Abstract
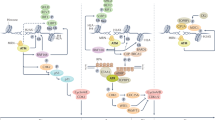
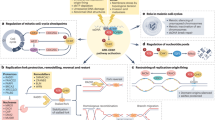
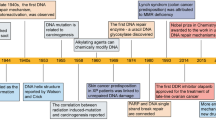
The chemical treatment of cancer started with the realization that DNA damaging agents such as mustard gas present notable antitumoural properties. Consequently, early drug development focused on genotoxic chemicals, some of which are still widely used in the clinic. However, the efficacy of such therapies is often limited by the side effects of these drugs on healthy cells. A refinement to this approach is to use compounds that can exploit the presence of DNA damage in cancer cells. Given that replication stress (RS) is a major source of genomic instability in cancer, targeting the RS-response kinase ataxia telangiectasia and Rad3-related protein (ATR) has emerged as a promising alternative. With ATR inhibitors now entering clinical trials, we here revisit the biology behind this strategy and discuss potential biomarkers that could be used for a better selection of patients who respond to therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Lord, C. J. & Ashworth, A. PARP inhibitors: synthetic lethality in the clinic. Science 355, 1152–1158 (2017).
Cimprich, K. A., Shin, T. B., Keith, C. T. & Schreiber, S. L. cDNA cloning and gene mapping of a candidate human cell cycle checkpoint protein. Proc. Natl Acad. Sci. USA 93, 2850–2855 (1996).
Jackson, S. P. & Bartek, J. The DNA-damage response in human biology and disease. Nature 461, 1071–1078 (2009).
Choi, M., Kipps, T. & Kurzrock, R. ATM mutations in cancer: therapeutic implications. Mol. Cancer Ther. 15, 1781–1791 (2016).
Gaillard, H., Garcia-Muse, T. & Aguilera, A. Replication stress and cancer. Nat. Rev. Cancer 15, 276–289 (2015).
Murga, M. et al. A mouse model of ATR-Seckel shows embryonic replicative stress and accelerated aging. Nat. Genet. 41, 891–898 (2009).
Murga, M. et al. Exploiting oncogene-induced replicative stress for the selective killing of Myc-driven tumors. Nat. Struct. Mol. Biol. 18, 1331–1335 (2011).
Saldivar, J. C., Cortez, D. & Cimprich, K. A. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat. Rev. Mol. Cell Biol. 18, 622–636 (2017).
Hansemann, D. Ueber asymmetrische Zelltheilung in Epithelkrebsen und deren biologische Bedeutung. Arch. Pathol. Anat. Physiol. Klin. Med. 119, 299–326 (1890).
Boveri, T. Zur Frage der Entstehung Maligner Tumoren. Gustav Fischer, Jena 1–64 (1914).
Bartkova, J. et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434, 864–870 (2005).
Gorgoulis, V. G. et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434, 907–913 (2005).
Bartkova, J. et al. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 444, 633–637 (2006).
Di Micco, R. et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 444, 638–642 (2006).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Macheret, M. & Halazonetis, T. D. DNA replication stress as a hallmark of cancer. Annu. Rev. Pathol. 10, 425–448 (2015).
Lee, S. E. et al. Saccharomyces Ku70, mre11/rad50 and RPA proteins regulate adaptation to G2/M arrest after DNA damage. Cell 94, 399–409 (1998).
Garvik, B., Carson, M. & Hartwell, L. Single-stranded DNA arising at telomeres in cdc13 mutants may constitute a specific signal for the RAD9 checkpoint. Mol. Cell. Biol. 15, 6128–6138 (1995).
Zou, L. & Elledge, S. J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 300, 1542–1548 (2003).
Cortez, D., Guntuku, S., Qin, J. & Elledge, S. J. ATR and ATRIP: partners in checkpoint signaling. Science 294, 1713–1716 (2001).
MacDougall, C. A., Byun, T. S., Van, C., Yee, M. C. & Cimprich, K. A. The structural determinants of checkpoint activation. Genes Dev. 21, 898–903 (2007).
Huertas, P. DNA resection in eukaryotes: deciding how to fix the break. Nat. Struct. Mol. Biol. 17, 11–16 (2010).
Kumagai, A., Lee, J., Yoo, H. Y. & Dunphy, W. G. TopBP1 activates the ATR-ATRIP complex. Cell 124, 943–955 (2006).
Mordes, D. A., Glick, G. G., Zhao, R. & Cortez, D. TopBP1 activates ATR through ATRIP and a PIKK regulatory domain. Genes Dev. 22, 1478–1489 (2008).
Bass, T. E. et al. ETAA1 acts at stalled replication forks to maintain genome integrity. Nat. Cell Biol. 18, 1185–1195 (2016).
Haahr, P. et al. Activation of the ATR kinase by the RPA-binding protein ETAA1. Nat. Cell Biol. 18, 1196–1207 (2016).
Jeon, Y. et al. TopBP1 deficiency causes an early embryonic lethality and induces cellular senescence in primary cells. J. Biol. Chem. 286, 5414–5422 (2011).
Miosge, L. A. et al. Systems-guided forward genetic screen reveals a critical role of the replication stress response protein ETAA1 in T cell clonal expansion. Proc. Natl Acad. Sci. USA 114, E5216–E5225 (2017).
Guo, Z., Kumagai, A., Wang, S. X. & Dunphy, W. G. Requirement for Atr in phosphorylation of Chk1 and cell cycle regulation in response to DNA replication blocks and UV-damaged DNA in Xenopus egg extracts. Genes Dev. 14, 2745–2756 (2000).
Liu, Q. et al. Chk1 is an essential kinase that is regulated by Atr and required for the G(2)/M DNA damage checkpoint. Genes Dev. 14, 1448–1459 (2000).
Kumagai, A. & Dunphy, W. G. Claspin, a novel protein required for the activation of Chk1 during a DNA replication checkpoint response in Xenopus egg extracts. Mol. Cell 6, 839–849 (2000).
Smits, V. A., Reaper, P. M. & Jackson, S. P. Rapid PIKK-dependent release of Chk1 from chromatin promotes the DNA-damage checkpoint response. Curr. Biol. 16, 150–159 (2006).
Tercero, J. A. & Diffley, J. F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412, 553–557 (2001).
Couch, F. B. et al. ATR phosphorylates SMARCAL1 to prevent replication fork collapse. Genes Dev. 27, 1610–1623 (2013).
Davies, S. L., North, P. S. & Hickson, I. D. Role for BLM in replication-fork restart and suppression of origin firing after replicative stress. Nat. Struct. Mol. Biol. 14, 677–679 (2007).
Pichierri, P., Rosselli, F. & Franchitto, A. Werner’s syndrome protein is phosphorylated in an ATR/ATM-dependent manner following replication arrest and DNA damage induced during the S phase of the cell cycle. Oncogene 22, 1491–1500 (2003).
Lossaint, G. et al. FANCD2 binds MCM proteins and controls replisome function upon activation of s phase checkpoint signaling. Mol. Cell 51, 678–690 (2013).
Murphy, A. K. et al. Phosphorylated RPA recruits PALB2 to stalled DNA replication forks to facilitate fork recovery. J. Cell Biol. 206, 493–507 (2014).
Cerbinskaite, A., Mukhopadhyay, A., Plummer, E. R., Curtin, N. J. & Edmondson, R. J. Defective homologous recombination in human cancers. Cancer Treat. Rev. 38, 89–100 (2012).
Chu, W. K. & Hickson, I. D. RecQ helicases: multifunctional genome caretakers. Nat. Rev. Cancer 9, 644–654 (2009).
Ge, X. Q., Jackson, D. A. & Blow, J. J. Dormant origins licensed by excess Mcm2-7 are required for human cells to survive replicative stress. Genes Dev. 21, 3331–3341 (2007).
Ibarra, A., Schwob, E. & Mendez, J. Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. Proc. Natl Acad. Sci. USA 105, 8956–8961 (2008).
Cortez, D., Glick, G. & Elledge, S. J. Minichromosome maintenance proteins are direct targets of the ATM and ATR checkpoint kinases. Proc. Natl Acad. Sci. USA 101, 10078–10083 (2004).
Yoo, H. Y., Shevchenko, A., Shevchenko, A. & Dunphy, W. G. Mcm2 is a direct substrate of ATM and ATR during DNA damage and DNA replication checkpoint responses. J. Biol. Chem. 279, 53353–53364 (2004).
Chen, Y. H. et al. ATR-mediated phosphorylation of FANCI regulates dormant origin firing in response to replication stress. Mol. Cell 58, 323–338 (2015).
Santocanale, C. & Diffley, J. F. A. Mec1- and Rad53-dependent checkpoint controls late-firing origins of DNA replication. Nature 395, 615–618 (1998).
Bagley, B. N. et al. A dominantly acting murine allele of Mcm4 causes chromosomal abnormalities and promotes tumorigenesis. PLoS Genet. 8, e1003034 (2012).
Pruitt, S. C., Bailey, K. J. & Freeland, A. Reduced Mcm2 expression results in severe stem/progenitor cell deficiency and cancer. Stem Cells 25, 3121–3132 (2007).
Shima, N. et al. A viable allele of Mcm4 causes chromosome instability and mammary adenocarcinomas in mice. Nat. Genet. 39, 93–98 (2007).
Huang, M., Zhou, Z. & Elledge, S. J. The DNA replication and damage checkpoint pathways induce transcription by inhibition of the Crt1 repressor. Cell 94, 595–605 (1998).
Zhao, X., Muller, E. G. & Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 2, 329–340 (1998).
Lopez-Contreras, A. J. et al. Increased Rrm2 gene dosage reduces fragile site breakage and prolongs survival of ATR mutant mice. Genes Dev. 29, 690–695 (2015).
Buisson, R., Boisvert, J. L., Benes, C. H. & Zou, L. Distinct but concerted roles of ATR, DNA-PK, and Chk1 in countering replication stress during S phase. Mol. Cell 59, 1011–1024 (2015).
Zhang, Y. W., Jones, T. L., Martin, S. E., Caplen, N. J. & Pommier, Y. Implication of checkpoint kinase-dependent up-regulation of ribonucleotide reductase R2 in DNA damage response. J. Biol. Chem. 284, 18085–18095 (2009).
D’Angiolella, V. et al. Cyclin F-mediated degradation of ribonucleotide reductase M2 controls genome integrity and DNA repair. Cell 149, 1023–1034 (2012).
Le, T. M. et al. ATR inhibition facilitates targeting of leukemia dependence on convergent nucleotide biosynthetic pathways. Nat. Commun. 8, 241 (2017).
Bester, A. C. et al. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 145, 435–446 (2011).
Ruiz, S. et al. Limiting replication stress during somatic cell reprogramming reduces genomic instability in induced pluripotent stem cells. Nat. Commun. 6, 8036 (2015).
Petermann, E., Woodcock, M. & Helleday, T. Chk1 promotes replication fork progression by controlling replication initiation. Proc. Natl Acad. Sci. USA 107, 16090–16095 (2010).
Boos, D. et al. Regulation of DNA replication through Sld3-Dpb11 interaction is conserved from yeast to humans. Curr. Biol. 21, 1152–1157 (2011).
Guo, C. et al. Interaction of Chk1 with Treslin negatively regulates the initiation of chromosomal DNA replication. Mol. Cell 57, 492–505 (2015).
Heffernan, T. P. et al. Cdc7-Dbf4 and the human S checkpoint response to UVC. J. Biol. Chem. 282, 9458–9468 (2007).
Toledo, L. I. et al. ATR prohibits replication catastrophe by preventing global exhaustion of RPA. Cell 155, 1088–1103 (2013).
Lee, J., Kumagai, A. & Dunphy, W. G. Positive regulation of Wee1 by Chk1 and 14-3-3 proteins. Mol. Biol. Cell 12, 551–563 (2001).
Peng, C. Y. et al. Mitotic and G2 checkpoint control: regulation of 14-3-3 protein binding by phosphorylation of Cdc25C on serine-216. Science 277, 1501–1505 (1997).
Sanchez, Y. et al. Conservation of the Chk1 checkpoint pathway in mammals: linkage of DNA damage to Cdk regulation through Cdc25. Science 277, 1497–1501 (1997).
Mailand, N. et al. Rapid destruction of human Cdc25A in response to DNA damage. Science 288, 1425–1429 (2000).
Brown, E. J. & Baltimore, D. ATR disruption leads to chromosomal fragmentation and early embryonic lethality. Genes Dev. 14, 397–402 (2000).
de Klein, A. et al. Targeted disruption of the cell-cycle checkpoint gene ATR leads to early embryonic lethality in mice. Curr. Biol. 10, 479–482 (2000).
Takai, H. et al. Aberrant cell cycle checkpoint function and early embryonic death in Chk1(−/−) mice. Genes Dev. 14, 1439–1447 (2000).
Fang, Y. et al. ATR functions as a gene dosage-dependent tumor suppressor on a mismatch repair-deficient background. EMBO J. 23, 3164–3174 (2004).
Fishler, T. et al. Genetic instability and mammary tumor formation in mice carrying mammary-specific disruption of Chk1 and p53. Oncogene 29, 4007–4017 (2010).
Gilad, O. et al. Combining ATR suppression with oncogenic Ras synergistically increases genomic instability, causing synthetic lethality or tumorigenesis in a dosage-dependent manner. Cancer Res. 70, 9693–9702 (2010).
Schoppy, D. W. et al. Oncogenic stress sensitizes murine cancers to hypomorphic suppression of ATR. J. Clin. Invest. 122, 241–252 (2012).
Monasor, A. et al. INK4a/ARF limits the expansion of cells suffering from replication stress. Cell Cycle 12, 1948–1954 (2013).
Tho, L. M., Libertini, S., Rampling, R., Sansom, O. & Gillespie, D. A. Chk1 is essential for chemical carcinogen-induced mouse skin tumorigenesis. Oncogene 31, 1366–1375 (2012).
Kawasumi, M. et al. Protection from UV-induced skin carcinogenesis by genetic inhibition of the ataxia telangiectasia and Rad3-related (ATR) kinase. Proc. Natl Acad. Sci. USA 108, 13716–13721 (2011).
Chen, C. F. et al. ATR mutations promote the growth of melanoma tumors by modulating the immune microenvironment. Cell Rep. 18, 2331–2342 (2017).
Tanaka, A. et al. Germline mutation in ATR in autosomal- dominant oropharyngeal cancer syndrome. Am. J. Hum. Genet. 90, 511–517 (2012).
Derenzini, E. et al. Constitutive activation of the DNA damage response pathway as a novel therapeutic target in diffuse large B cell lymphoma. Oncotarget 6, 6553–6569 (2015).
Krajewska, M. et al. ATR inhibition preferentially targets homologous recombination-deficient tumor cells. Oncogene 34, 3474–3481 (2015).
Sarmento, L. M. et al. CHK1 overexpression in T cell acute lymphoblastic leukemia is essential for proliferation and survival by preventing excessive replication stress. Oncogene 34, 2978–2990 (2015).
Hoglund, A. et al. Therapeutic implications for the induced levels of Chk1 in Myc-expressing cancer cells. Clin. Cancer Res. 17, 7067–7079 (2011).
Schulze, J. et al. Fos-dependent induction of Chk1 protects osteoblasts from replication stress. Cell Cycle 13, 1980–1986 (2014).
Verlinden, L. et al. The E2F-regulated gene Chk1 is highly expressed in triple-negative estrogen receptor /progesterone receptor /HER-2 breast carcinomas. Cancer Res. 67, 6574–6581 (2007).
Lopez-Contreras, A. J., Gutierrez-Martinez, P., Specks, J., Rodrigo-Perez, S. & Fernandez-Capetillo, O. An extra allele of Chk1 limits oncogene-induced replicative stress and promotes transformation. J. Exp. Med. 209, 455–461 (2012).
Sarkaria, J. N. et al. Inhibition of ATM and ATR kinase activities by the radiosensitizing agent, caffeine. Cancer Res. 59, 4375–4382 (1999).
Nishida, H. et al. Inhibition of ATR protein kinase activity by schisandrin B in DNA damage response. Nucleic Acids Res. 37, 5678–5689 (2009).
Toledo, L. I., Murga, M., Gutierrez-Martinez, P., Soria, R. & Fernandez-Capetillo, O. ATR signaling can drive cells into senescence in the absence of DNA breaks. Genes Dev. 22, 297–302 (2008).
Toledo, L. I. et al. A cell-based screen identifies ATR inhibitors with synthetic lethal properties for cancer-associated mutations. Nat. Struct. Mol. Biol. 18, 721–727 (2011).
Peasland, A. et al. Identification and evaluation of a potent novel ATR inhibitor, NU6027, in breast and ovarian cancer cell lines. Br. J. Cancer 105, 372–381 (2011).
Charrier, J. D. et al. Discovery of potent and selective inhibitors of ataxia telangiectasia mutated and Rad3 related (ATR) protein kinase as potential anticancer agents. J. Med. Chem. 54, 2320–2330 (2011).
Reaper, P. M. et al. Selective killing of ATM- or p53-deficient cancer cells through inhibition of ATR. Nat. Chem. Biol. 7, 428–430 (2011).
Fokas, E. et al. Targeting ATR in vivo using the novel inhibitor VE-822 results in selective sensitization of pancreatic tumors to radiation. Cell Death Dis. 3, e441 (2012).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02157792?term=NCT02157792&rank=02157791 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02487095?term=NCT02487095&rank=02487091 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02567422?term=NCT02567422&rank=02567421 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02589522?term=NCT02589522&rank=02589521 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02595892?term=NCT02595892&rank=02595891 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02595931?term=NCT02595931&rank=02595931 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02627443?term=NCT02627443&rank=02627441 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02723864?term=NCT02723864&rank=02723861 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT03022409?term=NCT03022409&rank=03022401 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT03309150?term=NCT03309150&rank=03309151 (2018).
Foote, K. M. et al. Discovery of 4-{4-[(3R)-3-methylmorpholin-4-yl]-6-[1-(methylsulfonyl)cyclopropyl]pyrimidin-2-y l}-1H-indole (AZ20): a potent and selective inhibitor of ATR protein kinase with monotherapy in vivo antitumor activity. J. Med. Chem. 56, 2125–2138 (2013).
Vendetti, F. P. et al. The orally active and bioavailable ATR kinase inhibitor AZD6738 potentiates the anti-tumor effects of cisplatin to resolve ATM-deficient non-small cell lung cancer in vivo. Oncotarget 6, 44289–44305 (2015).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT01955668?term=NCT01955668&rank=01955661 (2014).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02223923?term=NCT02223923&rank=02223921 (2017).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02264678?term=NCT02264678&rank=02264671 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02567409?term=NCT02567409&rank=02567401 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02630199?term=NCT02630199&rank=02630191 (2017).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT03328273?term=NCT03328273&rank=03328271 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT03330847?term=NCT03330847&rank=03330841 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT03334617?term=NCT03334617&rank=03334611 (2018).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT03188965?term=NCT03188965&rank=03188961 (2018).
Ramachandran, S. A. et al. Discovery of pyrazolopyrimidine derivatives as novel inhibitors of ataxia telangiectasia and rad3 related protein (ATR). Bioorg. Med. Chem. Lett. 27, 750–754 (2017).
Chen, H., Lisby, M. & Symington, L. S. RPA coordinates DNA end resection and prevents formation of DNA hairpins. Mol. Cell 50, 589–600 (2013).
Buisson, R. et al. Coupling of homologous recombination and the checkpoint by ATR. Mol. Cell 65, 336–346 (2017).
Eykelenboom, J. K. et al. ATR activates the S-M checkpoint during unperturbed growth to ensure sufficient replication prior to mitotic onset. Cell Rep. 5, 1095–1107 (2013).
Ruiz, S. et al. A genome-wide CRISPR screen identifies CDC25A as a determinant of sensitivity to ATR inhibitors. Mol. Cell 62, 307–313 (2016).
Lecona, E. & Fernandez-Capetillo, O. Replication stress and cancer: it takes two to tango. Exp. Cell Res. 329, 26–34 (2014).
Syljuasen, R. G. et al. Inhibition of human Chk1 causes increased initiation of DNA replication, phosphorylation of ATR targets, and DNA breakage. Mol. Cell. Biol. 25, 3553–3562 (2005).
Cottini, F. et al. Synthetic lethal approaches exploiting DNA damage in aggressive myeloma. Cancer Discov. 5, 972–987 (2015).
Muralidharan, S. V. et al. BET bromodomain inhibitors synergize with ATR inhibitors to induce DNA damage, apoptosis, senescence-associated secretory pathway and ER stress in Myc-induced lymphoma cells. Oncogene 35, 4689–4697 (2016).
Mayor-Ruiz, C., Dominguez, O. & Fernandez-Capetillo, O. TrapSeq: an RNA sequencing-based pipeline for the identification of gene-trap insertions in mammalian cells. J. Mol. Biol. 429, 2780–2789 (2017).
Buisson, R., Lawrence, M. S., Benes, C. H. & Zou, L. APOBEC3A and APOBEC3B activities render cancer cells susceptible to ATR inhibition. Cancer Res. 77, 4567–4578 (2017).
Nieto-Soler, M. et al. Efficacy of ATR inhibitors as single agents in Ewing sarcoma. Oncotarget 58759–58767 (2016).
Morgado-Palacin, I. et al. Targeting the kinase activities of ATR and ATM exhibits antitumoral activity in mouse models of MLL-rearranged AML. Sci. Signal. 9, ra91 (2016).
Kwok, M. et al. ATR inhibition induces synthetic lethality and overcomes chemoresistance in TP53- or ATM-defective chronic lymphocytic leukemia cells. Blood 127, 582–595 (2016).
Farres, J. et al. PARP-2 sustains erythropoiesis in mice by limiting replicative stress in erythroid progenitors. Cell Death Differ. 22, 1144–1157 (2015).
Alvarez, S. et al. Replication stress caused by low MCM expression limits fetal erythropoiesis and hematopoietic stem cell functionality. Nat. Commun. 6, 8548 (2015).
Austin, W. R. et al. Nucleoside salvage pathway kinases regulate hematopoiesis by linking nucleotide metabolism with replication stress. J. Exp. Med. 209, 2215–2228 (2012).
Ruzankina, Y. et al. Tissue regenerative delays and synthetic lethality in adult mice after combined deletion of Atr and Trp53. Nat. Genet. 41, 1144–1149 (2009).
Schmitt, A. et al. ATM deficiency is associated with sensitivity to PARP1- and ATR inhibitors in lung adenocarcinoma. Cancer Res. 77, 3040–3056 (2017).
Perkhofer, L. et al. ATM deficiency generating genomic instability sensitizes pancreatic ductal adenocarcinoma cells to therapy-induced DNA damage. Cancer Res. 77, 5576–5590 (2017).
Min, A. et al. AZD6738, a novel oral inhibitor of ATR, induces synthetic lethality with ATM deficiency in gastric cancer cells. Mol. Cancer Ther. 16, 566–577 (2017).
Menezes, D. L. et al. A synthetic lethal screen reveals enhanced sensitivity to ATR inhibitor treatment in mantle cell lymphoma with ATM loss-of-function. Mol. Cancer Res. 13, 120–129 (2015).
Williamson, C. T. et al. ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat. Commun. 7, 13837 (2016).
Middleton, F. K. et al. Common cancer-associated imbalances in the DNA damage response confer sensitivity to single agent ATR inhibition. Oncotarget 6, 32396–32409 (2015).
Mohni, K. N., Kavanaugh, G. M. & Cortez, D. ATR pathway inhibition is synthetically lethal in cancer cells with ERCC1 deficiency. Cancer Res. 74, 2835–2845 (2014).
Mohni, K. N. et al. A synthetic lethal screen identifies DNA repair pathways that sensitize cancer cells to combined ATR inhibition and cisplatin treatments. PLoS ONE 10, e0125482 (2015).
Huntoon, C. J. et al. ATR inhibition broadly sensitizes ovarian cancer cells to chemotherapy independent of BRCA status. Cancer Res. 73, 3683–3691 (2013).
Flynn, R. L. et al. Alternative lengthening of telomeres renders cancer cells hypersensitive to ATR inhibitors. Science 347, 273–277 (2015).
Deeg, K. I., Chung, I., Bauer, C. & Rippe, K. Cancer cells with alternative lengthening of telomeres do not display a general hypersensitivity to ATR inhibition. Front. Oncol. 6, 186 (2016).
Abu-Sanad, A. et al. Simultaneous inhibition of ATR and PARP sensitizes colon cancer cell lines to irinotecan. Front. Pharmacol. 6, 147 (2015).
Huehls, A. M., Wagner, J. M., Huntoon, C. J. & Karnitz, L. M. Identification of DNA repair pathways that affect the survival of ovarian cancer cells treated with a poly(ADP-ribose) polymerase inhibitor in a novel drug combination. Mol. Pharmacol. 82, 767–776 (2012).
Kim, H. et al. Targeting the ATR/CHK1 axis with PARP inhibition results in tumor regression in BRCA-mutant ovarian cancer models. Clin. Cancer Res. 23, 3097–3108 (2017).
Ogiwara, H. et al. Curcumin suppresses multiple DNA damage response pathways and has potency as a sensitizer to PARP inhibitor. Carcinogenesis 34, 2486–2497 (2013).
Yazinski, S. A. et al. ATR inhibition disrupts rewired homologous recombination and fork protection pathways in PARP inhibitor-resistant BRCA-deficient cancer cells. Genes Dev. 31, 318–332 (2017).
Murai, J. et al. SLFN11 blocks stressed replication forks independently of ATR. Mol. Cell 69, 371–384.e6 (2018).
Thomas, A. et al. Phase I study of ATR inhibitor M6620 in combination with topotecan in patients with advanced solid tumors. J. Clin. Oncol. https://doi.org/10.1200/JCO.2017.76.6915 (2017).
Hall, A. B. et al. Potentiation of tumor responses to DNA damaging therapy by the selective ATR inhibitor VX-970. Oncotarget 5, 5674–5685 (2014).
Li, C. C. et al. ATR-Chk1 signaling inhibition as a therapeutic strategy to enhance cisplatin chemosensitivity in urothelial bladder cancer. Oncotarget 7, 1947–1959 (2016).
O’Flanagan, C. H. et al. IGF-1R inhibition sensitizes breast cancer cells to ATM-related kinase (ATR) inhibitor and cisplatin. Oncotarget 7, 56826–56841 (2016).
Sangster-Guity, N., Conrad, B. H., Papadopoulos, N. & Bunz, F. ATR mediates cisplatin resistance in a p53 genotype-specific manner. Oncogene 30, 2526–2533 (2011).
Sultana, R. et al. Ataxia telangiectasia mutated and Rad3 related (ATR) protein kinase inhibition is synthetically lethal in XRCC1 deficient ovarian cancer cells. PLoS ONE 8, e57098 (2013).
Liu, S. et al. Inhibition of ATR potentiates the cytotoxic effect of gemcitabine on pancreatic cancer cells through enhancement of DNA damage and abrogation of ribonucleotide reductase induction by gemcitabine. Oncol. Rep. 37, 3377–3386 (2017).
Ma, J. et al. Mechanisms responsible for the synergistic antileukemic interactions between ATR inhibition and cytarabine in acute myeloid leukemia cells. Sci. Rep. 7, 41950 (2017).
Prevo, R. et al. The novel ATR inhibitor VE-821 increases sensitivity of pancreatic cancer cells to radiation and chemotherapy. Cancer Biol. Ther. 13, 1072–1081 (2012).
Josse, R. et al. ATR inhibitors VE-821 and VX-970 sensitize cancer cells to topoisomerase i inhibitors by disabling DNA replication initiation and fork elongation responses. Cancer Res. 74, 6968–6979 (2014).
Fujisawa, H. et al. VE-821, an ATR inhibitor, causes radiosensitization in human tumor cells irradiated with high LET radiation. Radiat. Oncol. 10, 175 (2015).
Cui, Y., Palii, S. S., Innes, C. L. & Paules, R. S. Depletion of ATR selectively sensitizes ATM-deficient human mammary epithelial cells to ionizing radiation and DNA-damaging agents. Cell Cycle 13, 3541–3550 (2014).
O’Driscoll, M., Ruiz-Perez, V. L., Woods, C. G., Jeggo, P. A. & Goodship, J. A. A splicing mutation affecting expression of ataxia-telangiectasia and Rad3-related protein (ATR) results in Seckel syndrome. Nat. Genet. 33, 497–501 (2003).
Sanjiv, K. et al. Cancer-specific synthetic lethality between ATR and CHK1 kinase activities. Cell Rep. 17, 3407–3416 (2016).
Dominguez-Kelly, R. et al. Wee1 controls genomic stability during replication by regulating the Mus81-Eme1 endonuclease. J. Cell Biol. 194, 567–579 (2011).
Rodriguez-Bravo, V., Guaita-Esteruelas, S., Florensa, R., Bachs, O. & Agell, N. Chk1- and claspin-dependent but ATR/ATM- and Rad17-independent DNA replication checkpoint response in HeLa cells. Cancer Res. 66, 8672–8679 (2006).
Muralidharan, S. V. et al. BET bromodomain inhibitors synergize with ATR inhibitors in melanoma in melanoma. Cell Death Dis. 8, e2982 (2017).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Seckel, H. Bird-headed dwarfs: studies in developmental anthropology including human proportions (Charles C. Thomas, Springfield, IL, 1960).
Ogi, T. et al. Identification of the first ATRIP-deficient patient and novel mutations in ATR define a clinical spectrum for ATR-ATRIP Seckel Syndrome. PLoS Genet. 8, e1002945 (2012).
Acknowledgements
Research was funded by Fundación Botín, by Banco Santander through its Santander Universities Global Division and by grants from the Spanish Ministry of Economy and Competitiveness (MINECO) (SAF2014-59498-R and SAF2014-57791-REDC); these projects were co-financed with European Regional Development funds, the Swedish Research council, Cancerfonden (CAN 2015/674) and the European Research Council (ERC-617840) to O.F.-C. and by a grant from MINECO (BFU2014-55168-JIN) that is co-funded with European Regional Development funds to E.L.
Reviewer information
Nature Reviews Cancer thanks A. Aguilera, D. Durocher and the anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
O.F.-C. and E.L. researched the data for the article, provided substantial contributions to discussions of its content, wrote the article and undertook review and/or editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lecona, E., Fernandez-Capetillo, O. Targeting ATR in cancer. Nat Rev Cancer 18, 586–595 (2018). https://doi.org/10.1038/s41568-018-0034-3
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-018-0034-3
This article is cited by
-
Blocking LBH expression causes replication stress and sensitizes triple-negative breast cancer cells to ATR inhibitor treatment
Oncogene (2024)
-
Targeting ATR in patients with cancer
Nature Reviews Clinical Oncology (2024)
-
Pathway-based signatures predict patient outcome, chemotherapy benefit and synthetic lethal dependencies in invasive lobular breast cancer
British Journal of Cancer (2024)
-
Enhancer-driven transcription of MCM8 by E2F4 promotes ATR pathway activation and glioma stem cell characteristics
Hereditas (2023)
-
Targeting DNA damage response pathways in cancer
Nature Reviews Cancer (2023)