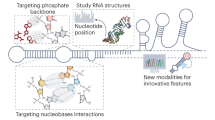
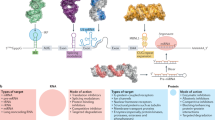
Abstract
Protein–RNA interactions have crucial roles in various cellular activities, which, when dysregulated, can lead to a range of human diseases. The identification of small molecules that target the interaction between RNA-binding proteins (RBPs) and RNA is progressing rapidly and represents a novel strategy for the discovery of chemical probes that facilitate understanding of the cellular functions of RBPs and of therapeutic agents with new mechanisms of action. In this Review, I present a current overview of targeting emerging RBPs using small-molecule inhibitors and recent progress in this burgeoning field. Small-molecule inhibitors that were reported for three representative emerging classes of RBPs, the microRNA-binding protein LIN28, the single-stranded or double-stranded RNA-binding Toll-like receptors and the CRISPR-associated (Cas) proteins, are highlighted from a medicinal-chemistry and chemical-biology perspective. However, although this field is burgeoning, challenges remain in the discovery and characterization of small-molecule inhibitors of RBPs.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout











Similar content being viewed by others
References
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Clamp, M. et al. Distinguishing protein-coding and noncoding genes in the human genome. Proc. Natl Acad. Sci. USA 104, 19428–19433 (2007).
Hopkins, A. L. & Groom, C. R. The druggable genome. Nat. Rev. Drug Discov. 1, 727–730 (2002).
Gerstberger, S., Hafner, M. & Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 15, 829–845 (2014).
Plaschka, C., Lin, P.-C. & Nagai, K. Structure of a pre-catalytic spliceosome. Nature 546, 617–621 (2017).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol. 19, 327–341 (2018).
Lukong, K. E., Chang, K.-w., Khandjian, E. W. & Richard, S. RNA-binding proteins in human genetic disease. Trends Genet. 24, 416–425 (2008).
Conlon, E. G. & Manley, J. L. RNA-binding proteins in neurodegeneration: mechanisms in aggregate. Genes Dev. 31, 1509–1528 (2017).
Angelbello, A. J. et al. Using genome sequence to enable the design of medicines and chemical probes. Chem. Rev. 118, 1599–1663 (2018).
Crooke, S. T., Witztum, J. L., Bennett, C. F. & Baker, B. F. RNA-targeted therapeutics. Cell Metab. 27, 714–739 (2018).
Kole, R., Krainer, A. R. & Altman, S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat. Rev. Drug Discov. 11, 125–140 (2012).
Rinaldi, C. & Wood, M. J. A. Antisense oligonucleotides: the next frontier for treatment of neurological disorders. Nat. Rev. Neurol. 14, 9–21 (2018).
Setten, R. L., Rossi, J. J. & Han, S.-p. The current state and future directions of RNAi-based therapeutics. Nat. Rev. Drug Discov. 18, 421–446 (2019).
Crooke, S. T. Vitraven — another piece in the mosaic. Antisense Nucleic Acid Drug Dev. 8, vii–viii (1998).
Crooke, S. T. & Geary, R. S. Clinical pharmacological properties of mipomersen (Kynamro), a second generation antisense inhibitor of apolipoprotein B. Br. J. Clin. Pharmacol. 76, 269–276 (2013).
Scoto, M., Finkel, R., Mercuri, E. & Muntoni, F. Genetic therapies for inherited neuromuscular disorders. Lancet Child Adolesc. Health 2, 600–609 (2018).
Benson, M. D. et al. Inotersen treatment for patients with hereditary transthyretin amyloidosis. N. Engl. J. Med. 379, 22–31 (2018).
Muntoni, F. et al. Golodirsen induces exon skipping leading to sarcolemmal dystrophin expression in Duchenne muscular dystrophy patients with mutations amenable to exon 53 skipping (S22.001). Neurology 90, S22.001 (2018).
Kim, J. et al. Patient-customized oligonucleotide therapy for a rare genetic disease. N. Engl. J. Med. 381, 1644–1652 (2019).
Sardh, E. et al. Phase 1 trial of an RNA interference therapy for acute intermittent porphyria. N. Engl. J. Med. 380, 549–558 (2019).
Warner, K. D., Hajdin, C. E. & Weeks, K. M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 17, 547–558 (2018).
Thomas, J. R. & Hergenrother, P. J. Targeting RNA with small molecules. Chem. Rev. 108, 1171–1224 (2008).
Connelly, C. M., Moon, M. H. & Schneekloth, J. S. Jr. The emerging role of RNA as a therapeutic target for small molecules. Cell Chem. Biol. 23, 1077–1090 (2016).
Disney, M. D. Targeting RNA with small molecules to capture opportunities at the intersection of chemistry, biology, and medicine. J. Am. Chem. Soc. 141, 6776–6790 (2019).
Mei, H.-Y. et al. Inhibition of an HIV-1 Tat-derived peptide binding to TAR RNA by aminoglycoside antibiotics. Bioorg. Med. Chem. Lett. 5, 2755–2760 (1995).
Mei, H.-Y. et al. Inhibitors of protein–RNA complexation that target the RNA: specific recognition of human immunodeficiency virus type 1 TAR RNA by small organic molecules. Biochemistry 37, 14204–14212 (1998).
Mei, H.-Y. et al. Discovery of selective, small-molecule inhibitors of RNA complexes — 1. The Tat protein/TAR RNA complexes required for HIV-1 transcription. Bioorg. Med. Chem. 5, 1173–1184 (1997).
Ratmeyer, L. S., Vinayak, R., Zon, G. & Wilson, W. D. An ethidium analog that binds with high specificity to a base-bulged duplex from the TAR RNA region of the HIV-1 genome. J. Med. Chem. 35, 966–968 (1992).
Moazed, D. & Noller, H. F. Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327, 389–394 (1987).
Mei, H.-Y. et al. Inhibition of self-splicing group I intron RNA: high-throughput screening assays. Nucleic Acids Res. 24, 5051–5053 (1996).
Mei, H.-Y., Cui, M., Lemrow, S. M. & Czarnik, A. W. Discovery of selective, small-molecule inhibitors of RNA complexes — II. Self-splicing group I intron ribozyme. Bioorg. Med. Chem. 5, 1185–1195 (1997).
von Ahsen, U., Davies, J. & Schroeder, R. Antibiotic inhibition of group I ribozyme function. Nature 353, 368–370 (1991).
Zapp, M. L., Stern, S. & Green, M. R. Small molecules that selectively block RNA binding of HIV-1 Rev protein inhibit Rev function and viral production. Cell 74, 969–978 (1993).
Wilson, W. D., Ratmeyer, L., Zhao, M., Strekowski, L. & Boykin, D. The search for structure-specific nucleic acid-interactive drugs: Effects of compound structure on RNA versus DNA interaction strength. Biochemistry 32, 4098–4104 (1993).
McConnaughie, A. W., Spychala, J., Zhao, M., Boykin, D. & Wilson, W. D. Design and synthesis of RNA-specific groove-binding cations: implications for antiviral drug design. J. Med. Chem. 37, 1063–1069 (1994).
Arambula, J. F., Ramisetty, S. R., Baranger, A. M. & Zimmerman, S. C. A simple ligand that selectively targets CUG trinucleotide repeats and inhibits MBNL protein binding. Proc. Natl Acad. Sci. USA 106, 16068–16073 (2009).
Li, J. et al. A ligand that targets CUG trinucleotide repeats. Chem. Eur. J. 22, 14881–14889 (2016).
Childs-Disney, J. L. et al. Induction and reversal of myotonic dystrophy type 1 pre-mRNA splicing defects by small molecules. Nat. Commun. 4, 2044 (2013).
Nakatani, K. Molecular design targeting repeat sequences in human genome. J. Synth. Org. Chem. Jpn 66, 1126–1133 (2008).
Mukherjee, S. et al. Structural insights into synthetic ligands targeting A–A pairs in disease-related CAG RNA repeats. Nucleic Acids Res. 47, 10906–10913 (2019).
Timchenko, L. T. et al. Identification of a (CUG)n triplet repeat RNA-binding protein and its expression in myotonic dystrophy. Nucleic Acids Res. 24, 4407–4414 (1996).
Miller, J. W. et al. Recruitment of human muscleblind proteins to (CUG)n expansions associated with myotonic dystrophy. EMBO J. 19, 4439–4448 (2000).
Velagapudi, S. P., Gallo, S. M. & Disney, M. D. Sequence-based design of bioactive small molecules that target precursor microRNAs. Nat. Chem. Biol. 10, 291–297 (2014).
Disney, M. D. et al. Inforna 2.0: A platform for the sequence-based design of small molecules targeting structured RNAs. ACS Chem. Biol. 11, 1720–1728 (2016).
Disney, M. D. & Angelbello, A. J. Rational design of small molecules targeting oncogenic noncoding RNAs from sequence. Acc. Chem. Res. 49, 2698–2704 (2016).
Costales, M. G., Matsumoto, Y., Velagapudi, S. P. & Disney, M. D. Small molecule targeted recruitment of a nuclease to RNA. J. Am. Chem. Soc. 140, 6741–6744 (2018).
Connelly, C. M., Boer, R. E., Moon, M. H., Gareiss, P. & Schneekloth, J. S. Discovery of inhibitors of microRNA-21 processing using small molecule microarrays. ACS Chem. Biol. 12, 435–443 (2017).
Warner, K. D. et al. Validating fragment-based drug discovery for biological RNAs: lead fragments bind and remodel the TPP riboswitch specifically. Chem. Biol. 21, 591–595 (2014).
Shi, Y. et al. Stabilization of lncRNA GAS5 by a small molecule and its implications in diabetic adipocytes. Cell Chem. Biol. 26, 319–330.e6 (2019).
Abulwerdi, F. A. et al. Selective small-molecule targeting of a triple helix encoded by the long noncoding RNA, MALAT1. ACS Chem. Biol. 14, 223–235 (2019).
Dai, Y. et al. Discovery of a branched peptide that recognizes the Rev response element (RRE) RNA and blocks HIV-1 replication. J. Med. Chem. 61, 9611–9620 (2018).
Matarlo, J. S. et al. The natural product butylcycloheptyl prodiginine binds pre-miR-21, inhibits dicer-mediated processing of pre-miR-21, and blocks cellular proliferation. Cell Chem. Biol. 26, 1133–1142.e4 (2019).
Yan, H. et al. Cyclic peptidomimetics as inhibitor for miR-155 biogenesis. Mol. Pharm. 16, 914–920 (2019).
Sundararaman, B. et al. Resources for the comprehensive discovery of functional RNA elements. Mol. Cell 61, 903–913 (2016).
Meisner, N.-C. et al. Identification and mechanistic characterization of low-molecular-weight inhibitors for HuR. Nat. Chem. Biol. 3, 508–515 (2007).
Chae, M.-J. et al. Chemical inhibitors destabilize HuR binding to the AU-rich element of TNF-α mRNA. Exp. Mol. Med. 41, 824–831 (2009).
Wu, X. et al. Identification and validation of novel small molecule disruptors of HuR-mRNA interaction. ACS Chem. Biol. 10, 1476–1484 (2015).
Wang, Z. et al. Rubipodanin A, the first natural N-desmonomethyl Rubiaceae-type cyclopeptide from Rubia podantha, indicating an important role of the N9-methyl group in the conformation and bioactivity. PLoS One 10, e0144950 (2015).
Lal, P. et al. Regulation of HuR structure and function by dihydrotanshinone-I. Nucleic Acids Res. 45, 9514–9527 (2017).
Kaur, K. et al. The fungal natural product azaphilone-9 binds to HuR and inhibits HuR-RNA interaction in vitro. PLoS One 12, e0175471 (2017).
Manzoni, L. et al. Interfering with HuR–RNA interaction: design, synthesis and biological characterization of tanshinone mimics as novel, effective HuR inhibitors. J. Med. Chem. 61, 1483–1498 (2018).
Clingman, C. C. et al. Allosteric inhibition of a stem cell RNA-binding protein by an intermediary metabolite. eLife 3, e02848 (2014).
Lan, L. et al. Natural product (−)-gossypol inhibits colon cancer cell growth by targeting RNA-binding protein Musashi-1. Mol. Oncol. 9, 1406–1420 (2015).
Gerard, M. et al. A 1536-well fluorescence polarization assay to screen for modulators of the MUSASHI family of RNA-binding proteins. Comb. Chem. High. Throughput Screen. 17, 596–609 (2014).
Minuesa, G. et al. Small-molecule targeting of MUSASHI RNA-binding activity in acute myeloid leukemia. Nat. Commun. 10, 2691 (2019).
Ohe, K. & Hagiwara, M. Modulation of alternative splicing with chemical compounds in new therapeutics for human diseases. ACS Chem. Biol. 10, 914–924 (2015).
Boer, R. E., Torrey, Z. R. & Schneekloth, J. S. Chemical modulation of pre-mRNA splicing in mammalian systems. ACS Chem. Biol. 15, 808–818 (2020).
Wang, E. et al. Targeting an RNA-binding protein network in acute myeloid leukemia. Cancer Cell 35, 369–384 (2019).
Han, T. et al. Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science 356, eaal3755 (2017).
Kaida, D. et al. Spliceostatin A targets SF3b and inhibits both splicing and nuclear retention of pre-mRNA. Nat. Chem. Biol. 3, 576–583 (2007).
Kotake, Y. et al. Splicing factor SF3b as a target of the antitumor natural product pladienolide. Nat. Chem. Biol. 3, 570–575 (2007).
Fan, L., Lagisetti, C., Edwards, C. C., Webb, T. R. & Potter, P. M. Sudemycins, novel small molecule analogues of FR901464, induce alternative gene splicing. ACS Chem. Biol. 6, 582–589 (2011).
Seiler, M. et al. H3B-8800, an orally available small-molecule splicing modulator, induces lethality in spliceosome-mutant cancers. Nat. Med. 24, 497–504 (2018).
Finci, L. I. et al. The cryo-EM structure of the SF3b spliceosome complex bound to a splicing modulator reveals a pre-mRNA substrate competitive mechanism of action. Genes Dev. 32, 309–320 (2018).
Alfadhli, A. et al. Analysis of small molecule ligands targeting the HIV-1 matrix protein-RNA binding site. J. Biol. Chem. 288, 666–676 (2013).
D’Agostino, V. G. et al. Screening approaches for targeting ribonucleoprotein complexes: a new dimension for drug discovery. SLAS Discov. 24, 314–331 (2019).
Boriack-Sjodin, P. A., Ribich, S. & Copeland, R. A. RNA-modifying proteins as anticancer drug targets. Nat. Rev. Drug Discov. 17, 435–453 (2018).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Liu, N. et al. N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions. Nature 518, 560–564 (2015).
Lovci, M. T., Bengtson, M. H. & Massirer, K. B. in RNA Processing: Disease and Genome-Wide Probing (ed Yeo., E. W.) 297–317 (Springer, 2016).
Hendrickson, D. G., Kelley, D. R., Tenen, D., Bernstein, B. & Rinn, J. L. Widespread RNA binding by chromatin-associated proteins. Genome Biol. 17, 28 (2016).
Skalska, L., Beltran-Nebot, M., Ule, J. & Jenner, R. G. Regulatory feedback from nascent RNA to chromatin and transcription. Nat. Rev. Mol. Cell Biol. 18, 331–337 (2017).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Auweter, S. D., Oberstrass, F. C. & Allain, F. H.-T. Sequence-specific binding of single-stranded RNA: is there a code for recognition? Nucleic Acids Res. 34, 4943–4959 (2006).
Valverde, R., Edwards, L. & Regan, L. Structure and function of KH domains. FEBS J. 275, 2712–2726 (2008).
Linder, P. & Jankowsky, E. From unwinding to clamping — the DEAD box RNA helicase family. Nat. Rev. Mol. Cell Biol. 12, 505–516 (2011).
Graumann, P. L. & Marahiel, M. A. A superfamily of proteins that contain the cold-shock domain. Trends Biochem. Sci. 23, 286–290 (1998).
Nakaminami, K., Karlson, D. T. & Imai, R. Functional conservation of cold shock domains in bacteria and higher plants. Proc. Natl Acad. Sci. USA 103, 10122–10127 (2006).
Cassandri, M. et al. Zinc-finger proteins in health and disease. Cell Death Discov. 3, 17071 (2017).
Saunders, L. R. & Barber, G. N. The dsRNA binding protein family: critical roles, diverse cellular functions. FASEB J. 17, 961–983 (2003).
Tian, B., Bevilacqua, P. C., Diegelman-Parente, A. & Mathews, M. B. The double-stranded-RNA-binding motif: interference and much more. Nat. Rev. Mol. Cell Biol. 5, 1013–1023 (2004).
Reich, D. P. & Bass, B. L. Mapping the dsRNA world. Cold Spring Harb. Perspect. Biol. 11, a035352 (2019).
Masliah, G., Barraud, P. & Allain, F. H.-T. RNA recognition by double-stranded RNA binding domains: a matter of shape and sequence. Cell. Mol. Life Sci. 70, 1875–1895 (2013).
Ray, D. et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 499, 172–177 (2013).
Castello, A. et al. Identification of RNA-binding domains of RNA-binding proteins in cultured cells on a system-wide scale with RBDmap. Nat. Protoc. 12, 2447–2464 (2017).
Basu, S. & Bahadur, R. P. A structural perspective of RNA recognition by intrinsically disordered proteins. Cell. Mol. Life Sci. 73, 4075–4084 (2016).
Lunde, B. M., Moore, C. & Varani, G. RNA-binding proteins: modular design for efficient function. Nat. Rev. Mol. Cell Biol. 8, 479–490 (2007).
Jankowsky, E. & Harris, M. Specificity and nonspecificity in RNA–protein interactions. Nat. Rev. Mol. Cell Biol. 16, 533–544 (2015).
Choi, Y. D. & Dreyfuss, G. Monoclonal antibody characterization of the C proteins of heterogeneous nuclear ribonucleoprotein complexes in vertebrate cells. J. Cell Biol. 99, 1997–1204 (1984).
Piñol-Roma, S., Choi, Y. D., Matunis, M. J. & Dreyfuss, G. Immunopurification of heterogeneous nuclear ribonucleoprotein particles reveals an assortment of RNA-binding proteins. Genes Dev. 2, 215–227 (1988).
Hentze, M. W. & Argos, P. Homology between IRE-BP, a regulatory RNA-binding protein, aconitase, and isopropylmalate isomerase. Nucleic Acids Res. 19, 1739–1740 (1991).
Rouault, T. A., Stout, C. D., Kaptain, S., Harford, J. B. & Klausner, R. D. Structural relationship between an iron-regulated RNA-binding protein (IRE-BP) and aconitase: Functional implications. Cell 64, 881–883 (1991).
Datar, K. V., Dreyfuss, G. & Swanson, M. S. The human hnRNP M proteins: identification of a methionine/arginine-rich repeat motif in ribonucleoproteins. Nucleic Acids Res. 21, 439–446 (1993).
Butter, F., Scheibe, M., Mörl, M. & Mann, M. Unbiased RNA–protein interaction screen by quantitative proteomics. Proc. Natl Acad. Sci. USA 106, 10626–10631 (2009).
König, J., Zarnack, K., Luscombe, N. M. & Ule, J. Protein–RNA interactions: new genomic technologies and perspectives. Nat. Rev. Genet. 13, 77–83 (2012).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Beckmann, B. M. et al. The RNA-binding proteomes from yeast to man harbour conserved enigmRBPs. Nat. Commun. 6, 10127 (2015).
Baltz, Alexander, G. et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 46, 674–690 (2012).
Matia-González, A. M., Laing, E. E. & Gerber, A. P. Conserved mRNA-binding proteomes in eukaryotic organisms. Nat. Struct. Mol. Biol. 22, 1027–1033 (2015).
Huang, R., Han, M., Meng, L. & Chen, X. Transcriptome-wide discovery of coding and noncoding RNA-binding proteins. Proc. Natl Acad. Sci. USA 115, E3879–E3887 (2018).
Jao, C. Y. & Salic, A. Exploring RNA transcription and turnover in vivo by using click chemistry. Proc. Natl Acad. Sci. USA 105, 15779–15784 (2008).
Bao, X. et al. Capturing the interactome of newly transcribed RNA. Nat. Methods 15, 213–220 (2018).
Queiroz, R. M. L. et al. Comprehensive identification of RNA–protein interactions in any organism using orthogonal organic phase separation (OOPS). Nat. Biotechnol. 37, 169–178 (2019).
Dreyfuss, G., Kim, V. N. & Kataoka, N. Messenger-RNA-binding proteins and the messages they carry. Nat. Rev. Mol. Cell Biol. 3, 195–205 (2002).
Hamy, F., Felder, E., Lipson, K. & Klimkait, T. Merged screening for human immunodeficiency virus Tat and Rev inhibitors. J. Biomol. Screen. 6, 179–187 (2001).
Iwata, M. et al. E7107, a new 7-urethane derivative of pladienolide D, displays curative effect against several human tumor xenografts. Cancer Res. 64, 691 (2004).
Eskens, F. A. L. M. et al. Phase I pharmacokinetic and pharmacodynamic study of the first-in-class spliceosome inhibitor E7107 in patients with advanced solid tumors. Clin. Cancer Res. 19, 6296–6304 (2013).
Bordeleau, M.-E. et al. Functional characterization of IRESes by an inhibitor of the RNA helicase eIF4A. Nat. Chem. Biol. 2, 213–220 (2006).
Moerke, N. J. et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 128, 257–267 (2007).
Roos, M. et al. A small-molecule inhibitor of Lin28. ACS Chem. Biol. 11, 2773–2781 (2016).
Lim, D., Byun, W. G., Koo, J. Y., Park, H. & Park, S. B. Discovery of a small-molecule inhibitor of protein–microRNA interaction using binding assay with a site-specifically labeled Lin28. J. Am. Chem. Soc. 138, 13630–13638 (2016).
Lightfoot, H. L., Miska, E. A. & Balasubramanian, S. Identification of small molecule inhibitors of the Lin28-mediated blockage of pre-let-7g processing. Org. Biomol. Chem. 14, 10208–10216 (2016).
Cheng, K., Wang, X. & Yin, H. Small-molecule inhibitors of the TLR3/dsRNA complex. J. Am. Chem. Soc. 133, 3764–3767 (2011).
Zhang, S. et al. Small-molecule inhibition of TLR8 through stabilization of its resting state. Nat. Chem. Biol. 14, 58–64 (2018).
Maji, B. et al. A high-throughput platform to identify small-molecule inhibitors of CRISPR-Cas9. Cell 177, 1067–1079.e19 (2019).
Mendell, J. R. et al. Single-dose gene-replacement therapy for spinal muscular atrophy. N. Engl. J. Med. 377, 1713–1722 (2017).
Hoy, S. M. Onasemnogene abeparvovec: first global approval. Drugs 79, 1255–1262 (2019).
Coovert, D. D. et al. The survival motor neuron protein in spinal muscular atrophy. Hum. Mol. Genet. 6, 1205–1214 (1997).
Hua, Y. et al. Peripheral SMN restoration is essential for long-term rescue of a severe spinal muscular atrophy mouse model. Nature 478, 123–126 (2011).
Ottesen, E. W., Singh, N. N., Luo, D. & Singh, R. N. High-affinity RNA targets of the Survival Motor Neuron protein reveal diverse preferences for sequence and structural motifs. Nucleic Acids Res. 46, 10983–11001 (2018).
Mercuri, E. et al. Nusinersen versus sham control in later-onset spinal muscular atrophy. N. Engl. J. Med. 378, 625–635 (2018).
Naryshkin, N. A. et al. SMN2 splicing modifiers improve motor function and longevity in mice with spinal muscular atrophy. Science 345, 688–693 (2014).
Kletzl, H. et al. The oral splicing modifier RG7800 increases full length survival of motor neuron 2 mRNA and survival of motor neuron protein: Results from trials in healthy adults and patients with spinal muscular atrophy. Neuromuscul. Disord. 29, 21–29 (2019).
Ratni, H. et al. Discovery of risdiplam, a selective survival of motor neuron-2 (SMN2) gene splicing modifier for the treatment of spinal muscular atrophy (SMA). J. Med. Chem. 61, 6501–6517 (2018).
Cheung, A. K. et al. Discovery of small molecule splicing modulators of survival motor neuron-2 (SMN2) for the treatment of spinal muscular atrophy (SMA). J. Med. Chem. 61, 11021–11036 (2018).
Palacino, J. et al. SMN2 splice modulators enhance U1–pre-mRNA association and rescue SMA mice. Nat. Chem. Biol. 11, 511–517 (2015).
Sivaramakrishnan, M. et al. Binding to SMN2 pre-mRNA-protein complex elicits specificity for small molecule splicing modifiers. Nat. Commun. 8, 1476 (2017).
Ambros, V. & Horvitz, H. Heterochronic mutants of the nematode Caenorhabditis elegans. Science 226, 409–416 (1984).
Batlle, E. & Clevers, H. Cancer stem cells revisited. Nat. Med. 23, 1124–1134 (2017).
Treiber, T. et al. A compendium of RNA-binding proteins that regulate microRNA biogenesis. Mol. Cell 66, 270–284.e13 (2017).
Tsialikas, J. & Romer-Seibert, J. LIN28: roles and regulation in development and beyond. Development 142, 2397–2404 (2015).
Ruby, J. G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Roush, S. & Slack, F. J. The let-7 family of microRNAs. Trends Cell Biol. 18, 505–516 (2008).
Lim, L. P. et al. The microRNAs of Caenorhabditis elegans. Genes Dev. 17, 991–1008 (2003).
Jonas, S. & Izaurralde, E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 16, 421–433 (2015).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524 (2014).
Treiber, T., Treiber, N. & Meister, G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell Biol. 20, 5–20 (2019).
Tsanov, K. M. et al. LIN28 phosphorylation by MAPK/ERK couples signalling to the post-transcriptional control of pluripotency. Nat. Cell Biol. 19, 60–67 (2017).
Wilbert, M. L. et al. LIN28 binds messenger RNAs at GGAGA motifs and regulates splicing factor abundance. Mol. Cell 48, 195–206 (2012).
O’Day, E. et al. An RNA-binding protein, Lin28, recognizes and remodels G-quartets in the microRNAs (miRNAs) and mRNAs it regulates. J. Biol. Chem. 290, 17909–17922 (2015).
Piskounova, E. et al. Lin28A and Lin28B inhibit let-7 microRNA biogenesis by distinct mechanisms. Cell 147, 1066–1079 (2011).
Chang, H.-M., Triboulet, R., Thornton, J. E. & Gregory, R. I. A role for the Perlman syndrome exonuclease Dis3l2 in the Lin28–let-7 pathway. Nature 497, 244–248 (2013).
Yamashita, S., Nagaike, T. & Tomita, K. Crystal structure of the Lin28-interacting module of human terminal uridylyltransferase that regulates let-7 expression. Nat. Commun. 10, 1960 (2019).
Heo, I. et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 138, 696–708 (2009).
Wang, L. et al. LIN28 zinc knuckle domain is required and sufficient to induce let-7 oligouridylation. Cell Rep. 18, 2664–2675 (2017).
Yermalovich, A. V. et al. Lin28 and let-7 regulate the timing of cessation of murine nephrogenesis. Nat. Commun. 10, 168 (2019).
Trang, P. et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 29, 1580–1587 (2010).
Roos, M. et al. Short loop-targeting oligoribonucleotides antagonize Lin28 and enable pre-let-7 processing and suppression of cell growth in let-7-deficient cancer cells. Nucleic Acids Res. 43, e9 (2014).
Mayr, F., Schütz, A., Döge, N. & Heinemann, U. The Lin28 cold-shock domain remodels pre-let-7 microRNA. Nucleic Acids Res. 40, 7492–7506 (2012).
Ustianenko, D. et al. LIN28 selectively modulates a subclass of let-7 microRNAs. Mol. Cell 71, 271–283.e5 (2018).
Nam, Y. et al. Molecular basis for interaction of let-7 microRNAs with Lin28. Cell 147, 1080–1091 (2011).
Loughlin, F. E. et al. Structural basis of pre-let-7 miRNA recognition by the zinc knuckles of pluripotency factor Lin28. Nat. Struct. Mol. Biol. 19, 84–89 (2012).
Albright, J. D. et al. Synthesis and anxiolytic activity of 6-(substituted-phenyl)-1,2,4-triazolo[4,3-b]pyridazines. J. Med. Chem. 24, 592–600 (1981).
Chen, Y. et al. LIN28/let-7/PD-L1 pathway as a target for cancer immunotherapy. Cancer Immunol. Res. 7, 487–497 (2019).
Park, S. O., Kim, J., Koh, M. & Park, S. B. Efficient parallel synthesis of privileged benzopyranylpyrazoles via regioselective condensation of β-keto aldehydes with hydrazines. J. Comb. Chem. 11, 315–326 (2009).
Lim, D., Byun, W. G. & Park, S. B. Restoring let-7 microRNA biogenesis using a small-molecule inhibitor of the protein–RNA interaction. ACS Med. Chem. Lett. 9, 1181–1185 (2018).
Wang, L. et al. Small-molecule inhibitors disrupt let-7 oligouridylation and release the selective blockade of let-7 processing by LIN28. Cell Rep. 23, 3091–3101 (2018).
Shumaker, D. K., Vann, L. R., Goldberg, M. W., Allen, T. D. & Wilson, K. L. TPEN, a Zn2+/Fe2+ chelator with low affinity for Ca2+, inhibits lamin assembly, destabilizes nuclear architecture and may independently protect nuclei from apoptosis in vitro. Cell Calcium 23, 151–164 (1998).
Oliver, C. L. et al. (−)-Gossypol acts directly on the mitochondria to overcome Bcl-2- and Bcl-XL-mediated apoptosis resistance. Mol. Cancer Ther. 4, 23–31 (2005).
Lorenz, D. A. et al. Expansion of cat-ELCCA for the discovery of small molecule inhibitors of the pre-let-7–Lin28 RNA–protein interaction. ACS Med. Chem. Lett. 9, 517–521 (2018).
Lorenz, D. A. & Garner, A. L. A click chemistry-based microRNA maturation assay optimized for high-throughput screening. Chem. Commun. 52, 8267–8270 (2016).
Garner, A. L. cat-ELCCA: catalyzing drug discovery through click chemistry. Chem. Commun. 54, 6531–6539 (2018).
Jacobsen, J. A., Major Jourden, J. L., Miller, M. T. & Cohen, S. M. To bind zinc or not to bind zinc: An examination of innovative approaches to improved metalloproteinase inhibition. Biochim. Biophys. Acta 1803, 72–94 (2010).
Root, K. et al. Aryl bis-sulfonamides bind to the active site of a homotrimeric isoprenoid biosynthesis enzyme IspF and extract the essential divalent metal cation cofactor. Chem. Sci. 9, 5976–5986 (2018).
Byun, W. G., Lim, D. & Park, S. B. Discovery of small-molecule modulators of protein–RNA interactions by fluorescence intensity-based binding assay. ChemBioChem 21, 818–824 (2020).
Liu, J. & Cao, X. Cellular and molecular regulation of innate inflammatory responses. Cell. Mol. Immunol. 13, 711–721 (2016).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Kerr, W. G. & Chisholm, J. D. The next generation of immunotherapy for cancer: small molecules could make big waves. J. Immunol. 202, 11–19 (2019).
Huck, B. R., Kötzner, L. & Urbahns, K. Small molecules drive big improvements in immuno-oncology therapies. Angew. Chem. Int. Ed. 57, 4412–4428 (2018).
Anwar, M. A., Shah, M., Kim, J. & Choi, S. Recent clinical trends in Toll-like receptor targeting therapeutics. Med. Res. Rev. 39, 1053–1090 (2019).
García, M. A. et al. Impact of protein kinase PKR in cell biology: from antiviral to antiproliferative action. Microbiol. Mol. Biol. Rev. 70, 1032–1060 (2006).
Dar, A. C., Dever, T. E. & Sicheri, F. Higher-order substrate recognition of eIF2α by the RNA-dependent protein kinase PKR. Cell 122, 887–900 (2005).
Banerjee, S. et al. OAS-RNase L innate immune pathway mediates the cytotoxicity of a DNA-demethylating drug. Proc. Natl Acad. Sci. USA 116, 5071–5076 (2019).
Aguado, L. C. et al. RNase III nucleases from diverse kingdoms serve as antiviral effectors. Nature 547, 114–117 (2017).
Malathi, K. et al. A transcriptional signaling pathway in the IFN system mediated by 2′-5′-oligoadenylate activation of RNase L. Proc. Natl Acad. Sci. USA 102, 14533–14538 (2005).
Kim, U., Wang, Y., Sanford, T., Zeng, Y. & Nishikura, K. Molecular cloning of cDNA for double-stranded RNA adenosine deaminase, a candidate enzyme for nuclear RNA editing. Proc. Natl Acad. Sci. USA 91, 11457–11461 (1994).
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Licht, K. & Jantsch, M. F. The other face of an editor: ADAR1 functions in editing-independent ways. BioEssays 39, 1700129 (2017).
Heraud-Farlow, J. E. & Kiebler, M. A. The multifunctional Staufen proteins: conserved roles from neurogenesis to synaptic plasticity. Trends Neurosci. 37, 470–479 (2014).
Takeda, K. & Akira, S. Toll-like receptors in innate immunity. Int. Immunol. 17, 1–14 (2005).
Anthoney, N., Foldi, I. & Hidalgo, A. Toll and Toll-like receptor signalling in development. Development 145, dev156018 (2018).
Marshak-Rothstein, A. Toll-like receptors in systemic autoimmune disease. Nat. Rev. Immunol. 6, 823–835 (2006).
Duffy, L. & O’Reilly, S. C. Toll-like receptors in the pathogenesis of autoimmune diseases: recent and emerging translational developments. Immunotargets Ther. 5, 69–80 (2016).
Zhang, L., Dewan, V. & Yin, H. Discovery of small molecules as multi-Toll-like receptor agonists with proinflammatory and anticancer activities. J. Med. Chem. 60, 5029–5044 (2017).
Yoo, E. et al. Determinants of activity at human Toll-like receptors 7 and 8: quantitative structure–activity relationship (QSAR) of diverse heterocyclic scaffolds. J. Med. Chem. 57, 7955–7970 (2014).
Zhang, Z. et al. Structural analysis reveals that Toll-like receptor 7 is a dual receptor for guanosine and single-stranded RNA. Immunity 45, 737–748 (2016).
Gosu, V., Son, S., Shin, D. & Song, K.-D. Insights into the dynamic nature of the dsRNA-bound TLR3 complex. Sci. Rep. 9, 3652 (2019).
Bevan, D. E. et al. Selection, preparation, and evaluation of small-molecule inhibitors of Toll-like receptor 4. ACS Med. Chem. Lett. 1, 194–198 (2010).
Tanji, H. et al. Toll-like receptor 8 senses degradation products of single-stranded RNA. Nat. Struct. Mol. Biol. 22, 109–115 (2015).
Tanji, H., Ohto, U., Shibata, T., Miyake, K. & Shimizu, T. Structural reorganization of the Toll-like receptor 8 dimer induced by agonistic ligands. Science 339, 1426–1429 (2013).
Liu, L. et al. Structural basis of Toll-like receptor 3 signaling with double-stranded RNA. Science 320, 379–381 (2008).
Choe, J., Kelker, M. S. & Wilson, I. A. Crystal structure of human Toll-like receptor 3 (TLR3) ectodomain. Science 309, 581–585 (2005).
Zheng, Z.-M. Circular RNAs and RNase L in PKR activation and virus infection. Cell Biosci. 9, 43 (2019).
Dalet, A., Gatti, E. & Pierre, P. Integration of PKR-dependent translation inhibition with innate immunity is required for a coordinated anti-viral response. FEBS Lett. 589, 1539–1545 (2015).
Liu, C.-X. et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell 177, 865–880.e21 (2019).
Grootjans, J., Kaser, A., Kaufman, R. J. & Blumberg, R. S. The unfolded protein response in immunity and inflammation. Nat. Rev. Immunol. 16, 469–484 (2016).
Mayo, C. B. et al. Structural basis of protein kinase R autophosphorylation. Biochemistry 58, 2967–2977 (2019).
Huang, H. et al. Dimeric structure of pseudokinase RNase L bound to 2-5A reveals a basis for interferon-induced antiviral activity. Mol. Cell 53, 221–234 (2014).
Han, Y. et al. Structure of human RNase L reveals the basis for regulated RNA decay in the IFN response. Science 343, 1244–1248 (2014).
Lee, K. P. K. et al. Structure of the dual enzyme Ire1 reveals the basis for catalysis and regulation in nonconventional RNA splicing. Cell 132, 89–100 (2008).
Feldman, H. C. et al. Structural and functional analysis of the allosteric inhibition of IRE1α with ATP-competitive ligands. ACS Chem. Biol. 11, 2195–2205 (2016).
Sun, L. et al. Discovery of 5-[5-fluoro-2-oxo-1,2- dihydroindol-(3Z)-ylidenemethyl]-2,4- dimethyl-1H-pyrrole-3-carboxylic acid (2-diethylaminoethyl)amide, a novel tyrosine kinase inhibitor targeting vascular endothelial and platelet-derived growth factor receptor tyrosine kinase. J. Med. Chem. 46, 1116–1119 (2003).
Jha, B. K. et al. Inhibition of RNase L and RNA-dependent protein kinase (PKR) by sunitinib impairs antiviral innate immunity. J. Biol. Chem. 286, 26319–26326 (2011).
Jammi, N. V., Whitby, L. R. & Beal, P. A. Small molecule inhibitors of the RNA-dependent protein kinase. Biochem. Biophys. Res. Commun. 308, 50–57 (2003).
Chen, H.-M., Wang, L. & D’Mello, S. R. A chemical compound commonly used to inhibit PKR, {8-(imidazol-4-ylmethylene)-6H-azolidino[5,4-g] benzothiazol-7-one}, protects neurons by inhibiting cyclin-dependent kinase. Eur. J. Neurosci. 28, 2003–2016 (2008).
Dabo, S. et al. Inhibition of the inflammatory response to stress by targeting interaction between PKR and its cellular activator PACT. Sci. Rep. 7, 16129 (2017).
Jinek, M. et al. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Liu, J.-J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Strecker, J. et al. Engineering of CRISPR-Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Nuñez, J. K., Harrington, L. B. & Doudna, J. A. Chemical and biophysical modulation of Cas9 for tunable genome engineering. ACS Chem. Biol. 11, 681–688 (2016).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Gerry, C. J. & Schreiber, S. L. Chemical probes and drug leads from advances in synthetic planning and methodology. Nat. Rev. Drug Discov. 17, 333–352 (2018).
Chen, J. S. & Doudna, J. A. The chemistry of Cas9 and its CRISPR colleagues. Nat. Rev. Chem. 1, 0078 (2017).
Pawluk, A. et al. Naturally occurring off-switches for CRISPR-Cas9. Cell 167, 1829–1838.e9 (2016).
Rauch, B. J. et al. Inhibition of CRISPR-Cas9 with bacteriophage proteins. Cell 168, 150–158.e10 (2017).
Shin, J. et al. Disabling Cas9 by an anti-CRISPR DNA mimic. Sci. Adv. 3, e1701620 (2017).
Hynes, A. P. et al. An anti-CRISPR from a virulent streptococcal phage inhibits Streptococcus pyogenes Cas9. Nat. Microbiol. 2, 1374–1380 (2017).
Gerard, B. et al. Application of a catalytic asymmetric Povarov reaction using chiral ureas to the synthesis of a tetrahydroquinoline library. ACS Comb. Sci. 14, 621–630 (2012).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017).
Abudayyeh, O. O. et al. RNA targeting with CRISPR–Cas13. Nature 550, 280–284 (2017).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schiöth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Sriram, K. & Insel, P. A. G protein-coupled receptors as targets for approved drugs: How many targets and how many drugs? Mol. Pharmacol. 93, 251–258 (2018).
Bagal, S. K. et al. Ion channels as therapeutic targets: a drug discovery perspective. J. Med. Chem. 56, 593–624 (2013).
Wu, P., Nielsen, T. E. & Clausen, M. H. FDA-approved small-molecule kinase inhibitors. Trends Pharmacol. Sci. 36, 422–439 (2015).
Nshogoza, G. et al. NMR fragment-based screening against tandem RNA recognition motifs of TDP-43. Int. J. Mol. Sci. 20, 3230 (2019).
Lama, L. et al. Development of human cGAS-specific small-molecule inhibitors for repression of dsDNA-triggered interferon expression. Nat. Commun. 10, 2261 (2019).
Kranzusch, P. J., Lee, A. S.-Y., Berger, J. M. & Doudna, J. A. Structure of human cGAS reveals a conserved family of second-messenger enzymes in innate immunity. Cell Rep. 3, 1362–1368 (2013).
Taylor, R. D., MacCoss, M. & Lawson, A. D. G. Rings in drugs. J. Med. Chem. 57, 5845–5859 (2014).
Wu, P., Clausen, M. H. & Nielsen, T. E. Allosteric small-molecule kinase inhibitors. Pharmacol. Ther. 156, 59–68 (2015).
Hudson, W. H. & Ortlund, E. A. The structure, function and evolution of proteins that bind DNA and RNA. Nat. Rev. Mol. Cell Biol. 15, 749–760 (2014).
Scott, D. E., Bayly, A. R., Abell, C. & Skidmore, J. Small molecules, big targets: drug discovery faces the protein–protein interaction challenge. Nat. Rev. Drug Discov. 15, 533–550 (2016).
Caudron-Herger, M. et al. R-DeeP: proteome-wide and quantitative identification of RNA-dependent proteins by density gradient ultracentrifugation. Mol. Cell 75, 184–199.e10 (2019).
Karatas, H. et al. High-affinity, small-molecule peptidomimetic inhibitors of MLL1/WDR5 protein–protein interaction. J. Am. Chem. Soc. 135, 669–682 (2013).
Stormo, G. D. & Zhao, Y. Determining the specificity of protein–DNA interactions. Nat. Rev. Genet. 11, 751–760 (2010).
Struntz, N. B. et al. Stabilization of the Max homodimer with a small molecule attenuates Myc-driven transcription. Cell Chem. Biol. 26, 711–723.e14 (2019).
Laver, J. D. et al. Synthetic antibodies as tools to probe RNA-binding protein function. Mol. Biosyst. 8, 1650–1657 (2012).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. A Cas9–guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014).
Liu, L., Yin, M., Wang, M. & Wang, Y. Phage AcrIIA2 DNA mimicry: structural basis of the CRISPR and anti-CRISPR arms race. Mol. Cell 73, 611–620.e3 (2019).
Liu, L. et al. The molecular architecture for RNA-guided RNA cleavage by Cas13a. Cell 170, 714–726.e10 (2017).
Acknowledgements
The author gratefully acknowledges financial support from AstraZeneca, Merck KGaA, Pfizer Inc. and the Max Planck Society. The author thanks L. Borgelt and J. Hwang for assistance in the preparation of figures and critical reading, H. Lightfoot and D. Lim for discussions on the LIN28–let-7 interaction, P. ‘t Hart and J. Imig for discussions on RNA-binding proteins and protein–RNA interactions, and P. Hommen for proofreading. The author would like to thank the anonymous reviewers for their thought-provoking comments and apologize to colleagues whose work was not cited owing to selected coverage.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- RNA-binding proteins
-
(RBPs). Proteins that contain various structural motifs that directly bind to ribonucleic acid. RBPs are a part of ribonucleoprotein complexes and regulate the metabolism of cellular RNAs.
- Ribonucleoprotein
-
A nuclear or cytoplasmic complex of RNAs and RNA-binding proteins.
- MicroRNA
-
(miRNA). Small non-coding RNA containing about 22 nucleotides, which negatively regulates gene targets.
- Non-coding RNA
-
(ncRNA). RNA that is not translated into a protein. Major classes of ncRNAs include ribosomal RNAs, transfer RNAs, long non-coding RNAs and small non-coding RNAs.
- Clustered regularly interspaced short palindromic repeats
-
(CRISPR). A series of short, repeating DNA sequences in the prokaryotic genome that are a hallmark of prokaryotic adaptive immune systems, in which CRISPR-associated (Cas) proteins recognize and bind to CRISPR RNA sequences as a guide to perform site-specific cleavage of external nucleic acids.
- Post-transcriptional regulators
-
Regulate RNA metabolism by various chemical changes catalysed by RNA-binding proteins on primary transcript of RNAs following transcription from a gene.
- Electrophoretic mobility shift assay
-
(EMSA). A method using a labelled nucleic acid to detect protein–nucleic-acid complexes, which migrate slower than the free nucleic acid in non-denaturing gel electrophoresis.
Rights and permissions
About this article
Cite this article
Wu, P. Inhibition of RNA-binding proteins with small molecules. Nat Rev Chem 4, 441–458 (2020). https://doi.org/10.1038/s41570-020-0201-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41570-020-0201-4
This article is cited by
-
Specificity, synergy, and mechanisms of splice-modifying drugs
Nature Communications (2024)
-
Small-molecule trapping of an RNA-binding protein blocks cancer cell growth
Nature Chemical Biology (2023)
-
Profiling dynamic RNA–protein interactions using small-molecule-induced RNA editing
Nature Chemical Biology (2023)
-
Chemoproteomic capture of RNA binding activity in living cells
Nature Communications (2023)
-
Remodeling oncogenic transcriptomes by small molecules targeting NONO
Nature Chemical Biology (2023)