Abstract
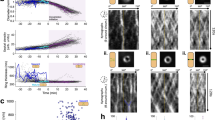
Mechanisms to control cell division are essential for cell proliferation and survival1. Bacterial cell growth and division require the coordinated activity of peptidoglycan synthases and hydrolytic enzymes2,3,4 to maintain mechanical integrity of the cell wall5. Recent studies suggest that cell separation is governed by mechanical forces6,7. How mechanical forces interact with molecular mechanisms to control bacterial cell division in space and time is poorly understood. Here, we use a combination of atomic force microscope imaging, nanomechanical mapping and nanomanipulation to show that enzymatic activity and mechanical forces serve overlapping and essential roles in mycobacterial cell division. We find that mechanical stress gradually accumulates in the cell wall, concentrated at the future division site, culminating in rapid (millisecond) cleavage of nascent sibling cells. Inhibiting cell wall hydrolysis delays cleavage; conversely, locally increasing cell wall stress causes instantaneous and premature cleavage. Cells deficient in peptidoglycan hydrolytic activity fail to locally decrease their cell wall strength and undergo natural cleavage, instead forming chains of non-growing cells. Cleavage of these cells can be mechanically induced by local application of stress with an atomic force microscope. These findings establish a direct link between actively controlled molecular mechanisms and passively controlled mechanical forces in bacterial cell division.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Cabeen, M. T. & Jacobs-Wagner, C. Bacterial cell shape. Nat. Rev. Microbiol. 3, 601–610 (2005).
Egan, A. J. & Vollmer, W. The physiology of bacterial cell division. Ann. NY Acad. Sci. 1277, 8–28 (2013).
Gray, A. N. et al. Coordination of peptidoglycan synthesis and outer membrane constriction during Escherichia coli cell division. eLife 4, e07118 (2015).
Van Der Hofstadt, M., Hüttener, M., Juárez, A. & Gomila, G. Nanoscale imaging of the growth and division of bacterial cells on planar substrates with the atomic force microscope. Ultramicroscopy 154, 29–36 (2015).
Chao, M. C. et al. Protein complexes and proteolytic activation of the cell wall hydrolase RipA regulate septal resolution in mycobacteria. PLoS Pathog. 9, e1003197 (2013).
Zhou, X. et al. Mechanical crack propagation drives millisecond daughter cell separation in Staphylococcus aureus. Science 348, 574–578 (2015).
Zhou, X., Halladin, D. K. & Theriot, J. A. Fast mechanically driven daughter cell separation is widespread in Actinobacteria. mBio 7, e00952-16 (2016).
Monteiro, J. M. et al. Cell shape dynamics during the staphylococcal cell cycle. Nat. Commun. 6, 8055 (2015).
Vijay, S., Anand, D. & Ajitkumar, P. Unveiling unusual features of formation of septal partition and constriction in mycobacteria–an ultrastructural study. J. Bacteriol. 194, 702–707 (2012).
Takade, A., Takeya, K., Taniguchi, H. & Mizuguchi, Y. Electron microscopic observations of cell division in Mycobacterium vaccae V1. J. Gen. Microbiol. 129, 2315–2320 (1983).
Dahl, J. L. Electron microscopy analysis of Mycobacterium tuberculosis cell division. FEMS Microbiol. Lett. 240, 15–20 (2004).
Koch, A. L. Biophysics of bacterial walls viewed as stress-bearing fabric. Microbiol. Rev. 52, 337–353 (1988).
Dufrêne, Y. F. Atomic force microscopy in microbiology: new structural and functional insights into the microbial cell surface. mBio 5, e01363-14 (2014).
Herman-Bausier, P. et al. Mechanical strength and inhibition of the Staphylococcus aureus collagen-binding protein Cna. mBio 7, e01529-16 (2016).
Oh, Y. J. et al. Curli mediate bacterial adhesion to fibronectin via tensile multiple bonds. Sci. Rep. 6, 33909 (2016).
Deng, Y., Sun, M. & Shaevitz, J. W. Direct measurement of cell wall stress stiffening and turgor pressure in live bacterial cells. Phys. Rev. Lett. 107, 158101 (2011).
Touhami, A., Jericho, M. H. & Beveridge, T. J. Atomic force microscopy of cell growth and division in Staphylococcus aureus. J. Bacteriol. 186, 3286–3295 (2004).
Dover, R. S., Bitler, A., Shimoni, E., Trieu-Cuot, P. & Shai, Y. Multiparametric AFM reveals turgor-responsive net-like peptidoglycan architecture in live streptococci. Nat. Commun. 6, 7193 (2015).
Eskandarian, H. A. et al. Division site selection linked to inherited cell surface wave troughs in mycobacteria. Nat. Microbiol. 2, 17094 (2017).
Turner, R. D., Hurd, A. F., Cadby, A., Hobbs, J. K. & Foster, S. J. Cell wall elongation mode in Gram-negative bacteria is determined by peptidoglycan architecture. Nat. Commun. 4, 1496 (2013).
Bailey, R. G. et al. The interplay between cell wall mechanical properties and the cell cycle in Staphylococcus aureus. Biophys. J. 107, 2538–2545 (2014).
Loskill, P. et al. Reduction of the peptidoglycan crosslinking causes a decrease in stiffness of the Staphylococcus aureus cell envelope. Biophys. J. 107, 1082–1089 (2014).
Santi, I., Dhar, N., Bousbaine, D., Wakamoto, Y. & McKinney, J. D. Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria. Nat. Commun. 4, 2470 (2013).
Dufrêne, Y. F., Martínez-Martín, D., Medalsy, I., Alsteens, D. & Müller, D. J. Multiparametric imaging of biological systems by force–distance curve-based AFM. Nat. Methods 10, 847–854 (2013).
Sun, S. X. & Jiang, H. Physics of bacterial morphogenesis. Microbiol. Mol. Biol. Rev. 75, 543–565 (2011).
Francius, G., Domenech, O., Mingeot-Leclercq, M. P. & Dufrêne, Y. F. Direct observation of Staphylococcus aureus cell wall digestion by lysostaphin. J. Bacteriol. 190, 7904–7909 (2008).
Wheeler, R. et al. Bacterial cell enlargement requires control of cell wall stiffness mediated by peptidoglycan hydrolases. mBio 6, e00660-15 (2015).
Ruggiero, A. et al. Structure and functional regulation of RipA, a mycobacterial enzyme essential for daughter cell separation. Structure 18, 1184–1190 (2010).
Hett, E. C. et al. A partner for the resuscitation-promoting factors of Mycobacterium tuberculosis. Mol. Microbiol. 66, 658–668 (2007).
Hett, E. C., Chao, M. C., Deng, L. L. & Rubin, E. J. A mycobacterial enzyme essential for cell division synergizes with resuscitation-promoting factor. PLoS Pathog. 4, e1000001 (2008).
Jiang, Y., Liang, X., Guo, M., Cao, Y. & Cai, S. Fracture mechanics modeling of popping event during daughter cell separation. Biomech. Model. Mechanobiol. 17, 1131–1137 (2018).
Thanky, N. R., Young, D. B. & Robertson, B. D. Unusual features of the cell cycle in mycobacteria: polar-restricted growth and the snapping-model of cell division. Tuberculosis 87, 231–236 (2007).
Smolyakov, G., Formosa-Dague, C., Severac, C., Duval, R. E. & Dague, E. High speed indentation measures by FV, QI and QNM introduce a new understanding of bionanomechanical experiments. Micron 85, 8–14 (2016).
Atilgan, E., Magidson, V., Khodjakov, A. & Chang, F. Morphogenesis of the fission yeast cell through cell wall expansion. Curr. Biol. 25, 2150–2157 (2015).
Meniche, X. et al. Subpolar addition of new cell wall is directed by DivIVA in mycobacteria. Proc. Natl Acad. Sci. USA 111, E3243–E3251 (2014).
Odermatt, P. D. et al. High-resolution correlative microscopy: bridging the gap between single molecule localization microscopy and atomic force microscopy. Nano Lett. 15, 4896–4904 (2015).
Acknowledgements
We thank E. J. Rubin for generously providing the RipA conditional depletion strain of M. smegmatis. This research was supported in part by grants to G.E.F. from the Swiss National Science Foundation (205321_134786 and 205320_152675), from the European Union FP7/2007-2013/ERC under grant agreement no. 307338-NaMic, H2020 - UE Framework Programme for Research & Innovation (2014-2020); ERC-2017-CoG; InCell; Project number 773091, from the Commission for Technology and Innovation under CTI no. 18330.1 PFNM-NM, and by a grant to J.D.M. from the Swiss National Science Foundation (310030B_176397). H.A.E. was supported by an EMBO Long Term Fellowship (191-2014) and an EMBO Advanced Long Term Fellowship (750-2016).
Author information
Authors and Affiliations
Contributions
P.D.O., G.E.F. and J.D.M. conceptualized the study. P.D.O., M.T.M.H. and H.A.E. developed imaging protocols. P.D.O., M.T.M.H., G.E.F. and J.D.M. designed experiments. P.D.O. and M.T.M.H. performed the experiments and analysed the data. A.P.N. participated in building the instrument and simulations. P.D.O., M.T.M.H., G.E.F. and J.D.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Physics thanks Yves Dufrene and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17, Table 1 and refs. 1 and 2.
Supplementary Video 1
A time sequence of M. smegmatis cells undergoing abrupt cleavage imaged in QNM with a low force setpoint. Frame rate: 5 min.
Supplementary Video 2
A time sequence of stiffness increase at the PCF imaged in QNM with a higher force setpoint. Frame rate: 5 min.
Supplementary Video 3
A time sequence of an alternative mechanism of separation of sibling cells when one sibling cell is deflated by piercing with the AFM cantilever tip. Sibling cells do not undergo rapid cell cleavage; instead, the surviving sibling gradually sheds the deflated sibling.
Rights and permissions
About this article
Cite this article
Odermatt, P.D., Hannebelle, M.T.M., Eskandarian, H.A. et al. Overlapping and essential roles for molecular and mechanical mechanisms in mycobacterial cell division. Nat. Phys. 16, 57–62 (2020). https://doi.org/10.1038/s41567-019-0679-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-019-0679-1
This article is cited by
-
Force spectroscopy of single cells using atomic force microscopy
Nature Reviews Methods Primers (2021)
-
Nanomechanical mechanisms of Lyme disease spirochete motility enhancement in extracellular matrix
Communications Biology (2021)
-
A biphasic growth model for cell pole elongation in mycobacteria
Nature Communications (2020)
-
Use the force
Nature Physics (2020)