Abstract
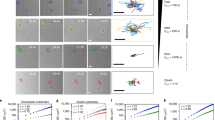
Cell migration over heterogeneous substrates during wound healing or morphogenetic processes leads to shape changes driven by different organizations of the actin cytoskeleton and by functional changes including lamellipodial protrusions and contractile actin cables. Cells distinguish between cell-sized positive and negative curvatures in their physical environment by forming protrusions at positive curvatures and actin cables at negative curvatures; however, the cellular mechanisms remain unclear. Here, we report that concave edges promote polarized actin structures with actin flow directed towards the cell edge, in contrast to well-documented retrograde flow at convex edges. Anterograde flow and contractility induce a tension anisotropy gradient. A polarized actin network is formed, accompanied by a local polymerization–depolymerization gradient, together with leading-edge contractile actin cables in the front. These cables extend onto non-adherent regions while still maintaining contact with the substrate through focal adhesions. The contraction and dynamic reorganization of this actin structure allows forward movements enabling cell migration over non-adherent regions on the substrate. These versatile functional structures may help cells sense and navigate their environment by adapting to external geometric and mechanical cues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data and custom written code are available upon request.
References
Giannone, G. et al. Lamellipodial actin mechanically links myosin activity with adhesion-site formation. Cell 128, 561–575 (2007).
Ravasio, A. et al. Gap geometry dictates epithelial closure efficiency. Nat. Commun. 6, 7683 (2015).
Wood, W. et al. Wound healing recapitulates morphogenesis in Drosophila embryos. Nat. Cell Biol. 4, 907–912 (2002).
Behrndt, M. et al. Forces driving epithelial spreading in zebrafish gastrulation. Science 338, 257–260 (2012).
Mogilner, A. On the edge: modeling protrusion. Curr. Opin. Cell Biol. 18, 32–39 (2006).
Gardel, M. L. et al. Traction stress in focal adhesions correlates biphasically with actin retrograde flow speed. J. Cell. Biol. 183, 999–1005 (2008).
Kiehart, D. P., Galbraith, C. G., Edwards, K. A., Rickoll, W. L. & Montague, R. A. Multiple forces contribute to cell sheet morphogenesis for dorsal closure in Drosophila. J. Cell. Biol. 149, 471–490 (2000).
Abreu-Blanco, M. T., Verboon, J. M., Liu, R., Watts, J. J. & Parkhurst, S. M. Drosophila embryos close epithelial wounds using a combination of cellular protrusions and an actomyosin purse string. J. Cell. Sci. 125, 5984–5997 (2012).
Friedl, P., Locker, J., Sahai, E. & Segall, J. E. Classifying collective cancer cell invasion. Nat. Cell Biol. 14, 777–783 (2012).
Klarlund, J. K. Dual modes of motility at the leading edge of migrating epithelial cell sheets. Proc. Natl Acad. Sci. USA 109, 15799–15804 (2012).
Omelchenko, T., Vasiliev, J. M., Gelfand, I. M., Feder, H. H. & Bonder, E. M. Rho-dependent formation of epithelial ‘leader’ cells during wound healing. Proc. Natl Acad. Sci. USA 100, 10788–10793 (2003).
Reffay, M. et al. Interplay of RhoA and mechanical forces in collective cell migration driven by leader cells. Nat. Cell Biol. 16, 217–223 (2014).
Parker, K. K. et al. Directional control of lamellipodia extension by constraining cell shape and orienting cell tractional forces. FASEB J. 16, 1195–1204 (2002).
James, J., Goluch, E. D., Hu, H., Liu, C. & Mrksich, M. Subcellular curvature at the perimeter of micropatterned cells influences lamellipodial distribution and cell polarity. Cell Motil. Cyto. 65, 841–852 (2008).
Rossier, O. M. et al. Force generated by actomyosin contraction builds bridges between adhesive contacts. EMBO J. 29, 1055–1068 (2010).
Peter, B. J. et al. BAR domains as sensors of membrane curvature: the amphiphysin BAR structure. Science 303, 495–499 (2004).
Bridges, A. A., Jentzsch, M. S., Oakes, P. W., Occhipinti, P. & Gladfelter, A. S. Micron-scale plasma membrane curvature is recognized by the septin cytoskeleton. J. Cell. Biol. 213, 23–32 (2016).
Rangamani, P. et al. Decoding information in cell shape. Cell 154, 1356–1369 (2013).
Elliott, H. et al. Myosin II controls cellular branching morphogenesis and migration in three dimensions by minimizing cell-surface curvature. Nat. Cell Biol. 17, 137–147 (2015).
Grasso, S., Hernández, J. A. & Chifflet, S. Roles of wound geometry, wound size, and extracellular matrix in the healing response of bovine corneal endothelial cells in culture. Am. J. Physiol. Cell Physiol. 293, 37 (2007).
Ravasio, A. et al. Regulation of epithelial cell organization by tuning cell-substrate adhesion. Integr. Biol. 7, 1228–1241 (2015).
Vedula, S. R. et al. Mechanics of epithelial closure over non-adherent environments. Nat. Commun. 6, 6111 (2015).
Giannone, G. et al. Periodic lamellipodial contractions correlate with rearward actin waves. Cell 116, 431–443 (2004).
Koestler, S. A., Auinger, S., Vinzenz, M., Rottner, K. & Small, J. V. Differentially oriented populations of actin filaments generated in lamellipodia collaborate in pushing and pausing at the cell front. Nat. Cell Biol. 10, 306–313 (2008).
Ponti, A., Machacek, M., Gupton, S. L., Waterman-Storer, C. M. & Danuser, G. Two distinct actin networks drive the protrusion of migrating cells. Science 305, 1782–1786 (2004).
Burnette, D. T. et al. A role for actin arcs in the leading-edge advance of migrating cells. Nat. Cell Biol. 13, 371–381 (2011).
Anon, E. et al. Cell crawling mediates collective cell migration to close undamaged epithelial gaps. Proc. Natl Acad. Sci. USA 109, 10891–10896 (2012).
Vedula, S. R. et al. Microfabricated environments to study collective cell behaviors. Methods Cell Biol. 120, 235–252 (2014).
Burnette, D. T. et al. A contractile and counterbalancing adhesion system controls the 3D shape of crawling cells. J. Cell. Biol. 205, 83–96 (2014).
Tee, Y. H. et al. Cellular chirality arising from the self-organization of the actin cytoskeleton. Nat. Cell Biol. 17, 445–457 (2015).
Charras, G. & Sahai, E. Physical influences of the extracellular environment on cell migration. Nat. Rev. Mol. Cell Biol. 15, 813–824 (2014).
Guetta-Terrier, C. et al. Protrusive waves guide 3D cell migration along nanofibers. J. Cell. Biol. 211, 683–701 (2015).
Gupta, M. et al. Adaptive rheology and ordering of cell cytoskeleton govern matrix rigidity sensing. Nat. Commun. 6, 7525 (2015).
Hotulainen, P. & Lappalainen, P. Stress fibers are generated by two distinct actin assembly mechanisms in motile cells. J. Cell. Biol. 173, 383–394 (2006).
Waterman-Storer, C. M., Desai, A., Bulinski, J. C. & Salmon, E. D. Fluorescent speckle microscopy, a method to visualize the dynamics of protein assemblies in living cells. Curr. Biol. 8, 1227–1230 (1998).
Mayer, M., Depken, M., Bois, J. S., Jülicher, F. & Grill, S. W. Anisotropies in cortical tension reveal the physical basis of polarizing cortical flows. Nature 467, 617–621 (2010).
Callan-Jones, A. C. & Voituriez, R. Actin flows in cell migration: from locomotion and polarity to trajectories. Curr. Opin. Cell Biol. 38, 12–17 (2016).
Salbreux, G., Prost, J. & Joanny, J. F. Hydrodynamics of cellular cortical flows and the formation of contractile rings. Phys. Rev. Lett. 103, 058102 (2009).
Brugues, A. et al. Forces driving epithelial wound healing. Nat. Phys. 10, 683–690 (2014).
Balaban, N. Q. et al. Force and focal adhesion assembly: a close relationship studied using elastic micropatterned substrates. Nat. Cell Biol. 3, 466–472 (2001).
Juelicher, F., Kruse, K., Prost, J. & Joanny, J. F. Active behavior of the cytoskeleton. Phys. Rep. 449, 3–28 (2007).
Tao, J., Li, Y., Vig, D. K. & Sun, S. X. Cell mechanics: a dialogue. Rep. Prog. Phys. 80, 036601 (2017).
Joanny, J. F., Kruse, K., Prost, J. & Ramaswamy, S. The actin cortex as an active wetting layer. Euro. Phys. J. E 36, 52 (2013).
Tinevez, J. Y. et al. Role of cortical tension in bleb growth. Proc. Natl Acad. Sci. USA 106, 18581–18586 (2009).
Wottawah, F. et al. Optical rheology of biological cells. Phys. Rev. Lett. 94, 098103 (2005).
Svitkina, T. M. & Borisy, G. G. Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia. J. Cell. Biol. 145, 1009–1026 (1999).
Serrels, B. et al. Focal adhesion kinase controls actin assembly via a FERM-mediated interaction with the Arp2/3 complex. Nat. Cell Biol. 9, 1046–1056 (2007).
Maiuri, P. et al. Actin flows mediate a universal coupling between cell speed and cell persistence. Cell 161, 374–386 (2015).
Guo, W. H. & Wang, Y. L. Retrograde fluxes of focal adhesion proteins in response to cell migration and mechanical signals. Mol. Biol. Cell 18, 4519–4527 (2007).
Bryant, D. M. et al. A molecular network for de novo generation of the apical surface and lumen. Nat. Cell Biol. 12, 1035–1045 (2010).
Seddiki, R. et al. Force-dependent binding of vinculin to α-catenin regulates cell–cell contact stability and collective cell behavior. Mol. Biol. Cell 29, 380–388 (2018).
Nelson, C. M. et al. Emergent patterns of growth controlled by multicellular form and mechanics. Proc. Natl Acad. Sci. USA 102, 11594–11599 (2005).
Cramer, L. P. Role of actin-filament disassembly in lamellipodium protrusion in motile cells revealed using the drug jasplakinolide. Curr. Biol. 9, 1095–1105 (1999).
Picone, R., Baum, B. & McKendry, R. Plasma microcontact patterning (PmuCP): a technique for the precise control of surface patterning at small-scale. Methods Cell Biol. 119, 73–90 (2014).
Mendoza, M. C., Besson, S. & Danuser, G. Quantitative fluorescent speckle microscopy (QFSM) to measure actin dynamics. Curr. Protoc. Cytom. 62, 2.18.1–2.18.26 (2012).
Acknowledgements
The authors thank D. Delacour, N. Gov, R.-M. Mège, group members from MBI and IJM for helpful discussions and critical reading of the manuscript. The authors would also like to thank MBI Microfabrication (G. Grenci and M. Asraf), MBI Science Communication Core (C.X. Wong and S. Wolf) and MBI Microscopy Core (F. Margadant) for continuous support. The authors are grateful for the ImagoSeine core facility of the Institut Jacques Monod, member of IBiSA and the France-BioImaging (ANR-10-INBS-04) infrastructure for their support. The authors thank A. Bershadsky, M. Sheetz and P. Kanchanawong for sharing of plasmid constructs. The authors are grateful to W. J. Nelson and F. Martin-Belmonte for their generous gift of MDCK cell lines. Financial support from the Mechanobiology Institute, the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007-2013) / ERC grant agreement numbers 617233 (B.L.) and 616480 (X.T.), USPC-NUS collaborative program, Agence Nationale de la Recherche (ANR) (“POLCAM” (ANR-17- CE13-0013), CODECIDE (ANR-17-CE-13-0022)), the LABEX ‘Who am I?’, and the Spanish Ministry of Economy and Competitiveness (BFU2015-65074-P) are gratefully acknowledged. T.C. acknowledges the USPC-NUS international PhD mobility programme. T.S. is supported by the Israel Science Foundation (grant number 921/15).
Author information
Authors and Affiliations
Contributions
B.L., R.V., T.C. and A.R. conceived the project. B.L. and R.V. supervised the project. T.C. performed the experiments. A.B. and X.T. helped with traction force experiments and data analysis. Y.T. established the laser ablation system and helped with the related experiments. T.C. performed image and data analysis with the help of H.T.O. The theoretical model was developed by A.C.-J. and R.V., while E.F. and T.S. performed the numerical simulation. B.C.L., A.R. and Y.T. contributed resources to the project. The article was written by T.C., A.C.J., R.V. and B.L., and read by all authors, who all contributed to the interpretation of the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Sections 1–4, Supplementary References 1–10, Supplementary Figures 1–6, Supplementary Table 1, Supplementary Video Captions 1–16.
Supplementary Video 1
MDCK cells migrate over a non-adherent circular island. Images were taken with wide field TIRF, mApple Actin shown in magenta and GFP MRLC shown in green. Scale bar, 10 µm.
Supplementary Video 2
Phase contrast video showing MDCK WT cells on fibronectin pattern (magenta line). Scale bar, 10 µm.
Supplementary Video 3
Live SIM video of MDCK GFP actin cell at negative curvature on fibronectin pattern (magenta line) shows anterograde flow. See Fig. 3a for colour code. Scale bar, 5 µm.
Supplementary Video 4
Photoconversion of radial fibre in mEos-actin (before conversion-green, after conversion-magenta) transfected MDCK cell at negative curvature on fibronectin pattern (white line). Images were acquired with a scanning confocal microscope showing anterograde flow of radial fibres. Scale bar, 2 µm.
Supplementary Video 5
Fluorescent speckle microscope of mEmerald-actin transfected MDCK cell at negative curvature on fibronectin pattern (magenta line) shows anterograde flow of speckles. Imaged with a wide-field TIRF microscope. Scale bar, 5 µm.
Supplementary Video 6
Fluorescent speckle microscope of mEmerald-actin transfected MDCK cell at positive curvature on fibronectin pattern (magenta line) shows retrograde flow of speckles. Imaged with a wide-field TIRF microscope. Scale bar, 5 µm.
Supplementary Video 7
Laser ablations of TFs in a MDCK cell expressing GFP-MRLC at negative curvature imaged with a scanning confocal microscope. Scale bar, 5 µm.
Supplementary Video 8
Myosin flow in control, blebbistatin treated and washout cells. MCDK cells were transfected with mApple Myosin IIA and imaged with a spinning disk confocal microscope with a super-resolution module. Video shows the same cell imaged under the three conditions. Scale bar, 5µm.
Supplementary Video 9
Photoconversion of a square ROI region (yellow box) in a MDCK cell transfected with mEos2-actin at negative curvature (white line shows edge of fibronectin pattern), imaged with scanning confocal microscope. Green channel shows unconverted fluorescence, magenta channel shows converted fluorescence. Scale bar, 5 µm.
Supplementary Video 10
Live SIM video of MDCK GFP actin cells showing actin dynamics in the centre region at negative curvature. Cells were treated with 200 µM CK666 for 30 min before washout to remove the underlying actin fibres (due to contractility induced anterograde flow of previously formed fibres and no new fibre formation with inhibition of actin polymerization) and reveal the dynamics of newly polymerized actin filaments. Edge of fibronectin pattern is shown with a magenta line. Scale bar, 5 µm.
Supplementary Video 11
1 nM jasplakinolide treatment induced the collapse of actin structure. MDCK cells were transfected with mApple Actin (magenta) and GFP Myosin IIA (green) and imaged with a spinning disk confocal microscope with a super-resolution module. Edge of fibronectin pattern is shown with a white line. Scale bar, 5µm.
Supplementary Video 12
Severance of the radial fibres with laser ablation induces recruitment of Arp2/3 and polymerization of actin at the rear of the ablation site. Scanning confocal images are shown for mApple actin (left) and mEmerald Arp-p34 transfected MDCK cell (right). See Fig. 5k for colour code. White line marks the edge of fibronectin pattern. Scale bar, 5 µm.
Supplementary Video 13
Laser ablation of actin cable out of the adherent pattern (left) induced a retraction of the actin structure whereas ablation of actin cable on the adherent pattern (right) had no effect. MDCK GFP MRLC cells were imaged with a scanning confocal microscope. Yellow line marks the edge of the fibronectin pattern (30 µm radius negative curvature). Scale bar, 10 µm.
Supplementary Video 14
MDCK monolayer could not migrate over non-adherent circle of 100 µm diameter. MDCK GFP MRLC (green) MDCK cells were transfected with mApple Actin (magenta) and were imaged with a wide-field TIRF microscope. Fibronectin pattern is shown in cyan. Scale bar, 20 µm.
Supplementary Video 15
MDCK α-catenin KD cells migrate over non-adherent circle of 30 µm diameter. Cells were transfected with mApple Actin (magenta) and mEmerald Myosin IIA (green) and were imaged with a wide-field TIRF microscope. Fibronectin pattern is shown in cyan. Scale bar, 10 µm.
Supplementary Video 16
Laser ablation of actin cable at the edge of an expanding MDCK monolayer on a continuous substrate resulted in transformation into lamellipodium when the edge was of positive curvature (left) and resulted in actin cable repair when the edge was of negative curvature (right). Cells were transfected with mApple Actin and imaged with a scanning confocal microscope. Scale bar, 10 µm.
Rights and permissions
About this article
Cite this article
Chen, T., Callan-Jones, A., Fedorov, E. et al. Large-scale curvature sensing by directional actin flow drives cellular migration mode switching. Nat. Phys. 15, 393–402 (2019). https://doi.org/10.1038/s41567-018-0383-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-018-0383-6
This article is cited by
-
How multiscale curvature couples forces to cellular functions
Nature Reviews Physics (2024)
-
Alteration of actin cytoskeletal organisation in fetal akinesia deformation sequence
Scientific Reports (2024)
-
Switch of cell migration modes orchestrated by changes of three-dimensional lamellipodium structure and intracellular diffusion
Nature Communications (2023)
-
Emergent collective organization of bone cells in complex curvature fields
Nature Communications (2023)
-
Mechanical stress driven by rigidity sensing governs epithelial stability
Nature Physics (2023)