Abstract
With the genetic portraits of all major human malignancies now available, we next face the challenge of characterizing the function of mutated genes, their downstream targets, interactions and molecular networks. Moreover, poorly understood at the functional level are also non-mutated but dysregulated genomes, epigenomes or transcriptomes. Breakthroughs in manipulative mouse genetics offer new opportunities to probe the interplay of molecules, cells and systemic signals underlying disease pathogenesis in higher organisms. Herein, we review functional screening strategies in mice using genetic perturbation and chemical mutagenesis. We outline the spectrum of genetic tools that exist, such as transposons, CRISPR and RNAi and describe discoveries emerging from their use. Genome-wide or targeted screens are being used to uncover genomic and regulatory landscapes in oncogenesis, metastasis or drug resistance. Versatile screening systems support experimentation in diverse genetic and spatio-temporal settings to integrate molecular, cellular or environmental context-dependencies. We also review the combination of in vivo screening and barcoding strategies to study genetic interactions and quantitative cancer dynamics during tumour evolution. These scalable functional genomics approaches are transforming our ability to interrogate complex biological systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ding, L. et al. Perspective on oncogenic processes at the end of the beginning of cancer genomics. Cell 173, 305–320.e10 (2018).
Nangalia, J. & Campbell, P. J. Genome sequencing during a patient’s journey through cancer. N. Engl. J. Med. 381, 2145–2156 (2019).
Schneider, G., Schmidt-Supprian, M., Rad, R. & Saur, D. Tissue-specific tumorigenesis: context matters. Nat. Rev. Cancer 17, 239–253 (2017).
Frese, K. K. & Tuveson, D. A. Maximizing mouse cancer models. Nat. Rev. Cancer 7, 654–658 (2007).
Weber, J. & Rad, R. Engineering CRISPR mouse models of cancer. Curr. Opin. Genet. Dev. 54, 88–96 (2019).
Kemp, C. J. Animal models of chemical carcinogenesis: driving breakthroughs in cancer research for 100 years. Cold Spring Harb. Protoc. 2015, 865–874 (2015). This paper is a historical perspective and methodological overview of chemical mutagenesis in animal models.
Warren, M. et al. Irradiated Blm-deficient mice are a highly tumor prone model for analysis of a broad spectrum of hematologic malignancies. Leukemia Res. 34, 210–220 (2010).
Sherborne, A. L. et al. Mutational analysis of ionizing radiation induced neoplasms. Cell Rep. 12, 1915–1926 (2015).
Lee, C. L. et al. Mutational landscape in genetically engineered, carcinogen-induced, and radiation-induced mouse sarcoma. JCI Insight 4, e128698 (2019).
Westcott, P. M. et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 517, 489–492 (2015).
Connor, F. et al. Mutational landscape of a chemically-induced mouse model of liver cancer. J. Hepatol. 69, 840–850 (2018).
McCreery, M. Q. et al. Evolution of metastasis revealed by mutational landscapes of chemically induced skin cancers. Nat. Med. 21, 1514–1520 (2015).
Nassar, D., Latil, M., Boeckx, B., Lambrechts, D. & Blanpain, C. Genomic landscape of carcinogen-induced and genetically induced mouse skin squamous cell carcinoma. Nat. Med. 21, 946–954 (2015).
Lange, S. et al. Analysis pipelines for cancer genome sequencing in mice. Nat. Protoc. 15, 266–315 (2020).
Lilue, J. et al. Sixteen diverse laboratory mouse reference genomes define strain-specific haplotypes and novel functional loci. Nat. Genet. 50, 1574–1583 (2018).
Hayward, W. S., Neel, B. G. & Astrin, S. M. Activation of a cellular onc gene by promoter insertion in ALV-induced lymphoid leukosis. Nature 290, 475–480 (1981).
Nusse, R. & Varmus, H. E. Many tumors induced by the mouse mammary tumor virus contain a provirus integrated in the same region of the host genome. Cell 31, 99–109 (1982).
Kool, J. & Berns, A. High-throughput insertional mutagenesis screens in mice to identify oncogenic networks. Nat. Rev. Cancer 9, 389–399 (2009).
Morishita, K. et al. Retroviral activation of a novel gene encoding a zinc finger protein in IL-3-dependent myeloid leukemia cell lines. Cell 54, 831–840 (1988).
van Lohuizen, M. et al. Identification of cooperating oncogenes in E mu-myc transgenic mice by provirus tagging. Cell 65, 737–752 (1991).
Suzuki, T. et al. New genes involved in cancer identified by retroviral tagging. Nat. Genet. 32, 166–174 (2002).
Theodorou, V. et al. MMTV insertional mutagenesis identifies genes, gene families and pathways involved in mammary cancer. Nat. Genet. 39, 759–769 (2007).
Girard, L. et al. Frequent provirus insertional mutagenesis of Notch1 in thymomas of MMTVD/myc transgenic mice suggests a collaboration of c-myc and Notch1 for oncogenesis. Genes Dev. 10, 1930–1944 (1996).
van der Lugt, N. M. et al. Proviral tagging in E mu-myc transgenic mice lacking the Pim-1 proto-oncogene leads to compensatory activation of Pim-2. EMBO J. 14, 2536–2544 (1995).
Mikkers, H. et al. High-throughput retroviral tagging to identify components of specific signaling pathways in cancer. Nat. Genet. 32, 153–159 (2002). This elegant study coins the concept of complementation tagging to screen for genes with functional homology.
Ranzani, M. et al. Lentiviral vector-based insertional mutagenesis identifies genes associated with liver cancer. Nat. Methods 10, 155–161 (2013).
Johansson, F. K. et al. Identification of candidate cancer-causing genes in mouse brain tumors by retroviral tagging. Proc. Natl Acad. Sci. USA 101, 11334 (2004).
Jorgensen, E. M. & Mango, S. E. The art and design of genetic screens: Caenorhabditis elegans. Nat. Rev. Genet. 3, 356–369 (2002).
St Johnston, D. The art and design of genetic screens: Drosophila melanogaster. Nat. Rev. Genet. 3, 176–188 (2002).
Thibault, S. T. et al. A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nat. Genet. 36, 283–287 (2004).
Ivics, Z., Hackett, P. B., Plasterk, R. H. & Izsvak, Z. Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells. Cell 91, 501–510 (1997). This landmark study reconstructs synthetic Sleeping Beauty transposons from ancient dormant transposable elements in fish genomes for transposition in mammalian cells.
Kawakami, K., Shima, A. & Kawakami, N. Identification of a functional transposase of the Tol2 element, an Ac-like element from the Japanese medaka fish, and its transposition in the zebrafish germ lineage. Proc. Natl Acad. Sci. USA 97, 11403–11408 (2000).
Ding, S. et al. Efficient transposition of the PiggyBac (PB) transposon in mammalian cells and mice. Cell 122, 473–483 (2005). This paper demonstrates efficient transposition of the PiggyBac system in mammalian cell lines and the mouse germ line.
Luo, G., Ivics, Z., Izsvák, Z. & Bradley, A. Chromosomal transposition of a Tc1/mariner-like element in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 95, 10769 (1998).
Dupuy, A. J., Fritz, S. & Largaespada, D. A. Transposition and gene disruption in the male germline of the mouse. Genesis 30, 82–88 (2001).
Fischer, S. E., Wienholds, E. & Plasterk, R. H. Regulated transposition of a fish transposon in the mouse germ line. Proc. Natl Acad. Sci. USA 98, 6759–6764 (2001).
Horie, K. et al. Efficient chromosomal transposition of a Tc1/mariner-like transposon Sleeping Beauty in mice. Proc. Natl Acad. Sci. USA 98, 9191–9196 (2001).
Collier, L. S., Carlson, C. M., Ravimohan, S., Dupuy, A. J. & Largaespada, D. A. Cancer gene discovery in solid tumours using transposon-based somatic mutagenesis in the mouse. Nature 436, 272–276 (2005).
Dupuy, A. J., Akagi, K., Largaespada, D. A., Copeland, N. G. & Jenkins, N. A. Mammalian mutagenesis using a highly mobile somatic Sleeping Beauty transposon system. Nature 436, 221–226 (2005). Together with Collier et al. (2005), this pioneering study demonstrates somatic mobilization of engineered Sleeping Beauty transposons for insertional mutagenesis and cancer gene discovery in mice.
Rad, R. et al. PiggyBac transposon mutagenesis: a tool for cancer gene discovery in mice. Science 330, 1104–1107 (2010). This is the first study to demonstrate PiggyBac transposon-based genetic screening in mice.
Wang, W. et al. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 105, 9290–9295 (2008).
Liang, Q., Kong, J., Stalker, J. & Bradley, A. Chromosomal mobilization and reintegration of Sleeping Beauty and PiggyBac transposons. Genesis 47, 404–408 (2009).
de Jong, J. et al. Chromatin landscapes of retroviral and transposon integration profiles. PLoS Genet. 10, e1004250 (2014).
Tipanee, J., Chai, Y. C., VandenDriessche, T. & Chuah, M. K. Preclinical and clinical advances in transposon-based gene therapy. Biosci. Rep. 37, BSR20160614 (2017).
Yoshida, J. et al. Chromatin states shape insertion profiles of the piggyBac, Tol2 and Sleeping Beauty transposons and murine leukemia virus. Sci. Rep. 7, 43613 (2017).
Landrette, S. F., Cornett, J. C., Ni, T. K., Bosenberg, M. W. & Xu, T. piggyBac transposon somatic mutagenesis with an activated reporter and tracker (PB-SMART) for genetic screens in mice. PLoS One 6, e26650 (2011).
Vassiliou, G. S. et al. Mutant nucleophosmin and cooperating pathways drive leukemia initiation and progression in mice. Nat. Genet. 43, 470–475 (2011).
Dupuy, A. J. et al. A modified Sleeping Beauty transposon system that can be used to model a wide variety of human cancers in mice. Cancer Res. 69, 8150–8156 (2009).
Starr, T. K. et al. A transposon-based genetic screen in mice identifies genes altered in colorectal cancer. Science 323, 1747–1750 (2009).
Rad, R. et al. A conditional piggyBac transposition system for genetic screening in mice identifies oncogenic networks in pancreatic cancer. Nat. Genet. 47, 47–56 (2015).
Genovesi, L. A. et al. Sleeping Beauty mutagenesis in a mouse medulloblastoma model defines networks that discriminate between human molecular subgroups. Proc. Natl Acad. Sci. USA 110, E4325–E4334 (2013).
Abbott, K. L. et al. The candidate cancer gene database: a database of cancer driver genes from forward genetic screens in mice. Nucleic Acids Res. 43, D844–D848 (2014).
Newberg, J. Y., Mann, K. M., Mann, M. B., Jenkins, N. A. & Copeland, N. G. SBCDDB: Sleeping Beauty Cancer Driver Database for gene discovery in mouse models of human cancers. Nucleic Acids Res. 46, D1011–D1017 (2018).
Rahrmann, E. P. et al. Forward genetic screen for malignant peripheral nerve sheath tumor formation identifies new genes and pathways driving tumorigenesis. Nat. Genet. 45, 756–766 (2013).
Tang, J. Z. et al. Transposon mutagenesis reveals cooperation of ETS family transcription factors with signaling pathways in erythro-megakaryocytic leukemia. Proc. Natl Acad. Sci. USA 110, 6091–6096 (2013).
de la Rosa, J. et al. A single-copy Sleeping Beauty transposon mutagenesis screen identifies new PTEN-cooperating tumor suppressor genes. Nat. Genet. 49, 730–741 (2017).
Takeda, H. et al. Transposon mutagenesis identifies genes and evolutionary forces driving gastrointestinal tract tumor progression. Nat. Genet. 47, 142–150 (2015).
March, H. N. et al. Insertional mutagenesis identifies multiple networks of cooperating genes driving intestinal tumorigenesis. Nat. Genet. 43, 1202–1209 (2011).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Mann, M. B. et al. Transposon mutagenesis identifies genetic drivers of Braf(V600E) melanoma. Nat. Genet. 47, 486–495 (2015).
Kas, S. M. et al. Insertional mutagenesis identifies drivers of a novel oncogenic pathway in invasive lobular breast carcinoma. Nat. Genet. 49, 1219–1230 (2017).
Keng, V. W. et al. A conditional transposon-based insertional mutagenesis screen for genes associated with mouse hepatocellular carcinoma. Nat. Biotechnol. 27, 264–274 (2009).
Berger, A. H., Knudson, A. G. & Pandolfi, P. P. A continuum model for tumour suppression. Nature 476, 163–169 (2011).
Luo, G. et al. Cancer predisposition caused by elevated mitotic recombination in Bloom mice. Nat. Genet. 26, 424–429 (2000).
Suzuki, T., Minehata, K., Akagi, K., Jenkins, N. A. & Copeland, N. G. Tumor suppressor gene identification using retroviral insertional mutagenesis in Blm-deficient mice. EMBO J. 25, 3422–3431 (2006). This elegant study uses a hypomorphic Blm allele to induce LOH of retrovirally inactivated tumour suppressor genes.
Weber, J. et al. PiggyBac transposon tools for recessive screening identify B-cell lymphoma drivers in mice. Nat. Commun. 10, 1415 (2019).
Wu, X. et al. Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482, 529–533 (2012).
Moriarity, B. S. et al. A Sleeping Beauty forward genetic screen identifies new genes and pathways driving osteosarcoma development and metastasis. Nat. Genet. 47, 615–624 (2015).
Koudijs, M. J. et al. High-throughput semiquantitative analysis of insertional mutations in heterogeneous tumors. Genome Res. 21, 2181–2189 (2011).
Friedel, R. H. et al. Clonal expansion analysis of transposon insertions by high-throughput sequencing identifies candidate cancer genes in a PiggyBac mutagenesis screen. PLoS One 8, e72338 (2013).
Mann, K. M. et al. Analyzing tumor heterogeneity and driver genes in single myeloid leukemia cells with SBCapSeq. Nat. Biotechnol. 34, 962–972 (2016).
Friedrich, M. J. et al. Genome-wide transposon screening and quantitative insertion site sequencing for cancer gene discovery in mice. Nat. Protoc. 12, 289–309 (2017).
Eser, S. et al. Selective requirement of PI3K/PDK1 signaling for Kras oncogene-driven pancreatic cell plasticity and cancer. Cancer Cell 23, 406–420 (2013).
Sutherland, K. D. & Berns, A. Cell of origin of lung cancer. Mol. Oncol. 4, 397–403 (2010).
Berquam-Vrieze, K. E. et al. Cell of origin strongly influences genetic selection in a mouse model of T-ALL. Blood 118, 4646–4656 (2011).
Bender, A. M. et al. Sleeping Beauty-mediated somatic mutagenesis implicates CSF1 in the formation of high-grade astrocytomas. Cancer Res. 70, 3557–3565 (2010).
Hong, I.-S. Stimulatory versus suppressive effects of GM-CSF on tumor progression in multiple cancer types. Exp. Mol. Med. 48, e242–e242 (2016).
Bard-Chapeau, E. A. et al. Transposon mutagenesis identifies genes driving hepatocellular carcinoma in a chronic hepatitis B mouse model. Nat. Genet. 46, 24–32 (2014).
Kodama, T. et al. Molecular profiling of nonalcoholic fatty liver disease-associated hepatocellular carcinoma using SB transposon mutagenesis. Proc. Natl Acad. Sci. USA 115, E10417–E10426 (2018).
Tschida, B. R. et al. Sleeping Beauty insertional mutagenesis in mice identifies drivers of steatosis-associated hepatic tumors. Cancer Res. 77, 6576–6588 (2017).
Riordan, J. D. et al. Chronic liver injury alters driver mutation profiles in hepatocellular carcinoma in mice. Hepatology 67, 924–939 (2018).
Rogers, L. M., Olivier, A. K., Meyerholz, D. K. & Dupuy, A. J. Adaptive immunity does not strongly suppress spontaneous tumors in a Sleeping Beauty model of cancer. J. Immunol. 190, 4393 (2013).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Lee, J. T. Epigenetic regulation by long noncoding RNAs. Science 338, 1435–1439 (2012).
Wartewig, T. et al. PD-1 is a haploinsufficient suppressor of T cell lymphomagenesis. Nature 552, 121–125 (2017).
Ratner, L., Waldmann, T. A., Janakiram, M. & Brammer, J. E. Rapid progression of adult T-cell leukemia-lymphoma after PD-1 inhibitor therapy. N. Engl. J. Med. 378, 1947–1948 (2018).
Chapeau, E. A. et al. Resistance mechanisms to TP53-MDM2 inhibition identified by in vivo piggyBac transposon mutagenesis screen in an Arf–/– mouse model. Proc. Natl Acad. Sci. USA 114, 3151–3156 (2017).
Perna, D. et al. BRAF inhibitor resistance mediated by the AKT pathway in an oncogenic BRAF mouse melanoma model. Proc. Natl Acad. Sci. USA 112, E536–E545 (2015).
Kas, S. M. et al. Transcriptomics and transposon mutagenesis identify multiple mechanisms of resistance to the FGFR inhibitor AZD4547. Cancer Res. 78, 5668–5679 (2018).
Morrissy, A. S. et al. Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529, 351–357 (2016).
Bertrand, K. C. et al. A functional genomics approach to identify pathways of drug resistance in medulloblastoma. Acta Neuropathol. Commun. 6, 146 (2018).
Molyneux, S. D. et al. Human somatic cell mutagenesis creates genetically tractable sarcomas. Nat. Genet. 46, 964–972 (2014).
Shultz, L. D., Ishikawa, F. & Greiner, D. L. Humanized mice in translational biomedical research. Nat. Rev. Immunol. 7, 118–130 (2007).
Bishop, J. M. Cellular oncogenes and retroviruses. Annu. Rev. Biochem. 52, 301–354 (1983).
Rayner, J. R. & Gonda, T. J. A simple and efficient procedure for generating stable expression libraries by cDNA cloning in a retroviral vector. Mol. Cell. Biol. 14, 880–887 (1994).
Whitehead, I., Kirk, H. & Kay, R. Expression cloning of oncogenes by retroviral transfer of cDNA libraries. Mol. Cell. Biol. 15, 704–710 (1995).
Goshima, N. et al. Human protein factory for converting the transcriptome into an in vitro-expressed proteome. Nat. Methods 5, 1011–1017 (2008).
Team, M. G. C. P. et al. The completion of the Mammalian Gene Collection (MGC). Genome Res. 19, 2324–2333 (2009).
Yang, X. et al. A public genome-scale lentiviral expression library of human ORFs. Nat. Methods 8, 659–661 (2011).
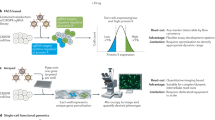
Sack, L. M. et al. Profound tissue specificity in proliferation control underlies cancer drivers and aneuploidy patterns. Cell 173, 499–514.e23 (2018). This ORF-based study screens the functional impact of over 16,000 genes on proliferation in vitro followed by a validation screen in an in situ breast cancer transplantation model.
Tsang, Y. H. et al. Functional annotation of rare gene aberration drivers of pancreatic cancer. Nat. Commun. 7, 10500 (2016).
Berger, A. H. et al. High-throughput phenotyping of lung cancer somatic mutations. Cancer Cell 32, 884 (2017).
Gao, H. et al. Forward genetic screens in mice uncover mediators and suppressors of metastatic reactivation. Proc. Natl Acad. Sci. USA 111, 16532–16537 (2014).
Sawey, E. T. et al. Identification of a therapeutic strategy targeting amplified FGF19 in liver cancer by oncogenomic screening. Cancer Cell 19, 347–358 (2011).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Martin, S. E. & Caplen, N. J. Applications of RNA interference in mammalian systems. Annu. Rev. Genomics Hum. Genet. 8, 81–108 (2007).
Boutros, M. & Ahringer, J. The art and design of genetic screens: RNA interference. Nat. Rev. Genet. 9, 554–566 (2008).
Watanabe, C., Cuellar, T. L. & Haley, B. Quantitative evaluation of first, second, and third generation hairpin systems reveals the limit of mammalian vector-based RNAi. RNA Biol. 13, 25–33 (2016).
Yeddula, N., Xia, Y., Ke, E., Beumer, J. & Verma, I. M. Screening for tumor suppressors: loss of ephrin receptor A2 cooperates with oncogenic KRas in promoting lung adenocarcinoma. Proc. Natl Acad. Sci. USA 112, E6476–E6485 (2015).
Beronja, S. et al. RNAi screens in mice identify physiological regulators of oncogenic growth. Nature 501, 185–190 (2013). This study performs a large-scale in vivo RNAi screen injecting more than 70,000 pooled shRNAs in utero to identify genes involved in epidermal growth and transformation and context-dependent effects of β-catenin.
Rudalska, R. et al. In vivo RNAi screening identifies a mechanism of sorafenib resistance in liver cancer. Nat. Med. 20, 1138–1146 (2014).
Dauch, D. et al. A MYC-aurora kinase A protein complex represents an actionable drug target in p53-altered liver cancer. Nat. Med. 22, 744–753 (2016).
Mullenders, J. & Bernards, R. Loss-of-function genetic screens as a tool to improve the diagnosis and treatment of cancer. Oncogene 28, 4409–4420 (2009).
Gargiulo, G., Serresi, M., Cesaroni, M., Hulsman, D. & van Lohuizen, M. In vivo shRNA screens in solid tumors. Nat. Protoc. 9, 2880–2902 (2014).
Zender, L. et al. An oncogenomics-based in vivo RNAi screen identifies tumor suppressors in liver cancer. Cell 135, 852–864 (2008). This study shows the feasibility of RNAi screens in mouse models using ex vivo manipulated liver progenitor cells.
Xue, W. et al. A cluster of cooperating tumor-suppressor gene candidates in chromosomal deletions. Proc. Natl Acad. Sci. USA 109, 8212–8217 (2012).
Gumireddy, K. et al. KLF17 is a negative regulator of epithelial–mesenchymal transition and metastasis in breast cancer. Nat. Cell Biol. 11, 1297–1304 (2009).
Murugaesu, N. et al. An in vivo functional screen identifies ST6GalNAc2 sialyltransferase as a breast cancer metastasis suppressor. Cancer Discov. 4, 304–317 (2014).
Scuoppo, C. et al. A tumour suppressor network relying on the polyamine–hypusine axis. Nature 487, 244–248 (2012).
Bric, A. et al. Functional identification of tumor-suppressor genes through an in vivo RNA interference screen in a mouse lymphoma model. Cancer Cell 16, 324–335 (2009).
Meacham, C. E., Ho, E. E., Dubrovsky, E., Gertler, F. B. & Hemann, M. T. In vivo RNAi screening identifies regulators of actin dynamics as key determinants of lymphoma progression. Nat. Genet. 41, 1133–1137 (2009).
Gargiulo, G. et al. In vivo RNAi screen for BMI1 targets identifies TGF-β/BMP–ER stress pathways as key regulators of neural- and malignant glioma-stem cell homeostasis. Cancer Cell 23, 660–676 (2013).
Vu, L. P. et al. Functional screen of MSI2 interactors identifies an essential role for SYNCRIP in myeloid leukemia stem cells. Nat. Genet. 49, 866–875 (2017).
Lujambio, A. & Banito, A. Functional screening to identify senescence regulators in cancer. Curr. Opin. Genet. Dev. 54, 17–24 (2019).
Braun, C. J. et al. Coordinated splicing of regulatory detained introns within oncogenic transcripts creates an exploitable vulnerability in malignant glioma. Cancer Cell 32, 411–426.e11 (2017).
Braun, C. J. & Hemann, M. T. Functional screens identify coordinators of RNA molecule birth, life, and death as targetable cancer vulnerabilities. Curr. Opin. Genet. Dev. 54, 105–109 (2019).
Ge, Y. et al. The splicing factor RBM25 controls MYC activity in acute myeloid leukemia. Nat. Commun. 10, 172 (2019).
Wucherpfennig, K. W. & Cartwright, A. N. Genetic screens to study the immune system in cancer. Curr. Opin. Immunol. 41, 55–61 (2016).
Zhou, P. et al. In vivo discovery of immunotherapy targets in the tumour microenvironment. Nature 506, 52–57 (2014). This elegant paper, using an in vivo RNAi screen in T cells, identifies regulatory switches controlling T cell function in immunosuppressive tumours.
Pallasch, C. P. et al. Sensitizing protective tumor microenvironments to antibody-mediated therapy. Cell 156, 590–602 (2014).
Knott, G. J. & Doudna, J. A. CRISPR–Cas guides the future of genetic engineering. Science 361, 866–869 (2018).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Zuo, E. et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos. Science 364, 289 (2019).
Lee, H. K., Smith, H. E., Liu, C., Willi, M. & Hennighausen, L. Cytosine base editor 4 but not adenine base editor generates off-target mutations in mouse embryos. Commun. Biol. 3, 19 (2020).
Cox, D. B. T. et al. RNA editing with CRISPR–Cas13. Science 358, 1019 (2017).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676.e14 (2018).
Pickar-Oliver, A. & Gersbach, C. A. The next generation of CRISPR–Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 20, 490–507 (2019).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Blasco, R. B. et al. Simple and rapid in vivo generation of chromosomal rearrangements using CRISPR/Cas9 technology. Cell Rep. 9, 1219–1227 (2014).
Maddalo, D. et al. In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system. Nature 516, 423–427 (2014).
Platt, R. J. et al. CRISPR–Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Sanchez-Rivera, F. J. et al. Rapid modelling of cooperating genetic events in cancer through somatic genome editing. Nature 516, 428–431 (2014).
Xue, W. et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 514, 380 (2014).
Chiou, S. H. et al. Pancreatic cancer modeling using retrograde viral vector delivery and in vivo CRISPR/Cas9-mediated somatic genome editing. Genes Dev. 29, 1576–1585 (2015).
Mazur, P. K. et al. Combined inhibition of BET family proteins and histone deacetylases as a potential epigenetics-based therapy for pancreatic ductal adenocarcinoma. Nat. Med. 21, 1163–1171 (2015).
Weber, J. et al. CRISPR/Cas9 somatic multiplex-mutagenesis for high-throughput functional cancer genomics in mice. Proc. Natl Acad. Sci. USA 112, 13982–13987 (2015). This paper presents the first proof-of-concept positive-selection CRISPRko screen in the soma of mice, through direct in vivo delivery of CRISPR mini-libraries.
Zuckermann, M. et al. Somatic CRISPR/Cas9-mediated tumour suppressor disruption enables versatile brain tumour modelling. Nat. Commun. 6, 7391 (2015).
Braun, C. J. et al. Versatile in vivo regulation of tumor phenotypes by dCas9-mediated transcriptional perturbation. Proc. Natl Acad. Sci. USA 113, E3892–E3900 (2016). This paper presents the first CRISPRa screen in mice exploiting ex vivo mutagenized B cell lymphoblastic leukaemia cells to identify mediators of resistance to the chemotherapy temozolomide.
Maresch, R. et al. Multiplexed pancreatic genome engineering and cancer induction by transfection-based CRISPR/Cas9 delivery in mice. Nat. Commun. 7, 10770 (2016).
Liu, F., Song, Y. & Liu, D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther. 6, 1258–1266 (1999).
Tietjen, G. T., Bracaglia, L. G., Saltzman, W. M. & Pober, J. S. Focus on fundamentals: achieving effective nanoparticle targeting. Trends Mol. Med. 24, 598–606 (2018).
Xu, C. et al. piggyBac mediates efficient in vivo CRISPR library screening for tumorigenesis in mice. Proc. Natl Acad. Sci. USA 114, 722–727 (2017).
Pan, D. et al. Biodistribution and toxicity studies of VSVG-pseudotyped lentiviral vector after intravenous administration in mice with the observation of in vivo transduction of bone marrow. Mol. Ther. 6, 19–29 (2002).
Rogers, Z. N. et al. Mapping the in vivo fitness landscape of lung adenocarcinoma tumor suppression in mice. Nat. Genet. 50, 483–486 (2018). This elegant study integrates in vivo CRISPRko screening and tumour barcoding to analyse GIs and quantitative cancer dynamics in lung adenocarcinoma.
Grimm, D. & Büning, H. Small but increasingly mighty: latest advances in AAV vector research, design, and evolution. Hum. Gene Ther. 28, 1075–1086 (2017).
Chu, V. T. et al. Efficient CRISPR-mediated mutagenesis in primary immune cells using CrispRGold and a C57BL/6 Cas9 transgenic mouse line. Proc. Natl Acad. Sci. USA 113, 12514 (2016).
Zhou, H. et al. In vivo simultaneous transcriptional activation of multiple genes in the brain using CRISPR–dCas9-activator transgenic mice. Nat. Neurosci. 21, 440–446 (2018).
Lino, C. A., Harper, J. C., Carney, J. P. & Timlin, J. A. Delivering CRISPR: a review of the challenges and approaches. Drug. Deliv. 25, 1234–1257 (2018).
Wang, D. et al. Adenovirus-mediated somatic genome editing of Pten by CRISPR/Cas9 in mouse liver in spite of Cas9-specific immune responses. Hum. Gene Ther. 26, 432–442 (2015).
Chow, R. D. et al. AAV-mediated direct in vivo CRISPR screen identifies functional suppressors in glioblastoma. Nat. Neurosci. 20, 1329 (2017).
Wang, G. et al. Mapping a functional cancer genome atlas of tumor suppressors in mouse liver using AAV–CRISPR-mediated direct in vivo screening. Sci. Adv. 4, eaao5508 (2018).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015). This paper presents the first CRISPRko transplantation screen in mice identifying suppressors of lung metastasis.
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Dong, M. B. et al. Systematic immunotherapy target discovery using genome-scale in vivo CRISPR screens in CD8 T cells. Cell 178, 1189–1204.e23 (2019).
Roth, T. L. et al. Pooled knockin targeting for genome engineering of cellular immunotherapies. Cell 181, 728–744.e21 (2020).
Zhu, S. et al. Guide RNAs with embedded barcodes boost CRISPR-pooled screens. Genome Biol. 20, 20–20 (2019).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Horlbeck, M. A. et al. Mapping the genetic landscape of human cells. Cell 174, 953–967.e922 (2018).
Costanzo, M. et al. Global genetic networks and the genotype-to-phenotype relationship. Cell 177, 85–100 (2019).
Ashworth, A. & Lord, C. J. Synthetic lethal therapies for cancer: what’s next after PARP inhibitors? Nat. Rev. Clin. Oncol. 15, 564–576 (2018).
Feldser, D. M. et al. Stage-specific sensitivity to p53 restoration during lung cancer progression. Nature 468, 572–575 (2010).
Junttila, M. R. et al. Selective activation of p53-mediated tumour suppression in high-grade tumours. Nature 468, 567–571 (2010).
Rad, R. et al. A genetic progression model of Braf(V600E)-induced intestinal tumorigenesis reveals targets for therapeutic intervention. Cancer Cell 24, 15–29 (2013).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Mueller, S. et al. Evolutionary routes and KRAS dosage define pancreatic cancer phenotypes. Nature 554, 62–68 (2018).
Buscail, L., Bournet, B. & Cordelier, P. Role of oncogenic KRAS in the diagnosis, prognosis and treatment of pancreatic cancer. Nat. Rev. Gastroenterol. Hepatol. 17, 153–168 (2020).
Rogers, Z. N. et al. A quantitative and multiplexed approach to uncover the fitness landscape of tumor suppression in vivo. Nat. Methods 14, 737–742 (2017).
Lastowska, M. et al. Identification of a neuronal transcription factor network involved in medulloblastoma development. Acta Neuropathol. Commun. 1, 35 (2013).
Vyazunova, I. et al. Sleeping Beauty mouse models identify candidate genes involved in gliomagenesis. PLoS One 9, e113489 (2014).
Koso, H. et al. Identification of FoxR2 as an oncogene in medulloblastoma. Cancer Res. 74, 2351 (2014).
Beckmann, P. J. et al. Sleeping Beauty insertional mutagenesis reveals important genetic drivers of central nervous system embryonal tumors. Cancer Res. 79, 905–917 (2019).
Collier, L. S. et al. Whole-body Sleeping Beauty mutagenesis can cause penetrant leukemia/lymphoma and rare high-grade glioma without associated embryonic lethality. Cancer Res. 69, 8429–8437 (2009).
Rangel, R. et al. Transposon mutagenesis identifies genes that cooperate with mutant Pten in breast cancer progression. Proc. Natl Acad. Sci. USA 113, E7749–E7758 (2016).
Suárez-Cabrera, C. et al. A transposon-based analysis reveals RASA1 is involved in triple-negative breast cancer. Cancer Res. 77, 1357–1368 (2017).
Chen, L. et al. Transposon insertional mutagenesis in mice identifies human breast cancer susceptibility genes and signatures for stratification. Proc. Natl Acad. Sci. USA 114, E2215–E2224 (2017).
Suárez-Cabrera, C. et al. The Ras-related gene ERAS is involved in human and murine breast cancer. Sci. Rep. 8, 13038 (2018).
van der Weyden, L. et al. Modeling the evolution of ETV6-RUNX1-induced B-cell precursor acute lymphoblastic leukemia in mice. Blood 118, 1041–1051 (2011).
Bergerson, R. J. et al. An insertional mutagenesis screen identifies genes that cooperate with Mll-AF9 in a murine leukemogenesis model. Blood 119, 4512–4523 (2012).
van der Weyden, L. et al. Increased tumorigenesis associated with loss of the tumor suppressor gene Cadm1. Molecular Cancer 11, 29 (2012).
van der Weyden, L. et al. Loss of RASSF1A synergizes with deregulated RUNX2 signaling in tumorigenesis. Cancer Res. 72, 3817–3827 (2012).
van der Weyden, L. et al. Jdp2 downregulates Trp53 transcription to promote leukaemogenesis in the context of Trp53 heterozygosity. Oncogene 32, 397–402 (2013).
Zanesi, N. et al. A sleeping Beauty screen reveals NF- κB activation in CLL mouse model. Blood 121, 4355–4358 (2013).
Been, R. A. et al. Genetic signature of histiocytic sarcoma revealed by a sleeping beauty transposon genetic screen in mice. PLoS One 9, e97280 (2014).
Giotopoulos, G. et al. A novel mouse model identifies cooperating mutations and therapeutic targets critical for chronic myeloid leukemia progression. J. Exp. Med. 212, 1551–1569 (2015).
van der Weyden, L. et al. Somatic drivers of B-ALL in a model of ETV6-RUNX1;Pax5+/– leukemia. BMC Cancer 15, 585 (2015).
Heltemes-Harris, L. M. et al. Sleeping Beauty transposon screen identifies signaling modules that cooperate with STAT5 activation to induce B-cell acute lymphoblastic leukemia. Oncogene 35, 3454–3464 (2016).
Loeb, K. R. et al. Insertional mutagenesis using the Sleeping Beauty transposon system identifies drivers of erythroleukemia in mice. Sci. Rep. 9, 5488 (2019).
Rahrmann, E. P. et al. Sleeping Beauty screen identifies RREB1 and other genetic drivers in human B-cell lymphoma. Mol. Cancer Res. 17, 567–582 (2019).
Guo, Y. et al. Comprehensive ex vivo transposon mutagenesis identifies genes that promote growth factor independence and leukemogenesis. Cancer Res. 76, 773–786 (2016).
Starr, T. K. et al. A Sleeping Beauty transposon-mediated screen identifies murine susceptibility genes for adenomatous polyposis coli (Apc)-dependent intestinal tumorigenesis. Proc. Natl Acad. Sci. USA 108, 5765–5770 (2011).
Morris, S. M. et al. Transposon mutagenesis identifies candidate genes that cooperate with loss of transforming growth factor-β signaling in mouse intestinal neoplasms. Int. J. Cancer 140, 853–863 (2017).
O’Donnell, K. A. et al. A Sleeping Beauty mutagenesis screen reveals a tumor suppressor role for Ncoa2/Src-2 in liver cancer. Proc. Natl Acad. Sci. USA 109, E1377–E1386 (2012).
Fan, Y. et al. Evaluating the landscape of gene cooperativity with receptor tyrosine kinases in liver tumorigenesis using transposon-mediated mutagenesis. J. Hepatol. 70, 470–482 (2019).
Kodama, T. et al. Transposon mutagenesis identifies genes and cellular processes driving epithelial–mesenchymal transition in hepatocellular carcinoma. Proc. Natl Acad. Sci. USA 113, E3384 (2016).
Dorr, C. et al. Transposon mutagenesis screen identifies potential lung cancer drivers and CUL3 as a tumor suppressor. Mol. Cancer Res. 13, 1238–1247 (2015).
Mann, K. M. et al. Sleeping Beauty mutagenesis reveals cooperating mutations and pathways in pancreatic adenocarcinoma. Proc. Natl Acad. Sci. USA 109, 5934–5941 (2012).
Perez-Mancera, P. A. et al. The deubiquitinase USP9X suppresses pancreatic ductal adenocarcinoma. Nature 486, 266–270 (2012).
Wu, J. et al. Insertional mutagenesis identifies a STAT3/Arid1b/β-catenin pathway driving neurofibroma initiation. Cell Rep. 14, 1979–1990 (2016).
Rahrmann, E. P. et al. Identification of PDE4D as a proliferation promoting factor in prostate cancer using a sleeping beauty transposon-based somatic mutagenesis screen. Cancer Res. 69, 4388 (2009).
Ahmad, I. et al. Sleeping Beauty screen reveals Pparg activation in metastatic prostate cancer. Proc. Natl Acad. Sci. USA 113, 8290–8295 (2016).
Karreth, FlorianA. et al. In vivo identification of tumor-suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell 147, 382–395 (2011).
Ni, T. K., Landrette, S. F., Bjornson, R. D., Bosenberg, M. W. & Xu, T. Low-copy piggyBac transposon mutagenesis in mice identifies genes driving melanoma. Proc. Natl Acad. Sci. USA 110, E3640–E3649 (2013).
Quintana, R. M. et al. A transposon-based analysis of gene mutations related to skin cancer development. J. Investig. Dermatol. 133, 239–248 (2013).
Takeda, H. et al. Sleeping Beauty transposon mutagenesis identifies genes that cooperate with mutant Smad4 in gastric cancer development. Proc. Natl Acad. Sci. USA 113, E2057–E2065 (2016).
Montero-Conde, C. et al. Transposon mutagenesis identifies chromatin modifiers cooperating with Ras in thyroid tumorigenesis and detects ATXN7 as a cancer gene. Proc. Natl Acad. Sci. USA 114, E4951–E4960 (2017).
Cadinanos, J. & Bradley, A. Generation of an inducible and optimized piggyBac transposon system. Nucleic Acids Res. 35, e87 (2007).
Yant, S. R., Huang, Y., Akache, B. & Kay, M. A. Site-directed transposon integration in human cells. Nucleic Acids Res. 35, e50 (2007).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Yusa, K., Zhou, L., Li, M. A., Bradley, A. & Craig, N. L. A hyperactive piggyBac transposase for mammalian applications. Proc. Natl Acad. Sci. USA 108, 1531–1536 (2011).
Kawakami, K., Largaespada, D. A. & Ivics, Z. Transposons as tools for functional genomics in vertebrate models. Trends Genet. 33, 784–801 (2017).
Keng, V. W. et al. Efficient transposition of Tol2 in the mouse germline. Genetics 183, 1565–1573 (2009).
Acknowledgements
The authors apologize to colleagues whose work was not acknowledged owing to space limitations. C.J.B. is funded by the Max-Eder Program of Deutsche Krebshilfe (70113377) and The Care for Rare Foundation. D.S. is supported by the German Research Foundation (SA1374/4-2; SFB 1321) and the European Research Council (Consolidator Grant 648521). R.R. receives funds from the Deutsche Krebshilfe (70112480), the German Research Foundation (RA 1629/2-1; SFB1243; SFB1321; SFB1335) and the European Research Council (Consolidator Grant 819642 PACA-MET and MSCA-ITN-ETN 861196).
Author information
Authors and Affiliations
Contributions
J.W., C.J.B., D.S. and R.R. all researched data for the article, provided a substantial contribution to discussions of the content and contributed to writing the article and to the review and/or editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Cancer thanks L. Dow, J. Jonkers and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Epitranscriptional
-
The post-transcriptional processing of RNA molecules, such as methylation of certain bases, which can influence RNA localization, stability, translation and decay.
- Nucleotide transversions
-
DNA point mutations that lead to purine to pyrimidine substitutions, or vice versa (A ↔ C, A ↔ T, C ↔ G, G ↔ T).
- Nucleotide transitions
-
DNA point mutations that lead to the exchange of two purines (A ↔ G) or two pyrimidines (C ↔ T).
- Saturation mutagenesis
-
The introduction of a large number of mutations within a specific genomic region or even a whole genome to identify all functional elements producing a specific phenotype.
- Slow transforming retroviruses
-
An oncogenic retrovirus that induces tumorigenesis through insertional mutagenesis. In contrast to acute transforming retroviruses, its genome does not contain viral oncogenes.
- Promoters
-
DNA sequences that, upon binding of transcription factor proteins, initiate transcription of downstream genes by an RNA polymerase.
- Enhancers
-
Cis-acting regulatory DNA elements that increase the activity of promoters and, thus, the transcription rate of genes (or other transcribed elements) in a cell type-specific but position-independent and orientation-independent manner.
- Hypomorphic
-
A mutation that causes a partial loss of gene function, such as reduced expression or lower substrate affinity.
- Gene-trapping elements
-
Regulatory cassettes consisting of splice acceptor sites and polyadenylation signals capable of terminating transcription prematurely from intronic positions.
- Local hopping
-
The preference of DNA transposon systems to reintegrate close to their excision site.
- Missense mutations
-
DNA point mutations leading to amino acid substitution.
- Silent mutations
-
DNA point mutations that do not lead to an amino acid change owing to the redundancy of the genetic code.
- Haploinsufficiency
-
The cause of dominant deleterious gene action in diploid organisms. For haploinsufficient tumour suppressors, monoallelic inactivation is sufficient to support oncogenesis. By contrast, for ‘classic’ tumour suppressors, inactivation of both alleles is required for tumorigenesis (recessive phenotype).
- Loss of heterozygosity
-
(LOH). The loss of the second (wild-type) allele for monoallelic mutations. In somatic cells, LOH can result from genetic deletions, mitotic recombination between non-sister chromatids or mitotic non-disjunction of chromosomes.
- Mitotic recombination
-
The DNA exchange between sister or non-sister homologous chromatids during mitosis.
- Cooperation
-
A process in which two or more entities (for example, cancer genes) act together for their mutual benefit (for example, promoting tumorigenesis).
- Insulator
-
A cis-acting regulatory DNA element preventing activation of gene promoters by distal enhancers (enhancer-blocking insulators) and/or euchromatin silencing through spread of neighbouring heterochromatin (barrier insulators).
- Hydrodynamic tail vein injection
-
(HTVI). A method for in vivo delivery of small compounds, such as DNA or nanoparticles, into hepatocytes by rapid injection of a large volume of liquid into the mouse tail vein, which induces cardiac congestion and a retrograde liquid flux into the liver.
- Non-homologous end joining
-
(NHEJ). An error-prone repair mechanism for DNA double-strand breaks.
- Homology-directed repair
-
(HDR). An error-free repair mechanism of DNA double-strand breaks, which requires an intact DNA template strand with significant homology to the region surrounding the to-be-repaired double-strand break.
- Cas9 nickase
-
(Cas9n). A Cas9 variant with only one functional endonuclease domain that is capable of inducing DNA single-strand but not double-strand breaks.
- Nonsense mutations
-
DNA point mutations resulting in a stop codon and subsequent premature termination of translation.
- Splice site mutations
-
DNA point mutations within splice sites that can disrupt RNA splicing, resulting in loss of exons (‘exon skipping’) or inclusion of introns (‘intron retention’).
- Dead Cas9
-
(dCas9). A catalytically inactive Cas9 variant that can bind but not cleave DNA owing to engineered deleterious mutations in both endonuclease domains.
- Synthetic lethality
-
An extreme form of negative genetic interaction, in which the loss of two genes leads to cell death, whereas the loss of each gene alone has milder effects or no effect on cellular fitness.
- Unique molecular identifiers
-
(UMIs). High-dimensional molecular barcodes for detection, quantification and tracking of RNA or DNA input molecules. Incorporation of UMIs into single guide RNA (sgRNA) libraries enables measurement of the behavioural consistency of identical sgRNAs within a screen.
- Cellular fitness
-
The ability of cancer cells to flourish within their niche and compete with neighbouring clones. The fitness of a given genotype can be different in different selective environments.
- Clonal sweep
-
A reduction in diversity through a beneficial variation (for example, mutation in a tumour subclone) that eliminates other variants owing to strong positive selection.
- Evolutionary constraints
-
Limitations or biases on the evolution of specific adaptative phenotypes, forcing, for example, cancer cells to evolve along specific molecular paths or sequences.
- Parallel evolution
-
The independent evolution of similar traits in lineages originating from a common ancestor, for example, acquisition of alterations in the same genes or pathways in subclones evolving from a common cell of origin within a cancer.
- Convergent evolution
-
The independent development of similar traits in separate lineages without a common ancestor, for example, acquisition of genetic or epigenetic alterations in the same genes or pathways in tumours from different patients.
Rights and permissions
About this article
Cite this article
Weber, J., Braun, C.J., Saur, D. et al. In vivo functional screening for systems-level integrative cancer genomics. Nat Rev Cancer 20, 573–593 (2020). https://doi.org/10.1038/s41568-020-0275-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-020-0275-9
This article is cited by
-
Genome-scale pan-cancer interrogation of lncRNA dependencies using CasRx
Nature Methods (2024)
-
Sleeping Beauty transposon mutagenesis in mouse intestinal organoids identifies genes involved in tumor progression and metastasis
Cancer Gene Therapy (2024)
-
High-throughput functional screen identifies YWHAZ as a key regulator of pancreatic cancer metastasis
Cell Death & Disease (2023)
-
A genome-wide in vivo CRISPR screen identifies essential regulators of T cell migration to the CNS in a multiple sclerosis model
Nature Neuroscience (2023)
-
Oncogenic context shapes the fitness landscape of tumor suppression
Nature Communications (2023)