Abstract
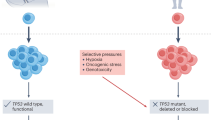
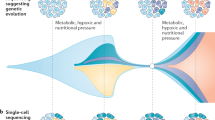
The emergence of treatment-resistant clones is a critical barrier to cure in patients with urothelial carcinoma. Setting the stage for the evolution of resistance, urothelial carcinoma is characterized by extensive mutational heterogeneity, which is detectable even in patients with early stage disease. Chemotherapy and immunotherapy both act as selective pressures that shape the evolutionary trajectory of urothelial carcinoma throughout the course of the disease. A detailed understanding of the dynamics of evolutionary drivers is required for the rational development of curative therapies. Herein, we describe the molecular basis of the clonal evolution of urothelial carcinomas and the use of genomic approaches to predict treatment responses. We discuss various mechanisms of resistance to chemotherapy with a focus on the mutagenic effects of the DNA dC->dU-editing enzymes APOBEC3 family of proteins. We also review the evolutionary mechanisms underlying resistance to immunotherapy, such as the loss of clonal tumour neoantigens. By dissecting treatment resistance through an evolutionary lens, the field will advance towards true precision medicine for urothelial carcinoma.
Key points
-
The emergence of therapy-resistant clones is a critical barrier to cure in patients with urothelial carcinoma.
-
The bulk of mutations in cancer driver genes accumulate early in the evolution of urothelial carcinoma in a pattern characteristic of DNA dC->dU editing enzyme APOBEC3 (APOBEC3)-mediated mutagenesis.
-
APOBEC3-mediated mutagenesis (stochastically) and cisplatin-based chemotherapy (deterministically) shape the evolutionary trajectory of urothelial carcinoma; early branching evolution is subject to clonal selection of APOBEC3-associated mutations following chemotherapy.
-
A high mutational load translates into neoantigenesis, evoking a robust antitumour immune response and higher sensitivity to immune-checkpoint inhibition.
-
As urothelial carcinomas evolve, they exploit several mechanisms of immune evasion, including neoantigen loss, major histocompatibility complex loss, downregulation of immunostimulatory signals, and activation of inhibitory signals.
-
Evolution-targeted therapeutic strategies, such as adaptive therapy and targeting of temporal collateral sensitivity, should be combined with precise tailoring of chemotherapeutic and immunotherapeutic interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Aragon-Ching, J. B. Challenges and advances in the diagnosis, biology, and treatment of urothelial upper tract and bladder carcinomas. Urol. Oncol. 35, 462–464 (2017).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
NIH. Cancer stat facts: bladder cancer. National Cancer Institute Surveillance, Epidemiology and End Results Program https://seer.cancer.gov/statfacts/html/urinb.html (2017).
Sridhar, S. S. Evolving treatment of advanced urothelial cancer. J. Oncol. Pract. 13, 309–315 (2017).
Sternberg, C. N. et al. Randomized phase III trial of high-dose-intensity methotrexate, vinblastine, doxorubicin, and cisplatin (MVAC) chemotherapy and recombinant human granulocyte colony-stimulating factor versus classic MVAC in advanced urothelial tract tumors: European organization for research and treatment of cancer protocol no. 30924. J. Clin. Oncol. 19, 2638–2646 (2001).
von der Maase, H. et al. Gemcitabine and cisplatin versus methotrexate, vinblastine, doxorubicin, and cisplatin in advanced or metastatic bladder cancer: results of a large, randomized, multinational, multicenter, phase III study. J. Clin. Oncol. 18, 3068–3077 (2000).
Bellmunt, J. et al. Randomized phase III study comparing paclitaxel/cisplatin/gemcitabine and gemcitabine/cisplatin in patients with locally advanced or metastatic urothelial cancer without prior systemic therapy: EORTC intergroup Study 30987. J. Clin. Oncol. 30, 1107–1113 (2012).
Bamias, A. et al. Prospective, open-label, randomized, phase III study of two dose-dense regimens MVAC versus gemcitabine/cisplatin in patients with inoperable, metastatic or relapsed urothelial cancer: a Hellenic cooperative oncology group study (HE 16/03). Ann. Oncol. 24, 1011–1017 (2013).
Alfred Witjes, J. et al. Updated 2016 EAU guidelines on muscle-invasive and metastatic bladder cancer. Eur. Urol. 71, 462–475 (2017).
Spiess, P. E. et al. Bladder cancer, version 5.2017, NCCN clinical practice guidelines in oncology. J. Natl Compr. Canc Netw 15, 1240–1267 (2017).
Kamat, A. M. et al. Society for Immunotherapy of Cancer consensus statement on immunotherapy for the treatment of bladder carcinoma. J. Immunother. Cancer 5, 68 (2017).
Balar, A. V. et al. Atezolizumab as first-line treatment in cisplatin-ineligible patients with locally advanced and metastatic urothelial carcinoma: a single-arm, multicentre, phase 2 trial. Lancet 389, 67–76 (2017).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Apolo, A. B. et al. Avelumab, an anti-programmed death-ligand 1 antibody, in patients with refractory metastatic urothelial carcinoma: results from a multicenter, phase Ib study. J. Clin. Oncol. 35, 2117–2124 (2017).
Massard, C. et al. Safety and efficacy of durvalumab (MEDI4736), an anti-programmed cell death ligand-1 immune checkpoint inhibitor, in patients with advanced urothelial bladder cancer. J. Clin. Oncol. 34, 3119–3125 (2016).
Sharma, P. et al. Nivolumab monotherapy in recurrent metastatic urothelial carcinoma (CheckMate 032): a multicentre, open-label, two-stage, multi-arm, phase 1/2 trial. Lancet Oncol. 17, 1590–1598 (2016).
Bellmunt, J. et al. Pembrolizumab as second-line therapy for advanced urothelial carcinoma. N. Engl. J. Med. 376, 1015–1026 (2017).
Zibelman, M., Ramamurthy, C. & Plimack, E. R. Emerging role of immunotherapy in urothelial carcinoma-advanced disease. Urol. Oncol. 34, 538–547 (2016).
Holohan, C., Van Schaeybroeck, S., Longley, D. B. & Johnston, P. G. Cancer drug resistance: an evolving paradigm. Nat. Rev. Cancer 13, 714–726 (2013).
Hölzel, M., Bovier, A. & Tüting, T. Plasticity of tumour and immune cells: a source of heterogeneity and a cause for therapy resistance? Nat. Rev. Cancer 13, 365–376 (2013).
Hughes, D. & Andersson, D. I. Evolutionary consequences of drug resistance: shared principles across diverse targets and organisms. Nat. Rev. Genet. 16, 459–471 (2015).
Gatenby, R. & Brown, J. The evolution and ecology of resistance in cancer therapy. Cold Spring Harb. Perspect. Med. 8, a033415 (2017).
Thomsen, M. B. et al. Spatial and temporal clonal evolution during development of metastatic urothelial carcinoma. Mol. Oncol. 10, 1450–1460 (2016).
Amirouchene-Angelozzi, N., Swanton, C. & Bardelli, A. Tumor evolution as a therapeutic target. Cancer Discov. 7, 805–817 (2017).
Greaves, M. Darwin and evolutionary tales in leukemia. The Ham-Wasserman lecture. Hematology Am. Soc. Hematol. Educ. Program 2009, 3–12 (2009).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Beckman, R. A. Efficiency of carcinogenesis: is the mutator phenotype inevitable? Semin. Cancer Biol. 20, 340–352. (2010).
Yeang, C. H. & Beckman, R. A. Long range personalized cancer treatment strategies incorporating evolutionary dynamics. Biol. Direct 11, 56 (2016).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e21 (2017).
Fedele, C., Tothill, R. W. & McArthur, G. A. Navigating the challenge of tumor heterogeneity in cancer therapy. Cancer Discov. 4, 146–148 (2014).
Baym, M. et al. Spatiotemporal microbial evolution on antibiotic landscapes. Science 353, 1147–1151 (2016).
Al Mamun, A. A. et al. Identity and function of a large gene network underlying mutagenic repair of DNA breaks. Science 338, 1344–1348 (2012).
Good, B. H., McDonald, M. J., Barrick, J. E., Lenski, R. E. & Desai, M. M. The dynamics of molecular evolution over 60,000 generations. Nature 551, 45–50 (2017).
Chen, H., Lin, F., Xing, K. & He, X. The reverse evolution from multicellularity to unicellularity during carcinogenesis. Nat. Commun. 6, 6367 (2015).
Blanes, A., Rubio, J., Sanchez-Carrillo, J. J. & Diaz-Cano, S. J. Coexistent intraurothelial carcinoma and muscle-invasive urothelial carcinoma of the bladder: clonality and somatic down-regulation of DNA mismatch repair. Hum. Pathol. 40, 988–997 (2009).
Nordentoft, I. et al. Mutational context and diverse clonal development in early and late bladder cancer. Cell Rep. 7, 1649–1663 (2014).
Robertson, A. G. et al. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 171, p540–556.e25 (2017).
Faltas, B. M. et al. Clonal evolution of chemotherapy-resistant urothelial carcinoma. Nat. Genet. 48, 1490–1499 (2016).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Warrick, J. I. et al. Tumor evolution and progression in multifocal and paired non-invasive/invasive urothelial carcinoma. Virchows Arch. 466, 297–311 (2015).
Lamy, P. et al. Paired exome analysis reveals clonal evolution and potential therapeutic targets in urothelial carcinoma. Cancer Res. 76, 5894–5906 (2016).
Cha, E. K. et al. Branched evolution and intratumor heterogeneity of urothelial carcinoma of the bladder. J. Clin. Oncol. 32 (suppl. 4), (2014).
Prandi, D. et al. Unraveling the clonal hierarchy of somatic genomic aberrations. Genome Biol. 15, 439 (2014).
Dentro, S. C., Wedge, D. C. & Van Loo, P. Principles of reconstructing the subclonal architecture of cancers. Cold Spring Harb. Perspect. Med. 7, a026625 (2017).
Venkatesan, S., Birkbak, N. J. & Swanton, C. Constraints in cancer evolution. Biochem. Soc. Trans. 45, 1–13 (2017).
Meads, M. B., Gatenby, R. A. & Dalton, W. S. Environment-mediated drug resistance: a major contributor to minimal residual disease. Nat. Rev. Cancer 9, 665–674 (2009).
Waclaw, B. et al. A spatial model predicts that dispersal and cell turnover limit intratumour heterogeneity. Nature 525, 261–264 (2015).
Antal, T., Krapivsky, P. L. & Nowak, M. A. Spatial evolution of tumors with successive driver mutations. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 92, 022705 (2015).
Liu, D. et al. Mutational patterns in chemotherapy resistant muscle-invasive bladder cancer. Nat. Commun. 8, 2193 (2017).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Findlay, J. M. et al. Differential clonal evolution in oesophageal cancers in response to neo-adjuvant chemotherapy. Nat. Commun. 7, 11111 (2016).
Swanton, C., McGranahan, N., Starrett, G. J. & Harris, R. S. APOBEC enzymes: mutagenic fuel for cancer evolution and heterogeneity. Cancer Discov. 5, 704–712 (2015).
Moris, A., Murray, S. & Cardinaud, S. AID and APOBECs span the gap between innate and adaptive immunity. Front. Microbiol. 5, 534 (2014).
Salter, J. D., Bennett, R. P. & Smith, H. C. The APOBEC protein family: united by structure, divergent in function. Trends Biochem. Sci. 41, 578–594 (2016).
Helleday, T., Eshtad, S. & Nik-Zainal, S. Mechanisms underlying mutational signatures in human cancers. Nat. Rev. Genet. 15, 585–598 (2014).
Roberts, S. A. et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, 970–976 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Burns, M. B., Temiz, N. A. & Harris, R. S. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat. Genet. 45, 977–983 (2013).
Chen, Z. et al. Heat shock proteins stimulate APOBEC-3-mediated cytidine deamination in the hepatitis B virus. J. Biol. Chem. 292, 13459–13479 (2017).
Stenglein, M. D. et al. APOBEC3 proteins mediate the clearance of foreign DNA from human cells. Nat. Struct. Mol. Biol. 17, 222–229 (2010).
Sharma, S. et al. The double-domain cytidine deaminase APOBEC3G is a cellular site-specific RNA editing enzyme. Sci. Rep. 6, 39100 (2016).
Sharma, S. et al. APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages. Nat. Commun. 6, 6881 (2015).
Kanu, N. et al. DNA replication stress mediates APOBEC3 family mutagenesis in breast cancer. Genome Biol. 17, 185 (2016).
Ohba, K. et al. In vivo and in vitro studies suggest a possible involvement of HPV infection in the early stage of breast carcinogenesis via APOBEC3B induction. PLoS ONE 9, e97787 (2014).
Talluri, S. et al. Apobec family of proteins mediate genomic changes in multiple myeloma by inducing DNA damage and C>T Mutations. Blood 130 (Suppl. 1), 4334 (2017).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci Transl Med 7, 283ra254 (2015).
de Bruin, E. C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
McGranahan, N. & Swanton, C. et al. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168, 613–628 (2017).
Boichard, A., Tsigelny, I. F. & Kurzrock, R. High expression of PD-1 ligands is associated with kataegis mutational signature and APOBEC3 alterations. Oncoimmunology 6, e1284719 (2017).
Mullane, S. A. et al. Correlation of apobec mRNA expression with overall survival and pd-l1 expression in urothelial carcinoma. Sci. Rep. 6, 27702 (2016).
Kazanov, M. D. et al. APOBEC-induced cancer mutations are uniquely enriched in early-replicating, gene-dense, and active chromatin regions. Cell Rep. 13, 1103–1109 (2015).
Hoopes, J. et al. APOBEC3A and APOBEC3B preferentially deaminate the lagging strand template during DNA REplication. Cell Rep.14, 1273–1282 (2016).
Seplyarskiy, V. B. et al. APOBEC-induced mutations in human cancers are strongly enriched on the lagging DNA strand during replication. Genome Res. 26, 174–182 (2016).
Nowarski, R. & Kotler, M. APOBEC3 cytidine deaminases in double-strand DNA break repair and cancer promotion. Cancer Res. 73, 3494–3498 (2013).
Smith, H. C., Bennett, R. P., Kizilyer, A., McDougall, W. M. & Prohaska, K. M. Functions and regulation of the APOBEC family of proteins. Semin. Cell Dev. Biol.23, 258–268 (2012).
Avesson, L. & Barry, G. The emerging role of RNA and DNA editing in cancer. Biochim. Biophys. Acta 1845, 308–316 (2014).
Okeoma, C. M., Lovsin, N., Peterlin, B. M. & Ross, S. R. APOBEC3 inhibits mouse mammary tumour virus replication in vivo. Nature 445, 927–930 (2007).
Middlebrooks, C. D. et al. Association of germline variants in the APOBEC3 region with cancer risk and enrichment with APOBEC-signature mutations in tumors. Nat. Genet. 48, 1330–1338 (2016).
Lipinski, K. A. et al. Cancer evolution and the limits of predictability in precision cancer medicine. Trends Cancer 2, 49–63 (2016).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Zhao, B., Hemann, M. T. & Lauffenburger, D. A. Modeling tumor clonal evolution for drug combinations design. Trends Cancer 2, 144–158 (2016).
Galluzzi, L. et al. Systems biology of cisplatin resistance: past, present and future. Cell Death Dis. 5, e1257 (2014).
Bellmunt, J. et al. Gene expression of ERCC1 as a novel prognostic marker in advanced bladder cancer patients receiving cisplatin-based chemotherapy. Ann. Oncol. 18, 522–528 (2007).
Font, A. et al. BRCA1 mRNA expression and outcome to neoadjuvant cisplatin-based chemotherapy in bladder cancer. Ann. Oncol. 22, 139–144 (2011).
Van Allen, E. M. et al. Somatic ERCC2 mutations correlate with cisplatin sensitivity in muscle-invasive urothelial carcinoma. Cancer Discov. 4, 1140–1153 (2014).
Desai, N. B. et al. Genomic characterization of response to chemoradiation in urothelial bladder cancer. Cancer 122, 3715–3723 (2016).
Plimack, E. R. et al. Defects in DNA repair genes predict response to neoadjuvant cisplatin-based chemotherapy in muscle-invasive bladder cancer. Eur. Urol. 68, 959–967 (2015).
Teo, M. Y. et al. DNA damage response and repair gene alterations are associated with improved survival in patients with platinum-treated advanced urothelial carcinoma. Clin. Cancer Res. 23, 3610–3618 (2017).
Roos, W. P. et al. DNA damage and the balance between survival and death in cancer biology. Nat. Rev. Cancer 16, 20–33 (2016).
Choi, W. et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 25, 152–165 (2014).
Seiler, R. et al. Impact of molecular subtypes in muscle-invasive bladder cancer on predicting response and survival after neoadjuvant chemotherapy. Eur. Urol. 72, 544–554 (2017).
Bertheau, P. et al. Exquisite sensitivity of TP53 mutant and basal breast cancers to a dose-dense epirubicin-cyclophosphamide regimen. PLoS Med. 4, e90 (2007).
Jovanovic, B. et al. A randomized phase II neoadjuvant study of cisplatin, paclitaxel with or without everolimus in patients with stage II/III triple-negative breast cancer (TNBC): responses and long-term outcome correlated with increased frequency of DNA damage response gene mutations, TNBC subtype, AR status, and Ki67. Clin. Cancer Res. 23, 4035–4045 (2011).
Dasari, S. & Tchounwou, P. B. Cisplatin in cancer therapy: molecular mechanisms of action. Eur. J. Pharmacol. 740, 364–378 (2014).
Massari, F. et al. Emerging concepts on drug resistance in bladder cancer: implications for future strategies. Crit. Rev. Oncol. Hematol. 96, 81–90 (2015).
Galluzzi, L. et al. Molecular mechanisms of cisplatin resistance. Oncogene 31, 1869–1883 (2012).
Munn, D. H. & Mellor, A. L. IDO in the tumor microenvironment: inflammation, counter-regulation, and tolerance. Trends Immunol. 37, 193–207 (2016).
Maleki Vareki, S. et al. Indoleamine 2,3-dioxygenase mediates immune-independent human tumor cell resistance to olaparib, gamma radiation, and cisplatin. Oncotarget 5, 2778–2791 (2014).
Afonso, J. et al. CD147 and MCT1-potential partners in bladder cancer aggressiveness and cisplatin resistance. Mol. Carcinog. 54, 1451–1466 (2015).
Tanaka, N. et al. Acquired platinum resistance involves epithelial to mesenchymal transition through ubiquitin ligase FBXO32 dysregulation. JCI Insight 1, e83654 (2016).
Baras, A. S. et al. Identification and validation of protein biomarkers of response to neoadjuvant platinum chemotherapy in muscle invasive urothelial carcinoma. PLoS ONE 10, e0131245 (2015).
Skowron, M. A. et al. Phenotype plasticity rather than repopulation from CD90/CK14+ cancer stem cells leads to cisplatin resistance of urothelial carcinoma cell lines. J. Exp. Clin. Cancer Res. 34, 144 (2015).
Kurtova, A. V. et al. Blocking PGE2-induced tumour repopulation abrogates bladder cancer chemoresistance. Nature 517, 209–213 (2015).
Dunn, G. P., Bruce, A. T., Ikeda, H., Old, L. J. & Schreiber, R. D. Cancer immunoediting: from immunosurveillance to tumor escape. Nat. Immunol. 3, 991–998 (2002).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Liu, X. S. & Mardis, E. R. Applications of immunogenomics to cancer. Cell 168, 600–612 (2017).
Le, D. T. et al. PD-1 Blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Choudhury, N. J. et al. Low T cell receptor diversity, high somatic mutation burden, and high neoantigen load as predictors of clinical outcome in muscle-invasive bladder cancer. Eur. Urol. Focus 2, 445–452 (2016).
Brown, S. D. et al. Neo-antigens predicted by tumor genome meta-analysis correlate with increased patient survival. Genome Res. 24, 743–750 (2014).
Faltas, B. et al. Generating a neoantigen map of advanced urothelial carcinoma by whole exome sequencing. J. Clin. Oncol. 34, (suppl. 2), (2016).
Haraksingh, R. R. & Snyder, M. P. Impacts of variation in the human genome on gene regulation. J. Mol. Biol. 425, 3970–3977 (2013).
Turajlic, S. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Anagnostou, V. et al. Evolution of neoantigen landscape during immune checkpoint blockade in non-small cell lung cancer. Cancer Discov. 7, 264–276 (2017).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Kirsch, I., Vignali, M. & Robins, H. T cell receptor profiling in cancer. Mol. Oncol. 9, 2063–2070 (2015).
Snyder, A. et al. Contribution of systemic and somatic factors to clinical response and resistance to PD-L1 blockade in urothelial cancer: an exploratory multi-omic analysis. PLoS Med. 14, e1002309 (2017).
Nghiem, P. T. et al. PD-1 Blockade with Pembrolizumab in advanced merkel-cell carcinoma. N. Engl. J. Med. 374, 2542–2552 (2016).
Sweis, R. F. et al. Molecular drivers of the non-T cell-inflamed tumor microenvironment in urothelial bladder cancer. Cancer Immunol. Res. 4, 563–568 (2016).
Kataoka, K. et al. Aberrant PD-L1 expression through 3′-UTR disruption in multiple cancers. Nature 534, 402–406 (2016).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 6, 202–216 (2016).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
George, S. et al. Loss of PTEN is associated with resistance to anti-PD-1 checkpoint blockade therapy in metastatic uterine leiomyosarcoma. Immunity 46, 197–204 (2017).
Van Allen, E. M. et al. Long-term benefit of PD-L1 blockade in lung cancer associated with JAK3 activation. Cancer Immunol. Res. 3, 855–863 (2015).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
McGranahan, N. et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271.e11 (2017).
Wang, X. et al. Suppression of type I IFN signaling in tumors mediates resistance to anti-PD-1 treatment that can be overcome by radiotherapy. Cancer Res. 77, 839–850 (2017).
Gao, J. et al. Loss of IFN-γ pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167, 397–404.e9 (2016).
Gettinger, S. et al. Impaired HLA class I antigen processing and presentation as a mechanism of acquired resistance to immune checkpoint inhibitors in lung cancer. Cancer Discov. 7, 1420–1435 (2017).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Korpal, M. et al. Evasion of immunosurveillance by genomic alterations of PPARγ/RXRα in bladder cancer. Nat. Commun. 8, 103 (2017).
Kardos, J. et al. Claudin-low bladder tumors are immune infiltrated and actively immune suppressed. JCI Insight 1, e85902 (2016).
Hendrickx, W. et al. Identification of genetic determinants of breast cancer immune phenotypes by integrative genome-scale analysis. Oncoimmunology 6, e1253654 (2017).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Jiménez-Sánchez, A. et al. Heterogeneous tumor-immune microenvironments among differentially growing metastases in an ovarian cancer patient. Cell 170, 927–938 (2017). e920.
Reuben, A. et al. Genomic and immune heterogeneity are associated with differential responses to therapy in melanoma. NPJ Genom. Med. https://doi.org/10.1038/s41525-017-0013-8 (2017).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548 (2017).
Murtaza, M. et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature 497, 108–112 (2013).
Russo, M. et al. Tumor heterogeneity and lesion-specific response to targeted therapy in colorectal cancer. Cancer Discov. 6, 147–153 (2016).
Wang, S., Gao, D. & Chen, Y. The potential of organoids in urological cancer research. Nat. Rev. Urol. 14, 401–414 (2017).
Pauli, C. et al. Personalized in vitro and in vivo cancer models to guide precision medicine. Cancer Discov. 7, 462–477 (2017).
Barlow, L., Meyer, R., Shelkey, E., Faltas, B. & Rubin, M. Integrin signaling modulation demonstrates potential therapeutic strategy in bladder cancer using three-dimensional organoid culture. Cancer Res 77 (suppl. 13), (2017).
Enriquez-Navas, P. M. et al. Exploiting evolutionary principles to prolong tumor control in preclinical models of breast cancer. Sci. Transl. Med. 8, 327ra324 (2016).
Das Thakur, M. et al. Modelling vemurafenib resistance in melanoma reveals a strategy to forestall drug resistance. Nature 494, 251–255 (2013).
Zhao, B. et al. Exploiting temporal collateral sensitivity in tumor clonal evolution. Cell 165, 234–246 (2016).
Szabados, B. et al. Response rate to chemotherapy after immune checkpoint inhibition in metastatic urothelial cancer. Eur. Urol. 73, 149–152 (2017).
Beckman, R. A. & Loeb, L. A. Evolutionary dynamics and significance of multiple subclonal mutations in cancer. DNA Repair. 56, 7–15 (2017).
Li, M. et al. First-in-class small molecule inhibitors of the single-strand DNA cytosine deaminase APOBEC3G. ACS Chem. Biol. 7, 506–517 (2012).
Olson, M. E. et al. Small molecule APOBEC3G DNA cytosine deaminase inhibitors based on a 4-amino-1,2,4-triazole-3-thiol scaffold. ChemMedChem 8, 112–117 (2013).
Petrylak, D. P. et al. A multicohort phase I study of ramucirumab (R) plus pembrolizumab (P): interim safety and clinical activity in patients with urothelial carcinoma. J. Clin. Oncol. 35 (suppl. 6), (2017).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Langer, C. J. et al. Carboplatin and pemetrexed with or without pembrolizumab for advanced, non-squamous non-small-cell lung cancer: a randomised, phase 2 cohort of the open-label KEYNOTE-021 study. Lancet Oncol. 17, 1497–1508 (2016).
Galsky, M. D. et al. Phase 2 Trial of gemcitabine, cisplatin, plus ipilimumab in patients with metastatic urothelial cancer and impact of dna damage response gene mutations on outcomes. Eur. Urol. 73, 751–759 (2018).
Petrylak, D. P. et al. Ramucirumab plus docetaxel versus placebo plus docetaxel in patients with locally advanced or metastatic urothelial carcinoma after platinum-based therapy (RANGE): a randomised, double-blind, phase 3 trial. Lancet 390, 2266–2277 (2017).
Apollo, A. B. et al. A phase II study of cabozantinib in patients (pts) with relapsed or refractory metastatic urothelial carcinoma (mUC). J. Clin. Oncol. 34 (suppl. 15), (2016).
Long, M. et al. Ibrutinib treatment improves T cell number and function in CLL patients. J. Clin. Invest. 127, 3052–3064 (2017).
Jiao, S. et al. PARP inhibitor upregulates PD-L1 expression and enhances cancer-associated immunosuppression. Clin. Cancer Res. 23, 3711–3720 (2017).
Sharma, P. et al. Efficacy and safety of nivolumab plus ipilimumab in metastatic urothelial carcinoma: first results from the phase I/II CheckMate 032 study. J. Immunother Cancer 4 (suppl. 2), (2016).
Chism, D. D. Urothelial carcinoma of the bladder and the rise of immunotherapy. J. Natl Compr. Canc Netw 15, 1277–1284 (2017).
Smith, D. C. et al. Epacadostat plus pembrolizumab in patients with advanced urothelial carcinoma: preliminary phase I/II results of ECHO-202/KEYNOTE-037. J. Clin. Oncol. 35 (suppl. 15), 4503 (2017).
Luke, J. J. et al. Preliminary antitumor and immunomodulatory activity of BMS-986205, an optimized indoleamine 2,3-dioxygenase 1 (IDO1) inhibitor, in combination with nivolumab in patients with advanced cancers [abstract 041]. 32nd Annual Meeting of the Society for Immunotherapy of Cancer (2017).
Hundal, J. et al. Cancer immunogenomics: computational neoantigen identification and vaccine design. Cold Spring Harb. Symp. Quant. Biol. 81, 105–111 (2016).
Zhang, X., Sharma, P. K., Peter Goedegebuure, S. & Gillanders, W. E. Personalized cancer vaccines: targeting the cancer mutanome. Vaccine 35, 1094–1100 (2017).
Woller, N. et al. Viral infection of tumors overcomes resistance to PD-1-immunotherapy by broadening neoantigenome-directed T cell responses. Mol. Ther. 23, 1630–1640 (2015).
Zamarin, D. et al. Localized oncolytic virotherapy overcomes systemic tumor resistance to immune checkpoint blockade immunotherapy. Sci Transl Med 6, 226ra232 (2014).
Acknowledgements
The authors thank S. Maleki Vareki, Cancer Research Laboratory Program, Lawson Health Research Institute, London, Ontario, Canada, for his diligent proofreading of this manuscript.
Author information
Authors and Affiliations
Contributions
Both authors made a substantial contribution to all aspects of the preparation of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
ClinicalTrial.gov: https://clinicaltrials.gov/
Rights and permissions
About this article
Cite this article
Vlachostergios, P.J., Faltas, B.M. Treatment resistance in urothelial carcinoma: an evolutionary perspective. Nat Rev Clin Oncol 15, 495–509 (2018). https://doi.org/10.1038/s41571-018-0026-y
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-018-0026-y
This article is cited by
-
Integrative score based on CDK6, PD-L1 and TMB predicts response to platinum-based chemotherapy and PD-1/PD-L1 blockade in muscle-invasive bladder cancer
British Journal of Cancer (2024)
-
Immune inactivation by neuropilin-1 predicts clinical outcome and therapeutic benefit in muscle-invasive bladder cancer
Cancer Immunology, Immunotherapy (2022)
-
SRT1720 inhibits the growth of bladder cancer in organoids and murine models through the SIRT1-HIF axis
Oncogene (2021)
-
Targeting natural splicing plasticity of APOBEC3B restricts its expression and mutagenic activity
Communications Biology (2021)
-
Common germline-somatic variant interactions in advanced urothelial cancer
Nature Communications (2020)