Abstract
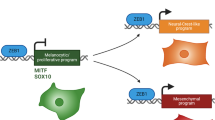
Malignant melanoma is notorious for its inter- and intratumour heterogeneity, based on transcriptionally distinct melanoma cell phenotypes. It is thought that these distinct phenotypes are plastic in nature and that their transcriptional reprogramming enables heterogeneous tumours both to undergo different stages of melanoma progression and to adjust to drug exposure during treatment. Recent advances in genomic technologies and the rapidly expanding availability of large gene expression datasets have allowed for a refined definition of the gene signatures that characterize these phenotypes and have revealed that phenotype plasticity plays a major role in the resistance to both targeted therapy and immunotherapy. In this Review we discuss the definition of melanoma phenotypes through particular transcriptional states and reveal the prognostic relevance of the related gene expression signatures. We review how the establishment of phenotypes is controlled and which roles phenotype plasticity plays in melanoma development and therapy. Because phenotype plasticity in melanoma bears a great resemblance to epithelial–mesenchymal transition, the lessons learned from melanoma will also benefit our understanding of other cancer types.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Pollock, P. M. et al. High frequency of BRAF mutations in nevi. Nat. Genet. 33, 19–20 (2003).
Ding, L. et al. Clonal architectures and driver mutations in metastatic melanomas. PLOS ONE 9, e111153 (2014).
Gartner, J. J. et al. Comparative exome sequencing of metastatic lesions provides insights into the mutational progression of melanoma. BMC Genomics 13, 505 (2012).
Nikolaev, S. I. et al. Exome sequencing identifies recurrent somatic MAP2K1 and MAP2K2 mutations in melanoma. Nat. Genet. 44, 133–139 (2011).
Sanborn, J. Z. et al. Phylogenetic analyses of melanoma reveal complex patterns of metastatic dissemination. Proc. Natl Acad. Sci. USA 112, 10995–11000 (2015).
Turajlic, S. et al. Whole genome sequencing of matched primary and metastatic acral melanomas. Genome Res. 22, 196–207 (2012).
Shain, A. H. & Bastian, B. C. From melanocytes to melanomas. Nat. Rev. Cancer 16, 345–358 (2016).
Cheli, Y. et al. Hypoxia and MITF control metastatic behaviour in mouse and human melanoma cells. Oncogene 31, 2461–2470 (2012).
Falletta, P. et al. Translation reprogramming is an evolutionarily conserved driver of phenotypic plasticity and therapeutic resistance in melanoma. Genes Dev. 31, 18–33 (2017).
Ferguson, J., Smith, M., Zudaire, I., Wellbrock, C. & Arozarena, I. Glucose availability controls ATF4-mediated MITF suppression to drive melanoma cell growth. Oncotarget 8, 32946–32959 (2017).
Kim, I. S. et al. Microenvironment-derived factors driving metastatic plasticity in melanoma. Nat. Commun. 8, 14343 (2017). This study elegantly demonstrates the requirement of a switch to a proliferative and differentiated phenotype for metastatic tumour growth.
Landsberg, J. et al. Melanomas resist T cell therapy through inflammation-induced reversible dedifferentiation. Nature 490, 412–416 (2012).
Hoek, K. S. et al. Metastatic potential of melanomas defined by specific gene expression profiles with no BRAF signature. Pigment Cell Res. 19, 290–302 (2006). This study lays the groundwork for the definition of invasive and proliferative phenotypes and for the phenotype-switch model.
Verfaillie, A. et al. Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state. Nat. Commun. 6, 6683 (2015).
Widmer, D. S. et al. Systematic classification of melanoma cells by phenotype-specific gene expression mapping. Pigment Cell Melanoma Res. 25, 343–353 (2012).
Hoek, K. S. et al. In vivo switching of human melanoma cells between proliferative and invasive states. Cancer Res. 68, 650–656 (2008).
Sensi, M. et al. Human cutaneous melanomas lacking MITF and melanocyte differentiation antigens express a functional Axl receptor kinase. J. Invest. Dermatol. 131, 2448–2457 (2011). This study is the first to demonstrate the existence of MITF low –AXL high melanoma cell subpopulations.
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016). This study demonstrates the inter- and intratumour heterogeneity of MITF low –AXL high melanoma cell subpopulations at the single-cell level.
Rambow, F. et al. Toward minimal residual disease-directed therapy in melanoma. Cell 174, 843–855 (2018). This study, which confirms the findings from reference 108, demonstrates that during initial drug response phase in vivo, cells with different phenotype signatures co-emerge, and that resistance originates from a minimal residual disease state that contains both MITF low and MITF high cell populations.
Tsoi, J. et al. Multi-stage differentiation defines melanoma subtypes with differential vulnerability to drug-induced iron-dependent oxidative stress. Cancer Cell 33, 890–904 (2018). This study establishes that, transcriptionally, melanoma differentiation can be described as four progressive stepwise states and that dedifferentiation occurs in response to either MAPKi therapy or immunotherapy based on ACT.
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016). This is the first analysis of transcriptomes related to immunotherapy resistance; it identifies a transcriptional signature that is expressed in innately immunotherapy-resistant tumours and that overlaps with MAPKi resistance signatures.
Bittner, M. et al. Molecular classification of cutaneous malignant melanoma by gene expression profiling. Nature 406, 536–540 (2000).
Wellbrock, C. & Arozarena, I. Microphthalmia-associated transcription factor in melanoma development and MAP-kinase pathway targeted therapy. Pigment Cell Melanoma Res. 28, 390–406 (2015).
Jeffs, A. R. et al. A gene expression signature of invasive potential in metastatic melanoma cells. PLOS ONE 4, e8461 (2009).
Rambow, F. et al. New functional signatures for understanding melanoma biology from tumor cell lineage-specific analysis. Cell Rep. 13, 840–853 (2015).
Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell 161, 1681–1696 (2015).
Ennen, M. et al. MITF-high and MITF-low cells and a novel subpopulation expressing genes of both cell states contribute to intra- and intertumoral heterogeneity of primary melanoma. Clin. Cancer Res. 23, 7097–7107 (2017).
Eichhoff, O. M. et al. The immunohistochemistry of invasive and proliferative phenotype switching in melanoma: a case report. Melanoma Res. 20, 349–355 (2010).
Haqq, C. et al. The gene expression signatures of melanoma progression. Proc. Natl Acad. Sci. USA 102, 6092–6097 (2005).
Harbst, K. et al. Molecular and genetic diversity in the metastatic process of melanoma. J. Pathol. 233, 39–50 (2014).
Alexaki, V. I. et al. GLI2-mediated melanoma invasion and metastasis. J. Natl Cancer Inst. 102, 1148–1159 (2010).
Pinner, S. et al. Intravital imaging reveals transient changes in pigment production and Brn2 expression during metastatic melanoma dissemination. Cancer Res. 69, 7969–7977 (2009).
Dissanayake, S. K. et al. Wnt5A regulates expression of tumor-associated antigens in melanoma via changes in signal transducers and activators of transcription 3 phosphorylation. Cancer Res. 68, 10205–10214 (2008).
O’Connell, M. P. et al. The orphan tyrosine kinase receptor, ROR2, mediates Wnt5A signaling in metastatic melanoma. Oncogene 29, 34–44 (2010).
Weeraratna, A. T. et al. Wnt5a signaling directly affects cell motility and invasion of metastatic melanoma. Cancer Cell 1, 279–288 (2002). This paper is the first description of a switch to an invasive phenotype induced by WNT5A, a marker of the invasive phenotype signature.
Bald, T. et al. Ultraviolet-radiation-induced inflammation promotes angiotropism and metastasis in melanoma. Nature 507, 109–113 (2014).
Riesenberg, S. et al. MITF and c-Jun antagonism interconnects melanoma dedifferentiation with pro-inflammatory cytokine responsiveness and myeloid cell recruitment. Nat. Commun. 6, 8755 (2015).
Carreira, S. et al. Mitf regulation of Dia1 controls melanoma proliferation and invasiveness. Genes Dev. 20, 3426–3439 (2006).
Wellbrock, C. & Marais, R. Elevated expression of MITF counteracts B-RAF-stimulated melanocyte and melanoma cell proliferation. J. Cell Biol. 170, 703–708 (2005).
Lauss, M. et al. Genome-wide DNA methylation analysis in melanoma reveals the importance of CpG methylation in MITF regulation. J. Invest. Dermatol. 135, 1820–1828 (2015).
Cheng, P. F. et al. Methylation-dependent SOX9 expression mediates invasion in human melanoma cells and is a negative prognostic factor in advanced melanoma. Genome Biol. 16, 42 (2015).
Morrison, S. J., White, P. M., Zock, C. & Anderson, D. J. Prospective identification, isolation by flow cytometry, and in vivo self-renewal of multipotent mammalian neural crest stem cells. Cell 96, 737–749 (1999).
Cheli, Y. et al. CD271 is an imperfect marker for melanoma initiating cells. Oncotarget 5, 5272–5283 (2014).
Held, M. A. et al. Characterization of melanoma cells capable of propagating tumors from a single cell. Cancer Res. 70, 388–397 (2010).
Quintana, E. et al. Phenotypic heterogeneity among tumorigenic melanoma cells from patients that is reversible and not hierarchically organized. Cancer Cell 18, 510–523 (2010).
Boiko, A. D. et al. Human melanoma-initiating cells express neural crest nerve growth factor receptor CD271. Nature 466, 133–137 (2010).
Restivo, G. et al. low neurotrophin receptor CD271 regulates phenotype switching in melanoma. Nat. Commun. 8, 1988 (2017). In this paper the authors demonstrate the requirement for phenotype plasticity for effective melanoma initiation and metastatic growth, using an inducible NGFR mouse model.
Caramel, J. et al. A switch in the expression of embryonic EMT-inducers drives the development of malignant melanoma. Cancer Cell 24, 466–480 (2013).
Wardwell-Ozgo, J. et al. HOXA1 drives melanoma tumor growth and metastasis and elicits an invasion gene expression signature that prognosticates clinical outcome. Oncogene 33, 1017–1026 (2014).
Ferretti, R., Bhutkar, A., McNamara, M. C. & Lees, J. A. BMI1 induces an invasive signature in melanoma that promotes metastasis and chemoresistance. Genes Dev. 30, 18–33 (2016).
Pastushenko, I. et al. Identification of the tumour transition states occurring during EMT. Nature 556, 463–468 (2018).
Khoja, L. et al. Prevalence and heterogeneity of circulating tumour cells in metastatic cutaneous melanoma. Melanoma Res. 24, 40–46 (2014).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Tabassum, D. P. & Polyak, K. Tumorigenesis: it takes a village. Nat. Rev. Cancer 15, 473–483 (2015).
Chapman, A. et al. Heterogeneous tumor subpopulations cooperate to drive invasion. Cell Rep. 8, 688–695 (2014). This is the first study to demonstrate the cooperativity of different melanoma phenotypes in vivo, which leads to enhanced melanoma invasiveness.
Maurus, K. et al. The AP-1 transcription factor FOSL1 causes melanocyte reprogramming and transformation. Oncogene 36, 5110–5121 (2017).
Hong, X. et al. Molecular signatures of circulating melanoma cells for monitoring early response to immune checkpoint therapy. Proc. Natl Acad. Sci. USA 115, 2467–2472 (2018).
Widmer, D. S. et al. Hypoxia contributes to melanoma heterogeneity by triggering HIF1α-dependent phenotype switching. J. Invest. Dermatol. 133, 2436–2443 (2013).
Arozarena, I. & Wellbrock, C. Targeting invasive properties of melanoma cells. FEBS J. 284, 2148–2162 (2017).
Bertolotto, C. et al. A SUMOylation-defective MITF germline mutation predisposes to melanoma and renal carcinoma. Nature 480, 94–98 (2011).
McGill, G. G., Haq, R., Nishimura, E. K. & Fisher, D. E. c-Met expression is regulated by Mitf in the melanocyte lineage. J. Biol. Chem. 281, 10365–10373 (2006).
Hoek, K. S. et al. Novel MITF targets identified using a two-step DNA microarray strategy. Pigment Cell Melanoma Res. 21, 665–676 (2008).
Loftus, S. K. et al. Comparison of melanoblast expression patterns identifies distinct classes of genes. Pigment Cell Melanoma Res. 22, 611–622 (2009).
Bautista, N. C., Cohen, S. & Anders, K. H. Benign melanocytic nevus cells in axillary lymph nodes. A prospective incidence and immunohistochemical study with literature review. Am. J. Clin. Pathol. 102, 102–108 (1994).
Patterson, J. W. Nevus cell aggregates in lymph nodes. Am. J. Clin. Pathol. 121, 13–15 (2004).
Chitty, J. L. et al. Recent advances in understanding the complexities of metastasis. F1000Res. 7, 1169 (2018).
Roesch, A. et al. Overcoming intrinsic multidrug resistance in melanoma by blocking the mitochondrial respiratory chain of slow-cycling JARID1B(high) cells. Cancer Cell 23, 811–825 (2013).
Wellbrock, C. et al. Oncogenic BRAF regulates melanoma proliferation through the lineage specific factor MITF. PLOS ONE 3, e2734 (2008).
Wellbrock, C., Weisser, C., Geissinger, E., Troppmair, J. & Schartl, M. Activation of p59(Fyn) leads to melanocyte dedifferentiation by influencing MKP-1-regulated mitogen-activated protein kinase signaling. J. Biol. Chem. 277, 6443–6454 (2002).
Bogunovic, D. et al. Immune profile and mitotic index of metastatic melanoma lesions enhance clinical staging in predicting patient survival. Proc. Natl Acad. Sci. USA 106, 20429–20434 (2009).
Simmons, J. L., Pierce, C. J., Al-Ejeh, F. & Boyle, G. M. MITF and BRN2 contribute to metastatic growth after dissemination of melanoma. Sci. Rep. 7, 10909 (2017).
Winnepenninckx, V. et al. Gene expression profiling of primary cutaneous melanoma and clinical outcome. J. Natl Cancer Inst. 98, 472–482 (2006).
Harbst, K. et al. Molecular profiling reveals low- and high-grade forms of primary melanoma. Clin. Cancer Res. 18, 4026–4036 (2012).
Jonsson, G. et al. Gene expression profiling-based identification of molecular subtypes in stage IV melanomas with different clinical outcome. Clin. Cancer Res. 16, 3356–3367 (2010).
Miskolczi, Z. et al. Collagen abundance controls melanoma phenotypes through lineage-specific microenvironment sensing. Oncogene 37, 3166–3182 (2018).
Taylor, K. L. et al. Differentiated melanocyte cell division occurs in vivo and is promoted by mutations in Mitf. Development 138, 3579–3589 (2011).
Kwa, M. Q., Herum, K. M. & Brakebusch, C. Cancer-associated fibroblasts: how do they contribute to metastasis? Clin. Exp. Metastasis 36, 71–86 (2019).
Seip, K. et al. Fibroblast-induced switching to the mesenchymal-like phenotype and PI3K/mTOR signaling protects melanoma cells from BRAF inhibitors. Oncotarget 7, 19997–20015 (2016).
Kaur, A. et al. sFRP2 in the aged microenvironment drives melanoma metastasis and therapy resistance. Nature 532, 250–254 (2016).
Kaur, A. et al. Remodeling of the collagen matrix in aging skin promotes melanoma metastasis and affects immune cell motility. Cancer Discov. 9, 64–81 (2019).
Wulf, H. C., Sandby-Moller, J., Kobayasi, T. & Gniadecki, R. Skin aging and natural photoprotection. Micron 35, 185–191 (2004).
Smith, M. P. et al. The immune microenvironment confers resistance to MAPK pathway inhibitors through macrophage-derived TNFα. Cancer Discov. 4, 1214–1229 (2014).
Huergo-Zapico, L. et al. NK-cell editing mediates epithelial-to-mesenchymal transition via phenotypic and proteomic changes in melanoma cell lines. Cancer Res. 78, 3913–3925 (2018).
Ohanna, M. et al. Senescent cells develop a PARP-1 and nuclear factor-κB-associated secretome (PNAS). Genes Dev. 25, 1245–1261 (2011).
Wiedemann, G. M. et al. Microphthalmia-associated transcription factor (MITF) regulates immune cell migration into melanoma. Transl Oncol. 12, 350–360 (2019).
Vazquez, F. et al. PGC1α expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell 23, 287–301 (2013).
Hendrix, M. J. et al. Tumor cell vascular mimicry: novel targeting opportunity in melanoma. Pharmacol. Ther. 159, 83–92 (2016).
Maniotis, A. J. et al. Vascular channel formation by human melanoma cells in vivo and in vitro: vasculogenic mimicry. Am. J. Pathol. 155, 739–752 (1999).
Mihic-Probst, D. et al. Tumor cell plasticity and angiogenesis in human melanomas. PLOS ONE 7, e33571 (2012).
Feige, E. et al. Hypoxia-induced transcriptional repression of the melanoma-associated oncogene MITF. Proc. Natl Acad. Sci. USA 108, E924–E933 (2011).
O’Connell, M. P. et al. Hypoxia induces phenotypic plasticity and therapy resistance in melanoma via the tyrosine kinase receptors ROR1 and ROR2. Cancer Discov. 3, 1378–1393 (2013).
Nakazawa, M. S., Keith, B. & Simon, M. C. Oxygen availability and metabolic adaptations. Nat. Rev. Cancer 16, 663–673 (2016).
Pascual, G. et al. Targeting metastasis-initiating cells through the fatty acid receptor CD36. Nature 541, 41–45 (2017).
Silva, I. P. & Long, G. V. Systemic therapy in advanced melanoma: integrating targeted therapy and immunotherapy into clinical practice. Curr. Opin. Oncol. 29, 484–492 (2017).
Wellbrock, C. & Arozarena, I. The complexity of the ERK/MAP-kinase pathway and the treatment of melanoma skin cancer. Front. Cell Dev. Biol. 4, 33 (2016).
Arozarena, I. & Wellbrock, C. Overcoming resistance to BRAF inhibitors. Ann. Transl Med. 5, 387 (2017).
Ahmed, F. & Haass, N. K. Microenvironment-driven dynamic heterogeneity and phenotypic plasticity as a mechanism of melanoma therapy resistance. Front. Oncol. 8, 173 (2018).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma. N. Engl. J. Med. 372, 2521–2532 (2015).
Schadendorf, D. et al. Pooled analysis of long-term survival data from phase II and phase III trials of ipilimumab in unresectable or metastatic melanoma. J. Clin. Oncol. 33, 1889–1894 (2015).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Zhu, J. et al. Resistance to cancer immunotherapy mediated by apoptosis of tumor-infiltrating lymphocytes. Nat. Commun. 8, 1404 (2017).
Fallahi-Sichani, M. et al. Adaptive resistance of melanoma cells to RAF inhibition via reversible induction of a slowly dividing de-differentiated state. Mol. Syst. Biol. 13, 905 (2017).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Su, Y. et al. Single-cell analysis resolves the cell state transition and signaling dynamics associated with melanoma drug-induced resistance. Proc. Natl Acad. Sci. USA 114, 13679–13684 (2017). References 105, 106 and 107 use single-cell analysis to investigate phenotype plasticity during BRAFi treatment and demonstrate the existence of individual cells that are primed for a transient and reversible resistance, which is induced through transcriptional changes.
Smith, M. P. et al. Inhibiting drivers of non-mutational drug tolerance is a salvage strategy for targeted melanoma therapy. Cancer Cell 29, 270–284 (2016).
Smith, M. P. et al. Effect of SMURF2 targeting on susceptibility to MEK inhibitors in melanoma. J. Natl Cancer Inst. 105, 33–46 (2013). Reference 109 identifies the role of MITF in the adaptive resistance to MAPKi, and reference 108 demonstrates that, in patients, MITF high melanoma cells are enriched during an initial drug tolerance phase, which is non-mutational and reversible.
Nazarian, R. et al. Melanomas acquire resistance to B-RAF(V600E) inhibition by RTK or N-RAS upregulation. Nature 468, 973–977 (2010).
Villanueva, J. et al. Acquired resistance to BRAF inhibitors mediated by a RAF kinase switch in melanoma can be overcome by cotargeting MEK and IGF-1R/PI3K. Cancer Cell 18, 683–695 (2010).
Sun, C. et al. Reversible and adaptive resistance to BRAF(V600E) inhibition in melanoma. Nature 508, 118–122 (2014).
Konieczkowski, D. J. et al. A melanoma cell state distinction influences sensitivity to MAPK pathway inhibitors. Cancer Discov. 4, 816–827 (2014).
Muller, J. et al. Low MITF/AXL ratio predicts early resistance to multiple targeted drugs in melanoma. Nat. Commun. 5, 5712 (2014).
Boshuizen, J. et al. Cooperative targeting of melanoma heterogeneity with an AXL antibody-drug conjugate and BRAF/MEK inhibitors. Nat. Med. 24, 203–212 (2018).
Van Allen, E. M. et al. The genetic landscape of clinical resistance to RAF inhibition in metastatic melanoma. Cancer Discov. 4, 94–109 (2014).
Yan, Y. et al. Genomic features of exceptional response in vemurafenib +/− cobimetinib-treated patients with BRAF V600-mutated metastatic melanoma. Clin. Cancer Res. https://doi.org/10.1158/1078-0432.CCR-18-0720 (2019).
Menon, D. R. et al. A stress-induced early innate response causes multidrug tolerance in melanoma. Oncogene 34, 4545 (2015).
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010).
Smith, M. P. & Wellbrock, C. Molecular pathways: maintaining MAPK inhibitor sensitivity by targeting nonmutational tolerance. Clin. Cancer Res. 22, 5966–5970 (2016).
Hugo, W. et al. Non-genomic and immune evolution of melanoma acquiring MAPKi resistance. Cell 162, 1271–1285 (2015).
Kong, X. et al. Cancer drug addiction is relayed by an ERK2-dependent phenotype switch. Nature 550, 270–274 (2017).
Obenauf, A. C. et al. Therapy-induced tumour secretomes promote resistance and tumour progression. Nature 520, 368–372 (2015).
Smith, M. P. et al. Targeting endothelin receptor signalling overcomes heterogeneity driven therapy failure. EMBO Mol. Med. 9, 1011–1029 (2017). This is the first study to demonstrate that interphenotype communications maintain phenotype heterogeneity and that MITF high cells support the growth of AXL high cells through EDN1.
Jones, P. A., Issa, J. P. & Baylin, S. Targeting the cancer epigenome for therapy. Nat. Rev. Genet. 17, 630–641 (2016).
Rabbie, R., Ferguson, P., Molina-Aguilar, C., Adams, D. J. & Robles-Espinoza, C. D. Melanoma subtypes: genomic profiles, prognostic molecular markers and therapeutic possibilities. J. Pathol. 247, 539–551 (2019).
Shain, A. H. et al. Exome sequencing of desmoplastic melanoma identifies recurrent NFKBIE promoter mutations and diverse activating mutations in the MAPK pathway. Nat. Genet. 47, 1194–1199 (2015).
Eroglu, Z. et al. High response rate to PD-1 blockade in desmoplastic melanomas. Nature 553, 347–350 (2018).
Das Thakur, M. et al. Modelling vemurafenib resistance in melanoma reveals a strategy to forestall drug resistance. Nature 494, 251–255 (2013). This study demonstrates that BRAFi-resistant melanomas become drug dependent and therefore that discontinuous dosing, which exploits the fitness disadvantage displayed by drug-resistant cells, prevents the onset of resistance.
Valpione, S. et al. Rechallenge with BRAF-directed treatment in metastatic melanoma: A multi-institutional retrospective study. Eur. J. Cancer 91, 116–124 (2018).
Vinal, D., Martinez, D. & Espinosa, E. Efficacy of rechallenge with BRAF inhibition therapy in patients with advanced BRAFV600 mutant melanoma. Clin. Transl Oncol. https://doi.org/10.1007/s12094-018-02028-0 (2019).
Smith, M. P. et al. A PAX3/BRN2 rheostat controls the dynamics of BRAF mediated MITF regulation in MITFhigh/AXLlow melanoma. Pigment Cell Melanoma Res. 32, 280–291 (2018).
Haq, R. et al. Oncogenic BRAF regulates oxidative metabolism via PGC1α and MITF. Cancer Cell 23, 302–315 (2013).
Haq, R. et al. BCL2A1 is a lineage-specific antiapoptotic melanoma oncogene that confers resistance to BRAF inhibition. Proc. Natl Acad. Sci. USA 110, 4321–4326 (2013).
Luke, J. J., Flaherty, K. T., Ribas, A. & Long, G. V. Targeted agents and immunotherapies: optimizing outcomes in melanoma. Nat. Rev. Clin. Oncol. 14, 463–482 (2017).
Boni, A. et al. Selective BRAFV600E inhibition enhances T cell recognition of melanoma without affecting lymphocyte function. Cancer Res. 70, 5213–5219 (2010).
Zhao, F. et al. Paracrine Wnt5a-β-catenin signaling triggers a metabolic program that drives dendritic cell tolerization. Immunity 48, 147–160 (2018).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Brinckerhoff, C. E. Cancer stem cells (CSCs) in melanoma: there’s smoke, but is there fire? J. Cell. Physiol. 232, 2674–2678 (2017).
Perego, M. et al. Heterogeneous phenotype of human melanoma cells with in vitro and in vivo features of tumor-initiating cells. J. Invest. Dermatol. 130, 1877–1886 (2010).
Roesch, A. et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell 141, 583–594 (2010).
Spinella, F. et al. Endothelin-1 and endothelin-3 promote invasive behavior via hypoxia-inducible factor-1α in human melanoma cells. Cancer Res. 67, 1725–1734 (2007).
Cheli, Y. et al. Mitf is the key molecular switch between mouse or human melanoma initiating cells and their differentiated progeny. Oncogene 30, 2307–2318 (2011).
Nishimura, E. K. et al. Key roles for transforming growth factor β in melanocyte stem cell maintenance. Cell Stem Cell 6, 130–140 (2010).
Webster, M. R. et al. Wnt5A promotes an adaptive, senescent-like stress response, while continuing to drive invasion in melanoma cells. Pigment Cell Melanoma Res. 28, 184–195 (2015).
Richard, G. et al. ZEB1-mediated melanoma cell plasticity enhances resistance to MAPK inhibitors. EMBO Mol. Med. 8, 1143–1161 (2016).
Ramsdale, R. et al. The transcription cofactor c-JUN mediates phenotype switching and BRAF inhibitor resistance in melanoma. Sci. Signal. 8, ra82 (2015).
Titz, B. et al. JUN dependency in distinct early and late BRAF inhibition adaptation states of melanoma. Cell Discov. 2, 16028 (2016).
Acknowledgements
C.W. acknowledges support by Cancer Research UK (grant number C11591/A16416). I.A. is supported by Miguel Servet contract CP15/00176 from the Instituto de Salud Carlos III-FEDER. We apologize to all colleagues whose work could not be cited here due to space limitations.
Peer review information
Nature Reviews Cancer thanks M. Herlyn, G. Merlino, E. Perez-Guijarro and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Both authors contributed equally to writing the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- ‘Proliferative’ phenotype
-
Distinct cell population with enhanced proliferation in culture, linked to expression of microphthalmia-associated transcription factor (MITF) and its target genes, which regulate the cell cycle and pigmentation.
- ‘Invasive’ phenotype
-
Distinct cell population that displays increased matrigel invasion, linked to the absence of microphthalmia-associated transcription factor (MITF) expression and the expression of mesenchymal markers.
- Epithelial–mesenchymal transition
-
(EMT). A non-mutational process that generates epithelial-derived phenotypes, which have lost their cell polarity and gained migratory and invasive properties.
- Mesenchymal–epithelial transition
-
(MET). The reverse process from epithelial–mesenchymal transition, representing the transition from motile, mesenchymal cells to planar arrays of polarized epithelial cells.
- Melanocyte lineage
-
Neural crest-derived cells that produce the pigment melanin and are located in the basal layer of the epidermis as well as in the eye, inner ear, meninges, heart and bones.
- Pigmentation
-
Deposition of the pigment melanin, which is produced by differentiated melanocytes, in keratinocytes in the skin.
- Transition states
-
Intermediate, hybrid states of stable phenotype states that are characterized by distinct transcriptional and epigenetic landscapes, gene regulatory networks, transcription factors and signalling pathways.
- Neural-crest derivatives
-
Diverse cell lineages derived from neural-crest cells, which include melanocytes, craniofacial cartilage and bone, smooth muscle, peripheral and enteric neurons and glial cells.
- Angiotropism
-
A process, also termed extravascular migratory metastasis, whereby tumour cells migrate along the external surfaces of vascular channels, without intravasation.
- Adoptive cell transfer therapy
-
(ACT). A type of immunotherapy in which T cells, usually taken from a cancer patient’s own blood or tumour tissue, are propagated in vitro and given back to a patient so as to target the tumour(s).
- Melanocyte differentiation antigen
-
(MDA). Surface antigens that act as markers of early, intermediate and mature stages in melanocyte differentiation and are expressed in melanoma subsets.
- Vasculogenic mimicry
-
Ability of metastatic tumour cells to mimic a true vascular endothelium and form microvascular channels without the presence of endothelial cells.
- Ferroptosis
-
A form of programmed cell death based on the accumulation of lipid peroxidation products and lethal reactive oxygen species derived from iron metabolism.
Rights and permissions
About this article
Cite this article
Arozarena, I., Wellbrock, C. Phenotype plasticity as enabler of melanoma progression and therapy resistance. Nat Rev Cancer 19, 377–391 (2019). https://doi.org/10.1038/s41568-019-0154-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-019-0154-4
This article is cited by
-
Roles and mechanisms of aberrant alternative splicing in melanoma — implications for targeted therapy and immunotherapy resistance
Cancer Cell International (2024)
-
Mcl-1 mediates intrinsic resistance to RAF inhibitors in mutant BRAF papillary thyroid carcinoma
Cell Death Discovery (2024)
-
ZEB1 controls a lineage-specific transcriptional program essential for melanoma cell state transitions
Oncogene (2024)
-
BRAF — a tumour-agnostic drug target with lineage-specific dependencies
Nature Reviews Clinical Oncology (2024)
-
Predicting tumor repopulation through the gene panel derived from radiation resistant colorectal cancer cells
Journal of Translational Medicine (2023)