. Clin. Oncol. 36, 773–779 (2018). Combination immune checkpoint inhibition with nivolumab plus ipilimumab enabled a high response rate in patients with MSI-H/dMMR metastatic CRCs (also in those with BRAF V600E mutations), and an indirect comparison of different trial cohorts suggests that this combination might improve the PFS and OS compared with single-agent ICI.
Lenz, H. J. J. et al. Durable clinical benefit with nivolumab (NIVO) plus low-dose ipilimumab (IPI) as first-line therapy in microsatellite instability-high/mismatch repair deficient (MSI-H/dMMR) metastatic colorectal cancer (mCRC) [abstract LBA18_PR]. Ann. Oncol. 29, mdy424.019 (2018).
Chalabi, M. et al. Neoadjuvant ipilimumab plus nivolumab in early stage colon cancer [abstract LBA37_PR]. Ann. Oncol. 29, mdy424.047 (2018).
Chen, E. X. et al. The CCTG CO.26 trial: a phase II randomized study of durvalumab plustremelimumab and best supportive care (BSC) versus BSC alone in patients with advanced colorectal carcinoma (CRC) refractory to standard therapies. J. Clin. Oncol. 35, (TPS3621 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02870920 (2019).
Giannakis, M. et al. Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep. 15, 857–865 (2016).
van Rooij, N. et al. Tumor exome analysis reveals neoantigen-specific T cell reactivity in an ipilimumab-responsive melanoma. J. Clin. Oncol. 31, e439–e442 (2013).
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Llosa, N. J. et al. The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov. 5, 43–51 (2015).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Domingo, E. et al. Somatic POLE proofreading domain mutation, immune response, and prognosis in colorectal cancer: a retrospective, pooled biomarker study. Lancet Gastroenterol. Hepatol. 1, 207–216 (2016).
Fabrizio, D. A. et al. Beyond microsatellite testing: assessment of tumor mutational burden identifies subsets of colorectal cancer who may respond to immune checkpoint inhibition. J. Gastrointest. Oncol. 9, 610–617 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03150706 (2019).
Guerra, J. et al. POLE somatic mutations in advanced colorectal cancer. Cancer Med. 6, 2966–2971 (2017).
Chan, T. A. et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann. Oncol. 30, 44–56 (2018).
Goodman, A. M. et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Ther. 16, 2598–2608 (2017).
Kirilovsky, A. et al. Rational bases for the use of the Immunoscore in routine clinical settings as a prognostic and predictive biomarker in cancer patients. Int. Immunol. 28, 373–382 (2016).
Galon, J. et al. MSI status plus immunoscore to select metastatic colorectal cancer patients for immunotherapies [abstract 12P]. Ann. Oncol. 29, mdy493.011 (2018).
Sharma, P., Hu-Lieskovan, S., Wargo, J. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Shin, D. S. et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Discov. 7, 188–201 (2017). Homozygous loss-of-function mutation in JAK1 identified as a likely mechanism of primary resistance to ICI in a patient with metastatic colon cancer with a high TMB and no response to pembrolizumab.
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Zhou, G. et al. Blockade of LAG3 enhances responses of tumor-infiltrating T cells in mismatch repair-proficient liver metastases of colorectal cancer. Oncoimmunology 7, e1448332 (2018).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
Skoulidis, F. et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov. 8, 822–835 (2018).
Chen, P. L. et al. Analysis of immune signatures in longitudinal tumor samples yields insight into biomarkers of response and mechanisms of resistance to immune checkpoint blockade. Cancer Discov. 6, 827–837 (2016).
Hamilton, S. R. BRAF mutation and microsatellite instability status in colonic and rectal carcinoma: context really does matter. J. Natl Cancer Inst. 105, 1075–1077 (2013).
Rajagopalan, H. et al. Tumorigenesis: RAF/RAS oncogenes and mismatch-repair status. Nature 418, 934 (2002).
Lochhead, P. et al. Microsatellite instability and BRAF mutation testing in colorectal cancer prognostication. J. Natl Cancer Inst. 105, 1151–1156 (2013).
Samowitz, W. S. et al. Poor survival associated with the BRAF V600E mutation in microsatellite-stable colon cancers. Cancer Res. 65, 6063–6069 (2005).
Gonsalves, W. I. et al. Patient and tumor characteristics and BRAF and KRAS mutations in colon cancer, NCCTG/Alliance N0147. J. Natl Cancer Inst. 106, dju106 (2014).
Clarke, C. N. & Kopetz, E. S. BRAF mutant colorectal cancer as a distinct subset of colorectal cancer: clinical characteristics, clinical behaviour, and response to targeted therapies. J. Gastrointest. Oncol. 6, 660–667 (2015).
Vedeld, H. M. et al. CpG island methylator phenotype identifies high risk patients among microsatellite stable BRAF mutated colorectal cancers. Int. J. Cancer 141, 967–976 (2017).
Tie, J. et al. Optimizing targeted therapeutic development: analysis of a colorectal cancer population with the BRAF V600E mutation. Int. J. Cancer 128, 2075–2084 (2011).
Roth, A. D. et al. Prognostic role of KRAS and BRAF in stage II and III resected colon cancer: results of the translational study on the PETACC-3, EORTC 40993, SAKK 60–00 trial. J. Clin. Oncol. 28, 466–474 (2010).
Taieb, J. et al. Prognostic effect of BRAF and KRAS mutation in patients with stage III colon cancer treated with leucovorin, fluorouracil, and oxaliplatin with or without cetuximab: a post hoc analysis of the PETACC-8 trial. JAMA Oncol. 2, 643–653 (2016). Retrospective analyses of MSI status, BRAF V600E and KRAS mutations in patients with stage III colon cancer included in the PETACC-8 clinical trial showing that mutations in both genes are independently associated with inferior outcomes in MSS cancers, suggesting that these markers need to be used for patient stratification in future trials involving adjuvant treatments.
Seppala, T. T. et al. Combination of microsatellite instability and BRAF mutation status for subtyping colorectal cancer. Br. J. Cancer 112, 1966–1975 (2015).
Smeby, J. et al. CMS-dependent prognostic impact of KRAS and BRAF V600E mutations in primary colorectal cancer. Ann. Oncol. 29, 1227–1234 (2018).
Vogelstein, B. et al. Genetic alterations during colorectal-tumor development. N. Engl. J. Med. 319, 525–532 (1988).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Fearon, E. R. Molecular genetics of colorectal cancer. Annu. Rev. Pathol. 6, 479–507 (2011).
Kambara, T. et al. BRAF mutation is associated with DNA methylation in serrated polyps and cancers of the colorectum. Gut 53, 1137–1144 (2004).
Rad, R. et al. A genetic progression model of BRAF V600E-induced intestinal tumorigenesis reveals targets for therapeutic intervention. Cancer Cell 24, 15–29 (2013).
Seligmann, J. F. et al. Investigating the poor outcomes of BRAF-mutant advanced colorectal cancer: analysis from 2530 patients in randomised clinical trials. Ann. Oncol. 28, 562–568 (2017).
Cremolini, C. et al. FOLFOXIRI plus bevacizumab versus FOLFIRI plus bevacizumab as first-line treatment of patients with metastatic colorectal cancer: updated overall survival and molecular subgroup analyses of the open-label, phase 3 TRIBE study. Lancet Oncol. 16, 1306–1315 (2015).
Yaeger, R. et al. BRAF mutation predicts for poor outcomes after metastasectomy in patients with metastatic colorectal cancer. Cancer 120, 2316–2324 (2014).
Guren, T. K. et al. Cetuximab in treatment of metastatic colorectal cancer: final survival analyses and extended RAS data from the NORDIC-VII study. Br. J. Cancer 116, 1271–1278 (2017).
Tol, J., Nagtegaal, I. D. & Punt, C. J. BRAF mutation in metastatic colorectal cancer. N. Eng. J. Med. 361, 98–99 (2009).
Renaud, S. et al. KRAS and BRAF mutations are prognostic biomarkers in patients undergoing lung metastasectomy of colorectal cancer. Br. J. Cancer 112, 720–728 (2015).
Schirripa, M. et al. BRAF and RAS mutations as prognostic factors in metastatic colorectal cancer patients undergoing liver resection. Br. J. Cancer 112, 1921–1928 (2015).
Margonis, G. A. et al. Association of BRAF mutations with survival and recurrence in surgically treated patients with metastatic colorectal liver cancer. JAMA Surg. 153, e180996 (2018).
Sanz-Garcia, E., Argiles, G., Elez, E. & Tabernero, J. BRAF mutant colorectal cancer: prognosis, treatment, and new perspectives. Ann. Oncol. 28, 2648–2657 (2017).
Morris, V. K. et al. Progression-free survival remains poor over sequential lines of systemic therapy in patients with BRAF-mutated colorectal cancer. Clin. Colorectal Cancer 13, 164–171 (2014).
Richman, S. D. et al. KRAS and BRAF mutations in advanced colorectal cancer are associated with poor prognosis but do not preclude benefit from oxaliplatin or irinotecan: results from the MRC FOCUS trial. J. Clin. Oncol. 27, 5931–5937 (2009).
Loupakis, F. et al. FOLFOXIRI plus bevacizumab as first-line treatment in BRAF mutant metastatic colorectal cancer. Eur. J. Cancer 50, 57–63 (2014).
di Nicolantonio, F. et al. Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J. Clin. Oncol. 26, 5705–5712 (2008).
Loupakis, F. et al. KRAS codon 61, 146, and BRAF mutations predict resistance to cetuximab plus irinotecan in KRAS codon 12 and 13 wild-type metastatic colorectal cancer. Br. J. Cancer 101, 715–721 (2009).
Maughan, T. S. et al. Addition of cetuximab to oxaliplatin-based first-line combination chemotherapy for treatment of advanced colorectal cancer: results of the randomised phase 3 MRC COIN trial. Lancet 377, 2103–2114 (2011).
Tveit, K. M. et al. Phase III trial of cetuximab with continuous or intermittent fluorouracil, leucovorin, and oxaliplatin (Nordic FLOX) versus FLOX alone in first-line treatment of metastatic colorectal cancer: the NORDIC-VII study. J. Clin. Oncol. 30, 1755–1762 (2012).
Bokemeyer, C. et al. Addition of cetuximab to chemotherapy as first-line treatment for KRAS wild-type metastatic colorectal cancer: pooled analysis of the CRYSTAL and OPUS randomised clinical trials. Eur. J. Cancer 48, 1466–1475 (2012).
Douillard, J. Y. et al. Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N. Eng. J. Med. 369, 1023–1034 (2013).
Stintzing, S. et al. Mutations within the EGFR signaling pathway: influence on efficacy in FIRE-3 — a randomized phase III study of FOLFIRI plus cetuximab or bevacizumab as first-line treatment for wild-type (WT) KRAS (exon 2) metastatic colorectal cancer (mCRC) patients. J. Clin. Oncol. 32, 445 (2014).
Karapetis, C. S. et al. PIK3CA, BRAF, and PTEN status and benefit from cetuximab in the treatment of advanced colorectal cancer — results from NCIC CTG/AGITG CO.17. Clin. Cancer Res. 20, 744–753 (2014).
Peeters, M. et al. Massively parallel tumor multigene sequencing to evaluate response to panitumumab in a randomized phase III study of metastatic colorectal cancer. Clin. Cancer Res. 19, 1902–1912 (2013).
Peeters, M. et al. Updated analysis of KRAS/NRAS and BRAF mutations in study 20050181 of panitumumab (pmab) plus FOLFIRI for second-line treatment (tx) of metastatic colorectal cancer (mCRC). J. Clin. Oncol. 32, 3568 (2014).
Seymour, M. et al. Panitumumab and irinotecan versus irinotecan alone for patients with KRAS wild-type, fluorouracil-resistant advanced colorectal cancer (PICCOLO): a prospectively stratified randomised trial. Lancet Oncol. 14, 749–759 (2013).
Pietrantonio, F. et al. Predictive role of BRAF mutations in patients with advanced colorectal cancer receiving cetuximab and panitumumab: a meta-analysis. Eur. J. Cancer 51, 587–594 (2015).
Rowland, A. et al. Meta-analysis of BRAF mutation as a predictive biomarker of benefit from anti-EGFR monoclonal antibody therapy for RAS wild-type metastatic colorectal cancer. Br. J. Cancer 112, 1888–1894 (2015).
Kopetz, S. et al. Phase II pilot study of vemurafenib in patients with metastatic BRAF-mutated colorectal cancer. J. Clin. Oncol. 33, 4032–4038 (2015).
Hyman, D. M. et al. Vermurafenib in multiple nonmelanoma cancers with BRAF V600 mutations. N. Eng. J. Med. 373, 726–736 (2015).
Prahallad, A. et al. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 483, 100–103 (2012).
Corcoran, R. B. et al. EGFR-mediated re-activation of MAPK signaling contributes to insensitivity of BRAF mutant colorectal cancers to RAF inhibition with vemurafenib. Cancer Discov. 2, 227–235 (2012).
Mao, M. et al. Resistance to BRAF inhibition in BRAF-mutant colon cancer can be overcome with PI3K inhibition or demethylating agents. Clin. Cancer Res. 19, 657–667 (2013).
Yaeger, R. et al. Pilot trial of combined BRAF and EGFR inhibition in BRAF-mutant metastatic colorectal cancer patients. Clin. Cancer Res. 21, 1313–1320 (2015).
Van Cutsem, E. et al. Updated results of the MEK inhibitor trametinib (T), BRAF inhibitor dabrafenib (D), and anti-EGFR antibody panitumumab (P) in patients (pts) with BRAF V600E mutated (BRAFm) metastatic colorectal cancer (mCRC) [abstract LBA-07]. Ann. Oncol. 26, iv119 (2015).
van Geel, R. M. J. M. et al. A phase Ib dose-escalation study of encorafenib and cetuximab with or without alpelisib in metastatic BRAF-mutant colorectal cancer. Cancer Discov. 7, 610–619 (2017).
Tabernero, J. et al. Phase 2 results: encorafenib (enco) and cetuximab (cetux) with or without alpelisib (alp) in patients with advanced BRAF-mutant colorectal cancer (BRAFm CRC). J. Clin. Oncol. 34, 3544 (2016).
Corcoran, R. B. et al. Combined BRAF and MEK inhibition with dabrafenib and trametinib in BRAF V600-mutant colorectal cancer. J. Clin. Oncol. 33, 4023–4031 (2015).
Flaherty, K. T. et al. Combined BRAF and MEK inhibition in melanoma with BRAF V600 mutations. N. Eng. J. Med. 367, 1694–1703 (2012).
Corcoran, R. B. et al. Combined BRAF, EGFR, and MEK inhibition in patients with BRAF V600E-mutant colorectal cancer. Cancer Discov. 8, 428–443 (2018). Prospective phase I study demonstrating improved initial response rates to targeted combination therapy with dabrafenib plus panitumumab in patients with BRAF V600E-mutant metastatic CRC by adding the MEK inhibitor trametinib.
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01750918 (2019).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02928224 (2019).
Van Cutsem, E. et al. BEACON CRC study safety lead-in: assessment of the BRAF inhibitor encorafenib + MEK inhibitor binimetinib + anti-epidermal growth factor receptor antibody cetuximab for BRAF V600E metastatic colorectal cancer [abstract O-027]. Ann. Oncol. 29, mdy149.026 (2018).
Popovici, V. et al. Identification of a poor-prognosis BRAF-mutant-like population of patients with colon cancer. J. Clin. Oncol. 30, 1288–1295 (2012).
Vecchione, L. et al. A vulnerability of a subset of colon cancers with potential clinical utility. Cell 165, 317–330 (2016).
Cremolini, C. et al. Vinorelbine in BRAF V600E mutated metastatic colorectal cancer: a prospective multicentre phase II clinical study. ESMO Open 2, e000241 (2017).
Barras, D. et al. BRAF V600E mutant colorectal cancer subtypes based on gene expression. Clin. Cancer Res. 23, 104–115 (2017).
Pietrantonio, F. et al. MET-driven resistance to dual EGFR and BRAF blockade may be overcome by switching from EGFR to MET inhibition in BRAF-mutated colorectal cancer. Cancer Discov. 6, 963–971 (2016).
Oddo, D. et al. Emergence of MET hyper-amplification at progression to MET and BRAF inhibition in colorectal cancer. Br. J. Cancer 117, 347–352 (2017).
Hazar-Rethinam, M. et al. Convergent therapeutic strategies to overcome the heterogeneity of acquired resistance in BRAF V600E colorectal cancer. Cancer Discov. 8, 417–427 (2018). Analysis of liquid biopsy samples from patients treated with BRAF V600E-targeted combination therapies showing that the development of acquired resistance is driven by multiple alterations in the MAPK pathway and clonal outgrowth in preclinical models is abrogated by ERK inhibition.
Jones, J. C. et al. Non-V600 BRAF mutations define a clinically distinct molecular subtype of metastatic colorectal cancer. J. Clin. Oncol. 35, 2624–2630 (2017).
Dankner, M. Targeted therapy for colorectal cancers with non-V600 BRAF mutations: perspectives for precision oncology. JCO Precis. Oncol. https://doi.org/10.1200/PO.18.00195 (2018).
Wan, P. T. et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004).
Cremolini, C. et al. BRAF codons 594 and 596 mutations identify a new molecular subtype of metastatic colorectal cancer at favorable prognosis. Ann. Oncol. 26, 2092–2097 (2015).
Schirripa, M. et al. Clinico-pathological and molecular characterisation of BRAF mutant metastatic colorectal cancer (mCRC): are all mutations created equal? J. Clin. Oncol. 36, 3590 (2018).
Shimada, Y. et al. Clinical significance of BRAF non-V600E mutations in colorectal cancer: a retrospective study of two institutions. J. Surg. Res. 232, 72–81 (2018).
Shinozaki, E. et al. Clinical significance of BRAF non-V600E mutations on the therapeutic effects of anti-EGFR monoclonal antibody treatment in patients with pretreated metastatic colorectal cancer: the Biomarker Research for anti-EGFR monoclonal Antibodies by Comprehensive Cancer genomics (BREAC) study. Br. J. Cancer 117, 1450–1458 (2017).
De Roock, W. et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol. 11, 753–762 (2010).
Ross, J. S. et al. The HER-2 receptor and breast cancer: ten years of targeted anti-HER-2 therapy and personalized medicine. Oncologist 14, 320–368 (2009).
Richman, S. D. et al. HER2 overexpression and amplification as a potential therapeutic target in colorectal cancer: analysis of 3256 patients enrolled in the QUASAR, FOCUS and PICCOLO colorectal cancer trials. J. Pathol. 238, 562–570 (2016).
Siena, S. et al. Targeting the human epidermal growth factor receptor 2 (HER2) oncogene in colorectal cancer. Ann. Oncol. 29, 1108–1119 (2018).
Bertotti, A. et al. A molecularly annotated platform of patient-derived xenografts (“xenopatients”) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov. 1, 508–523 (2011).
Yonesaka, K. et al. Activation of ERBB2 signaling causes resistance to the EGFR-directed therapeutic antibody cetuximab. Sci. Transl Med. 3, 99ra86 (2011).
Valtorta, E. et al. Assessment of a HER2 scoring system for colorectal cancer: results from a validation study. Mod. Pathol. 28, 1481–1491 (2015).
Sartore-Bianchi, A. et al. Dual-targeted therapy with trastuzumab and lapatinib in treatment-refractory, KRAS codon 12/13 wild-type, HER2-positive metastatic colorectal cancer (HERACLES): a proof-of-concept, multicentre, open-label, phase 2 trial. Lancet Oncol. 17, 738–746 (2016). Prospective trial showing an ORR of 30% in patients with HER2-positive KRAS wild-type treatment refractory metastatic CRCs receiving dual HER2 inhibition with trastuzumab plus lapatinib.
Martin, V. et al. HER2 gene copy number status may influence clinical efficacy to anti-EGFR monoclonal antibodies in metastatic colorectal cancer patients. Br. J. Cancer 108, 668–675 (2013).
Jeong, J. H. et al. HER2 amplification and cetuximab efficacy in patients with metastatic colorectal cancer harboring wild-type RAS and BRAF. Clin. Colorectal Cancer 16, e147–e152 (2017).
Raghav, K. et al. Validation of HER2 amplification as a predictive biomarker for anti-epidermal growth factor receptor antibody therapy in metastatic colorectal cancer. JCO Precis. Oncol. https://doi.org/10.1200/PO.18.00226 (2019).
Clark, J. W., Niedzwiecki, D., Hollis, D. & Mayer, R. Phase-II trial of 5-fluororuacil (5-FU), leucovorin (LV), oxaliplatin (Ox), and trastuzumab (T) for patients with metastatic colorectal cancer (CRC) refractory to initial therapy. Onkologie 26, (46 (2003).
Ramanathan, R. K. et al. Low overexpression of HER-2/neu in advanced colorectal cancer limits the usefulness of trastuzumab (Herceptin) and irinotecan as therapy. A phase II trial. Cancer Invest. 22, 858–865 (2004).
Hainsworth, J. D. et al. Targeted therapy for advanced solid tumors on the basis of molecular profiles: results from MyPathway, an open-label, phase IIa multiple basket study. J. Clin. Oncol. 36, 536–542 (2018).
Gluck, W. L. et al. Prolonged response of widely metastatic HER2-positive colon cancer to trastuzumab therapy. Colorectal Cancer 6, 57–61 (2017).
Ross, J. S. et al. Targeting HER2 in colorectal cancer: the landscape of amplification and short variant mutations in ERBB2 and ERBB3. Cancer 124, 1358–1373 (2018).
Martinelli, E. et al. Sequential HER2 blockade as effective therapy in chemorefractory, HER2 gene-amplified, RAS wild-type, metastatic colorectal cancer: learning from a clinical case. ESMO Open 3, e000299 (2018).
Siravegna, G. et al. Radiologic and genomic evolution of individual metastases during HER2 blockade in colorectal cancer. Cancer Cell 34, 148–162 (2018). Analysis of liquid biopsy samples from patients with HER2-positive metastatic CRC during treatment with HER2-targeted therapy identified an association between emerging alterations in ERBB2, RAS and PIK3CA and acquired resistance.
Haslem, D. S., Ji, H. P., Ford, J. M. & Nadauld, L. D. Precision oncology strategy in trastuzumab-resistant human epidermal growth factor receptor 2-positive colon cancer: case report of durable response to ado-trastuzumab emtansine. JCO Precis. Oncol. https://doi.org/10.1200/PO.16.00055 (2017).
Parikh, A., Atreya, C., Korn, W. M. & Venook, A. P. Prolonged response to HER2-directed therapy in a patient with HER2-amplified, rapidly progressive metastatic colorectal cancer. J. Natl Compr. Canc. Netw. 15, 3–8 (2017).
Siena, S. et al. HER2 amplification as a ‘molecular bait’ for trastuzumab-emtansine (T-DM1) precision chemotherapy to overcome anti-HER2 resistance in HER2 positive metastatic colorectal cancer: the HERACLES-RESCUE trial. J. Clin. Oncol. 34 (Suppl. 4), TPS774 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02564900 (2019).
Yoshino, T. et al. Updated results of phase I study of trastuzumab deruxtecan (DS-8201a) in HER2-expressing advanced colorectal cancer [abstract 563P]. Ann. Oncol. 29, mdy281.109 (2018).
Loree, J. M. et al. Molecular landscape of ERBB2/ERBB3 mutated colorectal cancer. J. Natl Cancer Inst. 110, 1409–1417 (2018).
Bertotti, A. et al. The genomic landscape of response to EGFR blockade in colorectal cancer. Nature 526, 263–267 (2015).
Kavuri, S. M. et al. HER2 activating mutations are targets for colorectal cancer treatment. Cancer Discov. 5, 832–841 (2015).
Hyman, D. M. et al. HER kinase inhibition in patients with HER2- and HER3-mutant cancers. Nature 554, 189–194 (2018).
US Food & Drug Association. FDA approves larotrectinib for solid tumors with NTRK gene fusions. FDA.gov https://www.fda.gov/drugs/fda-approves-larotrectinib-solid-tumors-ntrk-gene-fusions-0 (2018).
Stransky, N., Cerami, E., Schalm, S., Kim, J. L. & Lengauer, C. The landscape of kinase fusions in cancer. Nat. Commun. 5, 4846 (2014).
Madison, R. et al. Kinase fusions in colorectal cancers: a unique biologic subset [abstract 457PD]. Ann. Oncol. 29, viii152 (2018).
Pietrantonio, F. et al. ALK, ROS1, and NTRK rearrangements in metastatic colorectal cancer. J. Natl Cancer Inst. 109, djx089 (2017).
Pietrantonio, F. et al. RET fusions in a small subset of advanced colorectal cancers at risk of being neglected. Ann. Oncol. 29, 1394–1401 (2018). A case series of patients with metastatic CRCs harbouring RET rearrangements shows enrichment with MSI-H, RAS/BRAF wild-type tumours and inferior OS compared with RET-negative cancers.
Medico, E. et al. The molecular landscape of colorectal cancer cell lines unveils clinically actionable kinase targets. Nat. Commun. 6, 7002 (2015).
Cremolini, C. et al. Negative hyper-selection of metastatic colorectal cancer patients for anti-EGFR monoclonal antibodies: the PRESSING case-control study. Ann. Oncol. 28, 3009–3014 (2017).
Lee, S. J. et al. NTRK1 rearrangement in colorectal cancer patients: evidence for actionable target using patient-derived tumor cell line. Oncotarget 6, 39028–39035 (2015).
Drilon, A. et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N. Engl. J. Med. 378, 731–739 (2018). Prospective basket trial showing an ORR of 75% to the selective TRK inhibitor larotrectinib in adults and children with treatment-refractory TRK fusion-positive cancers, including disease regression in three of four patients with colon cancer.
Amatu, A. et al. Novel CAD-ALK gene rearrangement is drugable by entrectinib in colorectal cancer. Br. J. Cancer 113, 1730–1734 (2015).
Sartore-Bianchi, A. et al. Sensitivity to entrectinib associated with a novel LMNA-NTRK1 gene fusion in metastatic colorectal cancer. J. Natl Cancer Inst. 108, djv306 (2016).
Demetri, G. D. et al. Efficacy and safety of entrectinib in patients with NTRK fusion-positive (NTRK-fp) tumors: pooled analysis of STARTRK-2, STARTRK-1 and ALKA-372-001 [abstract LBA17]. Ann. Oncol. 29, mdy424.017 (2018).
Yakirevich, E. et al. Oncogenic ALK fusion in rare and aggressive subtype of colorectal adenocarcinoma as a potential therapeutic target. Clin. Cancer Res. 22, 3831–3840 (2016).
Santos, C., Sanz-Pamplona, R. & Salazar, R. RET-fusions: a novel paradigm in colorectal cancer. Ann. Oncol. 29, 1340–1343 (2018).
Yaeger, R. et al. Clinical sequencing defines the genomic landscape of metastatic colorectal cancer. Cancer Cell 33, 125–136 (2018).
Shaw, A. T. et al. Avelumab (anti-PD-L1) in combination with crizotinib or lorlatinib in patients with previously treated advanced NSCLC: phase 1b results from JAVELIN Lung 101. J. Clin. Oncol. 36, 9008 (2018).
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012).
The Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Shinmura, K. et al. RSPO fusion transcripts in colorectal cancer in Japanese population. Mol. Biol. Rep. 41, 5375–5384 (2014).
de Lau, W., Peng, W. C., Gros, P. & Clevers, H. The R-spondin/Lgr5/Rnf43 module: regulator of Wnt signal strength. Genes Dev. 28, 305–316 (2014).
Giannakis, M. et al. RNF43 is frequently mutated in colorectal and endometrial cancers. Nat. Genet. 46, 1264–1266 (2014).
Kahn, M. Can we safely target the WNT pathway? Nat. Rev. Drug. Discov. 13, 513–532 (2014).
Fevr, T., Robine, S., Louvard, D. & Huelsken, J. Wnt/β-catenin is essential for intestinal homeostasis and maintenance of intestinal stem cells. Mol. Cell. Biol. 27, 7551–7559 (2007).
Reya, T. et al. A role for Wnt signalling in self-renewal of haematopoietic stem cells. Nature 423, 409–414 (2003).
Madan, B. et al. Wnt addiction of genetically defined cancers reversed by PORCN inhibition. Oncogene 35, 2197–2207 (2016).
Li, C. et al. Identification of RSPO2 fusion mutations and target therapy using a porcupine inhibitor. Sci. Rep. 8, 14244 (2018).
Storm, E. E. et al. Targeting PTPRK-RSPO3 colon tumours promotes differentiation and loss of stem-cell function. Nature 529, 97–100 (2016).
Janku, F. et al. Phase I study of WNT974, a first-in-class Porcupine inhibitor, in advanced solid tumors. Mol. Cancer Ther. 14, C45 (2015).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02278133 (2017).
Rodon, J. et al. Biomarker analyses from a phase I study of WNT974, a first-in-class Porcupine inhibitor, in patients (pts) with advanced solid tumors. Cancer Res. 78, CT175 (2018).
Grasso, C. S. et al. Genetic mechanisms of immune evasion in colorectal cancer. Cancer Discov. 8, 730–749 (2018).
Taieb, J. et al. Prognostic value of BRAF and KRAS mutations in MSI and MSS stage III colon cancer. J. Natl Cancer Inst. 109, djw272 (2017).
Hu, J. et al. Coexistence of MSI with KRAS mutation is associated with worse prognosis in colorectal cancer. Medicine 95, e5649 (2016).
Siraj, A. K. et al. A very low incidence of BRAF mutations in Middle Eastern colorectal carcinoma. Mol. Cancer 13, 168 (2014).
Lin, C. C. et al. The prognostic role of microsatellite instability, codon-specific KRAS, and BRAF mutations in colon cancer. J. Surg. Oncol. 110, 451–457 (2014).
Imamura, Y. et al. Analyses of clinicopathological, molecular, and prognostic associations of KRAS codon 61 and codon 146 mutations in colorectal cancer: cohort study and literature review. Mol. Cancer 13, 135 (2014).
Ogura, T. et al. Clinicopathological characteristics and prognostic impact of colorectal cancers with NRAS mutations. Oncol. Rep. 32, 50–56 (2014).
Wangefjord, S. et al. Sex differences in the prognostic significance of KRAS codons 12 and 13, and BRAF mutations in colorectal cancer: a cohort study. Biol. Sex. Differ. 4, 17 (2013).
Samadder, N. J. et al. Associations between colorectal cancer molecular markers and pathways with clinicopathologic features in older women. Gastroenterology 145, 348–356 (2013).
Eklof, V. et al. The prognostic role of KRAS, BRAF, PIK3CA and PTEN in colorectal cancer. Br. J. Cancer 108, 2153–2163 (2013).
Phipps, A. I. et al. KRAS-mutation status in relation to colorectal cancer survival: the joint impact of correlated tumour markers. Br. J. Cancer 108, 1757–1764 (2013).
Nash, G. M. et al. KRAS mutation and microsatellite instability: two genetic markers of early tumor development that influence the prognosis of colorectal cancer. Ann. Surg. Oncol. 17, 416–424 (2010).
Samowitz, W. S. et al. Microsatellite instability and survival in rectal cancer. Cancer Causes Control 20, 1763–1768 (2009).
Barault, L. et al. Hypermethylator phenotype in sporadic colon cancer: study on a population-based series of 582 cases. Cancer Res. 68, 8541–8546 (2008).
Andreyev, H. J. et al. Kirsten ras mutations in patients with colorectal cancer: the ‘RASCAL II’ study. Br. J. Cancer 85, 692–696 (2001).
Kadowaki, S. et al. Prognostic value of KRAS and BRAF mutations in curatively resected colorectal cancer. World J. Gastroenterol. 21, 1275–1283 (2015).
Sinicrope, F. A. et al. Association of DNA mismatch repair and mutations in BRAF and KRAS with survival after recurrence in stage III colon cancers: a secondary analysis of 2 randomized clinical trials. JAMA Oncol. 3, 472–480 (2017).
Ogino, S. et al. KRAS mutation in stage III colon cancer and clinical outcome following intergroup trial CALGB 89803. Clin. Cancer Res. 15, 7322–7329 (2009).
Popovici, V. et al. Context-dependent interpretation of the prognostic value of BRAF and KRAS mutations in colorectal cancer. BMC Cancer 13, 439 (2013).
Mouradov, D. et al. Survival in stage II/III colorectal cancer is independently predicted by chromosomal and microsatellite instability, but not by specific driver mutations. Am. J. Gastroenterol. 108, 1785–1793 (2013).
Blons, H. et al. Prognostic value of KRAS mutations in stage III colon cancer: post hoc analysis of the PETACC8 phase III trial dataset. Ann. Oncol. 25, 2378–2385 (2014).
Sinicrope, F. A. et al. Analysis of molecular markers by anatomic tumor site in stage III colon carcinomas from adjuvant chemotherapy trial NCCTG N0147 (Alliance). Clin. Cancer Res. 21, 5294–5304 (2015).
Okuno, M. et al. RAS mutation is associated with unsalvageable recurrence following hepatectomy for colorectal cancer liver metastases. Ann. Surg. Oncol. 25, 2457–2466 (2018).
Cercek, A. et al. Clinical features and outcomes of patients with colorectal cancers harboring NRAS mutations. Clin. Cancer Res. 23, 4753–4760 (2017).
Dienstmann, R. et al. Analysis of mutant allele fractions in driver genes in colorectal cancer — biological and clinical insights. Mol. Oncol. 11, 1263–1272 (2017).
Passot, G. et al. Is hepatectomy justified for patients with RAS mutant colorectal liver metastases? An analysis of 524 patients undergoing curative liver resection. Surgery 161, 332–340 (2017).
Summers, M. G. et al. BRAF and NRAS locus-specific variants have different outcomes on survival to colorectal cancer. Clin. Cancer Res. 23, 2742–2749 (2017).
Adenis, A. et al. Survival, safety, and prognostic factors for outcome with Regorafenib in patients with metastatic colorectal cancer refractory to standard therapies: results from a multicenter study (REBECCA) nested within a compassionate use program. BMC Cancer 16, 412 (2016).
Margonis, G. A. et al. Codon 13 KRAS mutation predicts patterns of recurrence in patients undergoing hepatectomy for colorectal liver metastases. Cancer 122, 2698–2707 (2016).
Modest, D. P. et al. Outcome according to KRAS-, NRAS- and BRAF-mutation as well as KRAS mutation variants: pooled analysis of five randomized trials in metastatic colorectal cancer by the AIO colorectal cancer study group. Ann. Oncol. 27, 1746–1753 (2016).
Schirripa, M. et al. Role of NRAS mutations as prognostic and predictive markers in metastatic colorectal cancer. Int. J. Cancer 136, 83–90 (2015).
Petrelli, F., Coinu, A., Cabiddu, M., Ghilardi, M. & Barni, S. KRAS as prognostic biomarker in metastatic colorectal cancer patients treated with bevacizumab: a pooled analysis of 12 published trials. Med. Oncol. 30, 650 (2013).
Smith, J. C. et al. KRAS mutations are associated with inferior clinical outcome in patients with metastatic colorectal cancer, but are not predictive for benefit with cediranib. Eur. J. Cancer 49, 2424–2432 (2013).
Tejpar, S. et al. Association of KRAS G13D tumor mutations with outcome in patients with metastatic colorectal cancer treated with first-line chemotherapy with or without cetuximab. J. Clin. Oncol. 30, 3570–3577 (2012).
Tol, J. et al. Chemotherapy, bevacizumab, and cetuximab in metastatic colorectal cancer. N. Engl. J. Med. 360, 563–572 (2009).
Palomba, G. et al. Prognostic impact of KRAS, NRAS, BRAF, and PIK3CA mutations in primary colorectal carcinomas: a population-based study. J. Transl Med. 14, 292 (2016).
Bencsikova, B. et al. Efficacy of bevacizumab and chemotherapy in the first-line treatment of metastatic colorectal cancer: broadening KRAS-focused clinical view. BMC Gastroenterol. 15, 37 (2015).
Saridaki, Z. et al. BRAF V600E mutation analysis in patients with metastatic colorectal cancer (mCRC) in daily clinical practice: correlations with clinical characteristics, and its impact on patients’ outcome. PLOS ONE 8, e84604 (2013).
Sjoquist, K. M. et al. Personalizing survival predictions in advanced colorectal cancer: the ARCAD nomogram project. J. Natl Cancer Inst. 110, 638–648 (2017). Analysis of pooled data from 26 randomized trials including 22,674 patients with metastatic CRC confirming that both KRAS mutations (HR for OS = 1.35; P <0.001) and BRAF mutation (HR = 2.21; P <0.001) are associated with inferior OS and PFS in multivariable models.
Wang, Y. et al. Distinct impacts of KRAS, NRAS and BRAF mutations on survival of patients with metastatic colorectal cancer. J. Clin. Oncol. 36, 3513 (2018).
Tosi, F. et al. Effect of KRAS and BRAF mutations on survival of metastatic colorectal cancer after liver resection: a systematic review and meta-analysis. Clin. Colorectal Cancer 16, e153–e163 (2017).
Brudvik, K. W. et al. RAS mutation clinical risk score to predict survival after resection of colorectal liver metastases. Ann. Surg. 269, 120–126 (2019).
Karapetis, C. S. et al. K-Ras mutations and benefit from cetuximab in advanced colorectal cancer. N. Engl. J. Med. 359, 1757–1765 (2008).
Arnold, D. et al. Prognostic and predictive value of primary tumour side in patients with RAS wild-type metastatic colorectal cancer treated with chemotherapy and EGFR directed antibodies in six randomized trials. Ann. Oncol. 28, 1713–1729 (2017). Retrospective analysis of pooled data from six randomized trials with standard therapy with or without anti-EGFR antibodies in patients with RAS wild-type metastatic CRCs showing that a right-sided primary tumour location might be a negative predictive factor for benefit from EGFR inhibition.
Tian, S. et al. A combined oncogenic pathway signature of BRAF, KRAS and PI3KCA mutation improves colorectal cancer classification and cetuximab treatment prediction. Gut 62, 540–549 (2013).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl Med. 6, 224ra24 (2014).
Morelli, M. P. et al. Characterizing the patterns of clonal selection in circulating tumor DNA from patients with colorectal cancer refractory to anti-EGFR treatment. Ann. Oncol. 26, 731–736 (2015).
Russo, M. et al. Tumor heterogeneity and lesion-specific response to targeted therapy in colorectal cancer. Cancer Discov. 6, 147–153 (2016).
Mohan, S. et al. Changes in colorectal carcinoma genomes under anti-EGFR therapy identified by whole-genome plasma DNA sequencing. PLOS Genet. 10, e1004271 (2014).
Pietrantonio, F. et al. Heterogeneity of acquired resistance to anti-EGFR monoclonal antibodies in patients with metastatic colorectal cancer. Clin. Cancer Res. 23, 2414–2422 (2017).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nat. Rev. Drug Discov. 13, 828–851 (2014).
Papke, B. & Der, C. J. Drugging RAS: know the enemy. Science 355, (1158–1163 (2017).
Patricelli, M. P. et al. Selective inhibition of oncogenic KRAS output with small molecules targeting the inactive state. Cancer Discov. 6, 316–329 (2016).
Wiesweg, M. et al. Impact of RAS mutation subtype on clinical outcome-a cross-entity comparison of patients with advanced non-small cell lung cancer and colorectal cancer. Oncogene 38, 2953–2966 (2019).
Tran, E. et al. T cell transfer therapy targeting mutant KRAS in cancer. N. Engl. J. Med. 375, 2255–2262 (2016).
Loboda, A. et al. EMT is the dominant program in human colon cancer. BMC Med. Genomics 4, 9 (2011).
Perez-Villamil, B. et al. Colon cancer molecular subtypes identified by expression profiling and associated to stroma, mucinous type and different clinical behavior. BMC Cancer 12, 260 (2012).
Schlicker, A. et al. Subtypes of primary colorectal tumors correlate with response to targeted treatment in colorectal cell lines. BMC Med. Genomics 5, 66 (2012).
Budinska, E. et al. Gene expression patterns unveil a new level of molecular heterogeneity in colorectal cancer. J. Pathol. 231, 63–76 (2013).
Sadanandam, A. et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat. Med. 19, 619–625 (2013).
De Sousa E. Melo, F. et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat. Med. 19, 614–618 (2013).
Marisa, L. et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLOS Med. 10, e1001453 (2013).
Roepman, P. et al. Colorectal cancer intrinsic subtypes predict chemotherapy benefit, deficient mismatch repair and epithelial-to-mesenchymal transition. Int. J. Cancer 134, 552–562 (2014).
Calon, A. et al. Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nat. Genet. 47, 320–329 (2015). TGFβ signalling enhances the stimulating effect of cancer-associated fibroblasts on tumour-initiating cells and disease progression in preclinical models can be halted by inhibiting the crosstalk between cancer cells and cancer-associated fibroblasts by inhibition of TGFβ signalling.
Bramsen, J. B. et al. Molecular-subtype-specific biomarkers improve prediction of prognosis in colorectal cancer. Cell Rep. 19, 1268–1280 (2017).
Isella, C. et al. Selective analysis of cancer-cell intrinsic transcriptional traits defines novel clinically relevant subtypes of colorectal cancer. Nat. Commun. 8, 15107 (2017).
Sveen, A. et al. Colorectal cancer consensus molecular subtypes translated to preclinical models uncover potentially targetable cancer-cell dependencies. Clin. Cancer Res. 24, 794–806 (2018).
Marisa, L. et al. Clinical utility of colon cancer molecular subtypes: validation of two main colorectal molecular classifications on the PETACC-8 phase III trial cohort. J. Clin. Oncol. 35, 3509 (2017).
Yamaguchi, S. et al. A validation study of stratification by the 55-gene classifier for assessing recurrence risk in stage II colon cancer: the 55 STAR study (UMIN23879). J. Clin. Oncol. 36, 3526 (2018).
Laurent-Puig, P. et al. Colon cancer molecular subtype intratumoral heterogeneity and its prognostic impact: an extensive molecular analysis of the PETACC-8 [abstract 60PD]. Ann. Oncol. 29, mdy269.058 (2018).
Fontana, E., Eason, K., Cervantes, A., Salazar, R. & Sadanandam, A. Context matters — consensus molecular subtypes of colorectal cancer as biomarkers for clinical trials. Ann. Oncol. 30, 520–527 (2019).
Piskol, R. et al. A clinical applicable gene expression classifier reveals intrinsic and extrinsic contributions to consensus molecular subtypes in primary and metastatic colon cancer. Clin. Cancer Res. https://doi.org/10.1158/1078-0432.CCR-18-3032 (2019).
Trumpi, K. et al. Neoadjuvant chemotherapy affects molecular classification of colorectal tumors. Oncogenesis 6, e357 (2017).
Dunne, P. D. et al. Cancer-cell intrinsic gene expression signatures overcome intratumoural heterogeneity bias in colorectal cancer patient classification. Nat. Commun. 8, 15657 (2017).
De Smedt, L. et al. Expression profiling of budding cells in colorectal cancer reveals an EMT-like phenotype and molecular subtype switching. Br. J. Cancer 116, 58–65 (2017).
Trinh, A. et al. Practical and robust identification of molecular subtypes in colorectal cancer by immunohistochemistry. Clin. Cancer Res. 23, 387–398 (2016).
Lenz, H. J. et al. Impact of consensus molecular subtype on survival in patients with metastatic colorectal cancer: results from CALGB/SWOG 80405 (Alliance). J. Clin. Oncol. https://doi.org/10.1200/JCO.18.02258 (2019).
Song, N. et al. Clinical outcome from oxaliplatin treatment in stage II/III colon cancer according to intrinsic subtypes: secondary analysis of NSABP C-07/NRG Oncology randomized clinical trial. JAMA Oncol. 2, 1162–1169 (2016).
Okita, A. et al. Consensus molecular subtypes classification of colorectal cancer as a predictive factor for chemotherapeutic efficacy against metastatic colorectal cancer. Oncotarget 9, 18698–18711 (2018).
Teufel, M. et al. Molecular subtypes and outcomes in regorafenib-treated patients with metastatic colorectal cancer (mCRC) enrolled in the CORRECT trial. J. Clin. Oncol. 33, 3558 (2015).
Linnekamp, J. F. et al. Consensus molecular subtypes of colorectal cancer are recapitulated in in vitro and in vivo models. Cell Death Differ. 25, 616–633 (2018).
Le, D. T. et al. A blueprint to advance colorectal cancer immunotherapies. Cancer Immunol. Res. 5, 942–949 (2017).
Garg, A. D. et al. Trial watch: immunogenic cell death induction by anticancer chemotherapeutics. Oncoimmunology 6, e1386829 (2017).
Nordlinger, B. et al. Perioperative FOLFOX4 chemotherapy and surgery versus surgery alone for resectable liver metastases from colorectal cancer (EORTC 40983): long-term results of a randomised, controlled, phase 3 trial. Lancet Oncol. 14, 1208–1215 (2013).
Dosset, M. et al. PD-1/PD-L1 pathway: an adaptive immune resistance mechanism to immunogenic chemotherapy in colorectal cancer. Oncoimmunology 7, e1433981 (2018).
Grothey, A. et al. Fluoropyrimidine (FP) + bevacizumab (BEV) + atezolizumab vs FP/BEV in BRAFwt metastatic colorectal cancer (mCRC): findings from Cohort 2 of MODUL — a multicentre, randomized trial of biomarker-driven maintenance treatment following first-line induction therapy [abstract LBA19]. Ann. Oncol. 29, mdy424.020 (2018).
Tauriello, D. V. F. et al. TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 554, 538–543 (2018). Expression of TGFβ in the tumour microenvironment is a mechanism of immune evasion and inhibition of TGFβ signalling in mice with CRC liver metastases promotes an antitumour cytotoxic T cell response and renders the tumours sensitive to ICIs.
Lan, Y. et al. Enhanced preclinical antitumor activity of M7824, a bifunctional fusion protein simultaneously targeting PD-L1 and TGF-β. Sci. Transl Med. 10, eaan5488 (2018).
Kopetz, S. et al. M7824 (MSB0011359C), a bifunctional fusion protein targeting PD-L1 and TGF-β, in patients with heavily pretreated CRC: preliminary results from a phase I trial. J. Clin. Oncol. 36, 764 (2018).
Ebert, P. J. R. et al. MAP kinase inhibition promotes T cell and anti-tumor activity in combination with PD-L1 checkpoint blockade. Immunity 44, 609–621 (2016).
Bendell, J. C. et al. A phase Ib study of safety and clinical activity of atezolizumab (A) and cobimetinib (C) in patients (pts) with metastatic colorectal cancer (mCRC). J. Clin. Oncol. 36, 560 (2018).
Eng, C. et al. Atezolizumab with or without cobimetinib versus regorafenib in previously treated metastatic colorectal cancer (IMblaze370): a multicentre, open-label, phase 3, randomised, controlled trial. Lancet Oncol. 20, 849–861 (2019).
Becht, E. et al. Immune and stromal classification of colorectal cancer is associated with molecular subtypes and relevant for precision immunotherapy. Clin. Cancer Res. 22, 4057–4066 (2016).
Meyer, C. et al. Frequencies of circulating MDSC correlate with clinical outcome of melanoma patients treated with ipilimumab. Cancer Immunol. Immunother. 63, 247–257 (2014).
Weber, R. et al. Myeloid-derived suppressor cells hinder the anti-cancer activity of immune checkpoint inhibitors. Front. Immunol. 9, 1310 (2018).
Kim, K. et al. Eradication of metastatic mouse cancers resistant to immune checkpoint blockade by suppression of myeloid-derived cells. Proc. Natl Acad. Sci. USA 111, 11774–11779 (2014).
Azuaje, F. Artificial intelligence for precision oncology: beyond patient stratification. NPJ Precis. Oncol. 3, 6 (2019).
Sorich, M. J. et al. Extended RAS mutations and anti-EGFR monoclonal antibody survival benefit in metastatic colorectal cancer: a meta-analysis of randomized, controlled trials. Ann. Oncol. 26, 13–21 (2015).
Lipson, E. et al. Durable cancer regression off-treatment and effective re-induction therapy with an anti-PD-1 antibody. Clin. Cancer Res. 19, 462–468 (2013).
Hochster, H. S. et al. Efficacy and safety of atezolizumab (atezo) and bevacizumab (bev) in a phase Ib study of microsatellite instability (MSI)-high metastatic colorectal cancer (mCRC). J. Clin. Oncol. 35, 673–673 (2017).
Acknowledgements
This work was supported by grants from the Norwegian Cancer Society (project numbers 6824048–2016 and 182759–2016), the Research Council of Norway (project number 250993), South-Eastern Norway Regional Health Authority (project numbers 2016123 and 2017102) and the NIH (project numbers R01 CA218230, R01 CA184843 and R01 CA187238).
Author information
Authors and Affiliations
Contributions
All authors made substantial contributions to all aspects of the preparation of this manuscript.
Corresponding author
Ethics declarations
Competing interests
A.S. and R.A.L. are co-inventors of a pending patent application (Attorney Docket Number: INVEN-35063/US-1/PRO) regarding the use of HSP90 inhibitors in relation to the consensus molecular subtypes of colorectal cancer. S.K. is a co-inventor of a pending patent application regarding a clinical classifier for the consensus molecular subtypes of colorectal cancer.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
The European Union Colossus Project: https://www.colossusproject.eu/researchers/
Supplementary Information
Rights and permissions
About this article
Cite this article
Sveen, A., Kopetz, S. & Lothe, R.A. Biomarker-guided therapy for colorectal cancer: strength in complexity. Nat Rev Clin Oncol 17, 11–32 (2020). https://doi.org/10.1038/s41571-019-0241-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-019-0241-1
This article is cited by
-
Edge-based relative entropy as a sensitive indicator of critical transitions in biological systems
Journal of Translational Medicine (2024)
-
Racial disparities in colorectal cancer clinicopathological and molecular tumor characteristics: a systematic review
Cancer Causes & Control (2024)
-
Aberrant methylation in neurofunctional gene serves as a hallmark of tumorigenesis and progression in colorectal cancer
BMC Cancer (2023)
-
Risk of retinal vein occlusion in colorectal cancer patients receiving anti-vascular endothelial growth factors – a population-based cohort study
BMC Cancer (2023)
-
The Oncology Biomarker Discovery framework reveals cetuximab and bevacizumab response patterns in metastatic colorectal cancer
Nature Communications (2023)