Abstract
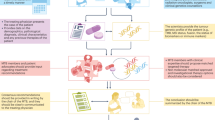
High-throughput methods to investigate tumour omic landscapes have quickly catapulted cancer specialists into the precision oncology era. The singular lesson of precision oncology might be that, for it to be precise, treatment must be personalized, as each cancer’s complex molecular and immune landscape differs from patient to patient. Transformative therapies include those that are targeted at the sequelae of molecular abnormalities or at immune mechanisms, and, increasingly, pathways previously thought to be undruggable have become druggable. Critical to applying precision medicine is the concept that the right combination of drugs must be chosen for each patient and used at the right stage of the disease. Multiple puzzles remain that complicate therapy choice, including evidence that deleterious mutations are common in normal tissues and non-malignant conditions. The host’s role is also likely to be key in determining treatment response, especially for immunotherapy. Indeed, maximizing the impact of immunotherapy will require omic analyses to match the right immune-targeted drugs to the individualized patient and tumour setting. In this Perspective, we discuss six key riddles that must be solved to optimize the application of precision oncology to otherwise lethal malignancies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Christie, A. Peril at End House (Collins Crime Club, 1932).
Strebhardt, K. & Ullrich, A. Paul Ehrlich’s magic bullet concept: 100 years of progress. Nat. Rev. Cancer 8, 473–480 (2008).
Ehrlich, P. Experimental researches on specific therapeutics. Am. J. Med. Sci. 139, 432 (1910).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Wetterstrand, K. A. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP). Genome.gov https://www.genome.gov/about-genomics/fact-sheets/DNA-Sequencing-Costs-Data (2019).
Bower, H. et al. Life expectancy of patients with chronic myeloid leukemia approaches the life expectancy of the general population. J. Clin. Oncol. 34, 2851–2857 (2016).
Westin, J. R. & Kurzrock, R. It’s about time: lessons for solid tumors from chronic myelogenous leukemia therapy. Mol. Cancer Ther. 11, 2549–2555 (2012).
Hungerford, D. A. & Nowell, P. C. A minute chromosome in human chronic granulocytic leukemia. Science 132, 1013–1035 (1960). This article describes, for the first time, the alteration coined as the ‘Philadelphia chromosome’, the prototype of genetic defects linked to cancer.
Rowley, J. D. A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 243, 290–293 (1973).
Kloetzer, W. et al. The human cellular abl gene product in the chronic myelogenous leukemia cell line K562 has an associated tyrosine protein kinase activity. Virology 140, 230–238 (1985).
Shtivelman, E., Lifshitz, B., Gale, R. P. & Canaani, E. Fused transcript of abl and bcr genes in chronic myelogenous leukaemia. Nature 315, 550–554 (1985).
Druker, B. J. et al. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nat. Med. 2, 561–566 (1996).
Druker, B. J. et al. Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N. Engl. J. Med. 344, 1031–1037 (2001). This publication presents clinical phase I data for imatinib in CML.
Braun, T. P., Eide, C. A. & Druker, B. J. Response and resistance to BCR-ABL1-targeted therapies. Cancer Cell 37, 530–542 (2020).
Westin, J. R., Kantarjian, H. & Kurzrock, R. Treatment of chronic myelogenous leukemia as a paradigm for solid tumors: how targeted agents in newly diagnosed disease transformed outcomes. Am. Soc. Clin. Oncol. Educ. Book https://doi.org/10.14694/EdBook_AM.2012.32.60 (2012).
Cohen, P., Cross, D. & Jänne, P. A. Kinase drug discovery 20 years after imatinib: progress and future directions. Nat. Rev. Drug Discov. 20, 551–569 (2021).
Drilon, A. et al. Efficacy of selpercatinib in RET fusion-positive non-small-cell lung cancer. N. Engl. J. Med. 383, 813–824 (2020).
Drilon, A. et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N. Engl. J. Med. 378, 731–739 (2018).
Shaw, A. T. et al. First-line lorlatinib or crizotinib in advanced ALK-positive lung cancer. N. Engl. J. Med. 383, 2018–2029 (2020).
Soria, J.-C. et al. Osimertinib in untreated EGFR-mutated advanced non-small-cell lung cancer. N. Engl. J. Med. 378, 113–125 (2018).
Shaw, A. T. et al. Ceritinib in ALK-rearranged non-small-cell lung cancer. N. Engl. J. Med. 370, 1189–1197 (2014).
Sawyers, C. L. et al. Imatinib induces hematologic and cytogenetic responses in patients with chronic myelogenous leukemia in myeloid blast crisis: results of a phase II study. Blood 99, 3530–3539 (2002).
Gerstung, M. et al. The evolutionary history of 2658 cancers. Nature 578, 122–128 (2020).
Christie, A. And Then There Were None (Harper-Collins, 2008).
Vogelstein, B. & Kinzler, K. W. The path to cancer — three strikes and you’re out. N. Engl. J. Med. 373, 1895–1898 (2015).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990). This is a comprehensive review of the genetic model of colorectal tumorigenesis.
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Ding, L. et al. Perspective on oncogenic processes at the end of the beginning of cancer genomics. Cell 173, 305–320 (2018).
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Kato, S., Lippman, S. M., Flaherty, K. T. & Kurzrock, R. The conundrum of genetic “drivers” in benign conditions. J. Natl Cancer Inst. 108, djw036 (2016).
Adashek, J. J., Kato, S., Lippman, S. M. & Kurzrock, R. The paradox of cancer genes in non-malignant conditions: implications for precision medicine. Genome Med. 12, 16 (2020).
Anglesio, M. S. et al. Cancer-associated mutations in endometriosis without cancer. N. Engl. J. Med. 376, 1835–1848 (2017).
Yamanishi, Y. et al. Regional analysis of p53 mutations in rheumatoid arthritis synovium. Proc. Natl Acad. Sci. USA 99, 10025–10030 (2002).
Angelo, L. S., Talpaz, M. & Kurzrock, R. Autocrine interleukin-6 production in renal cell carcinoma: evidence for the involvement of p53. Cancer Res. 62, 932–940 (2002).
Zhang, T. et al. p53 predominantly regulates IL-6 production and suppresses synovial inflammation in fibroblast-like synoviocytes and adjuvant-induced arthritis. Arthritis Res. Ther. 18, 271 (2016).
Hlevnjak, M. et al. CATCH: a prospective precision oncology trial in metastatic breast cancer. JCO Precis. Oncol. 5, 676–686 (2021).
Horak, P. et al. Comprehensive genomic and transcriptomic analysis for guiding therapeutic decisions in patients with rare cancers. Cancer Discov. 11, 2780–2795 (2021).
van Tilburg, C. M. et al. The pediatric precision oncology INFORM registry: clinical outcome and benefit for patients with very high-evidence targets. Cancer Discov. 11, 2764–2779 (2021). Along with Hlevnjak et al. (2021) and Horak et al. (2021), this study presents convincing prospective data for molecularly informed targeted therapies, showing that precision oncology generates a real-world benefit for patients with cancer.
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015).
Olafsson, S. et al. Somatic evolution in non-neoplastic IBD-affected colon. Cell 182, 672–684 (2020).
Kumar, R., Angelini, S., Snellman, E. & Hemminki, K. BRAF mutations are common somatic events in melanocytic nevi. J. Invest. Dermatol. 122, 342–348 (2004).
Allred, D. C. et al. Overexpression of HER-2/neu and its relationship with other prognostic factors change during the progression of in situ to invasive breast cancer. Hum. Pathol. 23, 974–979 (1992).
Rakovitch, E. et al. HER2/neu and Ki-67 expression predict non-invasive recurrence following breast-conserving therapy for ductal carcinoma in situ. Br. J. Cancer 106, 1160–1165 (2012).
Williams, K. E. et al. Molecular phenotypes of DCIS predict overall and invasive recurrence. Ann. Oncol. 26, 1019–1025 (2015).
Cappellen, D. et al. Frequent activating mutations of FGFR3 in human bladder and cervix carcinomas. Nat. Genet. 23, 18–20 (1999).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Lee-Six, H. et al. The landscape of somatic mutation in normal colorectal epithelial cells. Nature 574, 532–537 (2019).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Jaiswal, S. & Ebert, B. L. Clonal hematopoiesis in human aging and disease. Science 366, eaan4673 (2019).
Christie, A. The Man in the Mist (Illustrated London News Company, 1924).
Adashek, J. J., Subbiah, V. & Kurzrock, R. From tissue-agnostic to N-of-one therapies: (r)evolution of the precision paradigm. Trends Cancer Res. 7, 15–28 (2021).
Turski, M. L. et al. Genomically driven tumors and actionability across histologies: BRAF-mutant cancers as a paradigm. Mol. Cancer Ther. 15, 533–547 (2016).
Flaherty, K. T. et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N. Engl. J. Med. 363, 809–819 (2010).
Tiacci, E. et al. Targeting mutant BRAF in relapsed or refractory hairy-cell leukemia. N. Engl. J. Med. 373, 1733–1747 (2015).
Falchook, G. S. et al. Dabrafenib in patients with melanoma, untreated brain metastases, and other solid tumours: a phase 1 dose-escalation trial. Lancet 379, 1893–1901 (2012).
Kopetz, S. et al. PLX4032 in metastatic colorectal cancer patients with mutant BRAF tumors. J. Clin. Oncol. 28 (Suppl. 15), 3534 (2010).
Prahallad, A. et al. Unresponsiveness of colon cancer to BRAFV600E inhibition through feedback activation of EGFR. Nature 483, 100–103 (2012).
Corcoran, R. B. et al. EGFR-mediated re-activation of MAPK signaling contributes to insensitivity of BRAF mutant colorectal cancers to RAF inhibition with vemurafenib. Cancer Discov. 2, 227–235 (2012).
Kopetz, S. et al. Encorafenib, binimetinib, and cetuximab in BRAF V600E-mutated colorectal cancer. N. Engl. J. Med. 381, 1632–1643 (2019).
Kurzrock, R. et al. A novel c-abl protein product in Philadelphia-positive acute lymphoblastic leukaemia. Nature 325, 631–635 (1987).
Christie, A. The Mysterious Affair At Styles (John Lane Company, 1921).
Bailey, C. et al. Tracking cancer evolution through the disease course. Cancer Discov. 11, 916–932 (2021).
Ortmann, C. A. et al. Effect of mutation order on myeloproliferative neoplasms. N. Engl. J. Med. 372, 601–612 (2015). This seminal work sheds light on the effects of mutation order on clinical course and therapy response in myeloproliferative neoplasms, providing evidence that clonal evolution has to be considered as a decision-shaping factor in molecular oncology.
Bozic, I. & Wu, C. J. Delineating the evolutionary dynamics of cancer from theory to reality. Nat. Cancer 1, 580–588 (2020).
TRACERx Renal consortium. TRACERx renal: tracking renal cancer evolution through therapy. Nat. Rev. Urol. 14, 575–576 (2017).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012). This study describes spatial intratumoural heterogeneity, implicating a string of consequences for precision oncology approaches.
Chen, J. et al. Genomic landscape of lung adenocarcinoma in East Asians. Nat. Genet. 52, 177–186 (2020). This work presents data showcasing groundbreaking differences in the genomic and transcriptomic analyses of East Asian patients with lung cancer compared with European patients with lung cancer, serving as an example for the impact of ethnicity in cancer research and therapy.
Dearden, S., Stevens, J., Wu, Y.-L. & Blowers, D. Mutation incidence and coincidence in non small-cell lung cancer: meta-analyses by ethnicity and histology (mutMap). Ann. Oncol. 24, 2371–2376 (2013).
Agboola, A. J. et al. Molecular characteristics and prognostic features of breast cancer in Nigerian compared with UK women. Breast Cancer Res. Treat. 135, 555–569 (2012).
Bollig-Fischer, A. et al. Racial diversity of actionable mutations in non-small cell lung cancer. J. Thorac. Oncol. 10, 250–255 (2015).
Mao, X. et al. Distinct genomic alterations in prostate cancers in Chinese and Western populations suggest alternative pathways of prostate carcinogenesis. Cancer Res. 70, 5207–5212 (2010).
Kadakia, K. C. & Salem, M. E. Role of immune checkpoint inhibitors in understudied populations. JCO Oncol. Pract. 17, 246–248 (2021).
zur Hausen, H. Papillomaviruses and cancer: from basic studies to clinical application. Nat. Rev. Cancer 2, 342–350 (2002).
Marur, S., D’Souza, G., Westra, W. H. & Forastiere, A. A. HPV-associated head and neck cancer: a virus-related cancer epidemic. Lancet Oncol. 11, 781–789 (2010).
Moody, S. et al. Mutational signatures in esophageal squamous cell carcinoma from eight countries with varying incidence. Nat. Genet. 53, 1553–1563 (2021).
Shigematsu, H. et al. Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J. Natl Cancer Inst. 97, 339–346 (2005).
Knerr, S., Wayman, D. & Bonham, V. L. Inclusion of racial and ethnic minorities in genetic research: advance the spirit by changing the rules? J. Law Med. Ethics 39, 502–512 (2011).
Spratt, D. E. et al. Racial/ethnic disparities in genomic sequencing. JAMA Oncol. 2, 1070–1074 (2016).
Li, C. H., Haider, S., Shiah, Y.-J., Thai, K. & Boutros, P. C. Sex differences in cancer driver genes and biomarkers. Cancer Res. 78, 5527–5537 (2018).
Conforti, F. et al. Cancer immunotherapy efficacy and patients’ sex: a systematic review and meta-analysis. Lancet Oncol. 19, 737–746 (2018).
Wang, S., Zhang, J., He, Z., Wu, K. & Liu, X.-S. The predictive power of tumor mutational burden in lung cancer immunotherapy response is influenced by patients’ sex. Int. J. Cancer 145, 2840–2849 (2019).
Li, C. H. et al. Sex differences in oncogenic mutational processes. Nat. Commun. 11, 4330 (2020).
Cullin, N., Azevedo Antunes, C., Straussman, R., Stein-Thoeringer, C. K. & Elinav, E. Microbiome and cancer. Cancer Cell 39, 1317–1341 (2021).
Fan, Y. & Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 19, 55–71 (2021).
Nejman, D. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science 368, 973–980 (2020).
Sepich-Poore, G. D. et al. The microbiome and human cancer. Science 371, eabc4552 (2021).
Pernigoni, N. et al. Commensal bacteria promote endocrine resistance in prostate cancer through androgen biosynthesis. Science 374, 216–224 (2021).
Kadosh, E. et al. The gut microbiome switches mutant p53 from tumour-suppressive to oncogenic. Nature 586, 133–138 (2020).
Greathouse, K. L. et al. Interaction between the microbiome and TP53 in human lung cancer. Genome Biol. 19, 123 (2018).
Hayase, E. & Jenq, R. R. Role of the intestinal microbiome and microbial-derived metabolites in immune checkpoint blockade immunotherapy of cancer. Genome Med. 13, 107 (2021).
Abid, M. B., Shah, N. N., Maatman, T. C. & Hari, P. N. Gut microbiome and CAR-T therapy. Exp. Hematol. Oncol. 8, 31 (2019).
Lee, K. A. et al. Cross-cohort gut microbiome associations with immune checkpoint inhibitor response in advanced melanoma. Nat. Med. 28, 535–544 (2022).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018). Along with Routy et al. (Science, 2018), this study that shows how the composition of the microbiota is able to modulate the response to checkpoint therapy.
Routy, B. et al. The gut microbiota influences anticancer immunosurveillance and general health. Nat. Rev. Clin. Oncol. 15, 382–396 (2018).
Baruch, E. N. et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 371, 602–609 (2021). This seminal work shows the reversal of therapy refractiveness using faecal microbiota transplant.
Davar, D. et al. Fecal microbiota transplant overcomes resistance to anti-PD-1 therapy in melanoma patients. Science 371, 595–602 (2021).
Elkrief, A. & Routy, B. First clinical proof-of-concept that FMT can overcome resistance to ICIs. Nat. Rev. Clin. Oncol. 18, 325–326 (2021).
Christie, A. Murder on the Orient Express (Collins Crime Club, 1934).
Coley, W. B. The treatment of malignat tumors by repeated inoculations of erysipelas. Am. J. Med. Sci. 105, 487–510 (1893).
Hernández-Ramírez, R. U., Shiels, M. S., Dubrow, R. & Engels, E. A. Cancer risk in HIV-infected people in the USA from 1996 to 2012: a population-based, registry-linkage study. Lancet HIV 4, e495–e504 (2017).
Galanina, N., Goodman, A. M., Cohen, P. R., Frampton, G. M. & Kurzrock, R. Successful treatment of HIV-associated Kaposi Sarcoma with immune checkpoint blockade. Cancer Immunol. Res. 6, 1129–1135 (2018).
Mortaz, E. et al. Cancers related to immunodeficiencies: update and perspectives. Front. Immunol. 7, 365 (2016).
Marcus, L., Lemery, S. J., Keegan, P. & Pazdur, R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin. Cancer Res. 25, 3753–3758 (2019).
André, T. et al. Pembrolizumab in microsatellite-instability-high advanced colorectal cancer. N. Engl. J. Med. 383, 2207–2218 (2020).
Jardim, D. L., Goodman, A., de Melo Gagliato, D. & Kurzrock, R. The challenges of tumor mutational burden as an immunotherapy biomarker. Cancer Cell 39, 154–173 (2021).
Kato, S. et al. Expression of TIM3/VISTA checkpoints and the CD68 macrophage-associated marker correlates with anti-PD1/PDL1 resistance: implications of immunogram heterogeneity. Oncoimmunology 9, 1708065 (2020).
Adashek, J. J., Goloubev, A., Kato, S. & Kurzrock, R. Missing the target in cancer therapy. Nat. Cancer 2, 369–371 (2021).
Boichard, A. et al. APOBEC-related mutagenesis and neo-peptide hydrophobicity: implications for response to immunotherapy. Oncoimmunology 8, 1550341 (2019).
Pham, T. V. et al. Role of ultraviolet mutational signature versus tumor mutation burden in predicting response to immunotherapy. Mol. Oncol. 14, 1680–1694 (2020).
Goodman, A. M. et al. MHC-I genotype and tumor mutational burden predict response to immunotherapy. Genome Med. 12, 45 (2020).
Zamora, A. E., Crawford, J. C. & Thomas, P. G. Hitting the target: how T cells detect and eliminate tumors. J. Immunol. 200, 392–399 (2018).
Valpione, S. et al. The T cell receptor repertoire of tumor infiltrating T cells is predictive and prognostic for cancer survival. Nat. Commun. 12, 4098 (2021).
Bassani-Sternberg, M. et al. Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry. Nat. Commun. 7, 13404 (2016).
Gibbons, D. L. et al. 57O Efficacy, safety and tolerability of MEDI4736 (durvalumab [D]), a human IgG1 anti-programmed cell death-ligand-1 (PD-L1) antibody, combined with gefitinib (G): a phase I expansion in TKI-naïve patients (pts) with EGFR mutant NSCLC. J. Thorac. Oncol. 11, S79 (2016).
Felip, E. et al. Ceritinib plus nivolumab (NIVO) in patients (pts) with anaplastic lymphoma kinase positive (ALK+) advanced non-small cell lung cancer (NSCLC). J. Clin. Oncol. 35 (Suppl. 15), 2502 (2017).
Nasser, N. J., Gorenberg, M. & Agbarya, A. First line Immunotherapy for non-small cell lung cancer. Pharmaceuticals 13, 373 (2020).
Cercek, A. et al. PD-1 blockade in mismatch repair-deficient, locally advanced rectal cancer. N. Engl. J. Med. 386, 2363–2376 (2022). In this study, the use of PD1 inhibition alone is sufficient to provoke durable responses in locally advanced rectal cancer, illustrating the potential use of checkpoint inhibition as a stand-alone therapy.
Christie, A. & Fraser, H. The A.B.C. Murders (Harper-Collins, 1936).
Hong, D. S. et al. KRASG12C inhibition with sotorasib in advanced solid tumors. N. Engl. J. Med. 383, 1207–1217 (2020).
Rodon, J. et al. Genomic and transcriptomic profiling expands precision cancer medicine: the WINTHER trial. Nat. Med. 25, 751–758 (2019).
Schettini, F. et al. Identification of cell surface targets for CAR-T cell therapies and antibody-drug conjugates in breast cancer. ESMO Open 6, 100102 (2021).
Wang, J., Dean, D. C., Hornicek, F. J., Shi, H. & Duan, Z. RNA sequencing (RNA-Seq) and its application in ovarian cancer. Gynecol. Oncol. 152, 194–201 (2019).
Rogawski, D. S., Vitanza, N. A., Gauthier, A. C., Ramaswamy, V. & Koschmann, C. Integrating RNA sequencing into neuro-oncology practice. Transl Res. 189, 93–104 (2017).
Sicklick, J. K. et al. Molecular profiling of cancer patients enables personalized combination therapy: the I-PREDICT study. Nat. Med. 25, 744–750 (2019).
Kato, S. et al. Real-world data from a molecular tumor board demonstrates improved outcomes with a precision N-of-one strategy. Nat. Commun. 11, 4965 (2020).
Sicklick, J. K. et al. Molecular profiling of advanced malignancies guides first-line N-of-1 treatments in the I-PREDICT treatment-naïve study. Genome Med. 13, 155 (2021). The I-PREDICT trial is the first precision medicine trial to provide matched individualized (N-of-1) combination therapies to patients; a higher degree of matching correlated with improvement in all outcome parameters.
Jänne, P. A. et al. Adagrasib in non-small-cell lung cancer harboring a KRASG12C mutation. N. Engl. J. Med. 387, 120–131 (2022). Together with Hong et al. (2020), this is important clinical work showing the response of the first KRASG12C inhibitors, a target long thought to be part of the ‘undruggable’ realm.
Pang, Y. et al. Report of canonical BCR-ABL1 fusion in glioblastoma. JCO Precis. Oncol. 5, 1348–1353 (2021).
Schwaederle, M. et al. Impact of precision medicine in diverse cancers: a meta-analysis of phase II clinical trials. J. Clin. Oncol. 33, 3817–3825 (2015).
Schneider, G., Schmidt-Supprian, M., Rad, R. & Saur, D. Tissue-specific tumorigenesis: context matters. Nat. Rev. Cancer 17, 239–253 (2017).
Sharma, P. et al. The next decade of immune checkpoint therapy. Cancer Discov. 11, 838–857 (2021).
Nakamura, Y. et al. Clinical utility of circulating tumor DNA sequencing in advanced gastrointestinal cancer: SCRUM-Japan GI-SCREEN and GOZILA studies. Nat. Med. 26, 1859–1864 (2020).
Tie, J. et al. Circulating tumor DNA analysis guiding adjuvant therapy in stage II colon cancer. N. Engl. J. Med. 386, 2261–2272 (2022).
Malhotra, H., Radich, J. & Garcia-Gonzalez, P. Meeting the needs of CML patients in resource-poor countries. Hematol. Am. Soc. Hematol. Educ. Program. 2019, 433–442 (2019).
Henke, O., Mapendo, P. J., Mkwizu, E. W. & le Coutre, P. Early molecular response in East African Philadelphia chromosome-positive chronic myeloid leukaemia patients treated with Imatinib and barriers to access treatment. Ecancermedicalscience 14, 1089 (2020).
Nasser, A. et al. Molecular response to imatinib in patients with chronic myeloid leukemia in Tanzania. Blood Adv. 5, 1403–1411 (2021).
Acknowledgements
The authors apologize to all their esteemed colleagues whose work could not be included owing to space constraints. This Perspective was inspired by D. R. Green’s work “The coming decade of cell death research: five riddles” (Cell 177, 1094–1107 (2019)). A.W. and L.B. are supported by the Claudia von Schilling foundation. A.W. was supported by the Torsten-Haferlach Leukaemia Diagnostics Foundation. P.L. and S.F. are supported by the Molecular Precision Oncology Program of the National Center for Tumour Diseases in Heidelberg. S.F. is supported by the German Cancer Consortium. The authors are grateful for critical discussions with D. R. Green, P. Vandenabeele, J. C. A. Melms, M. Bostock, M. Hlevnjak and J. P. Suppelna. The National Center for Tumor Diseases (NCT) Heidelberg is a partnership between the German Cancer Research Center (DKFZ) and Heidelberg University Hospital.
Author information
Authors and Affiliations
Contributions
A.W., L.B. and R.K. conceived and wrote the manuscript. S.F., P.J.J., A.S. and P.L. provided conceptual support and critically reviewed the manuscript draft.
Corresponding authors
Ethics declarations
Competing interests
A.W., L.B. and P.L. declare no competing interests. S.F. has received consulting or advisory honoraria from Bayer, Illumina and Roche, honoraria from Eli Lilly, PharmaMar and Roche, research funding from AstraZeneca, Pfizer, PharmaMar and Roche, and travel or accommodation expenses from Eli Lilly, Illumina, PharmaMar and Roche. P.J.J. has had a consulting or advisory role for and has received honoraria, research funding and/or travel/accommodation expenses from Ariad, Abbvie, Bayer, Boehringer Ingelheim, Novartis, Pfizer, Servier, Roche, Celgene (Bristol Myers Squibb), Pierre Fabre, Janssen (Johnson & Johnson) and MSD. A.S. has received research grants from Celgene, Roche and AbbVie, honoraria from Roche, Celgene, Pfizer, AstraZeneca, Novartis, MSD, Tesaro, Eli Lilly, GlaxoSmithKline, Seagen, Gilead Sciences, Bayer, Amgen, Pierre Fabre, streamedup!, promedicis, onkowissen.de, Metaplan and Connect Media, and travel support from Roche, Celgene and Pfizer. R.K. has received research funding from Biological Dynamics, Boehringer Ingelheim, Debiopharm, Foundation Medicine, Genentech, Grifols, Guardant Health, Incyte, Konica Minolta, Medimmune (AstraZeneca), Merck Serono, Omniseq, Pfizer, Sequenom, Takeda and TopAlliance, has received consultant and/or speaker fees and/or has been an advisory board member for Actuate Therapeutics, AstraZeneca, Bicara Therapeutics, Biological Dynamics, Daiichi Sankyo, Eisai, EOM Pharmaceuticals, Iylon, Merck, NeoGenomics, NEOMED, Pfizer, Prosperdtx, Roche, TD2/Volastra, Turning Point Therapeutics and X-Biotech, has an equity interest in CureMatch, CureMetrix and IDbyDNA, serves on the board of CureMatch and CureMetrix, and co-founded CureMatch.
Peer review
Peer review information
Nature Reviews Cancer thanks David Solit and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wahida, A., Buschhorn, L., Fröhling, S. et al. The coming decade in precision oncology: six riddles. Nat Rev Cancer 23, 43–54 (2023). https://doi.org/10.1038/s41568-022-00529-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-022-00529-3
This article is cited by
-
Multifunctional nanoparticle-mediated combining therapy for human diseases
Signal Transduction and Targeted Therapy (2024)
-
Präzisionsmedizin in der Onkologie
Die Innere Medizin (2024)
-
Molecular diagnostics tailoring personalized cancer therapy—an oncologist’s view
Virchows Archiv (2024)
-
The application of Aptamer in biomarker discovery
Biomarker Research (2023)
-
Are sex and gender considered in head and neck cancer clinical studies?
npj Precision Oncology (2023)